Abstract
Introduction
Berberis vulgaris extract is a well-known herbal remedy for treating numerous conditions including coronavirus disease 2019 (COVID-19). The HAB4a method of preparing the extract is a common method used in clinical practice and a further aim of this study was to identify the key alkaloid compounds that are present in an alcohol versus a glycerine-based extraction and determine the mechanism by which these prevent and treat COVID-19.
Results and Discussion
Identified alkaloid compounds included bervulcine, vulracine, berberrubine, lambertine, berbidine, obamegine, berbamine, berberine, and palmatine in the alcohol-based extract. The glycerine-based extract contained all except bervulcine, vulracine, and berbidine. Berberine and palmatine were the most abundant compounds in both extracts but more berberine was present in the alcohol-based extract and more palmatine in the glycerine-based extraction. Databases revealed compound-gene interactions with human targets involved in signalling and gene regulation networks that could synergistically change cell function. The SARS-CoV-2 spike 1 protein purification yielded about 5 mg of highly purified protein (˃95%) from 1 L of bacterial culture. Isothermal titration calorimetry (ITC) analysis of any potential interaction of palmatine and berberine with the purified spike 1 protein was negative. Thus, it is possible the extract components work in a synergistic manner to prevent and treat COVID-19.
Conclusions
Berberis vulgaris root extract alkaloid components differ when alcohol or glycerine is used. Berberine and palmatine are the most abundant alkaloids but differ in the amount relative to each other in the extracts. The extract components could affect changes in a variety of human genes.
Materials and Methods
Ultra-High-Performance Liquid Chromatography—High-Definition quadrupole time-of-flight Mass Spectrometry was used to identify alkaloids. Databases were used to determine the genes that interact with the identified compounds. SARS-CoV-2 spike protein was expressed and purified. Purified protein was used in ITC with berberine and palmatine.
Introduction
Berberis vulgaris is a thorny shrub from the Berberidaceae family of plants, it can reach a height of up to 3 m and is indigenous to the Asian and European continents. Known to be a versatile plant, it produces small red berries that are edible, but are mostly free of any alkaloids, thus making their value completely nutritional. The stem bark, root bark, and roots are known to have therapeutic value 1 and have been shown to possess anti-fungal properties, 2 antiviral, 3 antioxidant, and anti-inflammatory 4 properties. Many of these listed properties closely align with those that could potentially help prevent and treat COVID-19 caused by the SARS-CoV-2 virus.
SARS-CoV-2 virus contains a single-stranded RNA genome that codes for structural proteins such as the spike, nucleocapsid, membrane protein, and envelope small membrane protein. Of the structural proteins, the spike protein is critical in cell entry as it interacts with the human angiotensin-converting enzyme (ACE2) and facilitates cellular entry of the virus. 5 The structural proteins are produced along with other nonstructural proteins (15-16 in total) which include the key proteases, papain-like protease (PLpro), and the main chymotrypsin-like protease (Mpro). The proteases are responsible for cleaving the polyprotein synthesized from the RNA genome to produce the proteins required to form new virus particles to be released. An RNA-dependant polymerase is responsible for replicating the RNA genome which is packaged with the proteins to form virus particles.
The COVID-19 pandemic has severely impacted the world, with the World Health Organization reporting more than 767 million cases and 6.9 million deaths (https://covid19.who.int/ accessed June 19, 2023). Identifying remedies that can be used to prevent and treat the causative virus SARS-CoV-2 is required as society prepares to live with the virus. Compounds or mixtures of natural compounds that can interact with key human or viral proteins could be used to prevent infection, treat those already infected, and prevent severe disease. Berberis vulgaris has been traditionally used in herbal preparations, with its’ main indication for therapeutical use being gastrointestinal as well as urinary tract disorders and pathologies. 1 To a lesser degree in both herbal and homeopathic preparations, it is indicated in catarrhal conditions affecting the upper respiratory tract and employed to alleviate fevers. 6 With current scientific measures employed, it is evident that Protoberberine alkaloids found in the herbal extract Berberis vulgaris can intercalate DNA, inhibiting neuroreceptors and various enzymes, but also demonstrate antiviral, antifungal, amoebicidal, and antibacterial activity.1,3
A study by Nuralin and Gürü 7 conducted a supercritical carbon dioxide extraction of compounds from the fruit of Berberis vulgaris and identified antioxidants rutin and apigenin. Rutin was suggested to be a good treatment compound for COVID-19. Gholizadeh-Moghadam and colleagues 8 identified additional compounds in the fruit extracts, namely gallic acid, caffeic acid, chlorogenic acid, p-coumaric acid, cinnamic acid, and quercetin and indicated that Berberis vulgaris contained the highest amount of phenolic and flavonoids out of the 18 Berberis species fruit samples analyzed. In an analysis of the root and twig extracts of Berberis vulgaris, 9 berberine was identified as being present in a very high concentration and it was thought the antimicrobial activity was mainly due to the presence of this compound. In general, the key compound berberine has been thought to account for the varied effects of the Berberis vulgaris extracts. Clinical studies using berberine have been conducted for a range of diseases such as diabetes, cancer, polycystic ovarian syndrome to diarrhea to name a few ( 10 references therein). Berberine, however, is not well absorbed when taken orally. 10 A clinical study showed some promise in combining berberine with an artichoke extract to enhance its bioavailability. 11 Berberine has also been examined for its use as a COVID-19 treatment. In a clinical study, no significant improvement was seen in the patients treated with 300 mg berberine alone over 2 weeks. 12 As a singular compound berberine lacked efficacy, but there is possible merit in looking at a group of compounds that might be acting synergistically to prevent or treat COVID-19. In a study conducted by Imanshahidi and Hosseinzadeh, 13 it was indicated that along with berberine other key alkaloids found in Berberis vulgaris include berbamine and palmatine. Palmatine along with other root extract compounds such as berbamine from Berberis Integerrim inhibited the ACE-2 receptor and hence could be useful in preventing and treating SARS-CoV-2 infection. 14 Berbamine has been shown 15 to independently have antiviral activity against SARS-CoV-2.
The Berberis vulgaris extract is used as a remedy to prevent and treat COVID-19 in a complementary health practice in South Africa. The HAB4a method is used by the practitioner to extract compounds in both alcohol and glycerine. Therefore, this study reports the compounds extracted from dried Berberis vulgaris roots in both alcohol and glycerine using the HAB4a method. Ultra-high-performance liquid chromatography (UPLC) coupled to a high-definition quadrupole time-of-flight mass spectrometer (HD QTOF) was used to profile the ethanol and glycerine extracts. Berberine and palmatine were identified as compounds present in the extract along with a few other compounds. The potential human and viral targets these compounds are known to interact with are discussed.
Materials and Methods
Extract Preparation
Two Berberis vulgaris extracts were prepared from the crushed and dried bark of the plant for the UPLC—High-Definition Mass Spectrometry study. One extract was prepared in 70% ethanol alcohol and the second in 100% glycerine. Both samples of dry matter of Berberis vulgaris bark were weighed at 14 g and extract mediums were each measured at 90 mL, this gave a concentration of 150 mg/mL. The extracts were prepared under hygienic dispensary conditions, extracted in amber glass bottles out of direct sunlight, and stored at a temperature below 23 °C, ensuring no denaturing of the extract could occur. An extraction time of 10 days or 240 h was permitted to ensure a high-quality extract. The dry material used in the extract was purchased from Pharma Germania, and the batch number was identified as; 06202/1.
Ultra-High-Performance Liquid Chromatography—High-Definition Quadrupole Time-of-Flight Mass Spectrometry (UHPLC–HD-QTOF-MS)
A Waters Classic binary UPLC coupled in series to a Waters SYNAPT G1 HDMS mass spectrometer was used to produce chromatographic resolution and mass spectral data. Method development and optimization of the chromatographic separation was done utilizing a Waters HSS T3 C18 column (150 mm × 2.1 mm, 1.8 µm), and the column temperature was controlled at 60 °C. A binary solvent mixture was used consisting of water (Eluent A) containing 10 mM formic acid (pH 2.3) and acetonitrile (Eluent B) containing 10 mM formic acid. The initial conditions were 95%A at a flow rate of 0.4 mL/min and were maintained for 1 min, followed by a gradient to 1%A at 15 min. The conditions were kept constant for 2 min and then changed to the initial conditions. The runtime was 20 min and the injection volume was 1 µL. Samples were kept cool at 8 °C in the Waters Sample Manager during the analysis.
The SYNAPT G1 mass spectrometer was used in V-optics mode and operated with an electrospray probe to enable the detection of all ESI-compatible compounds. Leucine enkephalin (50 pg/mL) was used as a reference calibrant (Lock Mass) to obtain typical mass accuracies between 1 and 5 mDalton (mDa). The mass spectrometer was operated in both ESI positive and negative modes with a capillary voltage of 2.5 kV, the sampling cone at 30 V, and the extraction cone at 4.0 V. The scan time was 0.2 s covering the 50 to 1500 Dalton mass range with an interscan time of 0.02 s. The source temperature was 120 °C, and the desolvation temperature was set at 450 °C. Nitrogen gas was used as the nebulization gas at a flow rate of 550 L/h, and cone gas was added at 50 L/h. The software used to control the hyphenated system and do all data manipulation was MassLynx 4.1 (SCN 872).
Identifying Human and SARS-CoV-2 Known Gene Targets Linked to Compounds
Compounds identified in the Berberis vulgaris extracts were entered into the Comparative Toxicogenomics Database, MDI Biological Laboratory, Salisbury Cove, Maine, and NC State University, Raleigh, North Carolina. World Wide Web (URL: http://ctdbase.org/) [March-May, 2023] as a chemical-gene search query. PubChem was also used to obtain possible synonyms which were also used to search the database for potential gene interactions. To obtain basic information about the genes and gene products identified to interact with the compounds, links available on the entry were used to access the human gene equivalent entry in the Gene database of the NCBI Bethesda (MD): National Library of Medicine (US), National Center for Biotechnology Information; 2004 – [2023: 03–05]. Available from: https://www.ncbi.nlm.nih.gov/gene/. Additional literature searches were conducted using search strings “[Gene name] SARS-CoV-2 COVID-19” using multiple search engines. Data from the various databases was manually integrated to identify clear links to a possible mechanism by which the gene-compound interaction and resulting effect might affect COVID-19 disease or prevent infection.
SARS-CoV-2 Spike (S1) Protein Expression and Purification
The S1 protein was overexpressed in Escherichia coli T7 cells by isopropyl β-D-1-thiogalactopyranoside (IPTG) induction. The insoluble SARS-CoV-2 S1 protein was purified using established procedures involving centrifugation steps and nickel affinity chromatography. Briefly, E coli cells harboring pET-32(a) + plasmid DNA were grown at 37 °C in LB broth with ampicillin (100 μg/mL). The overnight culture was diluted 100-fold into fresh LB medium containing ampicillin (100 μg/mL). The culture was allowed to grow for 3 h at 37 °C. IPTG was added to a final concentration of 1.0 mM when the optical density, measured at 600 nm of the culture medium reached 0.6 to 0.7. The cultures were allowed to grow for 4 h. The cells were harvested by centrifugation at 4000 × g and stored at −20 °C overnight. The cells were then resuspended in a minimal volume of ice-cold extraction Buffer A (20 mM Tris, 1 M NaCl, 5 mM imidazole, 0.25% CHAPS, pH 8.0). MgCl2 and DNase I were added to the final concentrations of 10 mM and 10 U/μL, respectively. When the viscosity of the mixture had decreased, the cells were ruptured by sonication and homogenization. The cell homogenate was centrifuged at 10,000 × g for 30 min. The supernatant was discarded and the pellet which contained the majority of the S1 protein expressed as inclusion bodies was then dissolved in Buffer B (20 mM Tris, 1 M NaCl, 8 M urea, 5 mM imidazole, pH 8.0) to solubilize the S1 protein. The solubilized S1 protein was passed through a HiTrap chelating FPLC column (GE Healthcare), charged with Ni2+, and equilibrated with Buffer B. Thereafter the S1 protein was eluted with a 75 mL gradient of 150 to 600 mM imidazole in Buffer B. The S1 protein-containing fractions, as determined by 12% SDS-PAGE, were pooled and EDTA was added to a final concentration of 5 mM. Then the protein was refolded by stepwise dialysis into Buffer A supplemented with 5 mM DTT and 10% glycerol. Folded SARS-CoV-2 S1 protein was desalted into storage buffer 20 mM Tris-HCl, 1 M NaCl, 5 mM DTT, and 10% glycerol, pH 8.0 using PD-10 gel filtration column (Amersham, Pharmacia Biotech) and aliquoted into cryotubes, flash-frozen in liquid N2 and stored at −80 °C. The concentration of the purified SARS-CoV-2 S1 protein was determined using the calculated extinction coefficient of 90,650 M−1cm−1 at 280 nm (Chirascan™ Plus, Applied Photophysics).
Isothermal Titration Calorimetry—SARS CoV-2 Spike 1 With Palmatine and Berberine
Isothermal Titration Calorimetry (ITC) tests were performed using Affinity ITC (TA Instruments) at 25 °C. Prior, to loading the SARS-CoV-2 spike 1 protein (S1) (20 µM) into the sample cell, the protein was degassed to remove any bubbles. Then 10 mM of either Berberis hemisulfate salt or palmatine chloride was used in the titration syringe. Total volumes of 350 µL and 250 µL were loaded into the sample cell and titration syringe, respectively. Titrations were performed by injecting 10 µL of the ligand (Berberis hemisulfate salt or palmatine chloride) into the protein at 200-s intervals and stirring at 80 rpm. The ITC data were analyzed using NanoAnalyze (TA Instruments).
Results
Compounds Identified in an Ethanol and Glycerine Extract of Berberis vulgaris
Two extracts of the bark of Berberis vulgaris were prepared and assessed by UPLC–HD-QTOF-MS (Figure 1). The analysis was conducted in positive ionization mode covering a mass range of 50 to 1500 Dalton. Table 1 is a list of compounds identified in the 2 extracts, the compound identification remains tentative as no pure analytical standards were used. One extract was prepared in medical alcohol and the other in glycerine. The alcohol preparation is routinely used for the prevention and treatment of SARS-CoV-2 infection but in children the glycerine extract is the preferred option. Although ESI as an ionization technique is compound specific preferentially highlighting compounds that ionize under defined conditions, it was still evident that berberine and palmatine were the main compounds present in the extracts followed by lambertine. These 2 compounds however co-eluted and were detected as one peak. The alcohol-based extract produced more berberine than palmatine, but the glycerine extract yielded significantly more palmatine than berberine. MS/MS analysis of the ethanol extract at selected masses (336 Dalton for berberine and 352 Dalton for palmatine), displayed the concentration differences of these 2 compounds in the ethanol extract (Figure 2). Similarly, MS/MS analysis of the glycerine extract as described above for the ethanol extract clearly displayed the higher palmatine concentration in relation to the berberine concentration (Figure 3).

ESIPos BPI chromatograms of the Berberis vulgaris EtOH extract (red) and glycerine extract (green).
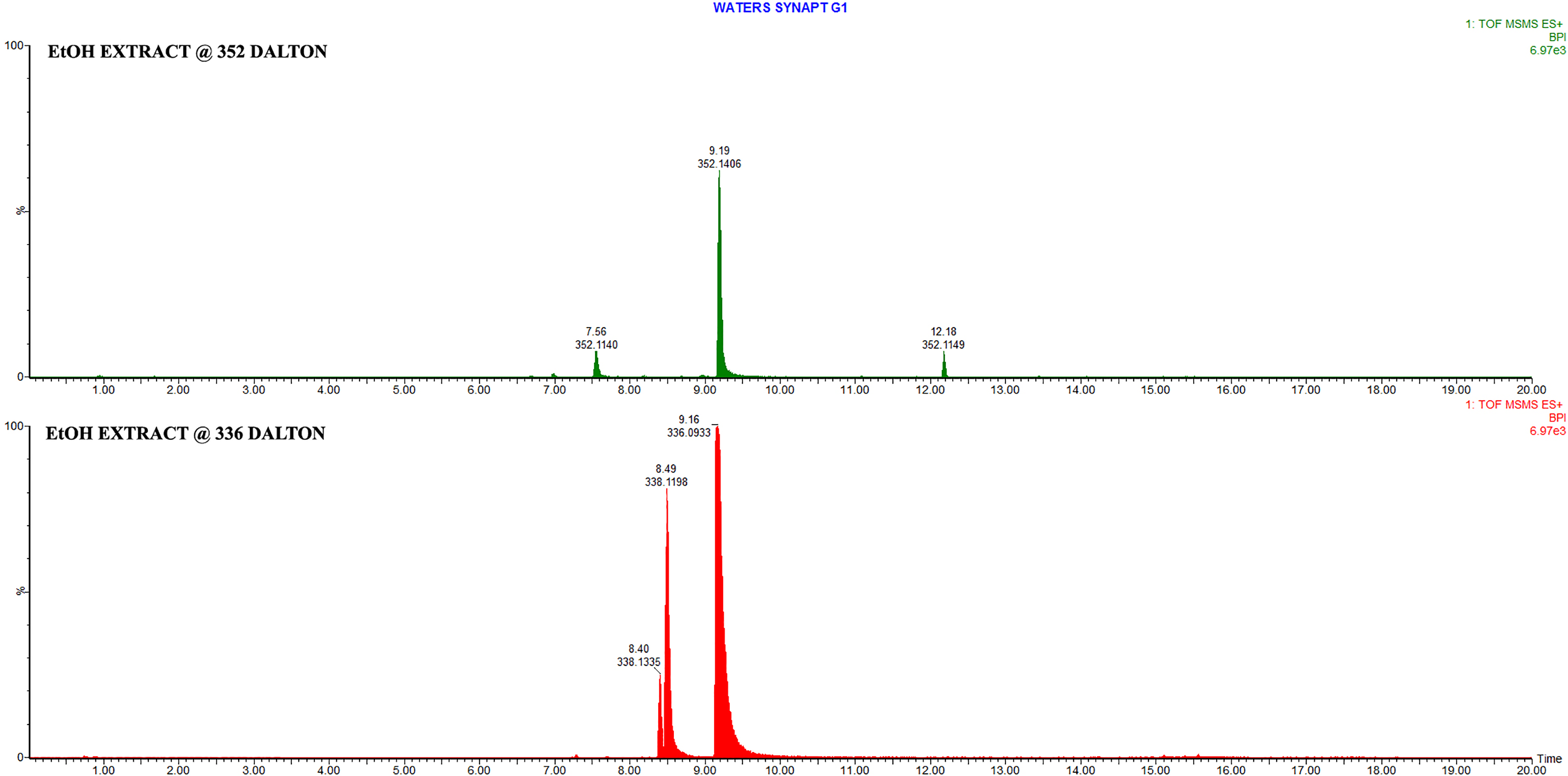
ESIPos MS/MS chromatograms of the Berberis vulgaris EtOH extract at 336 Da (Red) and 352 Da (green). Berberine Rt = 9.16 min and palmatine Rt = 9.19 min.
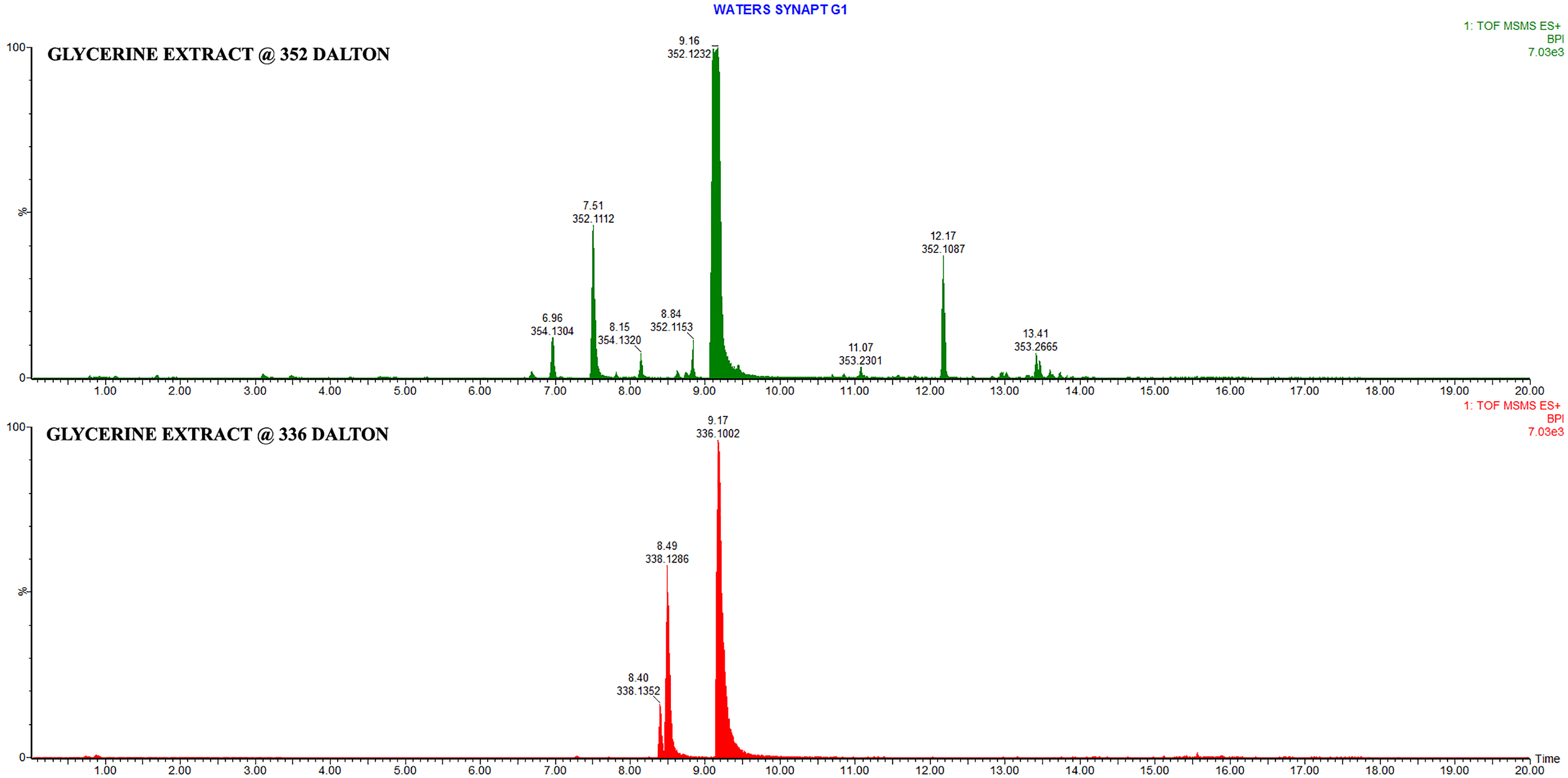
ESIPos MS/MS chromatograms of the Berberis vulgaris glycerine extract at 336 Da (Red) and 352 Da (green). Berberine Rt = 9.17 min and palmatine Rt = 9.19 min.
Compounds Detected in the Berberis vulgaris Extracts (Medical Alcohol and Glycerine).

The mass spectrum of each compound tentatively identified (Table 1) was submitted to a NIST 2014 Mass Spectral Library for possible identification. Only berberine and palmatine could be confirmed from the library search. MassFragment software from Waters was used to evaluate the fragmentation patterns of each compound with a known structure, MassFragment uses algorithms to detect product ion fragments. The method does not rely on the rule-based approach which can be limiting but is rather based on systematic bond disconnection of the precursor structure. By using MassFragment software and structures obtained from ChemSpider (http://www.chemspider.com/ accessed April 2023), 7 compounds could be positively identified. It was only bervulcine and vulracine that could not be evaluated with MassFragment as no confirmed structures could be obtained from ChemSpider or the Dictionary of Natural Products.
Known Chemical-Gene Interactions and the Potential Molecular Mechanism of Action for the Prevention and/ or Treatment of SARS-CoV-2
To rationalize how each compound could contribute to the prevention and treatment of SARS-CoV-2 infection, information was obtained from various databases to assess how the compounds might act on human and viral targets. On searching the CTB database no known interactions were identified for bervulcine, vervulcine, vulracine, berberrubine, lambertine, berbidine, or obamegine with any genes. Literature searches were also conducted to identify any potential gene-compound interactions cited in the literature. No relevant details were obtained in the literature for the compounds. Berbamine did yield results as it was subjected to in silico screening for its potential to bind to SARS-CoV-2 MPro, the proteolytic enzyme essential for the virus when producing proteins that assemble into new virions. It was found to have one of the highest binding energies of the compounds tested when compared to the known inhibitor. 16 It has also been shown experimentally to have an antiviral effect on cells infected with SARS-CoV-2 working within the concentration range of 0.1 to 10 μM. 15 The CTB database yielded 10 gene interactions, none of which showed direct links to the prevention and treatment of COVID-19. Genes identified included BCL2L1 and BID, both of which are linked to the BCL-2 family of proteins that are involved in inflammatory processes. Similarly, NFKB1A is a member of the NF-kappa B family of proteins which also play a role in inflammation. RELA and TNFAIP3 are linked to the NF-kappa B pathway. Activation of this pathway is associated with pro-inflammatory gene upregulation and inflammation. 17 Although many of the proteins with which berbamine interacts do seem to align with pathways that might have an impact, there was no direct published correlation found.
Berberine has been studied extensively and yielded the largest data set for all the compounds since it is generally known as an antioxidant and anti-inflammatory compound. It is thus not surprising it has been identified as a potential treatment for SARS-CoV-2.18–21 In these previous publications the numerous mechanisms by which berberine could be used to treat and prevent COVID-19, are described, and will not be repeated here. Maurya and colleagues 21 also conducted in silico screening of several compounds of which berberine was one, they found that the compound showed significant binding affinity to the spike glycoprotein as well as the ACE2 receptor, although it was not identified as the top-scoring compound out of the selection tested. There were interesting human genes identified that have been shown to interact with berberine, when searching the CTB database. In total, 247 genes were identified as having some sort of interaction with berberine. Some of these genes did have multiple records of the effect of the interaction on the gene or gene product. Of the 247 identified interactions the following items were determined to interrelate to SARS-CoV-2 infection and effects in a positive manner. Berberine resulted in increased expression of IL10 which is a cytokine that has potent anti-inflammatory properties. 22 This cytokine is said to prevent the host immune response to pathogens from reaching levels that damage normal tissue. 23 IL2 levels are inhibited and therefore reduced in critical COVID patients, 24 berberine increased the expression of IL2 mRNA. 22 Nicotinamide phosphoribosyl transferase (NAMPT) is an upstream regulator of interleukin 6. In a study conducted on patients whose serum tested positive for SARS-CoV-2 and displayed severe disease, NAMPT was upregulated. 25 Berberine decreases the expression of NAMPT. 26
The expression of KDM5B or JARID1B is increased with berberine. 27 The gene codes for a lysine demethylase which acts on the histones that structure DNA for storage, altering the patterns of structuring has effects on the accessibility of genes and hence their expression. It was found that microRNAs produced by SARS-CoV-2 target KDM5B transcripts to inhibit expression. 28 A different micro–RNA MIR19A is highly upregulated in COVID-19 infection 29 ; therefore, it stands to reason that the berberine effect of decreasing its expression 30 would work to potentially prevent or treat the COVID infection. The mir19-a micro–RNA is said to regulate inflammation and hence the targeting of this microRNA might be a reason SARS-CoV-2 produces an inflammatory cytokine storm, resulting in severe disease. 29
Berberine decreases the activity of metalloproteinases31,32 2 and 9, respectively. Drugs that currently target and inhibit both proteins have been shown to demonstrate significant antiviral activity toward SARS-CoV-2. 33
In a study looking at erythrocrine function during SARS-CoV-2 infection, it was found that the virus activates protein kinase C alpha affecting erythrocyte function. 34 Berberine decreases protein kinase C alpha activity and affects its localization. 35 Similarly, berberine decreases the activity and localization of protein kinase C epsilon 35 which has also been shown to be upregulated in SARS-CoV-2-infected individuals. 36 Protein kinase C's are involved in signaling pathways that are linked to inflammation.
Berberine increases the expression of SETD6 mRNA, 27 the protein product of this mRNA is a methyltransferase that modifies histones and alters the expression of specific genes related to the immune response. 37 In SARS-CoV-2 infection, SETD6 expression is downregulated. 38 Another methyltransferase SETDB1 has reduced levels of expression in children that develop more severe symptoms during infection with SARS-CoV-2 so it is presumed that increasing the levels would be protective. 39 Berberine is said to increase the expression of SETDB1. 27 Interestingly this gene was also identified to be upregulated in SARS-CoV-2 alpha variant infection but not in the beta and wild-type SARS-CoV-2 infections. 37 Another methyltransferase SETDB2 is also upregulated by berberine 27 and in SARS-CoV-2 infection, the levels of this protein have been shown to decrease. 40 The decrease in SETDB2 drives inflammation, therefore, increasing the levels can reverse this inflammatory response. 40
Transcription factors bind to DNA and alter gene expression. The expression of one transcription factor namely SREBF1 is decreased by berberine. 41 A study showed that the levels of SREBF1 are increased throughout SARS-CoV-2 infection. 42
Still looking at proteins that act on DNA, topoisomerases are enzymes that alter the topology of DNA (coiling) when the double helix is unwound. DNA unwinding must occur during replication and transcription of genes. The TOP1 protein activity is decreased by berberine. 43 TOP1 codes for DNA topoisomerase I. It has been found that inhibiting topoisomerase I suppresses the lethal inflammatory response associated with SARS-CoV-2 infection. 44
The well-known tumor suppressor protein p53 is also involved in the response to viral infections having an antiviral effect and has been proposed as a possible treatment option for SARS-CoV-2 infection. 45 Berberine increases the expression of the TP53 gene 46 which codes for p53.
Palmatine was the other compound found in high abundance in our extracts. Although not as extensively studied as berberine it yielded a total of 14 gene interactions on the CTB database. Two of the genes (ABCB1A and ABCB1B) are drug transport proteins found on cell membranes, these proteins act to remove drugs or foreign compounds from cells. In any treatment involving drugs, this is generally not desired because it decreases drug efficacy due to decreasing its concentration in cells, thus reducing the drug binding to targets within cells. Berberine seems to induce the active transport of drugs via these transporters, whereas palmatine seems to have the ability to bind the transport proteins and stimulate their ability to cleave ATP but not enhance active drug transport. 47 Since the mechanisms are relatively complex involving drug transport this difference that palmatine exhibits might have value in a combination therapy.
AHR gene expression is increased with palmatine. 48 This gene codes for the aryl hydrocarbon receptor. A detailed publication by George Anderson and colleagues 49 details AHR roles in SARS-CoV-2 entry and during disease progression. Although AHR has numerous roles the publication indicates that overall, it appears to have a negative impact on the outcomes especially for patients with underlying conditions. Palmatine increases the expression of the mRNA for this gene, but this does imply a link to the negative effects since from expression the protein must first be activated to exert effects via several signaling pathways. Interestingly other evidence indicates that AHR activation can reduce the number of ACE2 receptors. 50 Thus, it seems the role of AHR in SARS-CoV-2 infection is not entirely positive or negative.
Several chemokine coding genes (CCL2, CCL3, CCL4, CCL5, and CXCL8) are also negatively regulated by palmatine. 51 Chemokines generally signal the migration of leukocytes to inflammation sites by binding to G protein-coupled receptors found on these cells. 52 Interleukin 8 which is the product of the CXCL8 gene has been shown to have particularly high levels in patients that experience the cytokine storm associated with severe COVID outcomes. 53 CXCL8 is said to play a key role in the inflammation processes as well as the prothrombotic phenotype observed with SARS-CoV-2 infection and was proposed to be a good target for treatment. 54
An interleukin receptor that palmatine has been shown to affect is a modified form of interleukin 6 receptor (IL6R gene product). Palmatine reduces the expression of the IL6R gene. 51 Much like the chemokine's interleukins drive inflammation and hence the reduction in the receptors to which one of these interleukins bind could reduce the effect of the interleukin in inflammation.
VEGFA expression is decreased by palmatine. 51 The VEGFA levels are elevated in SARS-CoV-2 infection and particularly in severe COVID-19 that results in death. 55 VEGFA is a cytokine that similarly to the other chemokines is involved in driving inflammation.
Spike Protein ITC Studies With Berberine and Palmatine
The SARS-CoV-2 spike (S1) protein was purified to homogeneity ≥ 95%, as assessed by SDS-PAGE, with a molecular weight of 75 kDa and yields ranging between 5 mg and 10 mg per liter of bacterial culture (Figure 4). The ITC conditions used (20 µM SARS-CoV-2 S1 protein, 10 mM Berberis hemisulfate salt, and 10 mM Palmatine) for the binding of both Berberis hemisulfate salt and palmatine revealed no significant binding to SARS-CoV-2 S1 protein (data not shown).
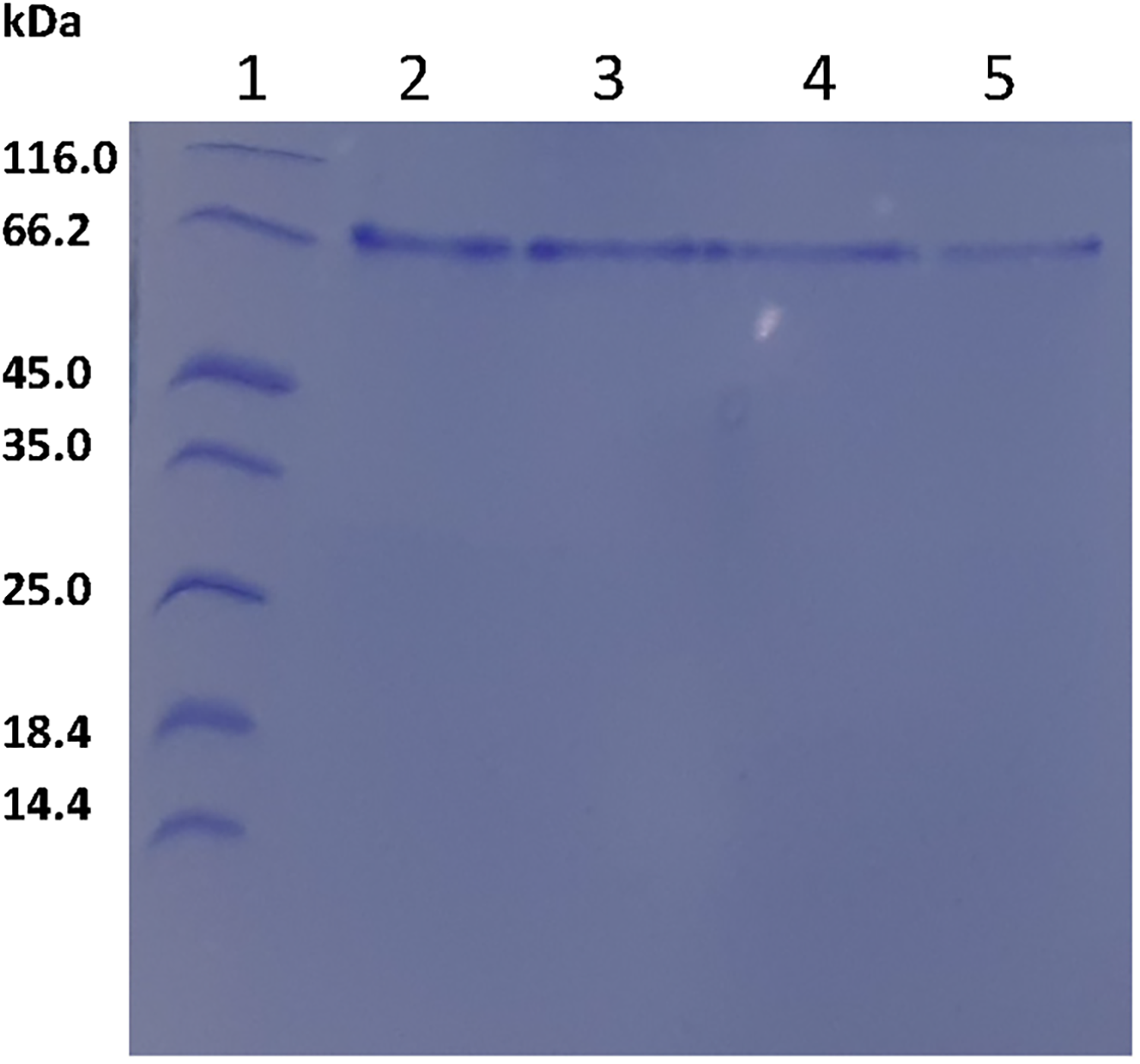

12% SDS-PAGE analysis of the purified recombinant SARS-CoV-2 Spike (S1) protein expressed in Escherichia coli T7 cells. Lane 1, protein molecular weight marker; lanes 2 to 5, purified SARS-CoV-2 S1 protein.
Discussion
Here for the first time, we report an analysis of the alkaloid compounds extracted from Berberis vulgaris roots using the HAB4a method of preparing a herbal extract. The method was carried out by a practitioner who routinely prepares the extracts for use in their practice. Compounds were identified to determine which main alkaloids were present and then use this information to further analyze the potential of the compounds in exerting a positive effect on the prevention and treatment of COVID-19. Berberrubine, bervulcine, berbamine, berberine, lambertine, and palmatine alkaloids were previously detected in root samples from Berberis vulgaris.13,56 Novel compounds vulracine, berbidine, and obamegine were also identified in the root extract (Table 1).
Berberine and palmatine were the main compounds detected in both the alcohol-based and glycerine-based extracts followed by lambertine. More berberine was present than palmatine in the alcohol-based extract, but the glycerine extract yielded significantly more palmatine than berberine. In general, it is assumed that extracts contain mainly berberine from the roots of Berberis vulgaris but data from this study shows that extraction parameters are key to ensuring this is always the case. Previously a study highlighted that different extraction parameters such as temperature and the use of ultrasound could alter the amount of berberine extracted from root samples. 4 Thus, it is not surprising that altering alcohol to glycerine would also produce variation in the amount of compounds present. The glycerine-based extract also failed to have bervulcine, vulracine, and berbidine present, so fewer compounds were extracted with glycerine than with the alcohol (Table 1). These data are valuable to practitioners conducting the HAB4a method because it demonstrates the composition of the extraction will differ based on the solvent used during extraction.
None of the compounds in the Berberis vulgaris extract have been shown experimentally to directly interact with SARS-CoV-2 proteins and hence affect the virus infecting and replicating in the body. In silico screening indicated berberine might bind the spike glycoprotein as well as the ACE-2 receptors. 21 The binding studies presented here, show that the 2 main components palmatine and berberine don’t directly interact with the spike protein of the virus. The human target to which the spike protein binds namely the ACE-2 receptor might, therefore, still be one of the proteins to which these compounds bind. The mechanism by which the main components berberine and palmatine prevent and treat COVID-19 appears to be by potentially acting on numerous human gene targets. The main targets are generally signaling molecules and then proteins that alter the packaging of DNA. These types of targets aim to change gene expression and hence the mix of proteins and pathways activated to prevent or overcome infection. It is also likely that the synergistic effect of the compounds in the extract is what yields positive outcomes and not just the action of one single compound on several targets. A previous study comparing berberine versus Berberis vulgaris root extract neuroprotective effect by evaluating SH-SY5Y cells demonstrated that both worked well to suppress apoptosis, but the extract functioned better than the pure berberine compound. 57 Coronil is also a combination herbal remedy designed as a supporting measure for COVID-19, it contains palmatine as one of several compounds. 58 There are numerous studies indicating the effects of the Berberis vulgaris extract on various organs and systems within the body ( 56 and references therein) these effects might all align to help prevent as well as treat the infection.
Conclusions
The HAB4a method of preparing a herbal extract yields compounds that have been detected previously from root extracts of Berberis vulgaris. The alcohol-based extraction yields more compounds and higher amounts of berberine than the glycerine-based extraction. Palmatine is present in higher amounts than berberine in glycerine-based extraction. Neither berberine nor palmatine bind the spike protein of SARS-CoV-2 but literature reveals that these compounds interact with several human gene targets in a manner that could act to prevent and treat SARS-CoV-2 infection.
Footnotes
Acknowledgments
This work is based on the research supported in part by the National Research Foundation of South Africa (Grant Number: 129409) and the University of South Africa.
Author Contributions
Samantha Gildenhuys Project leader which involved conceptualised with co-authors following initial discussions with Chad James. Collected and analyzed gene-compound interaction data and then compiled manuscript with co-author contributions Paul Anton Steenkamp conceptualized and conducted extract analysis to identify compounds and contributed to writing and editing the manuscript. Lucas Sekele conducted the purification of spike protein and ITC analysis. Salerwe Mosebi supervised spike protein purification and analysis and contributed to the writing and editing of the manuscript. Chad Clifton James was involved in project conceptualization, sourced and prepared the herbal extracts, and contributed to the writing and editing of the manuscript.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the NRF South Africa, (grant number 129409).
