Abstract
Introduction
Alcoholic liver disease (ALD), which is caused by prolonged heavy alcohol consumption, is one of the most prevalent types of chronic liver disease worldwide. 1 The stages of development generally include alcoholic fatty liver, alcoholic hepatitis, alcoholic liver fibrosis, and cirrhosis, which may eventually proceed to liver cancer. 2 Liver fibrosis is regarded as an important stage in the progression of ALD, with this process considered reversible. 3 The activation of hepatic stellate cells (HSC) plays a significant role in the development of alcoholic liver fibrosis. 4 Preventing HSC activation may reduce the deposition of extracellular matrix (ECM) and slow or even reverse liver fibrosis. 5
Ethanol and its metabolites are believed to play an instrumental role in the pathogenesis of ALD. 6 Enzymatically, ethanol is initially oxidized to acetaldehyde by ethanol dehydrogenase (ADH), with acetaldehyde being subsequently oxidized to acetate by acetaldehyde dehydrogenase (ALDH). Because acetaldehyde can induce the activation of HSCs, 7 drugs that inhibit this activation may be effective in the treatment of alcoholic liver fibrosis.
Several traditional Chinese medicines have been shown to be effective in the treatment of ALD. For example, Ginkgo biloba extract has been found to alleviate liver fibrosis and HSC activation by modulating nuclear factor kappa-B (NF-κB)/IκBα signaling, 8 and quercetin has been shown to inhibit the progression of liver fibrosis by modulating NF-кB/IкBα signaling. 9 These findings indicate that compounds derived from herbal medicines may be used to treat alcoholic liver fibrosis.
The pentacyclic triterpenoid asiatic acid (AA), which is extracted from the Chinese herb Centella asiatica, has been effective in the treatment of several diseases. 10 For example, AA was found to reduce alcohol-induced liver damage and CCl4-induced liver fibrosis in rats by regulating the NF-κB signal pathway11,12 and to attenuate liver fibrosis by blocking the TGF-β/Smad signaling pathway. 13 Few studies, however, have assessed the mechanism of action by which AA suppresses acetaldehyde-induced HSC activation. The present study, therefore, analyzed the theoretical mechanism of action of AA in acetaldehyde-induced HSC activation by evaluating the molecules targeted by AA in these cells through network pharmacology, molecular docking, and molecular dynamics simulations. The mechanism of action of AA in reducing alcoholic liver fibrosis was subsequently verified by in vitro experiments in HSCs.
Materials and Methods
Targets of AA and Alcoholic Liver Fibrosis
The Swiss target prediction database, 14 Comparative Toxicogenomics Database (CTD) and HERB database Genecards were screened to identify molecules targeted by AA in alcoholic liver fibrosis.15‐17 The UniProt protein database (https://www.uniprot.org/) was searched using the UniProtKB search function to standardize related target genes. Venn diagrams were subsequently drawn to identify potential targets for AA in the treatment of alcoholic liver fibrosis. 18
Construction of a Drug-Target-Disease Network
Core targets were imported into the STRING database to obtain a protein–protein interaction (PPI) network diagram. 19 The required minimum interaction score was set at 0.7. TSV files were downloaded from the STRING database and imported into Cytoscape 3.9.1 (https://cytoscape.org/),which was utilized for core target screening.
Gene Ontology and Kyoto Encyclopedia of Genes and Genomes Enrichment Analysis
The Metascape database was screened to identify cross-targets of AA associated with liver fibrosis, with analyses including gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analyses. 20 Potential mechanisms of action of AA were categorized by biological processes, cellular components (CCs), molecular functions (MFs), and key signaling pathways, with the results presented as bubble plots.
Molecular Docking
The crystal structure of the protein used for docking was obtained from the PDB database, 21 and the three-dimensional structure of the small molecule AA was constructed using the PubChem database. 22 Molecular docking was performed using AutoDock Vina 1.1.2 software. 23 Before starting molecular docking, the receptor protein was treated with PyMol 2.5.2 to remove water molecules, salt ions, and small molecules. A docking box measuring 22.5 Å per side, centered around the location of the active site, was constructed using the PyMol plugin center_of_mass.py. All processed small molecules and receptor proteins were converted using ADFRsuite 1.0 into the PDBQT format necessary for AutoDock Vina 1.1.2 docking. 24 The output docked conformations with higher scores were considered binding conformations for subsequent analysis.
Molecular Dynamics Simulation
The complex consisting of AA and the protein RELA was subjected to all-atomic molecular dynamics simulation using AMBER 18 software. 25 The charge on the small molecules was calculated prior to simulation using Gaussian 09's Antechamber module and Hartree-Fock (HF) SCF/6-31G*.26,27 Subsequently, small molecules and proteins were described using the GAFF2 small molecule and the ff14SB protein force fields, respectively.28,29 Using the LEaP module, hydrogen atoms were added to the system, a truncated octahedral TIP3P solvent box was added at a distance of 10 Å from the system, 30 and the system charge was neutralized by adding Na+ or Cl− to the system. Finally, the topology and parameter files used for the simulation were outputted.
Molecular dynamics simulation was performed using AMBER 18 software. 25 Prior to the simulation, the system was subjected to energy optimization, including a steepest descent method with 2500 steps and a conjugate gradient method with 2500 steps. After energy optimization was completed, the system temperature was slowly ramped up from 0 to 298.15 K within 200 ps at a fixed volume and a constant ramp rate. A 500 ps NVT (isothermal-isochoric) system simulation was performed at the system maintenance temperature of 298.15 K to enhance the distribution of solvent molecules in the solvent box. The 2 composite systems were simulated with 100 ns NPT (isothermal isobaric) tethering under periodic boundary conditions. The truncation distance of the non-bond was set at 10 Å, and long-range electrostatic interactions were calculated using the particle mesh Ewald (PME) method. 31 The hydrogen atom bond lengths were constrained by the SHAKE method and the Langevin algorithm was used for temperature control.32,33 The collision frequency γ was set at 2 ps−1, the system pressure was set at 1 atm, the integration steps were set at 2 fs, and the trajectories were saved at 10 ps intervals for subsequent analyses.
MM/GBSA Binding Free Energy Calculation
The binding free energy between protein and ligand was calculated by the MM/GBSA method.34‐36 An MD trajectory of 90 to 100 ns was used for calculations using the equation:
with
Cell Culture and Treatment
Rat HSC-T6 cells purchased from the Chinese Academy of Sciences were cultured in Dulbecco's modified eagle medium (Gibco) containing 10% fetal bovine serum (FBS; Siji Green), 100 U/mL penicillin, and 100 μg/mL streptomycin in an incubator maintained at 37 °C with 5% CO2. The alcohol-associated hepatic fibrosis cell model has been described, 39 and the cells were treated with 200 μM acetaldehyde (Da Mao) for 48 h. Based on the results of CCK-8 experiments, cells were treated with medium alone (control group), 200 μM acetaldehyde, 200 μM acetaldehyde plus low dose (20 μΜ) AA, and 200 μM acetaldehyde plus high dose (30 μΜ) AA for 48 h.
Determination of Safe Dosage Range of AA
Rat HSC-T6 cells were precultured at 37 °C in 5% CO2 for 24 h, and then 200 μM acetaldehyde and 0, 10, 20, 30, 40, 50, and 60 μM AA were added to each culture well, respectively. After 48 h, the cells were washed and resuspended in fresh medium, and 10 μL CCK-8 solution was added to each well (with care taken not to form bubbles in the hole, which would affect the reading of the OD value). If the substance to be tested is oxidizing or reductive, the fresh medium can be replaced before adding CCK-8 (remove the medium, wash the cells with the medium twice, and then add a new medium) to remove the drug effect. The cells were subsequently incubated for 1 to 4 h, and the absorbance of each well at 450 nm was determined using an enzyme labeling instrument.
Western Blotting
HSC-T6 cells were lysed in RIPA buffer containing 1% PMSF, and total proteins were extracted and quantified using BCA protein quantitative kits (Boster). Proteins were subjected to sodium dodecyl-sulfate polyacrylamide gel electrophoresis and transferred to PVDF membranes, which were incubated in 5% BSA for 1.5 h at room temperature. The membranes were subsequently incubated overnight at 4 °C with primary antibody, washed, and incubated with secondary antibody for 1 h at room temperature. The protein bands were developed with enhanced chemiluminescence solution (GlpBio), photographed, and processed with ImageJ software.
Quantitative Real-Time PCR
Total RNA was extracted from HSC-T6 cells using Trizol reagent (USA), and cDNA was synthesized from total RNA using AMV reverse transcriptase according to the manufacturer's protocol. α-SMA, COL1a1, actin, and p-NF-kappa B-p65 mRNAs were quantified using real-time PCR kits (Thermo Scientific) and primer sequences for α-SMA (forward, 5'-CGAAGCGCAGAGCAAGAGA-3'; reverse, 5'-CATGTCGTCCCAGTTGGTGAT-3'), COL1a1 (forward, 5'-GATCCTGCCGATGTCGCTAT-3'; reverse, 5'-TGTAGGCTACGCTGTTCTTGCA-3'), β-actin (forward, 5'-ACCACAGCTGAGAGGGAAATCG-3'; reverse, 5'-AGAGGTCTTTACGGATGTCAACG-3'), caspase-3 (forward, 5'-AACGGACCTGTGGACCTGAA-3'; reverse, 5'-TCAATACCGCAGTCCAGCTCT-3'), and BCL2 associated X (BAX) (forward, 5'-GATGGCTGGGGAGACACCTG-3'; reverse, 5'-GCGGCCCCAGTTGAAGTTG-3'). The amplification protocol consisted of an initial denaturation at 95 °C for 10 min and 40 cycles of denaturation at 95 °C for 15 s, annealing for 30 s at 60 °C and elongation for 30 s at 72 °C. The levels of all mRNAs were normalized relative to the level of β-actin mRNA using the 96−△Ct method. Three independent cDNA samples were assessed in each experiment, with all PCR experiments performed in triplicate.
Immunofluorescence Staining
HSC-T6 cells were cultured with either acetaldehyde, acetaldehyde plus AA, or medium alone, and were treated with 4% paraformaldehyde for 30 min before treatment with 3% BSA for 30 min. The cells were subsequently incubated with anti-p-NF-κB-p65 (1:200) primary antibody overnight at 4 °C, washed, and incubated with fluorescent secondary antibody for 1 h in the dark at room temperature. The nuclei were stained with DAPI for 10 min, and image analysis was performed using laser scanning confocal microscopy.
Statistical Analyses
Data were expressed as mean ± standard deviation (SD) and compared by either one-way ANOVA or t-tests. All statistical analyses were performed using Prism 8.0 software, with P < 0.05 considered statistically significant.
Results
Potential Targets of AA and Alcoholic Liver Fibrosis
Screening of the Swiss target prediction database, the CTD, and the HERB database identified 92 genes potentially interacting with AA. Analysis of the GeneCards database identified 710 targets related to alcoholic liver fibrosis. A Venn diagram of the 92 potential drug targets and the 710 potential disease targets identified 36 intersecting targets (Figure 1A).
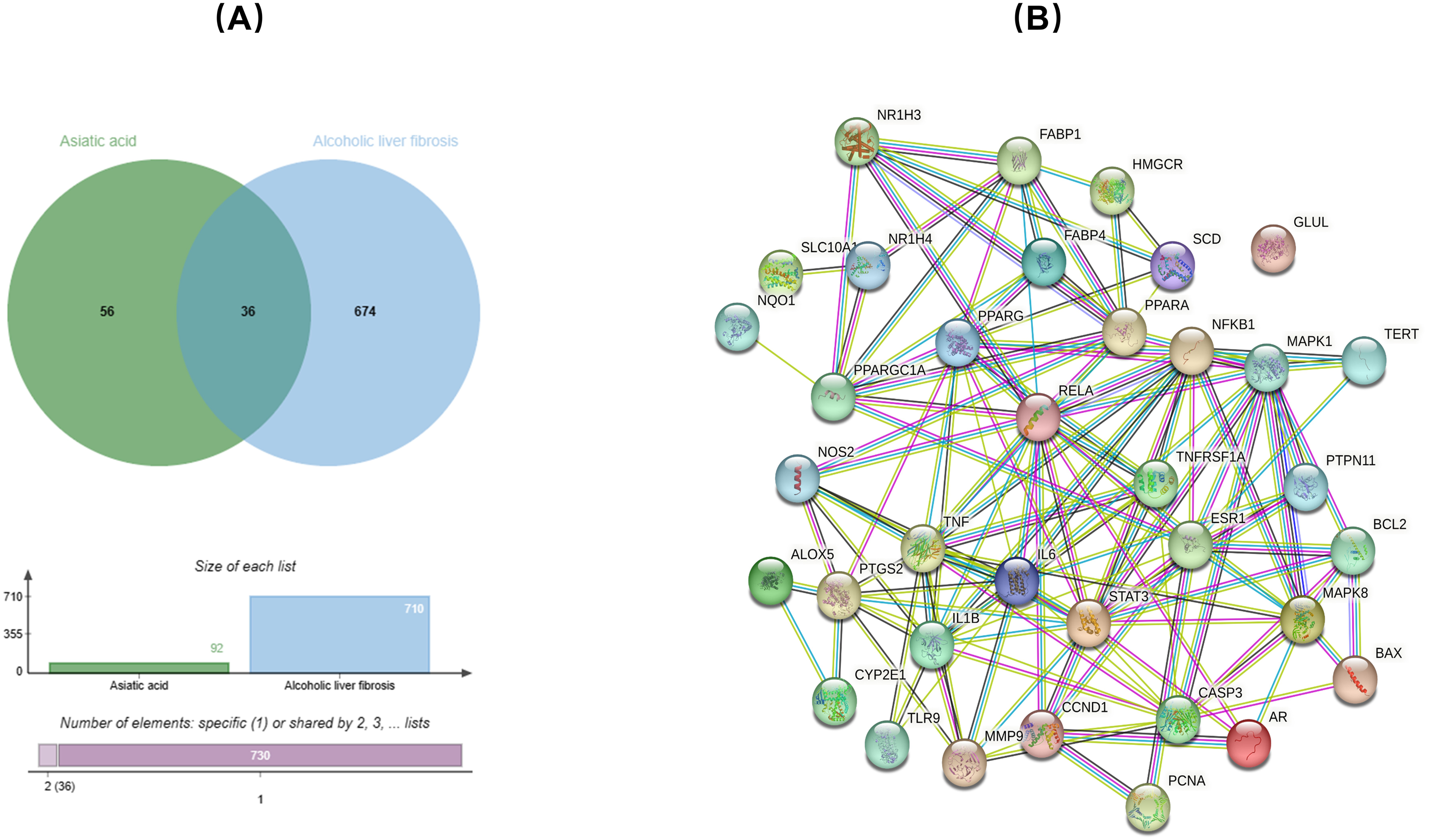
(A) Venn diagram of the intersection between asiatic acid (AA) and alcoholic liver fibrosis targets; (B) Protein interaction network of the intersecting targets.
PPI Network Diagram Results
A PPI network diagram was formed by importing the 36 intersecting targets into the STRING database. The minimum interacting fraction score was set at 0.7 (Figure 1B). TSV format files were downloaded from the STRING database and imported into Cytoscape 3.9.1 to determine core target degrees. These findings indicated that the top-ranked targets included signal transducer and activator of transcription 3 (STAT3), RELA, tumor necrosis factor (TNF), interleukin 6 (IL-6), and interleukin 1 beta (IL-1β) (Figure 2A-B).
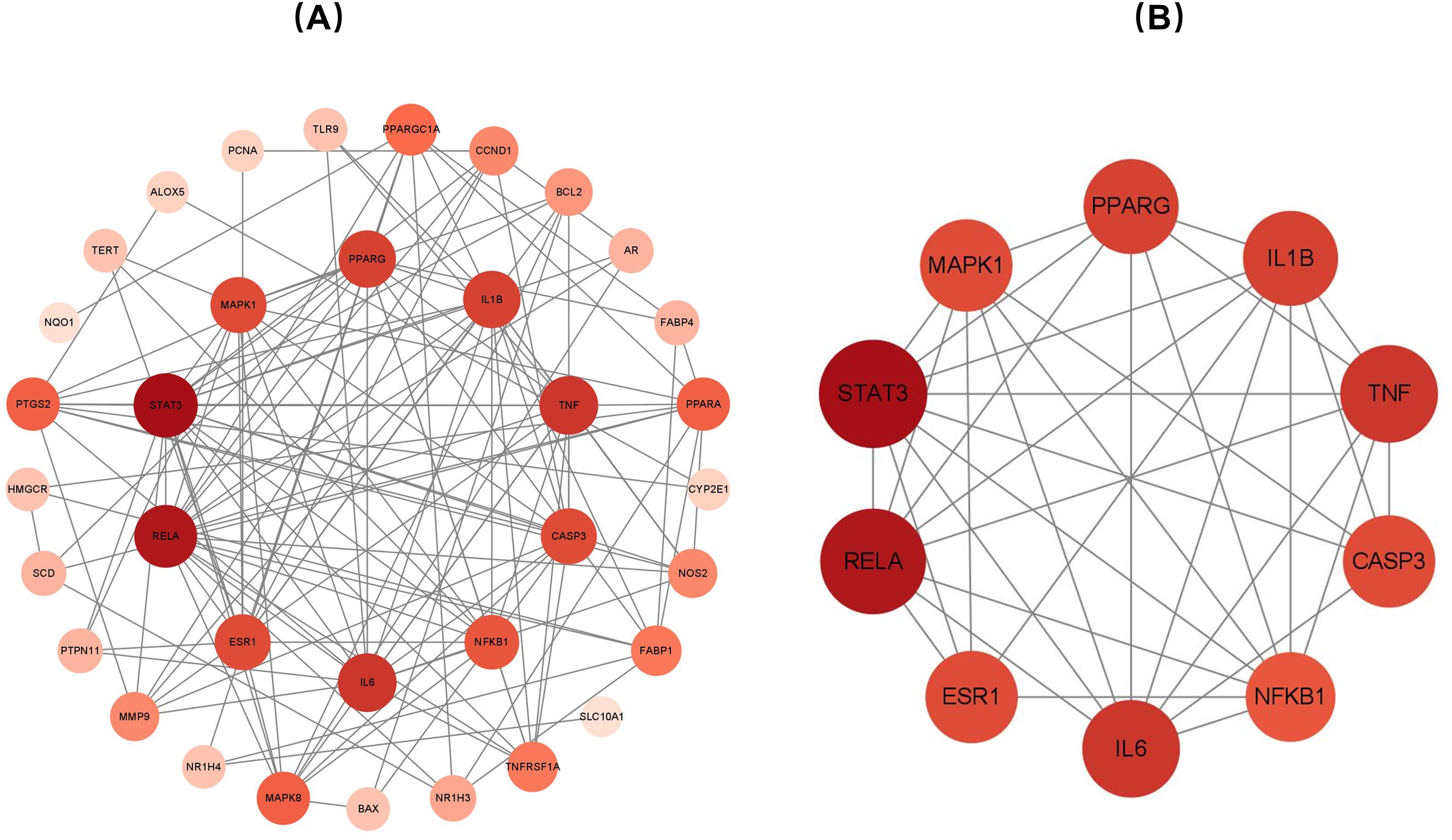
(A) Map of the intersection of target protein interactions. (B) Map of the top 10 target proteins interacting with asiatic acid (AA).
Results of GO and KEGG Enrichment Analyses
The relatedness of the 36 target genes to functional units, including biological processes, CCs and MFs, was assessed by GO and KEGG pathway enrichment analyses, with the filter set to an adjusted P-value < .05. GO enrichment analysis showed that these target genes played important roles in alcoholic liver fibrosis (Figure 3A), whereas KEGG pathway enrichment analysis of the top 10 pathways showed that ALD was the leading pathway (Figure 3B; bubble chart available at https://www.bioinformatics.com.cn). The NF-κB signaling pathway has been shown to be highly significant in ALD. Because RELA belongs to the NF-κB-related signaling pathway, molecular docking and molecular dynamics simulations of AA and RELA protein were performed to determine whether AA attenuated acetaldehyde-induced HSC activation through the RELA signaling pathway.

Results of (A) KEGG enrichment analysis and (B) GO analysis of target proteins intersecting with AA. Abbreviations: GO, gene ontology; KEGG, Kyoto Encyclopedia of Genes and Genomes; AA, asiatic acid.
Molecular Docking
The docking simulation technique is a convenient and effective means to explore the interactions of small molecules with protein targets. The present study used Vina 1.1.2 software to assess the docking of AA with the proteins IL1B, IL6, RELA, STAT3, and TNF-α. Evaluation of binding energy scores showed that AA bound strongly to IL-1β, IL-6, RELA, STAT3, and TNF-α proteins, with binding energies of −8.2, −7.4, −7.5, −6.4, and −3.3 kcal/mol, respectively. The binding of AA to RELA was further investigated by molecular dynamics simulations to assess the mechanism of action by which AA binds to RELA at the molecular level (Table 1, Figure 4A, B, C, D, E).

(A) Complexes formed by the small molecule AA and the proteins (A) IL-1β, (B) IL-6, (C) RELA, (D) STAT3, and (E) TNF-α based on docking. (left to right) global view, local 3D view, 2D interaction diagram. Abbreviations: 3D, three-dimensional; 2D, two-dimensional; STAT3, signal transducer and activator of transcription 3; TNF-α, tumor necrosis factor α; IL-6, interleukin 6; IL-1β, interleukin 1 beta; AA, asiatic acid.
Binding Energy of AA to IL-1β, IL-6, RELA, STAT3, and TNF-α Protein.

Abbreviations: STAT3, signal transducer and activator of transcription 3; TNF-α, tumor necrosis factor α; IL-6, interleukin 6; IL-1β, interleukin 1 beta.
Molecular Dynamics Simulation
The root-mean-square deviation (RMSD) of molecular dynamics simulations reflects the motion of the complex, with larger RMSDs and more intense fluctuations indicating more intense motion. Analysis showed that RELA protein fluctuated continuously during the simulations, suggesting that this protein was not sufficiently stable (Figure 5A). In contrast, the RMSD of AA showed that the latter fluctuated stably and narrowly (ie within 0.5 Å) during the early stage of the simulation. These findings indicated that AA was very closely and stably bound to RELA protein.

(A) Change in the RMSD of the complex over time during molecular dynamics simulation; (B) RMSF calculated based on molecular dynamics simulation trajectory. (C) Changes in the number of hydrogen bonds between small molecules and proteins during the process of molecular dynamics simulation. Abbreviations: RMSF, root mean square fluctuation; RMSD, root-mean-square deviation.
Root mean square fluctuation (RMSF) is an indicator of response to protein flexibility during molecular dynamics simulation. Usually, protein flexibility decreases after the drug binding, with the drug stabilizing the protein and affecting its enzymatic activity. Except for the local structure of the protein, the RMSF of RELA protein after binding small molecules was within 4 Å, whereas the RMSF of the empty protein (black line) was greater, indicating intrinsic structural instability during protein movement (Figure 5B). Taken together, these findings showed RELA protein could be stabilized by binding AA (Table 2).
Binding Free Energies and Energy Components Predicted by MM/GBSA (Kcal/mol).
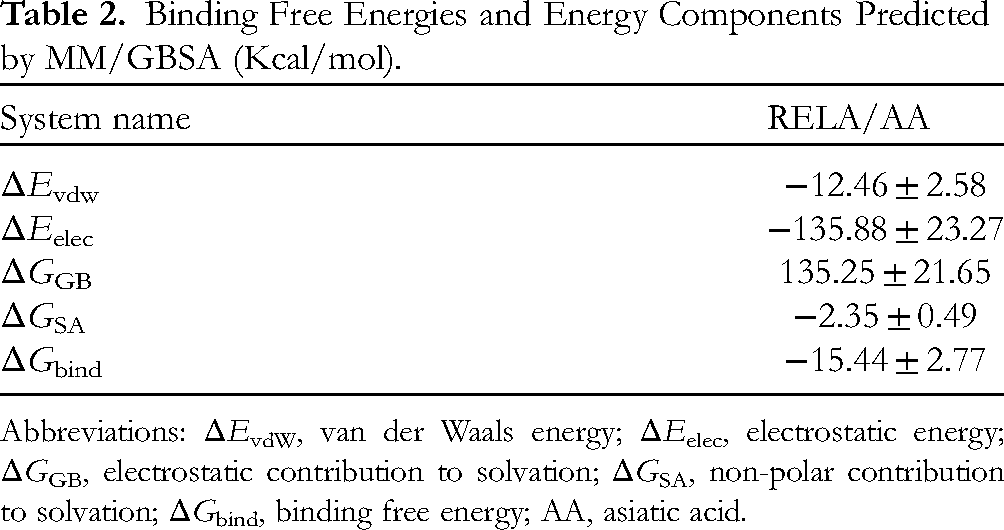
Abbreviations: ΔEvdW, van der Waals energy; ΔEelec, electrostatic energy; ΔGGB, electrostatic contribution to solvation; ΔGSA, non-polar contribution to solvation; ΔGbind, binding free energy; AA, asiatic acid.
Based on the trajectory of molecular dynamics simulations, the binding energy was calculated using the MM-GBSA method, which could more accurately reflect the binding effect of small molecules and target proteins. The binding energy of RELA to AA was found to be −15.44 ± 2.77 kcal/mol (Table 2), indicative of strong binding affinity. Energy decomposition showed that binding was primarily due to electrostatic energy, followed by van der Waals energy and nonpolar solvation-free energy (Figure 5C).
AA Attenuated Acetaldehyde-Induced HSC-T6 Activation
CCK-8 assays showed that an AA concentration of 40 µM was toxic to HSC-T6 cells, but AA at concentrations of 0 to 30 μM was determined to be nontoxic to the cells. AA was tested at concentrations of 0 to 30 µM (Figure 6A). Western blotting showed that AA could upregulate the expression of the proapoptotic proteins BAX and caspase 3 (Figure 6B). Western blotting and real-time quantitative polymerase chain reaction (RT-qPCR) showed that treatment of HSC-T6 cells with acetaldehyde significantly upregulated the expression of α-SMA and type I collagen protein and mRNA, but that these increases were attenuated by treatment with AA (Figure 6C, D). These results indicated that AA could inhibit the expression of α-SMA and type I collagen in HSC-T6 cells, thereby attenuating their acetaldehyde-induced activation.
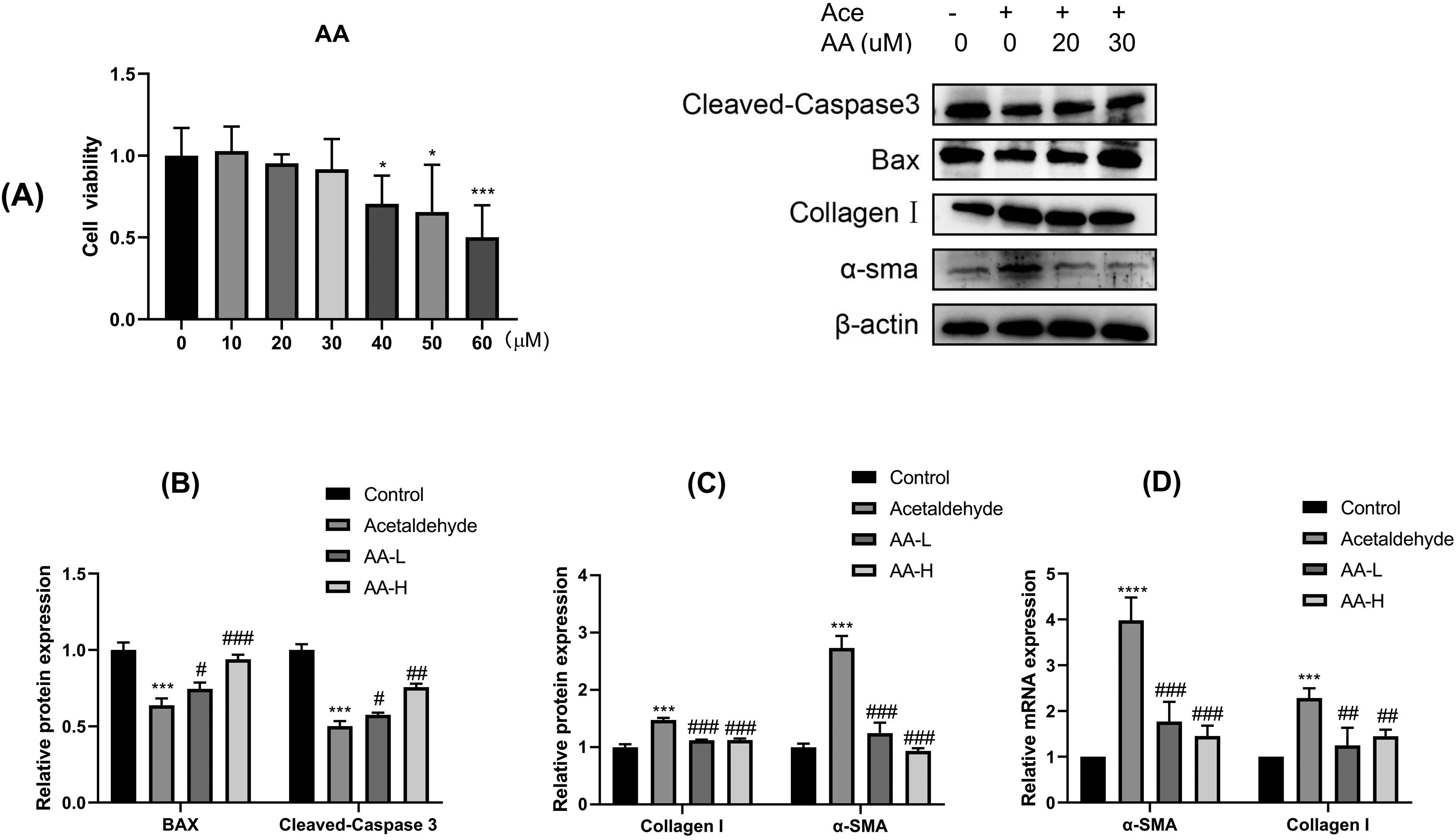
(A) CCK-8 kit assays showing safe doses of AA. (B) Western blotting assays showing the expression of the apoptotic proteins BAX and cleaved caspase 3; (C) Western blotting showing the expression of collagen type 1 and α-SMA; (D) RT-qPCR showing the expression of collagen type 1 and α-SMA mRNAs. Abbreviations: RT-qPCR, real-time quantitative polymerase chain reaction; AA, asiatic acid; BAX, BCL2 associated X.
AA May Reduce Alcoholic Liver Fibrosis by Modulating the NF-κB-P65 Signaling Pathway
The cumulative results of KEGG pathway enrichment analysis, molecular docking, and molecular dynamics showed that the NF-κB signaling pathway was associated with alcoholic liver fibrosis, suggested that AA may alleviate liver fibrosis by regulating the p-NF-κB-P65 signaling pathway. Immunofluorescence and Western blotting experiments showed that AA treatment of acetaldehyde-activated HSC-T6 cells significantly downregulated the expression of p-NF-κ B-p65 and IKB-α compared with acetaldehyde-activated cells (Figure 7A, B, C). These findings suggest that AA may attenuate the fibrotic expression of HSC-T6 cells by inhibiting the levels of phosphorylation of NF-κB-P65 and IKB-α.
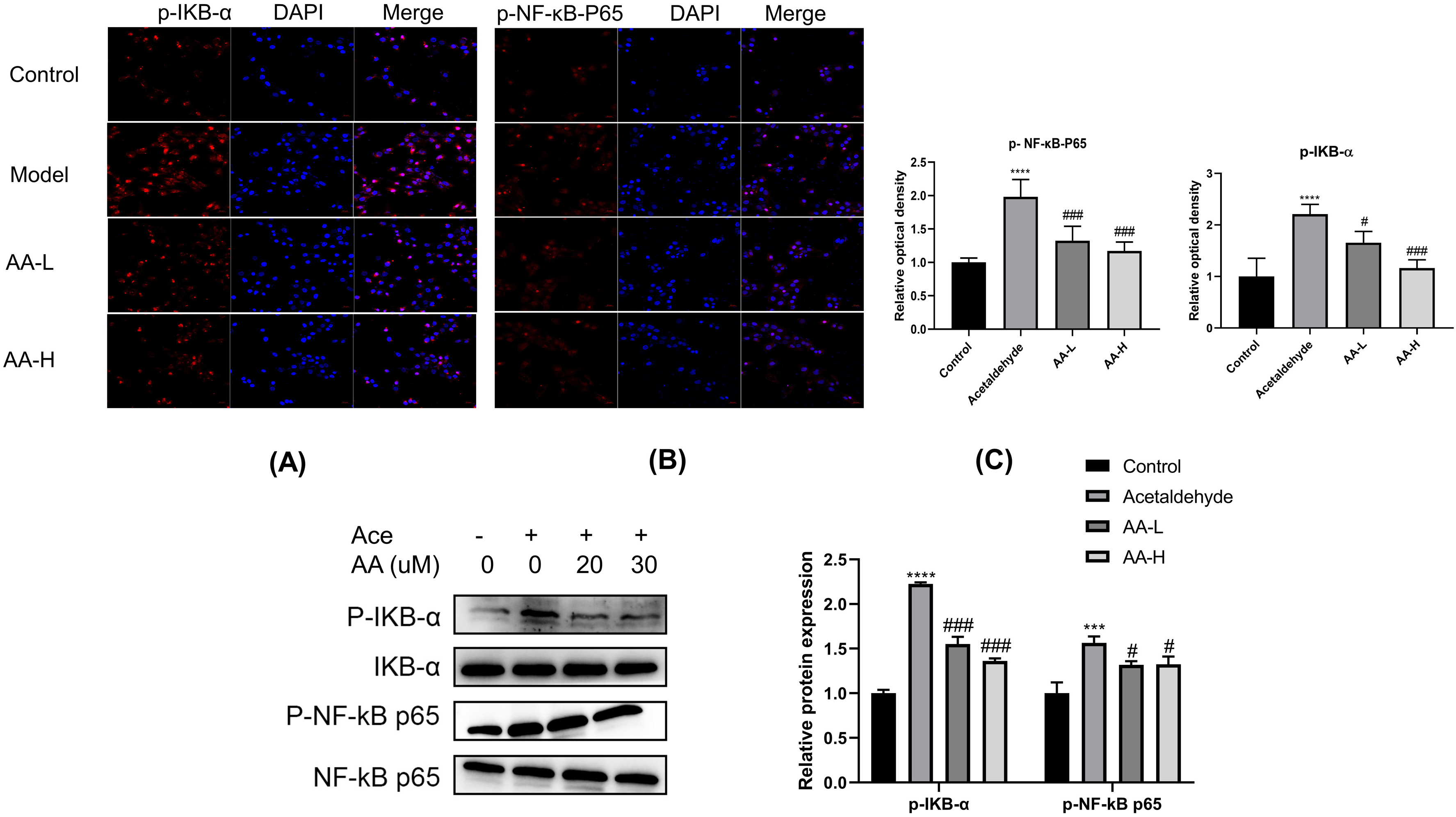
(A,B) Immunofluorescence experiments showing the expression of (A) p-IKB-α and (B) p-NF-κB-P65 in HSC-T6 cells. (C) Expression of total and phosphorylated IKB- α and NF-κB-P65 proteins in HSC-T6 cells. Abbreviations: HSC-T6, hepatic stellate cell-T6; NF-κB, nuclear factor kappa-B.
Discussion
ALD is a liver disease caused by acute or chronic alcohol intoxication. The liver is the principal organ in the body that metabolizes alcohol, with these metabolites found to induce liver damage. 40 Liver metabolites are considered one of the main causes of chronic ALD. 41 Although abstaining from alcohol is the most direct method of preventing or reversing the progression of ALD, alcoholics find it difficult to abstain from alcohol or curb their dependence on it. 42
Liver fibrosis is regarded as central to the progression of ALD, with this process considered reversible. No specific drugs are currently available to treat hepatic fibrosis, with western pharmaceutical agents being relatively ineffective and causing several adverse effects. 43 The most suitable treatment option for patients with advanced alcoholic cirrhosis is currently liver transplantation.
Natural products derived from herbal agents, including alkaloids, flavonoids, quinones, terpenoids, saponins, phenylpropanoids, and polysaccharides and possessing anti-fibrotic activity, may be effective in the treatment of hepatic fibrosis. 44 Several herbs and their formulations are also considered beneficial in the treatment of liver fibrosis, suggesting a new treatment strategy.
AA is a triterpenoid derived from Centella asiatica with antioxidant, anti-inflammatory and antifibrotic properties.45‐47 AA inhibits cardiac fibrosis in rats through modulation of the Nrf2/HO-1 and TGF-β1/Smads signaling pathways, 48 and ameliorates bleomycin-induced pulmonary fibrosis by suppressing fibrogenic and inflammatory signaling pathways. 49 AA has been found to attenuate CCl4-induced hepatic fibrosis by modulating the Nrf2/ARE, NF-κB/IκBα and JAK1/ STAT3 signaling pathways. In the present study, AA and alcoholic liver fibrosis targets were screened through a network of pharmacology-related databases, finding that the proteins STAT3, RELA, TNF, IL-6, and IL-1β were key targets. KEGG pathway enrichment analysis identified the closest signaling pathway as being the ALD signaling pathway. 50 In its inactive state, NF-κB protein consists of homodimers/heterodimers composed of p65 and p50 subunits in the cytoplasm that bind to the inhibitory protein IkB to form trimer complexes. The binding of TNF-α to its receptor on the cell membrane alters the conformation of the receptor, which transmits a signal to IKK kinase (IkB kinase), which phosphorylates IkB protein, resulting in its dissociation from the trimer.
Subsequently, NF-κB dimers in the cytoplasm enter the nucleus and bind to specific DNA sequences, promoting the transcription of related genes. The NF-κB signaling pathway is closely associated with the development and reversal of liver fibrosis. 51 The present study found that NF-κB was significantly involved in the pro-fibrotic effect on HSCs, with NF-κB activation triggering the downregulation of the TGF-β pseudoreceptor BAMBI, enhancing TGF-β signaling and cell activation. 52 TGF-β is regarded as the main cytokine responsible for HSCs activation and proliferation, inducing HSCs to produce type I collagen and other components of the ECM, thus promoting the development of liver fibrosis. 53 In addition, NF-κB inducing kinase (NIK) activation in bone marrow-derived macrophages, increase liver injury, and promote HSC activation and fibrosis, 54 Moreover, treatment with either NF-κB or proteasome inhibitors has been shown to reduce liver fibrosis in vivo in a mouse model of bile duct ligation. 55
In liver fibrosis, NF-κB can be activated by LPS, TNF, and IL-1β, enhancing TGF-β signaling and activating HSCs, 56 a major step in the pathogenesis of liver fibrosis. Inhibition of HSC activation and proliferation may therefore reverse liver fibrosis. 57 The present study found that AA attenuated acetaldehyde-induced activation and proliferation of HSC-T6 cells, and attenuated the expression of α-SMA and type 1 collagen. α-SMA has been identified as a marker of HSC activation and type 1 collagen as the major component of the ECM. Liver damage activates HSCs, accompanied by the elevated expression of α-SMA and the secretion of type 1 collagen, inducing collagen deposition and the development of liver fibrosis. 58 Network pharmacology-related studies have shown that AA can attenuate liver fibrosis, likely by regulating the protein NF-κB-P65 (RELA). In addition, molecular docking and molecular dynamics simulations showed that AA and NF-κB-P65 (RELA) bind to each other with high affinity. Experiments in HSC-T6 cells showed that AA could down-regulate the phosphorylation of NF-κB-P65 (RELA). Taken together, these findings suggest that AA can inhibit HSC activation and attenuate hepatic fibrosis by down-regulating NF-κB-P65 (RELA) phosphorylation.
Conclusions
The present study showed that AA was effective in inhibiting HSC-T6 activation and proliferation and that NF-κB was an important target of AA, playing a key role in its anti-fibrosis activity. These findings may provide new clues for the treatment of alcoholic liver fibrosis.
Supplemental Material
sj-docx-1-npx-10.1177_1934578X231194127 - Supplemental material for Network Pharmacology, Molecular Docking, Molecular Dynamics Simulation, and in vitro Experiments to Explore the Mechanism of Asiatic Acid Inhibiting Acetaldehyde-Induced Activation of Hepatic Stellate Cells
Supplemental material, sj-docx-1-npx-10.1177_1934578X231194127 for Network Pharmacology, Molecular Docking, Molecular Dynamics Simulation, and in vitro Experiments to Explore the Mechanism of Asiatic Acid Inhibiting Acetaldehyde-Induced Activation of Hepatic Stellate Cells by Tao Jiang, Xiaojin Chen, Wanzhi Yang, Ning Li, Jinhong Xu, Yang Lu, Sisi Zhang, Shuihong Yu and Yongxia Li in Natural Product Communications
Footnotes
Authors’ Contributions
YL, TJ, and XC designed the study. Material preparation, data collection and analysis were performed by NL, SY, WY, JX, YL, and SZ. The manuscript was written by TJ and XC.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by the Science Foundation of Anhui Province, China (grant No 2022AH052549), Anqing Medical and Health Self-Financing Science and Technology Plan Project (grant No 2022Z4004).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.

