Abstract
The potential targeting of xanthine oxidase (XO) for the treatment of gout has been explored through animal and human experiments. To identify potential XO inhibitors, a virtual screening method was utilized to examine fatty acids from Hedyotis pilulifera. Molecular docking results demonstrated that all possible configurations of the fatty acid 9,12,13-trihydroxyoctadeca-10E,15Z-dienoic acid exhibited a strong binding affinity to XO. In vitro experiment also showed that this compound inhibited XO activity compared to the control compound. Molecular dynamic simulation was used to further investigate the interaction between the ligand and protein. Additionally, ADMET studies revealed that this compound has highly favorable drug-like properties and pharmacokinetics.
Introduction
Computational biological development has opened up many new avenues in research to find active ingredients with potential biological effects to contribute to developing medicines for the health sciences. Application techniques such as in silico methods are increasingly proving outstanding in research, exploring the biological effects of potential active ingredients on the human body. 1 The application of in silico procedures supports identifying drug targets, evaluating compounds and protein interactions, pharmacokinetic analysis (ADME), and improving the drug properties of molecules.2–4 These techniques will be a great precursor to new drug development works.
Gout is a common disease characterized by episodes of acute arthritis with symptoms such as swelling, heat, redness, and severe and sudden pain in one or more joints, most commonly in the joint of the big toe. 5 This is a metabolic pathology caused by the deposition of monosodium urate crystals in the joints and tissues around the joints. Elevated blood uric acid levels form these crystals. 6 Many causes, such as increased uric acid production or diminished urinary uric acid elimination, can result in hyperuricemia. 7 Previous studies proved that xanthine oxidase (XO), in particular, is associated with increased uric acid production. XO is an enzyme that catalyzes the hydroxylation of purines, converts hypoxanthine into xanthin, and thereby synthesizes uric acid, which is released into the bloodstream and is the antioxidant capacity of the blood. 8 Chronic gout causes a lot of prolonged pain and nuisance and limits the mobility of the person; over time, this can lead to complications of muscle atrophy, stiffness, and joint deformity. Also, gout and hyperuricemia are associated with metabolic syndromes, kidney and cardiovascular diseases, hypertension, and diabetes mellitus. 9 Therefore, gout treatment is necessary and is being cared for by both patients and doctors. Today, under the development of the medicine and pharmaceutical industry, based on the mechanisms that cause hyperuric acidemia, many drugs have been released and contribute to the important regimen for gout treatment. Typical medications used for treating acute gout attacks include oral glucocorticoid anti-inflammatory drugs such as prednisone, and prednisolone; nonsteroid anti-inflammatory drugs: Indomethacin, ibuprofen, naproxen, and colchicine. 10 However, besides the effectiveness of treatment, it is worrying that the above drugs all have severe side effects. For glucocorticoids, side effects with short-term use, medium dose, and high dose include mood swings, hyperglycemia, fluid retention and hypertension, cushingoid features, weight gain, and gastrointestinal effects such as gastritis, gastrointestinal bleeding, ulcer formation, and bone and muscle effects. 11 Thus, the aim of this study is to find new molecules as hit compounds targeting XO for safety and effectiveness for patient treatment.
Medicines derived from medicinal herbs tend to be friendly and less toxic to the human body. In Vietnamese traditional medicine, Hedyotis pilulifera (Pit.) Tran Ngoc Ninh. (family Rubiaceae) is often used to treat gastrointestinal diseases and osteoarthritis pain, pharyngitis, and tonsillitis. 12 The main chemical components of this plant are glycosides and triterpene compounds, a group of substances with many good biological activities such as antibacterial, antitumor, and anti-inflammatory. 13 We isolated 3 fatty acids from Hedyotis pilulifera in a previous publication. 14 Interestingly, many studies have shown that certain fatty acids, such as oleic acid, arachidonic acid, and eicosapentaenoic acid, can inhibit XO activity. 15 Based on this evidence, we applied in silico methods combined with an in vitro inhibition study to analyze the mechanical interactions of fatty acids from Hedyotis pilulifera against XO activity.
Result
Virtual Screening
Our previous study isolated 3 fatty acids from Hedyotis pilulifera.
14
However, the bioactivity of these compounds has not been studied. Herein, we applied molecular docking to screen the fatty acids’ inhibitory activity on XO. Following the previous publication, the protocol was validated by redocking the cocrystal ligand to the 3D protein structure.
16
This docking protocol can be applied in this study as, according to the findings, the docking pose has a similar conformation to the crystal pose with an RMSD of 1.327 Å. Allopurinol is commonly used in gout treatment due to the competitive inhibitory activity of XO.
17
Therefore, we selected allopurinol as a reference inhibitor. The structures of the 3 compounds are shown in Figure 1. In the previous study, the stereoisomers of the 3 fatty acids were not determined; therefore, we created all of the stereoisomers of these compounds to validate their binding poses to XO. In particular, 9,12,13-trihydroxyoctadeca-10E,15Z-dienoic acid (

Chemical structures of fatty acids.
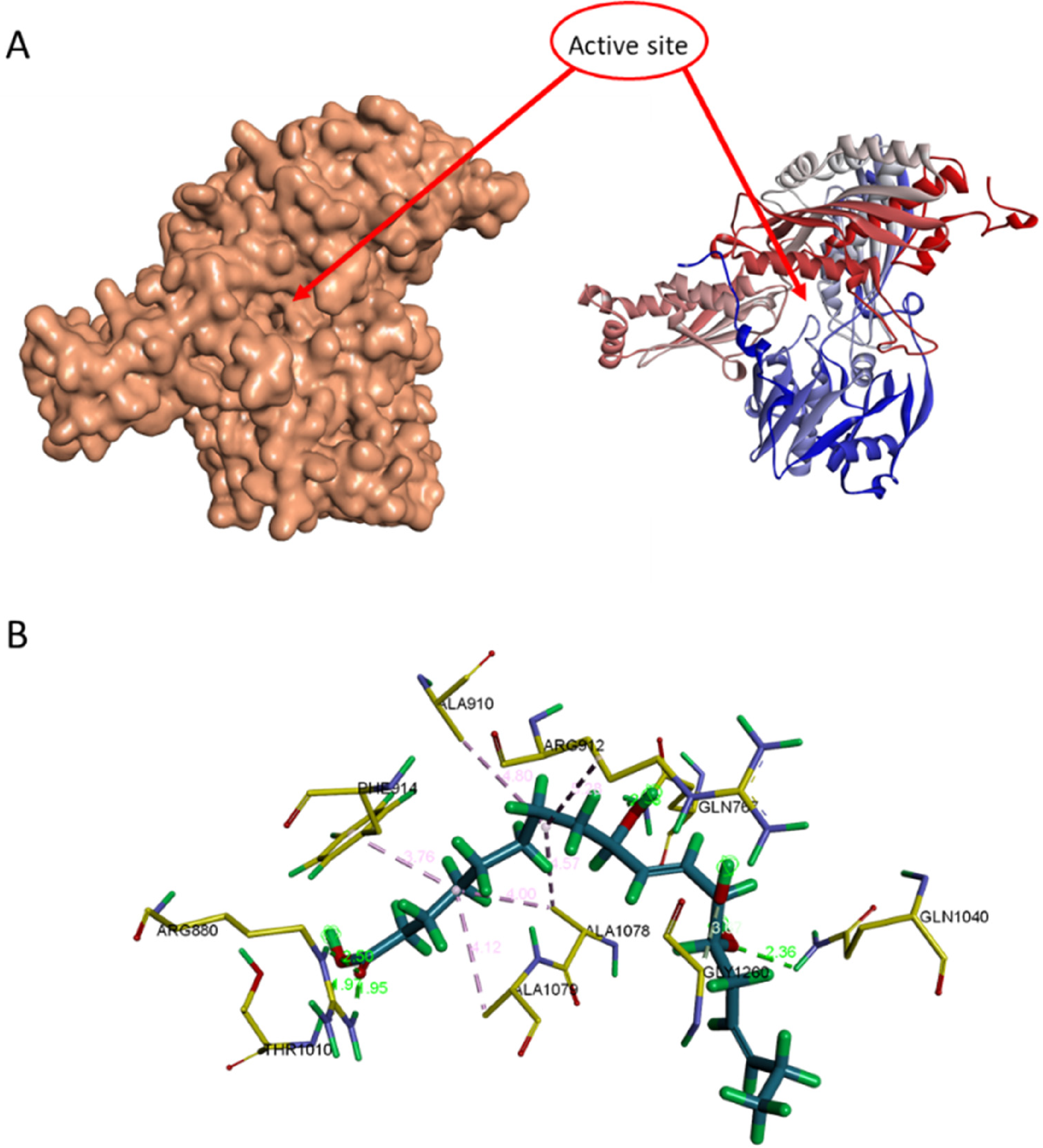
(A) The docking site of xanthine oxidase (XO) (B) interaction between compound
Docking Results of the Stereoisomers of the 3 Fatty Acids Selected for Virtual Screening.

Xanthine Oxidase Inhibitory Activity
Accordingly, in vitro experimental investigation followed the virtual screening to identify the potential hit compound(s) against XO. A range of different concentrations of compounds was used to assess their inhibitory activity, and allopurinol was utilized as the positive control. Compound
Molecular Docking Analysis
The highest score docking pose was shown and selected for dynamic simulation to further explain the biological interaction mechanism between compound
Bovine XO's overall sequence resembles that of human XO by approximately 90%. Most amino acids that interact with ursolic acid are conserved in both human and bovine. 18 The binding pose's lipophilic character and Van der Waals interaction are preserved because both enzymes’ amino acids at the catalytic region are the same. Hence, the molecular docking simulations suggested binding is applicable to the human enzyme.
Molecular Dynamic Simulation
Binding site stability
The molecular dynamic simulation selected the ligand's optimal docking pose, demonstrating general stability over 100 ns. Analysis of the RMSD of the protein carbon backbone in Figure 3A indicated that the protein rapidly attained a steady state after the 10th ns with an average RMSD value of approximately 0.5 nm. A similar result can also be seen for the complex, where the RMSD value changed slightly. Meanwhile, compound

Molecular dynamic simulation interaction between compound
Hydrogen bond analysis
During the 100 ns simulation, the ligand consistently formed hydrogen bonds with the protein, as shown in Figure 3D. Among the initial 10 ns, the protein and ligand formed 4 to 8 hydrogen bonds. However, from the 10th ns onward, the ligand only formed 2 to 4 hydrogen bonds with the protein. Nevertheless, the ligand-protein complex remained stable until the end of the simulation.
Binding free energy
At the final stage of simulation, the binding free energy of compound
Components of Binding Free Energy Calculation.
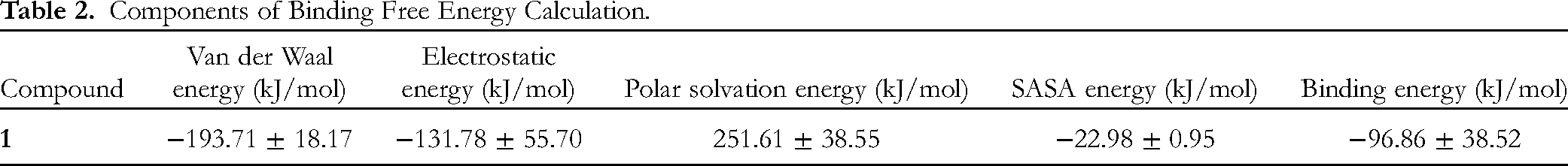
ADMET Studies
Lipinski's Rule of 5 is a principle used in drug discovery that assesses the pharmacokinetic and pharmacodynamic properties of a potential drug compound. The parameters, such as molecular weight (MW), log of octanol/water partition coefficient, number of hydrogen bond acceptors (nHBA), and number of hydrogen bond donors (nHBD), are considered to determine the oral bioavailability of compounds with optimal membrane permeability and hydrophobicity for therapeutic purposes. Acceptable values of nRotB and MR suggest oral bioavailability of drugs and good intestinal absorption.
According to the results presented in Table 3, compound
Drug Properties of Isolated Compounds Analyzed With SwissADME.

Abbreviations: MW, molecular weight; LogP, Log of octanol/water partition coefficient; nHBA, number of hydrogen bond acceptor(s); nHBD, number of hydrogen bond donor(s); MR, molar refractivity; nRotB, number of rotatable bonds; Log S, log of solubility; TPSA, total polar surface area.
Table 4 shows that compound
ADME Predictions of Isolated Compounds Computed by SwissADME.

Abbreviations: log Kp, Log of skin permeability; GI Abs, gastrointestinal absorption; BBB Per, blood–brain barrier permeability; P-gp, P-glycoprotein; CYP, cytochrome-P.
Discussion
Xanthine oxidase is an enzyme that plays a key role in the production of uric acid in the body. Uric acid is a waste product formed when purines are broken down. Excess uric acid can lead to several health problems, including gout and kidney stones. Many studies have shown that certain fatty acids can inhibit XO activity. In particular, polyunsaturated fatty acids such as oleic acid, arachidonic acid, eicosapentaenoic acid, and docosahexaenoic acid, found in fish oil, have been shown to have XO inhibitory effects. 15 These fatty acids reduce the production of reactive oxygen species (ROS) in the body, which is known to activate XO. By reducing ROS production, these fatty acids can inhibit XO activity and lower uric acid levels in the body. In addition, Kelley et al also showed that nitro-oleic acid could inhibit XO activity in a concentration-dependent manner. 22 In our study, compared with control compounds, we have shown that 9,12,13-trihydroxyoctadeca-10E,15Z-dienoic acid has the ability to inhibit XO activity. Although further studies are needed to determine the absolute configuration of this compound, the molecular docking results also indicated no significant difference in binding energies between the possible configurations of 9,12,13-trihydroxyoctadeca-10E,15Z-dienoic acid for XO. In addition, by selecting the configuration with the best binding energy, the binding mechanism between 9,12,13-trihydroxyoctadeca-10E,15Z-dienoic acid and XO is also clarified by simulation methods. Although further experimental studies are needed to modify the structure of 9,12,13-trihydroxyoctadeca-10E,15Z-dienoic acid to increase its biological activity and reduce the risk of toxicity to the body, this result shows the potential role for fatty acids in the treatment of gout targeting XO.
Conclusion
According to our research, the fatty acids tested, 9,12,13-trihydroxyoctadeca-10E,15Z-dienoic acid, had the highest affinity for XO. In vitro studies also demonstrated its inhibitory activity against XO, compared to the control compound, allopurinol. To understand more about its biological properties, ADMET studies and molecular interaction analysis were undertaken. These results suggest that the bioactive chemical we discovered has promise as a gout therapy. However, more study is required to strengthen this molecule's chemical characteristics and inhibitory activity.
Material and Method
Protein and Ligand Preparation
The Research Collaboratory for Structural Bioinformatics Protein Data Bank was applied to acquire the crystal structure of bovine XO complexed with hypoxanthine (PDB ID: 3NRZ). The remaining ligands and water molecules in the XO crystal structures were removed using Discovery Studio 2020. AutoDockTools version 1.5.6 23 was then utilized to add polar hydrogens and Kollman charges to the protein, after which the macromolecule was exported to a dockable pdbqt format for molecular docking. To compute the RMSD between 2 molecules, Pymol 2.4 was employed. The 3-dimensional structures of fatty acids derived from Hedyotis pilulifera were either downloaded from the PubChem database or generated by MarvinSketch 20.17 24 (ChemAxon). Eventually, employing Open Babel 3.1.1, all ligands were converted into a dockable pdbqt format.
Molecular Docking
AutoDock Vina 1.2.0 was utilized to find potential compounds that react with XO. 25 A grid box covering the protein's active site was generated using the parameters center_x, center_y, and center_z = 37.35, 19.85, 17.85, respectively, and the size of the grid box (30 × 30 × 30). Based on their docking scores, the compounds were ordered and reported in kcal/mol. Finally, Discovery Studio Visualizer 2020 was employed to visualize the molecular interactions between the proteins and the selected ligands.
In Vitro Assay
The previously published protocol was slightly modified to assess XO's inhibitory activity in 96-well plates using a spectrophotometric technique. 16 The experiment included 150 nmol/L of xanthine solution dissolved in phosphate buffer (pH 7.5), 70 nmol/L of phosphate buffer (pH 7.5), and 0.01 IU/mL of XO solution. Test compounds were prepared in dimethyl sulfoxide (DMSO) and diluted with phosphate buffer (pH 7.5). The absorbance was measured at 290 nm on a microplate spectrophotometer (BioTek Epoch). The blank was prepared in the same way, but without the XO solution. The negative control was 0.5% DMSO without an inhibitor, and the positive control was allopurinol.
The compounds were tested at 500, 100, 20, and 4 μg/mL doses and determined as the half-maximal inhibitory concentration (IC50) using SigmaPlot 12.5. All experiments were repeated 3 times. The XO inhibitory activity was expressed as the inhibitory percentage (I) calculated using the following formula:
Molecular Dynamics Simulations and Binding Free Energy Calculations
The simulation technique was implemented following our earlier studies. 26 The stability of the ligand in the complex with XO was further investigated using molecular dynamics simulations with GROMACS 2020.4. The complex's topology under the CHARMM36 forcefield was generated using the CHARMM-GUI server, 27 and it was then solvated in a truncated octahedral box containing TIP3P water molecules. NaCl (0.15 M) was added to the system. The requisite values of 300 K temperature and 1 bar pressure were attained to provide a completely stable system. Finally, the molecular dynamics simulation for the complex was run for 100 ns, while the trajectory data were saved every 10 ps for analysis.
In Silico Drug-Likeness and Toxicity
The Lipinski rule has been employed to predict a compound's drug-likeness. The SwissADME website, which provides details on pharmacokinetics properties, was utilized to calculate the ADME parameters of the ligands. 28 The DL-AOT prediction server was used to predict acute oral toxicity (T). 29
Footnotes
Acknowledgments
This research was supported by Hue University under the Core Research Program, Grant No. NCM.DHH.2023.01.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was funded by Hue University under the Core Research Program, Grant No. NCM.DHH.2023.01.
