Abstract
The increasing antimicrobial drug resistance of Salmonella typhimurium has been a major public health problem in many countries. The type III secretion system (TTSS) of S typhimurium is an attractive target for the development of new antibiotics against this bacterium. In this study, we determined whether berberrubine derived from Berberis vulgaris could inhibit TTSS expression in S typhimurium. Berberrubine inhibited TTSS expression at a concentration of 60.9 µM. Additionally, TTSS-related mRNA expression of sipA, sipB, sopA, and sopD was downregulated in the presence of berberrubine. Collectively, we anticipate that berberrubine can be a useful tool to prevent the risks of infections caused by pathogenic bacteria.
This is a visual representation of the abstract.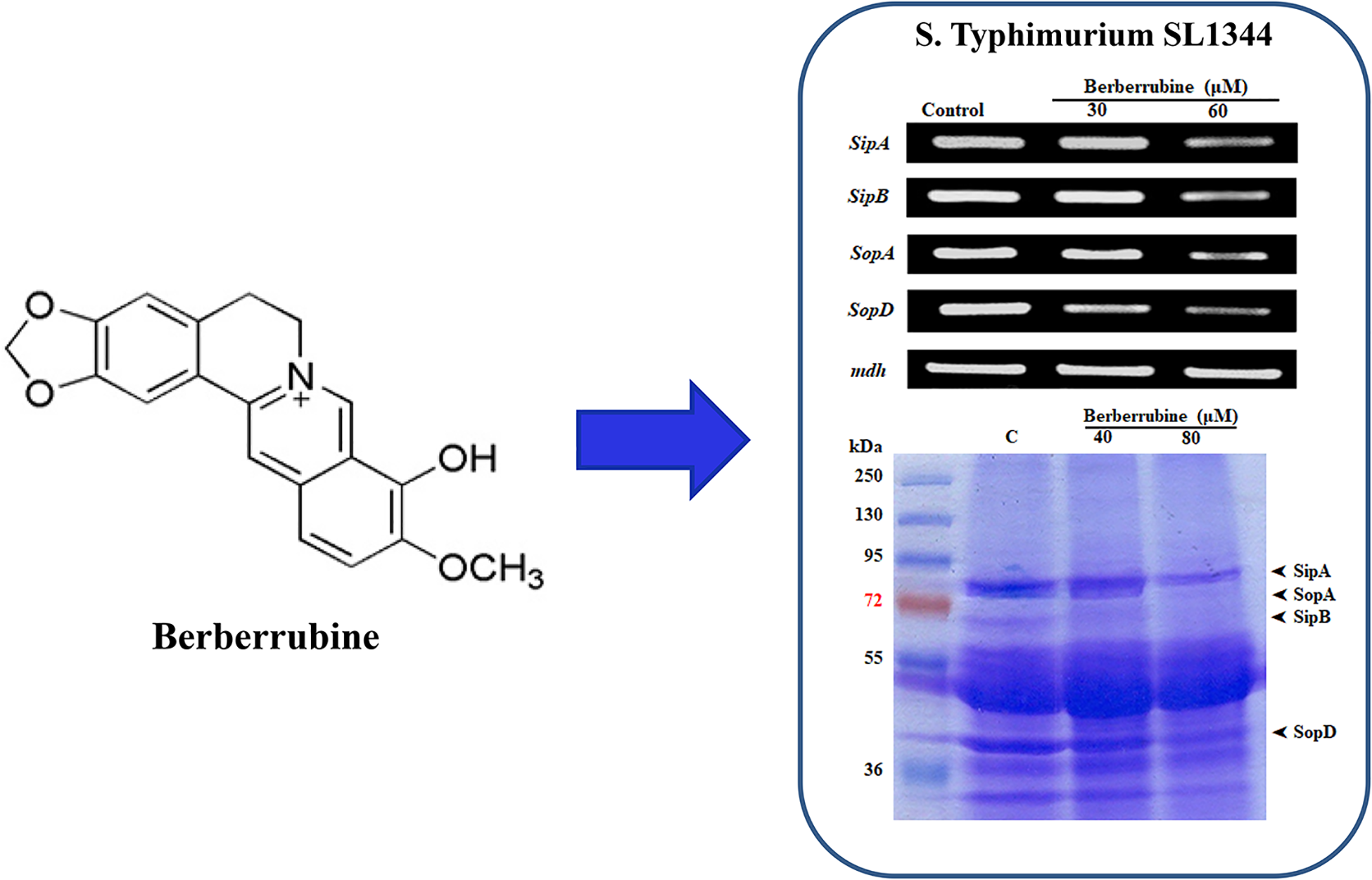
Introduction
Approximately 250 000 to 500 000 plant species are currently known to exist, but only 1% to 10% are used by humans. 1 Plants are the major sources of food, cosmetics, and medicine. Plant-derived secondary metabolites, including alkaloids, flavonoids, phenols, terpenoids, and steroids, have pharmacological properties, such as antibacterial and antifungal effects.2‐4
Globally, many humans are exposed to infectious diseases caused by pathogenic bacteria such as Salmonella, Escherichia coli, Yersinia, and Vibrio. Recently, several antibiotic-resistant bacteria have become a public health problem worldwide.5,6 Therefore, there is a need for strategies to develop new antibiotics that are safe, nontoxic, without selection pressure, and against which pathogenic bacteria are less resistant. Gram-negative bacteria have virulence factors called Type III secretion systems (TTSSs) that enable the translocation of bacterial effector proteins directly into the host cell cytosol. 7 Consequently, TTSSs are good targets for developing antibacterial agents because they can inhibit infection without suppressing bacterial growth. 8 Salmonella has 2 different virulence-related TTSSs that are encoded within pathogenicity islands: Salmonella pathogenicity island 1 (SPI-1) and Salmonella pathogenicity island 2 (SPI-2). SPI-1 and SPI-2 play fundamental roles in the pathogenesis within host cells. SPI-1 is essential in the invasion stage of host cell infection, while SPI-2 is the major factor contributing to apoptosis, proliferation, and bacterial survival within macrophages. 9
Several of the TTSS inhibitors known so far are of natural origin, such as plants, bacteria, and fungi. The caminosides were identified from Caminus sphaeroconia, and caminoside A suppresses the secretion of effector protein (EspB) in EPEC. Aurodox, guadinomines, cytosporone B, and fusaric acid are produced by Streptomyces goldiniensis, Streptomyces sp. K01-0509, Dothirorella sp., and the fungus Fusarium oxysporum, respectively. Furthermore, hopeaphenol, sanguinarine chloride, and thymol are natural products isolated from plants with TTSS inhibitory effects. Hopeaphenol was isolated from Anisoptera thurifera and A polyandra and suppressed TTSS in Y pseudotuberculosis, P aeruginosa, and C trachomatis. Sanguinarine chloride, produced by the bloodroot plant, Sanguinaria canadensis, was also found to inhibit the expression of TTSS against Salmonella enterica serovar typhimurium. Thymol, found in Thymus species, inhibited the secretion system of Salmonella.
Berberrubine, an active metabolite of berberine, can be isolated from Berberis vulgaris and possesses various pharmacological activities (Figure 1). It can also be synthesized from berberine chloride by vacuum pyrolysis at 200 °C with urea. 10 Berberrubine has also been reported to exhibit antifungal and antitumor activities.11‐13
The chemical structure of berberrubine.
In this study, we investigated whether berberrubine could inhibit TTSS activity, bacterial growth, and TTSS-related gene expression of Salmonella typhimurium. To our knowledge, this is the first study to show the TTSS inhibitory activity of berberrubine. Therefore, it is suggested that the results obtained from this study would be useful for the development of novel antivirulence agents.
Results and Discussion
Inhibition of TTSS-Mediated Hemolytic Activity by Berberrubine
The TTSS inhibitory activity was investigated to confirm the TTSS inhibition of berberrubine using a contact-dependent hemolysis assay. Berberrubine inhibited Salmonella typhimurium TTSS-mediated hemolysis of the red blood cells (Figure 2A).

Inhibition of type III secretion system (TTSS)-mediated hemolytic activity in vitro by berberrubine. (A) Berberrubine inhibits TTSS-mediated contact-dependent hemolysis. (B) The inhibition of TTSS-mediated hemolytic activity is not attributable to berberrubine-induced bacterial growth inhibition. The data represent the average values from three independent experiments with standard deviation. Control: Salmonella typhimurium without berberrubine treatment, Berberrubine: Salmonella typhimurium treated with berberrubine.
A bacterial growth inhibition assay and minimum inhibitory concentration (MIC) test were performed to confirm the potential of berberrubine to inhibit the growth of Salmonella typhimurium. It was assessed by cultivating the bacteria in either the presence or absence of berberrubine. The growth curve shows that 100 mM berberrubine did not significantly inhibit the growth of Salmonella typhimurium compared to that of the control (Figure 2B). The MICs of berberrubine against pathogenic bacteria are shown in Table 1. The MIC of berberrubine for Salmonella typhimurium was 251 mM. The MIC of nalidixic acid, which was used as a control in the MIC test, for Salmonella typhimurium was 31.4 mM. These results indicated that the inhibition of TTSS expression did not reduce bacterial growth. Moreover, TTSSs which are essential for the virulence of Salmonella spp. SPI-1 and SPI-2 encode effector proteins, such as SipA, SipB, and SipC, whereas SopA, SopB, and SopD are encoded by genes which are located outside SPI-1. 14 Because the TTSSs of Salmonella typhimurium are good targets for developing novel antibiotics, we determined whether berberrubine suppressed the TTSS-1 gene expression.
Minimum Inhibitory Concentrations for Antibacterial Activity of Berberrubine against Pathogenic Bacteria.

Suppression of TTSS Effector Gene and Protein Secretion by Berberrubine
Sodium dodecyl sulphate-polyacrylamide gel electrophoresis (SDS-PAGE) and Coomassie blue staining were examined to determine whether berberrubine suppresses TTSS effectors. Figure 3C shows that the effector protein (SipA, SipB, SopA, and SopD) secretion by S typhimurium SL1344 was suppressed in the presence of berberrubine. Furthermore, berberrubine exhibited dose-dependent inhibitory effects on TTSS-mediated protein secretion. The effect of berberrubine on the expression of TTSS effectors was estimated by reverse transcription polymerase chain reaction (RT-PCR). Berberrubine downregulated the mRNA expression of some effectors, including sipA, sipB, sopA, and sopD in S typhimurium SL1344 (Figure 3A and B). Figure 3 shows that the TTSS-related mRNA expression, such as that of sipA and sipB, was downregulated at a 60 mM concentration of berberrubine. The mRNA expression of sopA and sopD was slightly suppressed at a 30 mM concentration of berberrubine, which was significant compared to that in the presence of 60 mM of berberrubine. A prevalently control mdh was used to normalize the RT-PCR findings between samples. The RT-PCR data indicated that berberrubine inhibited mRNA expression. We also found that berberrubine inhibited the TTSS expression of Salmonella typhimurium at concentrations that did not reduce bacterial growth (Figures 2B and 3A). The IC50 of berberrubine for TTSS inhibition was 60.9 mM, but it also showed antibacterial activity at higher concentrations. In our previous study, we investigated the inhibitory effect of obovatol on the TTSS expression of Salmonella typhimurium by regulating the expression of TTSS effector genes and secreted proteins without affecting bacterial growth. 15 It was observed that obovatol had hemolytic properties (IC50: 19.8 mM). A comparison of the inhibitory activities of berberrubine and obovatol based on RT-PCR and SDS-PAGE findings demonstrated that berberrubine was more potent, as confirmed by the band intensities for SipA, SipB, SopA, and SopD (Figure 3C, second and fourth arrowheads). However, SopE2 expression was not confirmed. Nevertheless, berberrubine exhibited greater inhibitory activity at low concentrations by downregulating target proteins than obovatol. 15 Moreover, the band intensity of other effector proteins should be investigated in a concentration-dependent manner using berberrubine, since its novel mechanistic mode of action will shed light on overcoming the drug resistance of Salmonella typhimurium.
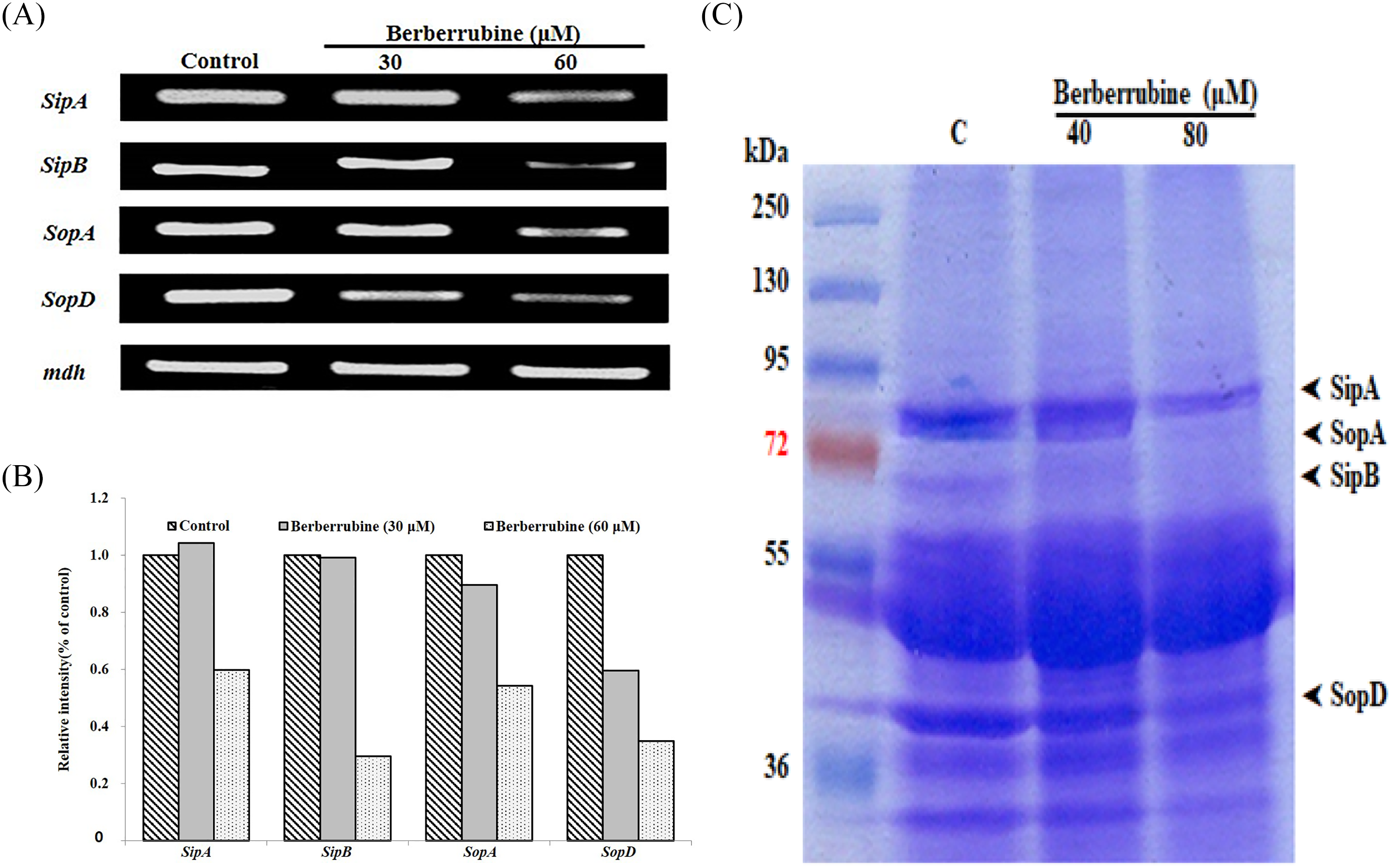
RT-PCR and western blot analyses of Salmonella TTSS-related mRNA and protein expression in the presence of berberrubine. (A) Effector gene expression in S typhimurium SL1344 is suppressed by berberrubine. mdh was used as an internal control. Control: untreated Salmonella typhimurium, Berberrubine: Salmonella typhimurium treated with berberrubine. (B) The relative intensity of TTSS-related mRNA expression. The data indicate a classical set of three independent experiments. (C) The pattern of berberrubine-induced inhibition of effector proteins. The secreted proteins were evaluated by staining the gel with Coomassie blue. Abbreviations: TTSS, type III secretion system; mdh, L-malate dehydrogenase; RT-PCR, reverse transcription polymerase chain reaction.
Molecular Docking Analysis
Table 2 exhibits the binding affinity of three different compounds, that is, berberine, berberrubine, and obovatol, against SipA, SopA, SopD, and SopB, which are potent proteins in the Salmonella typhimurium TTSS. Berberine was taken as a control for the molecular docking study to compare the binding affinity of berberrubine and obovatol with different proteins. Berberine exhibited a higher affinity for Sip A (−7.6 kcal/mol), whereas, in the case of SopB, berberrubine exhibited a higher binding property (−7.4 kcal/mol). In the case of SopD, berberine and berberrubine exhibited the same binding affinity (−6.3 kcal/mol), and for SopA, berberine and obovatol exhibited the same affinity score (−5.9 kcal/mol). Furthermore, hydrogen interaction is considered a strong binding force between the ligand and surrounding amino acids based on ligand–protein interaction. Figure 4 shows that berberrubine forms hydrogen interactions with Lys 16 in chain A, whereas berberine forms carbon–hydrogen interactions with Ser 30 and Thr 35, respectively. There was no evidence to demonstrate binding affinity between TTSS proteins and berberrubine. Moreover, it is difficult to identify the binding effectiveness of ligand molecules to the target proteins in molecular docking, but it is necessary to identify clearly the binding site of the ligand through site-specific mutagenicity of the target protein (TTSS) and analyze the structure–activity relationship based on the structure of berberine, berberrubine, and obovatol as ligands.
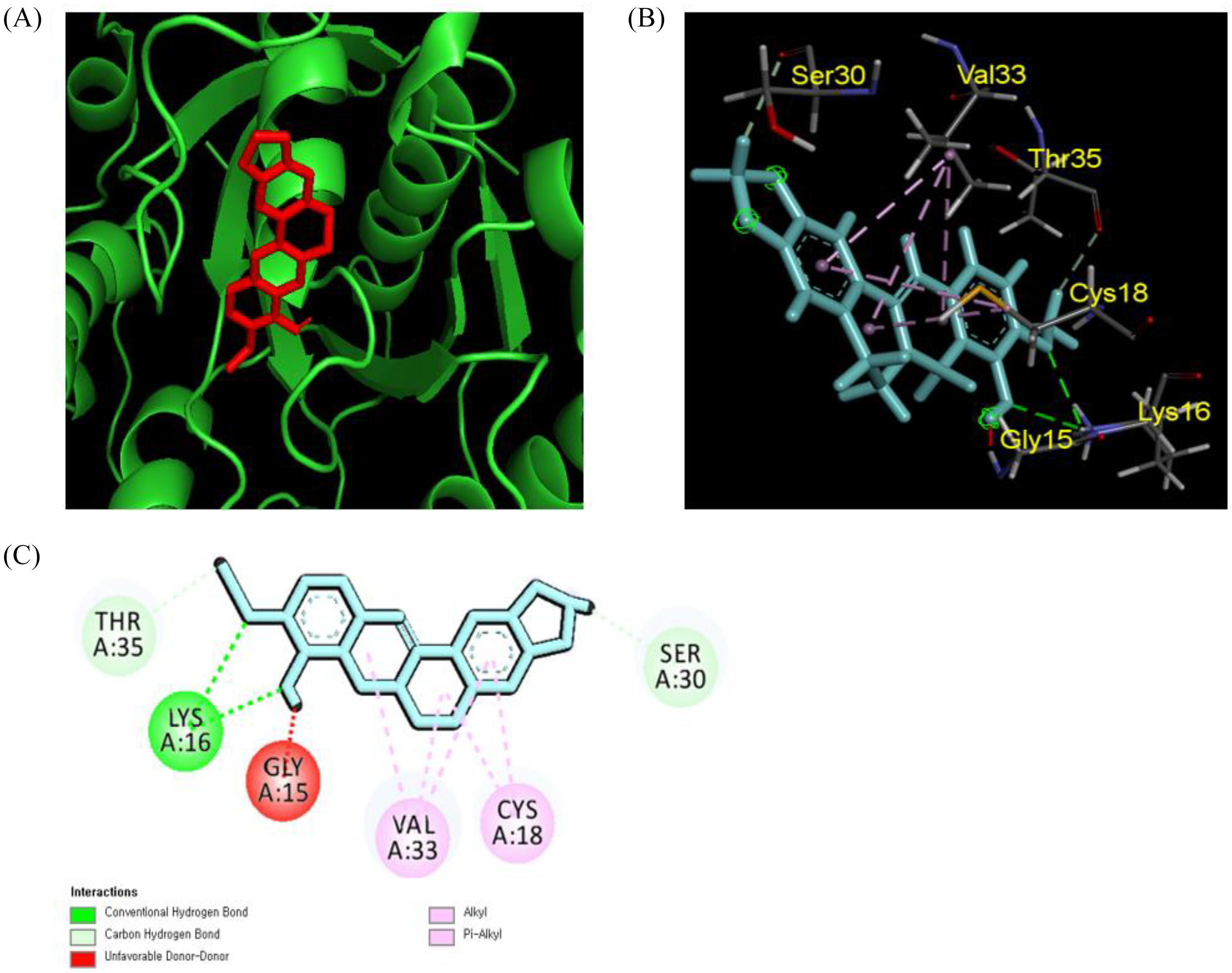
Molecular docking results of berberrubine with SopB (4DID). Berberrubine interaction with SopB protein (A); 3D molecular interaction of berberrubine (B); 2D representation of berberrubine with SopB (C).
Binding Affinity of Berberine, Berberrubine, and Obovatol for Salmonella Type III Secretion System (TTSS) Proteins.
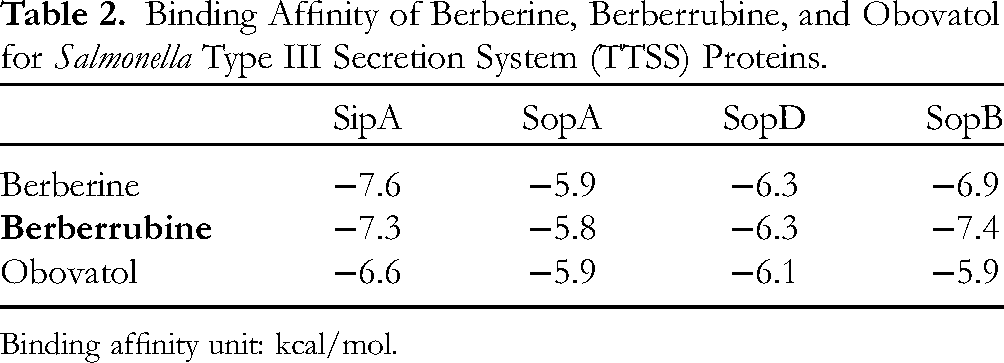

Binding affinity unit: kcal/mol.
Conclusions
The present study involved the search and development of inhibitors of the TTSS to obtain novel antivirulence drugs from natural resources. A hemolytic assay was used to identify TTSS inhibitors. Berberrubine was finally selected, having strong inhibitory activity against Salmonella typhimurium TTSS. This compound does not affect Salmonella typhimurium growth. We performed RT-PCR and molecular docking analysis to identify the mechanism of TTSS inhibition by berberrubine on Salmonella typhimurium. The inhibition of TTSS by berberrubine led to a decrease in effector proteins (SipA, SipC, SopA, SopB, and SopD) in Salmonella typhimurium. This suggests that the mechanism of action is related to the formation of a translocon pore in the host-cell membrane, promoting invasion into the host-cell cytosol, positioning the Salmonella-containing vacuole via a pathway involving Rho GTPase and myosin II, and activating serine/threonine protein kinase B for antiapoptotic signaling, among other effects. In conclusion, the results of this study suggest that berberrubine inhibits TTSS expression in Salmonella typhimurium. Further studies are needed to elucidate the mechanisms underlying TTSS inhibition and berberrubine toxicity. This study can help in the development of new TTSS-inhibiting antibiotics, as berberrubine can be a useful tool for the prevention of infections caused by Salmonella typhimurium.
Materials and Methods
Materials and Bacterial Strain
Berberrubine was purchased from Sigma-Aldrich, Inc., and brain heart infusion broth (BHI) from DIFCO. All other reagents were of the highest quality commercially available. The bacterial strain used in this study, Salmonella typhimurium SL1344, has been described previously by Wilmes-Riesenberg et al. 16 A TTSS-mediated hemolytic assay, bacterial growth curve analysis, and MIC test were performed.
TTSS-Mediated Hemolytic Assay
Salmonella typhimurium TTSS-mediated hemolytic activity was measured by the method adopted by Hudson et al. 17 Briefly, S typhimurium SL1344 was inoculated into the BHI medium and cultured by shaking at 25 °C for 14 hours. The cultures were then diluted 10-fold in fresh culture media. Berberrubine was added into each subculture and incubated at 37 °C for 1 hour. The bacteria were recovered by centrifugation, mixed with red blood cells prepared in a microcentrifuge tube, and centrifuged for 10 minutes at 6000g. The mixture was incubated for 4 hours at 37 °C and resuspended in fresh phosphate-buffered saline (PBS), and centrifuged at 600g for 5 minutes. The cell pellets were washed in PBS and then centrifuged again at 6000g for 5 minutes. Finally, the supernatant was collected to determine the TTSS-mediated hemolytic activity.
Bacterial Growth
Salmonella typhimurium SL1344 was inoculated into BHI broth and grown overnight (14-16) at 2 °C. The overnight culture was diluted 10-fold in fresh BHI media, and the test compounds were added. Later, the cultures were aerated by shaking at 37 °C for 4 hours. An aliquot (100 µL) of each culture was inoculated onto a BHI agar plate and incubated without shaking at 37 °C for 24 to 48 hours. The surviving colonies in each sample were counted using a click counter.
MIC Determination
The MICs of berberrubine were determined by a serial dilution method using various human pathogens. Salmonella typhimurium was inoculated in BHI broth and incubated at 37 °C for 14 hours. The cultures grown overnight were adjusted to ∼105 CFU/mL. The 96-microwell plates were inoculated with 200 µL of each bacterial strain and berberrubine. Later, a 2-fold serial dilution of berberrubine was performed in plates containing 100 µL of fresh media. The plates were incubated at 37 °C for 24 hours, and the MICs were determined.
RNA Extraction and RT-PCR
The berberrubine-treated bacterial cells were collected by centrifugation at 1000g. Total RNA was extracted from the bacteria by using a TRIzol reagent. cDNA was generated from the RNA by using the Superscript III Fist-Strand Synthesis system according to the manufacturer's directions. RT-PCR was performed using the following primer pairs: sipA, forward primer 5′-TGTTCGGCTATTATCAATCGTCT-3′ and reverse primer 5′-CGC AGC AAT CTT ACG CAC CT-3′; sipB, forward primer 5′-CTG ACT GGG CTG CGG TAT TCG TG-3′ and reverse primer 5′-CTG CGG TGG GAC TTG CGG TAA-3′; sopA, forward primer 5′-ATT CAG ACA CGG CGA TGA TG-3′ and reverse primer 5′-TGG CGT CCG TCA GGT GAT AAG CA-3′; sopD, forward primer 5′-TTA CTA TCA AGA TGG ACG CTT CT-3′ and reverse primer 5′-GTG CAT TTC CCG TCA CTT-3′; and mdh, forward primer 5′-ATG AAA GTC GCA GTC CTC GGC GCT GCT GGC GG-3′ and reverse primer 5′-ATA TCT TTT TTC AGC GTA TCC AGC AT-3′. 18 The PCR products were analyzed on 1% agarose gels containing ethidium bromide.
Inhibition of Protein Secretion
Overnight cultures of S typhimurium SL1344 (conditions: BHI broth, 14 hours, and 25 °C) were inoculated into fresh medium containing the designated berberrubine concentrations and incubated for 4 hours at 37 °C. The proteins secreted into the culture supernatant were precipitated with trichloroacetic acid at a concentration of 10% (v/v) and collected by centrifugation (3600g, 20 minutes, 4 °C). The protein pellets were washed with cold acetone, resuspended in loading buffer, separated by SDS-PAGE, and stained with Coomassie blue.
Molecular Docking of Berberrubine
To explore the binding affinity of berberrubine, and obovatol with different proteins, including SopB, SopA, SopD, and SipA, we performed molecular docking simulations using AutoDock Vina 19 through the DockingApp's interface. 20 All the water molecules and ligands were deleted from the initial structure of SopB, SopA, SopD, and SipA proteins, and polar hydrogen atoms were added before docking. The grid box was centered for each target on the native binding site. The grid box for SopB was set to 16 Å × 24 Å × 20 Å with its center at positions x = 25.85, y = −7.3, and z = 10.80, whereas for SopD the grid box was set at x = 55.23, y = 9.78, and z = 8. Similarly, for SopA and SipA, the grid box information was as follows; x = −10.17, y = −26.66, and z = 61.32; x = 17.66, y = 26.41, and z = 10.41, respectively. The molecular binding was then visualized using the Pymol and Discovery Studio. 21
Footnotes
Acknowledgements
The lab members are greatly appreciated for their critical comments.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Ethical Approval
This research was not applicable for ethical approval.
Statement of Human and Animal Rights
This article does not contain any studies with human or animal subjects.
Informed Consent
There are no human subjects in this article and informed consent is not applicable.
Trial Registration
Not applicable, because this article does not contain any clinical trials.
