Abstract
Introduction
Cancer poses a serious present threat to human health. According to statistics from the World Health Organization, in 2020 there were 19.29 million new cancer cases and 9.96 million deaths worldwide. 1 Cancer is considered the second largest cause of death globally. 2 Although anticancer drugs have made remarkable progress, many cancers remain resistant or even virtually intractable. 3 Many anti-cancer drugs also feature serious adverse reactions and high prices, both of which take a heavy toll on patients and their families. Therefore, the development of anticancer agents is an urgent need.
Bioactive peptides have been widely demonstrated to exert anticancer effects through multiple pathways.4,5 Antimicrobial peptides (AMPs) are a subset of biological peptides derived from both prokaryotes and eukaryotes. True to their namesake, they boast antibacterial, antiviral, and immunomodulatory properties.6‐8 However, they have also been considered as potential anticancer drugs. Numerous AMPs are found in amphibians, which are inseparably related to their living environment.9,10 AMPs are usually composed of 12 to 50 amino acid residues and have an overall positive charge. 11 This positive charge is one of the key features of AMPs. Maria Magana 12 and Jaspreet Kaur Boparai 13 found that AMPs often achieve their antibacterial effects by destroying the membrane structure through electrostatic interaction with bacterial membranes. The precise mechanisms of action include electroporation, barrel staves, disordered toroidal pores, toroidal pores, membrane thinning or thickening, and many others. Other properties of AMPs, which include structure and amphiphilicity, are also involved. Some AMPs achieve cell necrosis or apoptosis by interfering with mitochondrial performance, cell cycle, and other pathways. 6 This has been documented for a variety of malignancies, including bladder, hepatocellular, gastric, skin, and ovarian cancer.14‐16
In this study, we discovered a novel antimicrobial peptide, named brevinin-1EG, from the skin tissue of
Materials and Methods
Materials
U-2OS, HepG2, HT-29, A375, HeLa, and A549 cells used in this study were preserved by our laboratory. RPMI 1640 medium was used to grow HT-29, A375, and A549 cells. The rest of the cells were grown in Dulbecco's modified Eagle medium (DMEM).
Experimental Animals
Extraction of Skin Tissue RNA and Construction of cDNA Library
Frogs were executed under anesthesia. A small section of dorsal skin was removed and minced in 1 mL of TRIzol. Total RNA was extracted according to the instructions that came with the TRIzol reagent. A cDNA library was subsequently constructed by reverse transcription using the HiScript II 1st Strand cDNA Synthesis Kit (Vazyme). Oligo (dT)18 in the kit was replaced with specific primers of our own design (5'-ATTGACTCGAGTCGACATCGATTTTTTTTTTTTTTTTTT-3ʹ). The specific operation refers to the instruction manual.
PCR and Original TA Cloning Kit
Using the synthetic cDNA as our template, we amplified a polymerase chain reaction (PCR) product using the upstream primer (5'-ATTGACTCGAGTCGACATCGA-3ʹ) and the downstream primer (5'-ATGTTCACCACAAAGAAATCCA-3ʹ). The PCR cycle program was as follows: predenaturation at 95 °C for 2 min; 30 cycles of denaturation at 95 oC for 30 s, annealing at 52 °C for 30 s, and extension at 72 °C for 1 min. PCR products were detected using agarose gel electrophoresis and recovered using gel recovery kit (Zomanbio). The recovered products were ligated with pMD-19T, and the ligated products were transformed into
BLAST Alignment
The sequencing results were compared on the NCBI website by Basic Local Alignment Search Tool (BLAST) (https://blast.ncbi.nlm.nih.gov/Blast.cgi), and the amino acid sequences of the mature peptides were analyzed.
Bioinformatic Characterization of Brevinin-1EG
Bioinformatic characterization was performed with online analysis tools Heli Quest and I-TASSER, which predicted the basic properties and structural features of brevinin-1EG.
Synthesis of Brevinin-1EG
Brevinin-1EG was synthesized by GL Biochem (Shanghai) Limited using a solid-phase chemical synthesis method. Its molecular weight was calculated to be 2637.25 Da, and its purity was found to be > 95% by mass spectrometry and high-performance liquid chromatography.
Structural Testing
The structure of samples in different solutions was detected by translational circular dichroism. 17 We dissolved brevinin-1EG in 10 mM phosphate buffered saline (PBS), 30 mM sodium dodecyl sulfate (SDS), or 50% trifluoroethanol (TFE) solution and formulated it into a loading sample with a concentration of 100 μg/mL. The circular dichroism parameters were as follows: wavelength range 190 to 300 nm; bandwidth 1 nm; reaction time 1 s; scanning speed 100 nm/min. Finally, these data were analyzed using Colabfold online tools.
Hemolytic Activity Assay
Hemolysis experiments were slightly modified from existing methods.
18
Briefly, 200 μL of 4% (v/v) sheep red blood cells (Solarbio) were placed into an Eppendorf tube, along with either peptide solutions of different concentrations or Triton X-100. These tubes were allowed to incubate at 37 °C for 30 min. The above solution was centrifuged at 1000 rpm for 5 min, and the absorbance of the supernatant was measured at 540 nm. PBS and 1% TritonX-100 were used as the negative and positive control, respectively. Hemolysis was calculated using the following formula:
Antioxidant Assays
We examined the ability of brevinin-1EG to scavenge ABTS·+ and DPPH radical cations, which have demonstrated antioxidant activity. The specific experimental operation method can be found in the reference.
19
Each experiment consisted of 3 independent replicates. The following equation was used to measure the radical scavenging rate: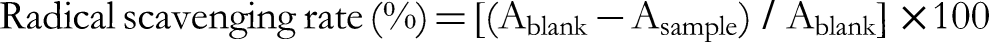
ABTS·+ Radical Scavenging Ability
A stock solution, prepared by mixing 0.2 mL of 7.4 M ABTS (Aladdin) and 0.2 mL of 2.6 M K2S2O, was left to incubate in the dark for 12 h. We then diluted it with PBS buffer to ensure that the absorbance at 734 nm was 0.7 ± 0.02. We mixed 190 μL of ABTS·+ working solution with different concentrations of 10 μL brevinin-1EG in a 96-well plate, which was incubated at 37 °C for 30 min in the dark. Finally, we detected the absorbance of the mixed solution at 734 nm.
DPPH Radical Scavenging Ability
We prepared a 0.1 mM DPPH (MedChemExpress) solution, diluted with absolute ethanol to an absorbance of 1.0 at 517 nm. DPPH solution (190 mL) and 10 μL of brevinin-1EG solution were added to each well of a 96-well plate and allowed to incubate in the dark at 37 °C for 30 min. After the reaction was complete, the absorbance of the solution at 517 nm was detected.
Antiproliferative Activity Assay
The antiproliferative activity of brevinin on human HeLa, melanoma, and liver cancer cells was measured by the CCK-8 kit (Bioss). 20 We seeded 5 × 103 cells into each well of a 96-well plate and cultured them in a medium containing 10% serum for 24 h. Different concentrations of brevinin-1EG were then added to each well and allowed to incubate for a further 24 h. We then added 10 µL of CCK-8 to each sample and incubated it for 2 h. Finally, the absorbance at 450 nm of each sample was measured using an ELISA plate reader.
Lactate Dehydrogenase (LDH) Detection
The content of total intracellular LDH was detected by the LDH cytotoxicity detection kit (Beyotime), following the manufacturer's instructions. We seeded 5 × 103 cells into each well of a 96-well plate. Once the cells had reached 80% to 90% confluency, different concentrations of brevinin-1EG were added to the treated cells for 24 h. In addition, we set background blank control wells, sample control wells, and sample maximum enzyme activity control wells. One hour before the scheduled detection time point, we added 20 μL LDH release reagent provided by the kit to the sample maximum enzyme activity control well.
Statistical Analysis
We analyzed the data using Student
Results
Cloning of cDNA Encoding the Premaster of Brevinin-1EG
The full-length cDNA encoding brevinin-1EG was successfully cloned from the skin tissue cDNA library. The DNA sequence consists of 216 nucleotides in total and the brevinin-1EG precursor of 71 amino acid residues (Figure 1). These include a signal peptide region, an acidic amino acid-rich spacer region, a translocase cleavage site, and a maturation region (consisting of 24 amino acids), whose sequence is FLPLLAGLAANFLPTIFCKISRKC.

The cDNA and deduced amino acid sequence of brevinin-1EG.
We discovered a peptide that had a “Rana box” (Cys18-(Xaa)4-Lys-Cys24) and 4 invariant residues, identified as Ala9, Cys18, Lys23, and Cys24. These are hallmark features of the brevinin family. 21 Thus, we introduce brevinin-1EG as the most newly discovered member of the brevinin-1 family.
Bioinformatics Analysis of Brevinin-1EG
To understand brevinin-1EG further, bioinformatics and secondary structure were further analyzed using Heli Quest, I-TASSER, and Colabfold online tools. The results show that brevinin-1EG carries a + 3 net charge, a hydrophobicity of 0.815, a hydrophobic moment of 0.279, and a hydrophobic surface composed of I, A, L, I, L, A, C, F, A (Figure 2A and B). What is more, the secondary structure of brevinin-1EG contains a large number of α-helix structures (Figure 2A to D).

Bioinformatics and structure analyses of brevinin-1EG. (A) Structural parameters of brevinin-1EG. (B-C) The α-helix-dominated structure prediction of brevinin-1EG and the arrow indicates the hydrophobic surface. (D) Secondary structure prediction of brevinin-1EG. (E) Circular dichroism analysis of brevinin-1EG in 10 mM PBS, 30 mM SDS, or 50% TFE solution.
Structure Prediction of Brevinin-1EG
The molecular weight of the synthetic peptide brevinin-1EG was 2637.25 Da, and its purity was greater than 95% (Figure 3). To confirm the online analysis results, circular dichroism was used to test the structure of brevinin-1EG dissolved in either 10 mM PBS, 30 mM SDS, or 50% TFE (Figure 2E). When brevinin-1EG was dissolved in 10 mM PBS, it exhibited a randomly coiled structure. In a 30 mM SDS and 50% TFE solution of brevinin-1EG, there were 2 negative Cotton effects at >200 nm and one positive Cotton effect at 192 nm, which is characteristic of α-helices. Thus, brevinin-1EG appears to be mainly dominated by an α-helical structure, which is consistent with the predicted structure.

High-performance liquid chromatogram (HPLC) and mass spectra (ms) of brevinin-1EG. (A) The purity of brevinine-1EG was >95%. (B) The retention time and peak area information of brevinin-1EG. (C) The molecular mass of brevinin-1EG is 2637.25.
Antiproliferative Activity
To detect the antiproliferative activities of the synthesized peptides, we treated U-2OS, HepG2, HT-29, A375, HeLa, and A549 cells with brevinin-1EG. Brevinin-1EG exhibited strong anti-proliferative activity against all 6 tested tumor cells, particularly against HepG2 cells, with an IC50 value of 15.88 μg/mL (Figure 4 and Table 1). The IC50 values of brevinin-1EG on U-2OS, HT-29, A375, HeLa, and A549 cells were 19.30, 19.75, 19.33, 21.21, and 21.46 μg/mL, respectively (Table 1).

The antproliferative activity of brevinin-1EG and its effect on lactate dehydrogenase (LDH) release in cancer cells. (A) U-2OS, HepG2, HT-29. (B) A375, HeLa, A549. (C) and (D) Brevinin-1EG induced the release of LDH in HepG2 and HeLa cells. Compared to the negative control group, *
The IC50 Value of Brevinin-1EG Calculated From Figure 4A and B.

LDH Detection
LDH released by cells can assess cell membrane integrity. We detected the release of LDH when different concentrations of brevinin-1EG acted on HepG2 and HeLa cells. The results indicate that brevinin can exert an anti-proliferative effect by destroying the membrane of cancer cells (Figure 4C and D).
Hemolytic Activity
The hemolytic activity of brevinin-1EG was tested using 4% (v/v) sheep red blood cell suspension. Brevinin-1EG has a certain hemolytic activity with an HC50 of 138.6 μg/mL (Figure 5A and B). However, when added to the blood cell suspension at the IC50 effective against tumor cells, brevinin-1EG caused hardly any hemolysis. Only about 34.44% hemolysis was observed at 5 times the IC50, supporting the safety of brevinin-1EG.
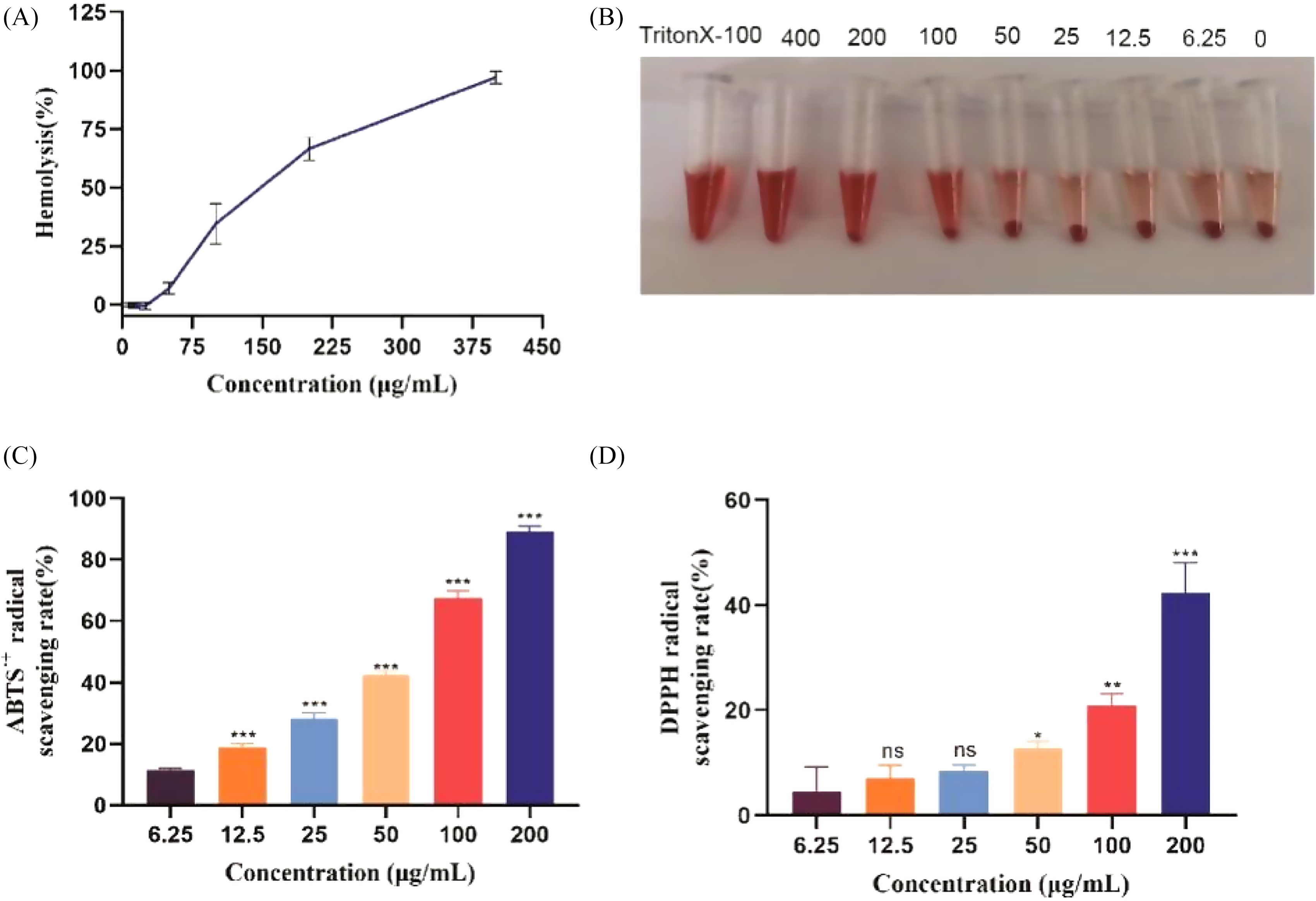
Hemolytic and antioxidant activities of brevinin-1EG. (A) and (B) Effects of 6.25, 12.5, 25, 50, 100, 200, and 400 μg/mL concentrations of brevinin-1EG on erythrocytes. (C) Brevinin-1EG on ABTS + radical scavenging activity. (D) Brevinin-1EG on DPPH radical scavenging activity. *
Antioxidant Activity In Vitro
Figure 5C and D shows the ABTS·+ and DPPH free radical scavenging kinetics of brevinin-1EG, which exhibits potent antioxidant activity in a concentration-dependent manner. Among these 2 free radicals, the ability of brevinin-1EG to scavenge ABTS·+ was stronger than that of DPPH. When the concentration of brevinin-1EG was 200 μg/mL, 89.25% of ABTS·+ and 42.37% of DPPH were removed after 5 min.
Discussion
Frogs are an important source of AMPs, which are an important part of their innate immunity and defense against microbial infections. 22 AMPs can be divided into cathelicidin, brevinin, and other families. 23 The brevinin family consists of brevinin-1 and brevinin−2, both of which have a “Rana box” and α-helical structure. We confirmed the antimicrobial peptide in this article as brevinin-1 and named it brevinin-1EG. Both the online tool and the circular dichroism test indicated the structure of brevinin-1EG to be an α-helix (Figure 2).
Curing cancer is one of the biggest challenges in biomedicine. At present, although a variety of drugs have been marketed, there is still a long way to go toward achieving a cure, and present treatments exact a heavy toll in terms of adverse reactions. A considerable number of research institutions and pharmaceutical companies are devoted to the research and development of new drugs. AMPs have been repeatedly shown to have utility as anticancer agents24,25 as a result of their ability to induce apoptosis and membrane lysis or pore formation. The strength of this anti-proliferative activity is related to their amino acids, charge, and structure. AMPs are able to target cancer cells effectively in comparison to nonmalignant cells due to the differences between healthy and cancer cell membranes, which include membrane electrical properties, and membrane flow velocity. 26 Some peptides can also hydrophobically interact with healthy cell membranes, resulting in toxicity.
In this study, we tested the anti-proliferative activity of brevinin-1EG on U-2OS, HepG2, HT-29, A375, HeLa, and A549 cells. We also tested LDH release in HepG2 and HeLa cells. Our results show that brevinin-1EG can kill a variety of cancer cells by attacking their membrane structure. Brevinin-1EG possesses a similar amino acid sequence and structure to brevinin-1RL1. Brevinin-1RL1 exerts anti-proliferative effects by inducing apoptosis and necrosis. 20 Therefore, we speculate that the anti-proliferative mechanism of brevinin-1EG is similar to that of brevinin-1RL1, though this, of course, requires confirmation.
Hemolytic activity is often an important indicator to measure the safety of peptides. 27 This study shows that the hemolytic activity of brevinin-1EG is concentration-dependent and has good safety at concentrations effective to kill cancer cells. This hemolytic activity can be further reduced by structural modifications. Studies have found that the hemolytic activity of brevinin-1 is related to its hydrophobicity, hydrophobic surface, and α-helix length. In addition, brevinin-1EG also has an antioxidant capacity, which can scavenge ABTS·+ and DPPH free radicals, with the former being greater than the latter. This may be affected by the absolute ethanol solution.
To summarize, we discovered a new member of the brevinin family in
Footnotes
Author Contribution
All authors conceived and designed the study. GW and WL contributed equally and should be considered as co-first authors, who performed the experiments, collected and analyzed the data, and wrote the manuscript. YT and ZC participated in the revision of the article and put forward important suggestions. YS wrote the manuscript and proofread the manuscript, and provided funding.
Data Availability
The original contributions presented in this study are included in the article. In addition, further inquiries may be addressed to the corresponding author.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Natural Science Foundation of China (grant number 31860607) and Yunnan Ten Thousand Talents Plan Young & Elite Talents Project and Major Science.
Ethical Approval
The Ethics committee of Kunming University of Science and Technology approved and supported the work.
Statement of Human and Animal Rights
We declare that the research strictly follows “Principles of Laboratory Animal Cares” and “National Experimental Animal Management Regulations.”
Informed Consent
No human trials were involved in this study.
