Abstract
Objective/Background
Natural products (NPs) derived from microorganisms are the basis of a plethora of clinically utilized medications, namely, antimicrobial remedies. Although these secondary metabolites have been extensively explored all over the planet, they remain understudied in the Middle East and North Africa (MENA) region.
Methods
A literature search was conducted to first find NPs that were isolated from environmental fungi and bacteria that inhabit the soils and seawater of the MENA region. Then, purified molecules with biological activity against pathogenic bacteria, biofilms, fungi, and parasites were described in terms of structure, function, and location. Moreover, the methods that could be used to ameliorate the discovery of novel NPs from this region were investigated.
Results
A multitude of antimicrobial molecules from various chemical classes were found to be derived from the environmental microbes of MENA. Although many were rediscovered, some represented novel structural scaffolds for novel families of antimicrobial agents. Additionally, the geographical distribution showed a high number of these NPs were unraveled in a restricted area leaving much of MENA untapped. Furthermore, as relatively traditional and low-efficiency methods were typically used in the discovery process, advanced high-throughput techniques were suggested to enhance this practice at the regional level.
Conclusion
MENA represents a fairly unexploited region where antimicrobial drug discovery could be performed comprehensively through the concomitant exploration of untouched geographical locations and advanced molecular techniques.
Introduction
Antimicrobial resistance (AMR) is a phenomenon where microorganisms progressively change and become non-responsive to drugs originally designed to kill them.1,2 The term AMR is interchangeably used to describe the resistance of bacteria to their antimicrobials. Nonetheless, it can be also applied to the decreased susceptibility that other microbes (viruses, fungi, and parasites) might display against their antimicrobial agents. 3 The different forms of AMR pose an issue all over the planet. For example, in the United States, more than 2.8 million people are infected by antibiotic-resistant bacteria yearly, of whom 35 000 individuals pass away. 4 On the other hand, in malaria-endemic countries, the spread of artemisinin-resistant strains is predicted to add 116 000 deaths to the annual count of more than 400 000.5‐7 Moreover, the global burden of AMR is suspected to increase, becoming the leading cause of death by 2050 with more than 10 million casualties being associated with antimicrobial-resistant infections each year.8,9 Furthermore, the overwhelming financial load of AMR is even more destructive with estimations that AMR-associated costs could surpass 1 trillion dollars per annum by 2050. 10
Treating infections caused by drug-resistant pathogens requires switching from one class of antimicrobial to the other. However, the emergence of multidrug-resistant (MDR), extensively drug-resistant, and pandrug-resistant (PDR) microorganisms with resistance to all classes of antimicrobials has rendered this strategy useless.11‐13 Alternatively, two main approaches arise, either the usage of combination therapy of different antimicrobials or the discovery of new classes of antimicrobial agents.14,15 The former is already showing significant inhibitions of the growth of pathogens in clinical settings.14,16,17 Nevertheless, long-term exposure to the antimicrobials used in these combinations can eventually induce resistance to these mixtures in vitro and in vivo.18‐22 Henceforth, the discovery journey must be the path to take, to ensure victory against PDR pathogens. Researchers can choose between multiple tactics to find a hit drug against superbugs: in silico screening/molecular docking, fragment and structure-based approaches, hybrid inhibitor design, drug repurposing, and exploring natural products (NPs). 23 Although the last appears the most primitive in concept, it has shown time after time to be essential in drug discovery. This could be seen by the fact that NPs, their derivatives, and their mimics represent around 50% of all approved drugs between 1981 and 2019. They also exemplify approximately 50% of all antimicrobial drugs discovered in that period. 24 Additionally, the return to the inspection of NPs for antimicrobial biological activities is further reinforced by the relative ineffectiveness of molecular and biochemical approaches. These techniques were unable to yield the expected increase in the number of newly discovered classes of anti-infective agents although they have been utilized in this type of research for several decades. 25
NPs are secondary metabolites (SMs) that originate from a variety of organisms such as plants, terrestrial and marine microorganisms, invertebrates, and less frequently vertebrates. 26 Those derived from microorganisms have shown great potential in antimicrobial drug discovery.27‐29 However, since research in the past has been limited to a narrow range of microbial NP producers, the chances of novel drug discovery have been relatively low.30, 31 Consequently, the search for untapped genera and species has been deemed worthy, especially in highly biodiverse areas.32‐34 The main path to finding these understudied microbes might be in targeting fairly unexploited regions where these untouched bugs might be thriving. The Middle East and North Africa (MENA) region is characterized by its immense biodiversity while being relatively untapped when it comes to drug discovery. Thus, in this review, we will be focusing on antimicrobial agents derived from microorganisms inhabiting the MENA region (Mauritania, Morocco, Algeria, Tunisia, Libya, Egypt, Sudan, Palestine, Jordan, Lebanon, Syria, Saudi Arabia, Yemen, Oman, United Arab Emirates, Qatar, Bahrain, Kuwait, Iraq, and Iran). We will also be describing their actions on various pathogens with indication of quantitative parameters when available. Moreover, we will be shedding light on their cytotoxic activity when applicable. Finally, we will be discussing the methods that can be used to enhance the opportunities of their discovery.
Pure Compounds
In the following section, we will be describing completely purified antimicrobial NPs isolated from fungi and bacteria found in the MENA region, residing in both terrestrial and marine niches.
Fungi
An unexpected number of chemically diverse SMs exhibiting antimicrobial activities emerged from marine fungi living in the MENA region, as shown in Table 1. Starting from the Red Sea, an inlet of the Indian Ocean, lying between Africa and Asia, where ditryptophenaline (Structure 1) was discovered at the coast of Ein El-Sukhna-Zafarana, Egypt. This compound was isolated from Aspergillus ochraceopetaliformis MN0-83316, a symbiote of the coral Galaxea fascicuralis and was previously isolated from Aspergillus flavus. 35 This diketopiperazine homodimer consists of two identical halves, each of which possesses 4 fused rings including an indoline function cis fused to a pyrrolidine ring and a fully substituted 2,5-dioxopiperazine ring with a benzyl at C5 and a methyl at N6. Ditryptophenaline demonstrated antimicrobial activity against Gram-positive (Bacillus subtilis subsp. spizizenii ATCC 6633) and Gram-negative (Escherichia coli ATCC 11775) bacteria, as well as against the fungus Candida parapsilosis ATCC 22019. The inhibition zones ranged from 8 to 9 mm when the compound concentration was set at 50 mmoles. 36
Compounds Derived from Fungi Inhabiting the MENA Region.

Abbreviation: MENA, Middle East and North Africa.
The Egyptian Red Sea is also the home to Halimeda opuntia, which was collected from the coast of Rass Mohamed, Egypt. This green alga shelters the fungus Aspergillus versicolor, which synthesizes several bioactive compounds. One instance is the previously isolated Siderin (Structure 3), 48 a coumarin derivative benzopyrone with 2 methoxy groups at C4 and C7, as well as a methyl at C5 that was formerly purified from the plant Toona ciliata. Another example is the novel tricyclic anthraquinone Isorhodoptilometrin-1-methyl ether (Structure 2), with a methoxy group at C1, hydroxy groups at C6, and C8, as well as a 2-hydroxypropyl attached at C3. Both of these compounds demonstrated antibacterial activity against Gram-positive pathogens B. subtilis, Bacillus cereus, and Staphylococcus aureus with zones of inhibition ranging between 11 and 15 mm. Thus, both these molecules were less potent than the control drug, Oxytetracycline, whose zones of inhibition against these pathogens ranged from 17 to 20 mm. Moreover, these compounds showed null to minimal cytotoxicity against the normal human cell line CFU-GM. 37
Another endophytic fungus of an alga was found off the coast of South Hurghada, Egypt. It was classified as Fusarium equiseti. This fungus inhabits the brown alga Padina pavonica and has shown the ability to secrete an abundance of previously discovered NPs. Examples comprise the nucleoside Cordycepin (3’-deoxyadenosine, Structure 5), originating from Cordyceps militari, 49 the macrolide 17-demethyl-2,11-dideoxy-rhizoxin (Structure 6), with a tetraoxatetracyclohenicos-14-ene-6,19-dione, which was first isolated from a Rhizopus species, 50 and w-Hydroxyemodin (Structure 4), a tricyclic anthraquinone with hydroxy groups at C1, C3, and C8, as well as a hydroxymethyl group at C6, known to be a constituent of Penicillium chrysogenum. 51 All of these compounds exhibited antifungal activity against the fungus Candida albicans (C. albicans). Moreover, they demonstrated antibacterial activity against S. aureus and Bacillus megaterium, while Cordycepin and w-Hydroxyemodin also inhibited the growth of B. subtilis. Besides, their potency against these bacterial pathogens was either inferior or equal to that of the control drug Oxytetracycline. 38
Red Sea marine fungi whose SMs proved to possess antimicrobial potential were also found in sponges. Hyrtios erectus found off the coast of the Saudi Arabian city of Yanbu houses Penicillium vinaceum. This fungus yielded a plethora of antimicrobial NPs such as the diketopiperazine derivative Brevianamide F (Structure 7), a hexahydropyrrolo[1,2-a]pyrazine-1,4-dione with an indol-3-ylmethyl substituent at position 3, which was previously isolated from Aspergillus fumigatus. 52 Another compound is Citreoisocoumarin (Structure 8), a coumarin derivative with hydroxy groups at C6 and C8, as well as 2-hydroxy-4-oxopentyl attached at C3, previously purified from a culture of Penicillium nalgiovense. 53 Both these molecules inhibited the growth of S. aureus ATCC 25923 with a 19-mm inhibition zone. The fungal culture also generated iso-α-cyclopiazonic acid (Structure 9); an ergoline derivative alkaloid previously obtained from A. flavus. 54 It showed the ability to inhibit the growth of E. coli ATCC 25922 with a 20-mm zone. Moreover, the researchers detected the known metabolite Terretrione A (Structure 10), previously found in a culture of Aspergillus terreus. 55 This compound possesses a 1,4-diazepane core structure demethylated at positions 1 and 4, with carbonyl groups at C2, C5, and C7 as well as benzyl at C6 and 2-methylpropyl at C3. Terretrione A, alongside Brevianamide F, inhibited the growth of C. albicans ATCC 14053 with a 25-27 mm inhibition zone. All the aforementioned NPs were less potent against these bacterial and fungal pathogens than their respective reference drugs: ampicillin, imipenem, and clotrimazole. 39
Similarly, other Terretriones were discovered from Penicillium sp. CYE-87 inhabiting the tunicate Didemnum sp. from the Suez Canal, Egypt. They included the previously isolated Terretrione C (Structure 11) 55 and the newly deciphered Terretrione D (Structure 12). They both resemble Terretrione A; however, C has a butan-2-yl at C3, while D lacks a methyl group at position 4 and has a propan-2-yl at C3. Both of these NPs exhibited antifungal activity against C. albicans ATCC 14053 with inhibition zones of 17-19 mm. Their minimal inhibitory concentration (MIC) was determined to be 32 μg/mL against the same strain. 40 Another Penicillium species inhabiting a tunicate of the Didemnum genus was isolated from the Egyptian Red Sea from a mangrove forest at Sharm El-Sheikh. This microorganism generated 2 novel fatty acid esters Penicilloitins A (Structure 13) and B (Structure 16). Additionally, it generated known alkaloids: the ergoline derivative α-cyclopiazonic acid (Structure 14), 54 and the indolamine tryptamine (Structure 15). All of these compounds inhibited the growth of E. coli ATCC 25922 with zones of 20-22 mm, while only Penicilloitin B inhibited S. aureus ATCC 25923 with a 19-mm zone. These SMs were less efficient in inhibiting the growth of these pathogens when compared to the reference drugs imipenem and ampicillin. 41
The fungi of the Mediterranean Sea also offer a rich source for the discovery of NPs. For example, Varicosporina ramulosa recovered from the coast of Abou-Keer, Alexandria, Egypt, generated 2 novel antimicrobial compounds. Dibutyl phthalate (Structure 17), a 1,2-diester of phthalic acid with 2 butan-1-ol, and, ergosta-5,7,22-trien-3β-ol, an ergosterol (Structure 18) related molecule. Both of these molecules inhibited the growth of E. coli, B. subtilis, and C. albicans, while only dibutyl phthalate inhibited the growth of Fusarium solani. 42 Moreover, multiple novel Trichorzianines were isolated from Trichoderma atroviride NF16 coming from the coast of Akhziv. These peptaibols are linear non-ribosomal peptides (NRPs) with an acetylated N-terminus, C-terminal amino alcohol, and a high proportion of α-aminoisobutyric acid (Aib). The structure of one of them TA 1938 is shown in Table 1 (Structure 19). These peptides demonstrated a good antimicrobial activity on a range of environmental bacteria with MICs ranging between 12.5 and 200 μg/mL, while they only showed modest antibacterial activity on Staphylococcus albus and B. subtilis with MICs ranging between MIC 50 and 200 μg/mL. 43
Additionally, fungi inhabiting sponges collected offshore Sdot-Yam showed a diversity of chemical compounds with antifungal abilities. First, Aspergillus insuetus OY-207 inhabiting a sponge belonging to the genus Psammocinia synthesized the novel meroterpenoid Insuetolide A (Structure 20), possessing a tricos-8-ene-7,14,18-trione central structure, which inhibited the growth of the fungus Neurospora crassa with a MIC value of 140 μM. This fungus also gave rise to two drimane sesquiterpenes: Strobilactone A (Structure 21) and its derivative (E,E)-6-(60,70-dihydroxy-20,40-octadienoyl)-strobilactone A, previously isolated from Aspergillus and Strobilurus species.56,57 They are characterized by a hexahydrobenzo-benzofuran-1-one and inhibited the growth of the fungus Neurospora crassa with MICs of 242 and 162 µM, respectively. 44 Second, Aspergillus tubingensis OY907 inhabiting the sponge Ircinia variabilis secreted the previously unexploited Tubingenoic anhydride A (Structure 23) and 2-carboxymethyl-3-hexylmaleic acid (Structure 22) anhydride, which was previously identified in cultures of Aspergillus FH-X-21. 58 Both these NPs inhibited the growth of Neurospora crassa with MICs of 330 and 207 μM, respectively. 45
Likewise, soil fungi demonstrated the ability to manufacture antimicrobial NPs, as seen for compounds originating from Daldinia cf concentrica, an endophyte of the olive tree Olea europaea, from Ha'Ela Valley, Judean Hills. These molecules include the antifungal aldehyde trans-2-octenal (Structure 24), previously isolated from O. europaea itself, 59 which inhibited the growth of the fungi Aspergillus niger, Botrytis cinerea, Alternaria alternata, and Penicillium digitatum. Additionally, this fungus secreted the antifungal ketone 4-heptanone (Structure 25), previously purified from Burkholderia ambifaria, 60 which inhibited the growth of B. cinerea. 46 Another example is noticed for the novel Preussilides A (Structure 26) and C (Structure 27), isolated from Preussia similis DSM 104666 inhabiting the medicinal plant Globularia alypum from Batna, Algeria. Preussilides are characterized by having a naphthalen-1-yl-4,6-dimethylhepta-24,6-trienoic acid core structure. These novel antifungal bicyclic polyketides showed activity against Mucor plumbeus (MIC 150 and 37.5 μg/mL) less efficiently than the control drug Nystatin (MIC 3.12 μg/mL). Preussilide C was also active against A. fumigatus DSM 819 (MIC 8.33 μg/mL), which was more effective than the control drug Cycloheximide (MIC 33.33 μg/mL) and Preussilide A (MIC 66.67 μg/mL). Moreover, both preussilides inhibited Sclerotinia sclerotiorum DSM 1946 with inhibition zones of 28 and 29 mm at 100 μg/paper disk. Both these compounds also exhibited moderate cytotoxicity against the normal mouse fibroblastic cell line L929, with IC50 values lower than 10 μM. 47
Bacteria
The profile of antimicrobial NPs purified from environmental bacteria was extensively diverse in respect to variation in both structure and function. The producing bacteria are spread out across the whole MENA region and inhabit both soil and water, as shown in Tables 2 and 3, respectively.
Compounds Derived from Soil bacteria Inhabiting the MENA Region.

Abbreviation: MENA, Middle East and North Africa.
Compounds Derived from Marine bacteria Inhabiting the MENA Region.
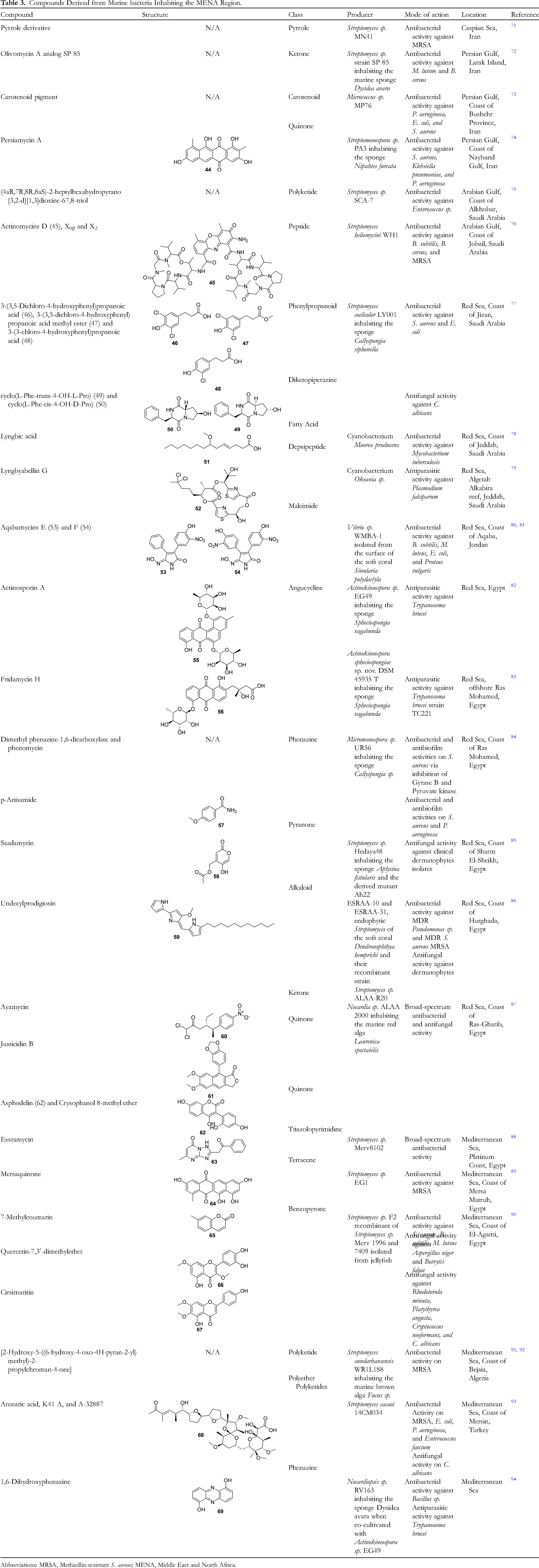
Abbreviations: MRSA, Methicillin-resistant S. aureus; MENA, Middle East and North Africa.
Terrestrial Bacteria
When it comes to soil prokaryotes, Tunisia has shown its potency as an invaluable source of antimicrobial SMs. A first example is Streptomyces sp. TN82, isolated from the Tunisian Sahara, which synthesizes a variety of NPs. Two were of interest: 3-Phenylpyrazin-2(1H)-one, a pyrazine which has never before been reported to occur naturally, and 3-O-methylviridicatin, a quinolin-2-one with a phenyl at position 4 and a methoxy group at position 3, previously isolated from a Penicillium sp. 95 3-Phenylpyrazin-2(1H)-one (Structure 28) and 3-O-methylviridicatin (Structure 29) both exhibited antibacterial activity against S. aureus ATCC 6538 (MIC of 4 and 4.5 μg/mL), Listeria monocytogenes ATCC 19117 (MIC of 1 and 9 μg/mL), and Salmonella typhimurium ATCC 14028 (MIC of 2 and 4.5 μg/mL). 3-Phenylpyrazin-2(1H)-one was more efficient than ampicillin against L. monocytogenes and S. typhimurium, but not against S. aureus while 3-O-methylviridicatin was less potent than the reference drug against all 3 pathogens. 61
A second instance is that of Streptomyces sp. TN256, another inhabitant of the Tunisian Sahara that secretes an assortment of antimicrobial NPs. This collection of metabolites has all been previously described and comprises: the alkene 1-nonadecene (Structure 32), 96 previously isolated from the plant Rosa damascena, the alkaloid derivative N-[2-(1H-indol-3-yl)-2 oxo-ethyl] acetamide (Structure 30), purified from cultures of Streptomyces anulatus, 97 the phthalate derivative di-(2-ethylhexyl) phthalate (Structure 31) obtained formerly from the plant Aloe vera Linne, 98 and the diketopiperazine Cyclo (L-Pro-L-Tyr) (Structure 33), a hexahydropyrrolo[1,2-a] pyrazine-1,4-dione with a hydroxylated benzyl, which was shown to be produced earlier by the actinomycete strain A8. 99 All of these four compounds inhibited the growth of Gram-positive (Micrococcus luteus LB 14110 and S. aureus ATCC 6538) and Gram-negative (E. coli ATCC 8739 and Salmonella enterica ATCC 43972) bacteria, as well as the fungus Fusarium sp., with zones of inhibition ranging between 10 and 21 mm in diameter. 62 Another Streptomyces found in the Tunisian Sahara at an oasis is Streptomyces misionensis V16R3Y1. This microorganism produced the diketopiperazine cyclo (L-Leu-L-Pro) (Structure 34), a hexahydropyrrolo[1,2-a]pyrazine-1,4-dione with 2-methyl propyl at position 3, which has been previously isolated from a Bacillus strain. 100 This compound demonstrated broad-spectrum yet moderate antibacterial and antifungal activity. 63 Another bacterial strain was isolated from a soil sample in south Tunisia and was deciphered to be Streptomyces sp. TN262. This strain generated two known macrolides, Cineromycin B (Structure 35) and its derivative 2,3-Dihydrocineromycin B (Structure 36), previously isolated from a Streptomyces species, 101 which are branded with a tetramethyl-1-oxacyclotetradeca-36,9-trien-2-one with two hydroxy groups at positions 5 and 8. They both inhibited the growth of M. luteus LB 14110 and S. aureus ATCC 6538 with 12 to 13 mm inhibition zones. Additionally, Streptomyces sp. TN262 produced Tryptophol (Structure 37), a known aromatic alcohol, 102 that showed antibacterial activity against S. enterica ATCC 43972 with a 12-mm zone. Moreover, it inhibited the previously mentioned Gram-positive pathogens, but with more potency than Cineromycins. Besides, Tryptophol inhibited the growth of Fusarium sp. with a 12-mm zone. 64
Another country in the Maghreb region where soil bacteria have provided a promising source of antimicrobial NPs is Algeria. One example is Streptomyces sp. HG29, which was isolated from Algerian Saharan soil. This strain synthesized two known macrolides Oligomycins A (Structure 38), 103 and E (Structure 39), 104 which are typically produced by other Streptomyces species. These compounds inhibited the growth of several species of Aspergillus, Fusarium, and Penicillium with MICs below 30 mg/mL and were comparable in potency to the utilized control drugs Amphotericin B and Itraconazole. 65 Another case is the bacterium Nocardiopsis sp. HR-4 isolated from salt-lake soil in the Algerian desert. This strain secreted two angucyclinones: the known ( − )-8-O-methyltetrangomycin (Structure 40) 105 and the novel ( − )-7-deoxy-8-O-methyltetrangomycin (Structure 41), which are characterized by a 2,4-dihydrobenzo[a]anthracene-1,7,12-trione core structure. Both of these compounds showed antibacterial activity against Gram-positive bacteria S. aureus ATCC 25923 and 43300, Enterococcus faecalis ATCC 29212, and M. luteus ATCC 4698, with zones of inhibition having diameters between 23 and 41 mm. 66
Moving east in the African continent, specifically to Egypt, a Streptomyces species labeled MAR01 was isolated from a soil sample from Burg-Arab, Alexandria. This strain produced a novel antibiotic named Meroparamycin (Structure 42), comprising a benzene ring connected to an amido group at C1 and a branched aliphatic ketone at C4. This compound showed antibacterial activity against B. subtilis ATCC 6633, S. aureus ATCC 29737, Staphylococcus epidermidis ATCC 12228, and M. luteus ATCC 9341 with MICs ranging from 3.13 to 12.5 mg/mL. It also inhibited the growth of the Gram-negative bacterium E. coli ATCC 10536 and the fungus C. albicans ATCC 10231 with MICs of 50 and 25 mg/mL, respectively. 67 In Saudi Arabia, a different Streptomyces sp., called MS-6–6, was found in a soil sample originating from the western region of the country. This strain was capable of producing a known macrolide antibiotic treponemycin, characterized by a macrocyclic lactone ring with a nitrile group and a diene function, previously isolated from Daldinia cf concentrica. 106 Treponemcyin (Structure 43) proved to be a broad-spectrum antibiotic inhibiting various bacteria with relatively low MIC values: less than 2.9 μg/mL for Gram-positive pathogens and less than 26.7 μg/mL for Gram-negative bacteria. However, this antibacterial agent attracted research for its anti-tuberculous activity as it inhibited the growth of Mycobacterium tuberculosis ATCC 25177, with a MIC of 4.17 μg/mL. Additionally, this molecule demonstrated antifungal capacity against C. albicans with a MIC of 13.3 μg/mL, and, although the cytotoxicity of the pure compound was not assessed, the crude extract of strain MS-6-6 was found to be non-toxic toward the normal human fetal lung fibroblast cell line (MRC-5), thus projecting the low toxicity of Treponemcyin. 68
Heading north of the Arabian Peninsula, to Iraq, the microbial population of rhizospheric soil of wheat plants from the capital city of Baghdad was studied. One of the isolated strains, Bacillus licheniformis BL1, synthesized a novel bacitracin derivative peptide antibiotic. This compound showed broad-spectrum antibacterial activity against MDR-UTI (urinary tract infection) causing clinical isolates including Klebsiella pneumoniae, Acinetobacter baumannii, Pseudomonas aeruginosa, and E. coli. 69 On the other hand, another terrestrial bacterium that led to the discovery of a pure novel antibacterial agent is Bacillus thuringiensis subsp. kurstaki Bn1 (Bt-Bn1) isolated from the insect Balaninus nucum, a common pest of the hazelnut tree in the Black Sea region of Turkey. The new compound was named Thuricin Bn1 and was characterized as being a Bacteriocin that inhibited the development of various plant phytopathogenic bacteria and spoilage microorganisms. 70
Marine Bacteria
The bacteria inhabiting the seas of the MENA region have shown a noteworthy capacity to produce functionally varied NPs. On the northern coast of Iran, from the Caspian Sea, the strain Streptomyces sp. MN41 was isolated. This bacterium synthesizes a previously uncharacterized pyrrole derivative, which exhibits bactericidal effects against Methicillin-resistant S. aureus (MRSA) ATCC 33591 with a MIC of 2.8 μg/mL and an MBC (minimal bactericidal concentration) of 5.62 μg/mL. 71 However, it is Iran's southern coast outlining the Persian Gulf that is the source of marine bacteria with powerful antimicrobial SMs. From Larak island, a Streptomyces sp. strain SP 85, inhabiting the marine sponge Dysidea avara, was found to secrete a novel olivomycin A derivative known as SP 85. This antibacterial molecule possesses a tricyclic chromophore/aglycon connected to a disaccharide at C6 and a trisaccharide at C2. It demonstrated inhibition against M. luteus and B. cereus, while showing low levels of cytotoxicity against the normal cell human umbilical vein endothelial cell line HUVECs. 72 From the shore of Bushehr Province, Micrococcus sp. MP76 was found. This strain produced a carotenoid pigment that inhibited the growth of S. aureus ATCC 25923, E. coli ATCC 25922, and P. aeruginosa ATCC 27853 with IC50 values of 3.4, 4.8, and 4.8 mg/mL, respectively. 73 Streptomonospora sp. PA3 inhabited a sponge that was recovered from the coast of Nayband. A novel aromatic polyketide was isolated from this bacterium and denoted as Persiamycin A (Structure 44). It is a tetracyclic anthraquinone with hydroxyl groups at C1, C3, C8, and C11, which inhibits the growth of S. aureus ATCC1337, K. pneumoniae ATCC1290, and P. aeruginosa ATCC1310, with inhibition zones of around 9 mm. 74
It has been shown progressively that the Saudi Arabian coast of the Arabian Gulf (Persian Gulf) shelters bacteria with antimicrobial metabolites. One example is that of Streptomyces sp. SCA-7 from the coast of the city of Alkhobar. This strain synthesized the novel polyketide (4aR,7R,8R,8aS)-2-heptylhexahydropyrano[3,2-d][1,3]dioxine-67,8-triol, which showed antibacterial activity against Enterococcus sp. comparable to that of streptomycin. 75 Another case is that of Streptomyces heliomycini WH1 from the coast of the city of Jobail, which produces multiple well-known actinomycins, namely D (Structure 45), X0β, and X2.107‐109 These compounds possess bicyclic chromopeptide lactones sharing the chromophoric phenoxazinone dicarboxylic acid to which are attached two pentapeptide lactones of nonribosomal origin and are known to be synthesized by soil-inhabiting microorganisms. Actinomycin X2 was the most potent among the three isolated compounds with MICs lower than 0.15 μM against Gram-positive pathogens B. subtilis ATCC 6051, B. cereus ATCC 14579, and S. aureus ATCC 6538 and ATCC 43300 (MRSA), even surpassing the antimicrobial efficacy of Ciprofloxacin. All of these compounds isolated from WH1 also demonstrated low levels of cytotoxicity on normal human embryo liver cells L02. 76
Nevertheless, the highest number of pure compounds have originated from bacteria that live in the Red Sea. A first instance is Streptomyces coelicolor LY001 inhabiting the sponge Callyspongia siphonella collected off the coast of Jizan, Saudi Arabia. This strain was able to synthesize three new natural 3-phenylpropanoic acid derivatives: 3-(3,5-dichloro-4-hydroxyphenyl) propanoic acid (Structure 46), 3-(3,5-dichloro-4-hydroxyphenyl) propanoic acid methyl ester (Structure 47), and 3-(3-chloro-4-hydroxyphenyl) propanoic acid (Structure 48). These phenylpropanoids all demonstrated antibacterial activity against S. aureus ATCC 25923 (MICs between 32 and 64 μg/mL) and E. coli ATCC 25922 (MICs between 16 and 32 μg/mL), yet they were all less potent than Ciprofloxacin. Additionally, S. coelicolor LY001synthesizes the known cyclo(L-Phe-trans-4-OH-L-Pro) (Structure 49), 110 and cyclo(L-Phe-cis-4-OH-D-Pro) (Structure 50), 111 which were previously isolated from bacterial endophytes of spider mites and sponges, respectively. These compounds possess a hexahydropyrrolo[1,2-a]pyrazine-1,4-dione with a benzyl at position 3 and a hydroxy group at position 7. These diketopiperazines inhibited the growth of C. albicans ATCC 14053 with MICs of 32 μg/mL, thus less efficiently than Ketoconazole. 77 The Saudi Red Sea also houses cyanobacteria with antimicrobial metabolites such as Moorea producens isolated from the coast of Jeddah which secretes lyngbic acid (Structure 51). This compound is also known as (S)-7-methoxytetradec-4(E)-enoic acid and was previously isolated from the same bacterial species. 112 It showed antibacterial activity against M. tuberculosis H37Rv ATCC 27294 inhibiting 65% of bacterial cells at a concentration of 12.5 μg/mL. 78 Additionally, an Okeania species isolated from the coast of the same city at the level of Algetah Alkabira reef produced Lyngbyabellin G (Structure 52). This depsipeptide was isolated from the same species in the past 113 and was characterized by having an esterized cycle containing 2 thiazole groups with a 4,4-dichloropentyl attached at position 12. Besides, it effectively inhibited the growth of the parasite Plasmodium falciparum strain FCR-3 with an IC50 of 1.1 μM. 79
From the Red Sea in south Jordan near Aqaba, the Vibrio sp. WMBA-1 was isolated from the surface of the soft coral Sinularia polydactyla. Although this strain synthesized seven novel nitro maleimides, 2 were of main interest as anti-infective agents: Aqabamycins E (Structure 53) and F (Structure 54). The latter has a hydroxylated nitro maleimide connected to two identical nitro-hydroxy-phenyl groups, one at C3 and the other at C4. The former has a similar structure except that the group at C4 is a non-substituted phenyl. 80 Both these compounds present antibacterial activity against B. subtilis, M. luteus, E. coli, and Proteus vulgaris with MICs below 12.5 μg/mL for Aqabamycin E and below 25 μg/mL for Aqabamycin F. 81
Nonetheless, the discovery of bioactive material from the Red Sea has been predominantly led by Egyptian researchers. Numerous pure antimicrobial NPs have been found from the Egyptian coast of the Red Sea. Actinosporin A (Structure 55), a novel O-glycosylated angucycline was isolated from Actinokineospora sp. EG49 inhabiting the sponge Spheciospongia vagabunda. This compound is a tetrahydroxylated tetracyclic anthraquinone to which is attached a trihydroxy-methyl-oxanyl at C4 through an O bond. 82 The same sponge species was recovered offshore Ras Mohamed, Egypt, and was found to contain Actinokineospora spheciospongiae sp. nov. DSM 45935 T. This bacterium was shown to produce the novel angucycline Fridamycin H (Structure 56), a similar compound to Actinosporin A with a monohydroxylated anthraquinone with a trihydroxy-methyl-oxanyl attached at C4 through an O bond and a hydroxy-methyl-butyric acid attached at C8. 83 Both Actinosporin A and Fridaymycin H exhibited antiparasitic activity against Trypanosoma brucei strain TC221 after 48 h with IC50 values of 15.47 and 7.18 µΜ, respectively. Neither compound showed cytotoxicity when tested against J774.1 macrophages.82,83 From the coast of Ras Mohamed, Micromonospora sp. UR56 inhabiting a sponge of the Callyspongia genus was isolated. This strain secretes multiple compounds with three phenazines proving their importance as antimicrobial agents. First, there are dimethyl phenazine-1,6-dicarboxylate and phenazine-1,6-dicarboxylic acid monomethyl ester (phenomycin), which were previously isolated from Streptomyces fervens var. phenomyceticus and Pseudomonas phenazinium, respectively.114,115 Both of these compounds demonstrated an ability to inhibit the growth and biofilm formation of S. aureus ATCC 9144. 84 Second, there is the known p-anisamide (Structure 57), also known as 4-methoxybenzamide. 116 This molecule proved to be efficient in inhibiting the growth and biofilm formation not only of S. aureus ATCC 9144, but also of P. aeruginosa ATCC27853. However, as antibacterials, these molecules were less efficient in inhibiting the growth of bacterial pathogens when compared to Gentamicin. 84
Other bacteria have also emerged from the Egyptian coast of the Red Sea. The coast of Sharm El-Sheikh is the place of origin of Streptomyces sp. Hedaya 48 inhabiting the sponge Aplysina fistularis. This strain produced the novel antimycotic agent Saadamycin {4-(hydroxymethyl)-5-hydroxy-2H-pyran-2-one} (Structure 58). This compound inhibited the growth of clinical dermatophyte isolates including isolates from the genera Aspergillus and Candida with MIC values below 5 μg/mL and MFC values below 10 μg/mL. This compound was proven to be more effective than the clinically utilized antifungal. Myconazole, whose MIC and MFC (minimal Fungicidal Concentration) values were in the mg/mL range. 85 Other Red Sea Streptomyces strains were isolated from their host, the soft coral Dendronephthya hemprichi, at the coast of Hurghada, Egypt, and were called ESRAA-10 and ESRAA-31. These bacteria synthesized Undecylprodigiosin (Structure 59), whose production was amplified in recombinant strain ALAA-R20 after successful gene transfer was performed on it. Undecylprodigiosin is a known alkaloid previously isolated from Streptomyces lividans, 117 characterized by 3 pyrrole rings connected to an undecane saturated chain. It demonstrated antibacterial activity against MDR S. aureus and Pseudomonas sp., as well as MDR dermatophytes with MICs ranging between 0.5 and 4.0 μg/mL. Moreover, Undecylprodigiosin showcased relatively low MBC and MFC values against these MDR pathogens (< 8 μg/mL), revealing its potential cidal action against the said microbes. 86 However, Red Sea bacteria do not only inhabit corals and sponges, but they also live as endophytes/epiphytes of algae such as in the case of Nocardia sp. ALAA 2000 inhabiting the marine red alga Laurenica spectabilis, which was collected off the coast of Ras-Gharib, Egypt. This strain can be considered as a factory for antimicrobial NPs synthesizing several powerful SMs. For example, this bacterium produced two peri-hydroxy-anthraquinones, Asphodelin (4,7’-bichrysophanol, Structure 62), 118 and Crysophanol 8-methyl ether, 119 which were previously isolated from the plants Asphodelus microcarpus and Eremurus chinensis, respectively. Also, it produced the arylnaphthalene lignanolide justicidin B (Structure 61), which was previously isolated from a Hoplophyllum (Rutaceae) species. 120 Besides, it gave birth to the novel compound 1,1-Dichloro-4-ethyl-5-(4-nitro-phenyl)-hexan-2-one, which was called Ayamycin by the authors. All four of these NPs demonstrated potent antimicrobial capacities on multiple representatives of Gram-positive and Gram-negative bacteria, as well as a wide range of fungi. Ayamycin (Structure 60) proved to be the most significant inhibitor of microbial growth, with MIC values below 0.5 μg/mL against all of the tested pathogens. 87
Egypt has another coastal line on its northern border. This Mediterranean Egyptian coast is the homeland of multiple bacteria with antimicrobial potential. Streptomyces sp. Merv8102 isolated off the Platinum coast in Egypt is a good example. It synthesized the first triazolopyrimidine antibiotic isolated from nature, named Essramycin (Structure 63). This compound inhibited the growth of Gram-positive bacteria with MIC values around 1 μg/mL, while it inhibited that of E. coli ATCC 10536 and P. aeruginosa ATCC 10145 with MIC values of 8 and 3.5 μg/mL, respectively, 88 although another research team debunked these results. 121 Another sample of putative microbes from the Egyptian Mediterranean coast is Streptomyces sp. EG1, isolated from the port city of Mersa Matruh. This strain is the producer of a novel tetracene derivative Mersaquinone (Structure 64), a tetracyclic anthraquinone with hydroxyl groups at C3, C8, C10, and C11, as well as a methyl group at C13. Mersaquinone meaningfully inhibited the growth of the MRSA strain TCH1516 with a MIC value of 3.36 μg/mL; nonetheless, this compound was less efficient than the reference drug Ciprofloxacin, whose MIC against MRSA was 0.93 µM. 89 Another impressive microbial factory whose origin is from the Egyptian Mediterranean coast was Streptomyces sp. F2. Although this recombinant strain was created in the laboratory, its parent strains, Streptomyces sp. Merv 1996 and 7409, were isolated from a jellyfish off the coast of El-Agami. Streptomyces sp. F2 generated 3 benzopyrone derivatives, which are known to be plant metabolites, and whose antimicrobial activities were investigated. The first was the heterocyclic compound 7-methylcouramin (Structure 65), which inhibited the growth of Gram-positive bacteria S. aureus ATCC 6538, B. subtilis ATCC 6051, and M. luteus ATCC 9341, with MIC values of 0.8, 3.6, and 2 μg/mL, respectively. The second was Quercetin-7,3′-dimethyl ether (Structure 66), also known as 2-(3,4-dihydroxyphenyl)-5-hydroxy-3,7-dimethoxychromen-4-one, which inhibited the growth of filamentous fungi A. niger and Botrytis fabae with MICs of 1 and 2.5 μg/mL, respectively. The third was Cirsimaritin (Structure 67) {5-hydroxy-2-(4-hydroxyphenyl)-6,7-dimethoxychromen-4-one}, which suppressed the growth of Rhodotorula minuta, Platythyrea angusta, Cryptococcus neoformans, and C. albicans with a MIC value of 1 μg/mL. 90
However, it is not only the Egyptian coast of the Mediterranean that offers bacteria with an extensive arsenal of antimicrobial NPs. Other countries’ coasts on the Mediterranean Sea basin also house such organisms. Algeria is a prime illustration where the strain Streptomyces sundarbansensis WR1L1S8, an endophyte of a marine brown alga of the Fucus genus was isolated off the coast of Bejaia. Although this strain made a plethora of antimicrobial agents, one particularly stood out as a novel anti-MRSA polyketide compound [2-hydroxy-5-((6-hydroxy-4-oxo-4H-pyran-2-yl)methyl)-2-propylchroman-4-one].91,92 This molecule interestingly inhibited the growth of MRSA ATCC 43300 without affecting that of the Methicillin-sensitive S. aureus (MSSA) ATCC 25923. Regardless, this molecule was less effective against MRSA than both Vancomycin and Gentamicin. 91 Another case is seen in Turkey where Streptomyces cacaoi 14CM034 was isolated off the coast of Mersin. This bacterium engendered 3 polyether compounds: two previously isolated from bacteria named Arenaric acid (Structure 68), 122 and K41 A, 123 while the third A-32887, although known synthetically, was never before discovered from a natural source. All of these polyketides exhibited antimicrobial activities against the Vancomycin-Resistant Enterococcus faecium DSM13590 (VRE), E. coli O157:H7, MRSA S. aureus DSM11729, P. aeruginosa ATCC27853, and C. albicans DSM5817. K41 A was the most effective against the Gram-positive pathogens (MICs range between 0.11 and 1.79 μg/mL), while A-32887 was the best anti-yeast compound. 93 Moreover, a Mediterranean Nocardiopsis sp. RV163, a symbiote of the sponge Dysidea avara, was able to produce 1,6-dihydroxyphenazine (Structure 69), when co-cultivated with the Red Sea inhabiting Actinokineospora sp. EG49. This molecule showcased an antibacterial activity against Bacillus sp. P25 (11 mm inhibition zone) and an antiparasitic activity against T. brucei (IC50 value of 19 μM). 94
Geological Distribution of Antimicrobial NPs in the MENA Region
In a first effort to improve the chances of discovering new antimicrobial SMs, we thought that excavating for environmental microbes in previously unexploited locations was necessary. Therefore, the research output of various groups across the MENA region, whether they resulted in obtaining pure compounds or not, were visualized on a map, as seen in Figure 1. It was then evident that the Red Sea represented an abundantly active zone when it comes to NP discovery, especially at the level of its west (Egyptian) coast, as seen in Figure 2. However, the rest of the region was sparsely examined, with entire countries left untouched. Nonetheless, these countries represent incomparable opportunities for NP discovery due to their elevated levels of biodiversity and their unique variability of niches. Examples include Mauritania, a desert-filled African country with a coast on the Atlantic. Another instance encompasses the heart of the Levant, specifically Lebanon and Syria, which possess coasts on the Mediterranean while being rich in mountain ranges as well as green forests. Hence, these countries, among other understudied areas, should be considered for the mining of new microbial genera, species, or strains to harvest novel classes of antimicrobial metabolites.

A map of the distribution of extracts, fractions, and compounds derived from environmental microbes inhabiting the MENA region. Abbreviation: MENA, Middle East and North Africa.

A map of the distribution of extracts, fractions, and compounds derived from environmental microbes inhabiting the MENA region with a focus on those living at the level of the red sea basin and coasts. Abbreviation: MENA, Middle East and North Africa.
Methods to Improve the Discovery of Antimicrobial NPs in the MENA Region
In a further effort to enhance the discovery of antimicrobial NPs from microorganisms inhabiting the MENA region, several modern approaches could be utilized. The three main methodologies are described below, highlighting for each the benefits as well as the disadvantages.
Metagenomics
Metagenomics is a culture-independent genomic analysis of microbial communities, which relies on the direct extraction of DNA from environmental samples. This environmental DNA (eDNA) can be subsequently sequenced and analyzed, enabling researchers to understand the microbial composition of the studied sample. Additionally, this metagenome reflects the genomically encoded capacities of the present microbes. Moreover, metagenomic libraries could be created by shotgun cloning of random parts of the eDNA into sufficient vectors, which can then be inserted into a suitable bacterial host. The metabolites of the generated clones are consequently screened for their antimicrobial biological activities. 124 Direct genomic cloning offers an opportunity to capture operons or genes that encode pathways that may lead to the synthesis of complex molecules, such as antibiotics. 125 The biosynthetic pathways of these novel bioactive metabolites can then be further dissected by interpreting metagenomic sequencing data. 124
The usage of such metagenomic procedures has revolutionized the drug discovery procedure in biological sciences. They significantly enhanced the development of numerous antimicrobial NPs originating from various environmental niches. 126 This improvement in discovery is the result not only of the functional screening of clones but also through information generated on microbial metabolism, which inspires new antimicrobials synthesis via heterologous expression. 127
Both novel and previously isolated antibiotics have been detected by functional screens from metagenomic libraries.128‐131 Among the previously unexploited compounds, turbomycin was one of the first antibiotics discovered by metagenomics. It was identified by accident in a clone that had hemolytic activity as part of a basic study directed toward rationalizing the prevalence of hemolysins among cultured soil bacteria. 128 Another example is that of violacein, a broad-spectrum antibiotic initially identified in a metagenomic clone of Chromobacterium violaceum. This antibacterial agent showed activity against S. aureus, Bacillus sp., and Streptococcus sp. 130
This bioinformatics tool offers an upper edge by saving time and reducing reliance on microbial culture in laboratories, making it possible to explore bioactive compounds from organisms that are resistant to cultivation. 132 Although the benefits of metagenomics research are evident, this approach also has notable limitations. It includes low resolution, bias classification of short target segments, and false functional confirmation. 124 Besides, metagenomics is expensive, labor-intensive, and requires sterling skills for wet-lab. 133
High-Throughput Screening
High-Throughput Screening (HTS) is a drug discovery technique that has gained widespread popularity over the past two decades, becoming a standard tool in the pharmaceutical industry. It is a method of screening and assaying a large number of biological modulators and effectors against selected and specific targets. 134 Due to the need to process thousands of assays per day, HTS has revolved around the combined field of multiple-well microplates and robotic processing. 135 HTS utilizes a variety of biological assays to find lead compounds. These include microbiological techniques like broth microdilution or biofilm formation assay which facilitate the discovery of new antibacterial, antifungal and antibiofilm agents. 136
Itoh et al developed a high-throughput strategy which led to the discovery of potent analogs of the antibiotic lysocin E. These compounds were demonstrated to be superior agents for the treatment of infectious diseases caused by Gram-positive bacteria, including MRSA. 137 On the other hand, Maraviroc was identified as the most promising hit against the Human Immunodeficiency Virus (HIV-1) after HTS of the Pfizer file collection. 138
The acts of miniaturization and automation of HTS help to reduce the use of reagents, minimize or remove labor-intensive steps, and greatly reduce the cost of research. 139 Nevertheless, the primary advantage of the HTS method remains in the speeding up of the discovery process. 140 Furthermore, this method plays an important role in the early stages of drug development, providing a qualitative and quantitative characterization of compound libraries and analytical support for preclinical and clinical absorption, distribution, metabolism, excretion (ADME) studies. HTS, therefore, facilitates the early elimination of unsuitable compounds. Yet, as HTS can simultaneously administrate multiple drugs together, a false apparent activity of a certain compound could be observed during screening, might it be an enhanced or a discouraged one. 141
Molecular Modeling
Drug discovery has developed from the classical screening of NPs to more sophisticated modern approaches, namely molecular modeling. 142 This method is a useful tool in many research applications, such as structure elucidation. It relies on theoretical and analytical approaches used to model the structure, properties, and dynamic behavior of molecules at the atomic level. 143 This branch of science applies the basic laws of physics and chemistry to the study of molecules. The ultimate goal is to create models and simulations that can assist in the various stages of a pipeline of discovery by predicting, rationalizing, and estimating the properties and interactions of molecules, thus enabling a more rational approach to the production of drugs. 144
How is molecular modeling relevant for NP research? Well, although several NPs currently in use have been discovered, the characterization of various drug targets has only been achieved recently. This flow of information contributed immensely to understanding the structure–activity relationships of NPs, as well as their relations to their targets. 142
Wang et al discovered compound ZINC05683641 as a newly available inhibitor against the New Delhi metallo-beta-lactamase 1 (NDM-1), an enzyme allowing bacteria to resist beta-lactam antibiotics, through molecular modeling. The binding mode and interaction mechanism of ZINC05683641 against NDM-1 were explored at atomic level. 145 Another study used molecular modeling with HIV-1 as a target molecule. Structures similar to known integrase inhibitors were identified which were capable of preventing viral replication. 146
Molecular modeling methods give significant advantages of saving time and expense by enabling thorough characterization and analysis of the expected drug-binding modes and the strength of protein-ligand interactions without the biological risks involved. 143 Molecular modeling is intended to play an ever more central role in the integration of information on the genetic, structural, biological, or chemical origin, increasing the responsibility for delivering effective medicines and making possible the eagerly awaited increase in the productivity of the pharmaceutical industry. 147 However, regardless that these methods are well established, the accuracy of the force fields used, simulation times, and scoring functions remain a limitation. 143
Conclusion
Taken together, these data showcase the presence of previously uncultured soil and marine dwelling microorganisms of the MENA region capable of synthesizing a plethora of SMs with antimicrobial activities. However, these microbes only represent a small fraction of the vast pool of novel potential NP manufacturers that inhabit untapped locations within MENA. Therefore, further research that explores unexploited niches and their microbial residents for the discovery of novel antimicrobials is needed and can be improved through the utilization of advanced molecular approaches.
Footnotes
Acknowledgements
The authors thank the graphic designer Asmaa Nabil Al-Ghrawi for her assistance with the figures.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The authors would like to acknowledge the National Council for Scientific Research of Lebanon (CNRS-L) for granting a doctoral fellowship to Bassel Awada and Dany Abi Chahine. The fellowships were offered in collaboration with the American University of Beirut (AUB) and Université Saint-Joseph de Beyrouth (USJ) for Bassel Awada and Dani Aby Chahine, respectively.
