Abstract
Objective
Spinal cord injury (SCI) is a debilitating disease that cannot be cured at present. Kaempferol (KPL) has proven neuroprotective, antioxidant, and antibacterial properties. However, the neuroprotective effect of KPL on SCI and its potential mechanism are still unclear.
Methods
Network pharmacology and molecular docking were used to determine the potential mechanisms of KPL on SCI. In vitro studies were used to validate the results. Hydrogen peroxide (H2O2)-exposed PC12 cells were pretreated with or without KPL. The safe concentrations of KPL, H2O2, and MK2206 in PC12 cells were determined via cell counting kit-8 assays. Then, terminal deoxynucleotidyl transferase dUTP nick end labeling staining and Annexin V-FITC/PI double staining were used to detect apoptotic cells. In addition, Western blotting was utilized to measure apoptosis-related gene and protein expression levels.
Results
Network pharmacology and molecular docking data demonstrated an interaction between KPL and Akt1 and identified apoptosis as a potential target of the KPL/Akt1 complex. The data verified that KPL could inhibit mitochondrial apoptosis in vivo by inducing Akt1 phosphorylation. Subsequently, MK2206, a pharmacological Akt1 inhibitor, was used to assess the role of Akt1 in this process, and we found that MK2206 attenuated the antiapoptotic effect of KPL on PC12 cells.
Conclusion
This study demonstrates that KPL could inhibit mitochondrial apoptosis by inducing the phosphorylation of Akt1, suggesting that KPL is a very promising candidate for SCI treatment.
Introduction
Spinal cord injury (SCI) is a nerve injury resulting in multiple functional deficits due to the destruction of spinal cord neurons and axons. There is currently no cure for SCI. 1 Data from the National SCI Statistical Center showed that the number of new SCI cases increased from 12,000 in 2012 2 to 18,000 in 2021. 3 A previous study reported that the largest obstacle to recovery after SCI is extensive neuronal death resulting from apoptosis. 4 Cell stress induced by injury activates the mitochondrial pathway, which can trigger apoptosis. 5 Mitochondrial dysfunction is involved in apoptosis, inflammation, and neurodegeneration.6,7 In the mitochondrial apoptotic pathway, death signaling activates the BH3-domain-containing Bcl-2 family members and indirect activation of BAX/BAK, thereby allowing unimpeded spontaneous activation of BAX/BAK in the mitochondrial outer membrane 8 ; subsequently, the caspase cascade is initiated, resulting in the activation of the downstream protein Caspase-3, which cleaves its substrate, causing cell death. 9
Kaempferol (KPL) is the principal component of the traditional Chinese herb Kaempferia galanga L. and is found in many traditional Chinese medicines, such as Epimedium brevicornu Maxim. 10 KPL is a tetrahydroxyflavone containing 4 hydroxy groups at positions 3, 5, 7, and 4′ (Figure 1), and it has a wide spectrum of pharmacological activities. A previous study indicated that at different concentrations, KPL has potential neuroprotective activity.11,12 KPL was found to decrease oxidative stress levels and apoptosis levels and increase the expression of Bcl-2. 13 Currently, there are no reports on the potential mechanisms of KPL in inhibiting apoptosis after SCI.

Two-dimensional chemical structure of kaempferol (KPL).
In this study, we investigated the underlying molecular mechanism of KPL after SCI by network pharmacology and molecular docking techniques. Hydrogen peroxide (H2O2)-exposed PC12 cells are often used in in vitro studies of nervous system diseases, such as SCI.14–16 Herein, we report the neuroprotective effect of KPL in PC12 cells exposed to H2O2, and we find that KPL inhibits apoptosis by activating a particular molecule.
Methods and Materials
Reagents and Chemicals
A cell counting kit-8 (CCK-8; Lot: 0241016) was obtained from DIYIOBIO, dimethyl sulfoxide from Sigma Aldrich, and KPL (CAS: 520-18-3, Lot: 22807, liquid chromatography-mass spectrometry, purity: 99.62%) from MedChemExpress. To detect apoptotic cells, terminal deoxynucleotidyl transferase dUTP nick end labeling (TUNEL) staining was performed using an In Situ Cell Death Detection Kit (11684795910, Roche). We also purchased an anti-Akt1 antibody (ab233755) and anti-Akt1 antibody (S473; ab81283) from Abcam, anti-BAX (60267-1-lg) and anti-Bcl-2 (26593-1-AP) antibodies from Proteintech, an anti-Cytochrome C antibody (cat: AF0146) from Affinity, an anti-β-actin antibody (cat: 21338) from Signalway Antibody, an anti-MK2206 antibody (lot: S107811) from Selleck, and an anticleaved caspase-3 antibody from Cell Signaling Technology.
Cell Culture and Drug Treatment
The PC12 cell line was purchased from Shanghai Cell Research Center. PC12 cells were maintained in RPMI 1640 medium with 10% fetal bovine serum (Gibco; Lot: 2176404) and 1% penicillin‒streptomycin (100×; New Cell & Molecular Biotech) at 37 °C in an incubator containing 5% CO2.
Cell Viability Assay
Cell viability was evaluated by CCK-8 assays. PC12 cells were incubated for 24 h after plating in 96-well plates. The cells were exposed to different concentrations of H2O2, KPL, or MK2206 for specific periods. Afterward, the cells in each well were treated with CCK-8 solution (10 μL) and fresh RPMI 1640 and further incubated for 25 min. The absorbance of each well was then measured at a wavelength of 450 nm on a microplate reader (Infinite M200 PRO; Tecan).
Apoptosis Assays
A TUNEL assay was performed to determine the cells' apoptosis rate. Briefly, H2O2-injured PC12 cells were pretreated with KPL, and TUNEL staining was performed according to the instructions of the In Situ Cell Death Detection Kit. The stained cells were observed using a light microscope (Olympus).
Flow cytometry analysis was performed as follows: the apoptosis rate was determined with an Annexin V-FITC/PI double staining apoptosis detection kit. The cells were washed with phosphate-buffered saline, fixed in 4% paraformaldehyde for 15 min, and then stained. The apoptosis rate was analyzed by flow cytometry with an LSRII flow cytometer.
Network Pharmacology
Screening of the Active Ingredients of KPL and the Targets of KPL and SCI
The active ingredients of KPL were determined by searching the Traditional Chinese Medicine Systems Pharmacology (TCMSP; http://lsp.nwu.edu.cn/tcmsp.php), BATMAN-TCM (http://bionet.ncpsb.org/batman-tcm), 17 and Symmap (http://www.symmap.org/) 18 databases. The oral bioavailability was set to ≥30%, and the drug-likeness was set to ≥0.18 to screen the active ingredients.19,20 “SCI,” “Spine Cord Injury,” and “spinal cord injury” were entered as keywords into the Genecards (https://www.genecards.org), DrugBank (https://go.drugbank.com/) and Therapeutic Target Database (TTD; http://db.idrblab.net/ttd/) databases to search for relevant genes. The obtained target information was input into UniProt (httpswww.uniprot.org) to obtain the corresponding standard gene names.
Protein–Protein Interaction Network
Intersecting target genes were entered into the multiple proteins option of the STRING online database. As described in our previous study, 21 parameters were set to the default settings. All search results were imported into Cytoscape 3.7.1 software to visualize the protein–protein interaction (PPI) network.
Gene Ontology Enrichment and Kyoto Encyclopedia of Genes and Genomes Pathway Analyses
The intersecting target genes were entered into the Metascape online database (http://metascape.org/gp/index), and the min overlap was set to 3, the P-value cutoff to .01, and the min enrichment to 1.5. All search results were imported into the LC-Bio Cloud Platform (https://www.omicstudio.cn), and graphs were generated.
Molecular Docking Validation
As described in our previous study, 21 in brief, two-dimensional (2D) docking was performed with Protein Plus software (https://proteins.plus/). The 3-dimensional (3D) structures of the proteins were downloaded from the PDB database. The ligand molecules were prepared, and the docking pocket was identified. Then, the data were imported into AutoDock Tools (version 1.5.6) for molecular docking analysis and visualized using PyMOL (Version 2.5.2).
Western Blot Analysis
PC12 cells were lysed in denaturing buffer from the Minute™ Total Protein Extraction Kit (Cat# SD-001, Invent Biotechnologies) to obtain protein extracts. The protein concentration of the supernatant was measured using a BCA protein assay kit (Beyotime Biotechnology) and a full-wavelength functional microplate reader (Infinite M200Pro, Tecan). The procedure was performed according to the manufacturer's instructions. The following antibodies were used: Bcl-2 (1:500), cleaved caspase-3 (1:1000), BAX (1:6000), Akt1 (1:1000), Akt1 (phospho-S473, 1:4000), and β-actin (1:6000).
Statistical Analysis
All statistical analyses were performed by SPSS (Version 25.0) and GraphPad Prism (version 6.0c). Two-tailed Student's t-test was used for pairwise comparisons between the 2 groups. One-way analysis of variance was used to compare more than 2 groups of data. P < .05 was regarded as statistically significant.
Results
Potential Molecular Mechanisms of KPL on SCI
To elucidate the molecular mechanisms of KPL after SCI, we performed network pharmacology. We found that KPL (MOL000422) targets 84 related genes in total, including 63 identified in the TCMSP database, 5 in the BATMAN database, and 67 in the SymMap database. After deduplication, 51 target genes of KPL were identified. SCI was found to affect the expression of 1808 related genes in total, including 1787 identified in the Genecards database, 29 in the Drugbank database, and 7 in the TTD. After deduplication, 15 target genes of SCI were identified. There were 46 key genes targeted by KPL after SCI (Figure 2A). These 46 key genes were included in the PPI network, and the key genes interacting with TOP10 were identified as AKT1, TNF, CASP3, JUN, PTGS2, PPARG, ESR1, MAPK8, HMOX1, and RELA (Figure 2B).

(A) Wayne’s diagram showing that a total of 1808 genes were associated with SCI and 84 with the drugs, with 46 being key genes. (B) PPI network.
Gene ontology (GO) enrichment analysis of the target genes was performed for molecular function (MF), biological process (BP), and cellular component (CC) terms. The MF terms mainly included processes related to protein homodimerization activity, transcription factor binding, protein serine/threonine/tyrosine kinase activity, heme binding, and DNA-binding transcription factor binding. The BP terms were mainly related to the cellular response to nitrogen compounds, response to hormones, and regulation of the apoptotic signaling pathway. The CC terms mainly included membrane rafts, the membrane, and the side of the membrane (Figure 3A to C). Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis illustrated that the mechanism of KPL in treating SCI mainly involves the pathway apoptosis–multiple species (Figure 3D). The target genes were associated with biological processes and signaling pathways related to apoptosis, and the gene with the strongest association with apoptosis was Akt1 (Figure 3E).

(A) MF terms, (B) CC terms, (C) BP terms, (D) the data in the bubble plots were subjected to KEGG pathway analysis, and (E) chord map of BP terms and KEGG pathways related to apoptosis.
Molecular docking is a technique used to simulate the binding of small molecule ligands to receptor proteins and can be used to determine the binding position of drugs acting on disease-related genes and the binding energy between them. Free binding between molecules shows that the binding energy is negative, and the smaller the negative value is, the more stable the binding. 22 The ligand KPL could bind to the protein 1UNP (encoded by Akt1) well (binding energy: −5.72 kcal/mol). Further analysis of binding to the surface of the receptor (Figure 4A) at the 2D (Figure 4B) and 3D structure levels (Figure 4C) was performed, and the results revealed that the formation of hydrogen bonds between KPL and VAL-7, LYS-8, TRP-99, and TRP-11 was responsible for the stable binding of the ligand to the receptor. These results demonstrate an interaction between KPL and Akt1.
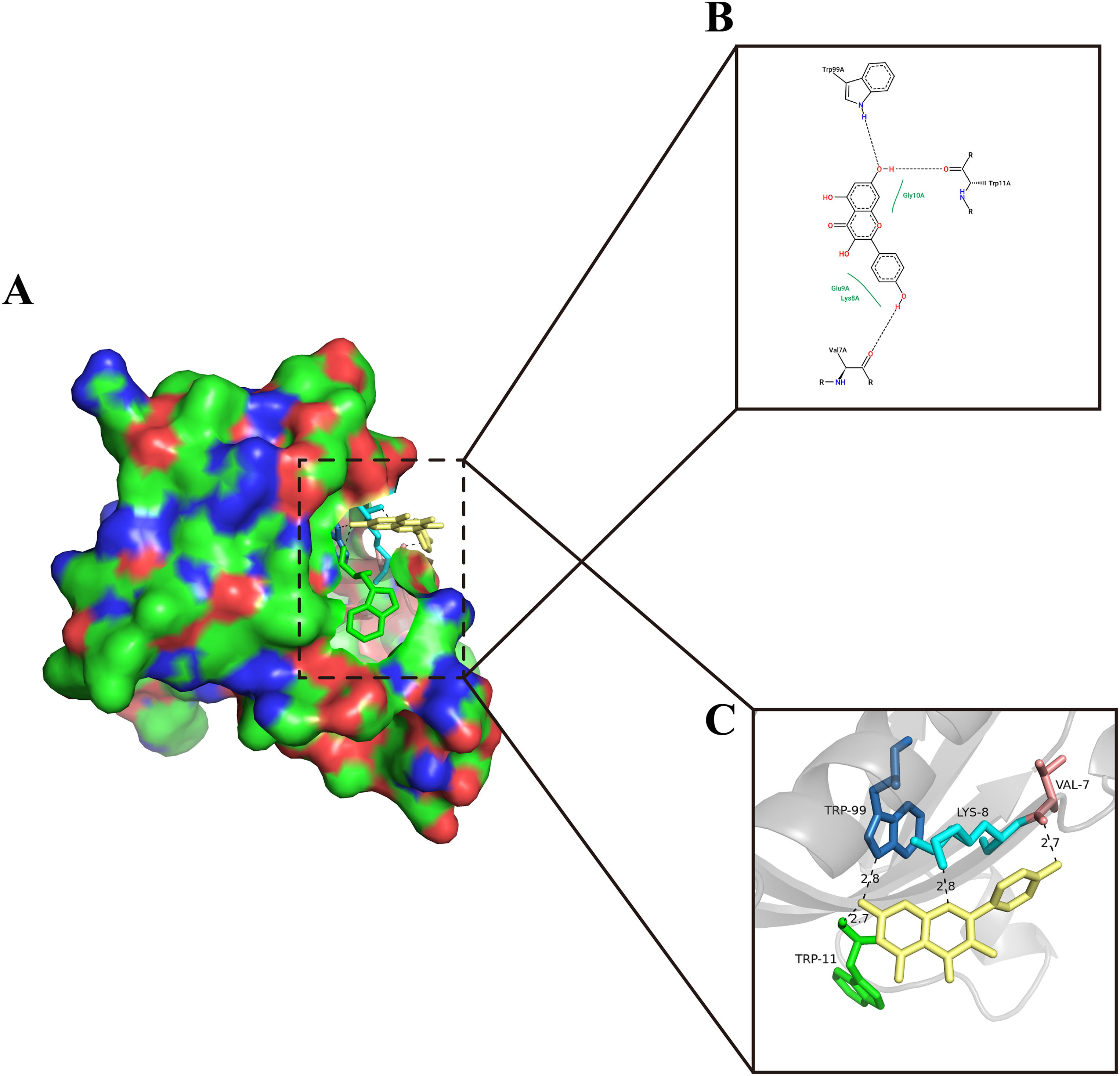
Molecular docking of the ligand KPL to the receptor 1UNP (encoded by Akt1). (A) The surface of the receptor and 3D structure of the ligands. (B) 2D structures of the ligands and receptors. (C) 3D structures of the ligands and receptors.
KPL Improves the Survival of PC12 Cells Exposed to H2O2
To clarify the molecular mechanism of KPL after SCI, we used PC12 cells, which are widely used as a neurobiological model. 23 The growth of PC12 cells treated with various concentrations of KPL was analyzed, and the results showed that cell viability was effectively increased when 10 μM KPL was added to 100 μL of culture medium (P < .0001); however, when the concentration of KPL reached 50 μM, cell viability significantly decreased (P < .0001; Figure 5A). Cells treated with 200 μM H2O2 exhibited ∼50% cell viability at 24 h (P < .0001; Figure 5B); moreover, the cells adopted a spherical shape, the connections between cells disappeared, the cells detached from each other, and the cell density decreased (Figure 5E). Subsequently, to elucidate whether KPL increased the viability of PC12 cells treated with H2O2, 200 μM H2O2 was added 30 min after the addition of 10 μM KPL. At 24 h after treatment, the viability and number of cells in the KPL-treated groups were higher than those in the untreated group, and the treated cells exhibited a clear and regular cell morphology and arrangement and significantly increased viability (P < .0001; Figure 5D and E).
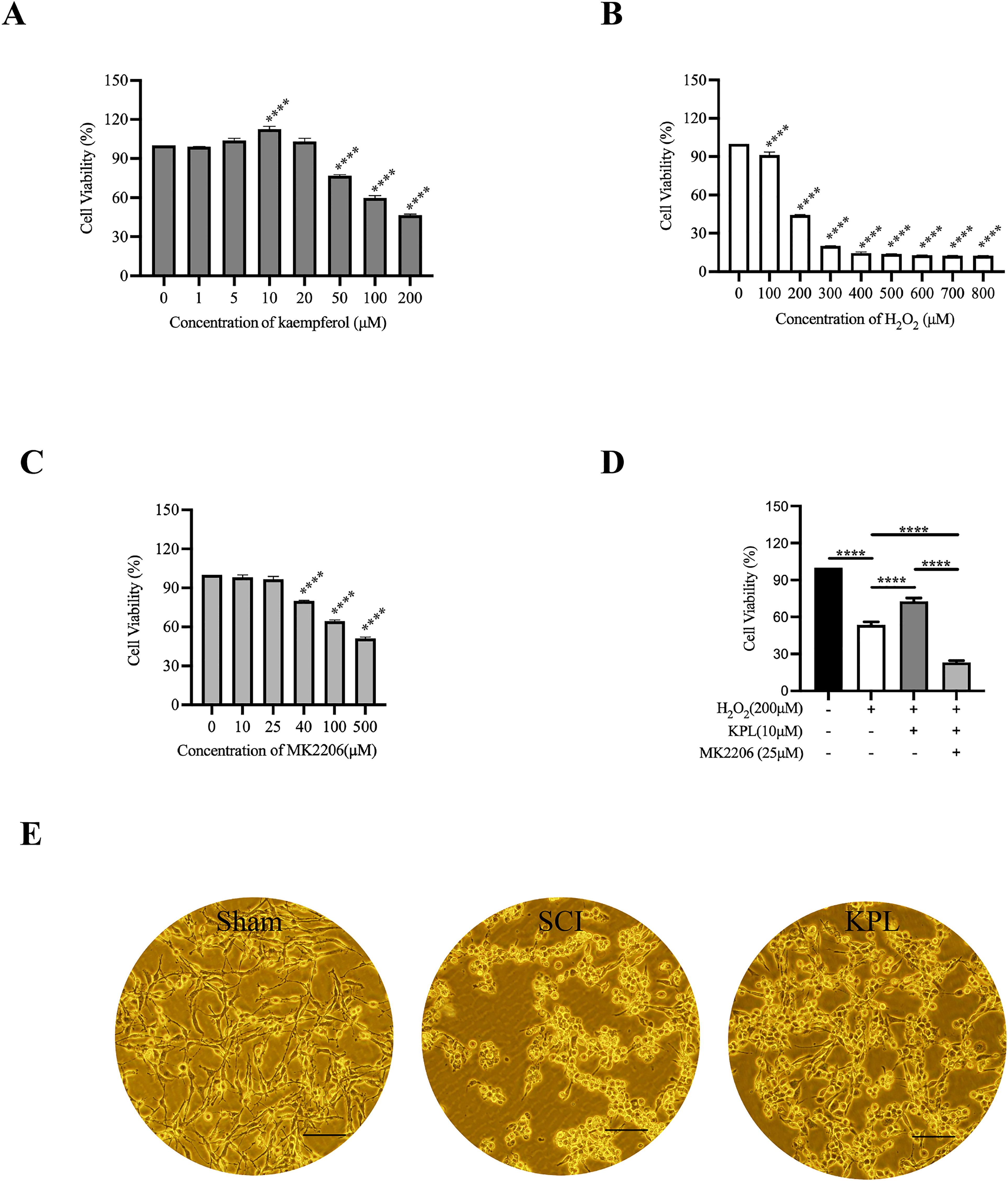
KPL increases the cell viability of PC12 cells following exposure to H2O2. (A) PC12 cells were treated with various concentrations of KPL (B), H2O2 (C), or MK2206 (D), or treated with 10 μM KPL, 25 μM MK2206, and 200 μM H2O2. (E) Cell morphology under a light microscope (magnification: 200×; bar: 100 μm). ****P < .0001.
KPL Inhibits Apoptosis in PC12 Cells Exposed to H2O2
We previously demonstrated that KPL improves the survival of PC12 cells treated with H2O2, possibly by inhibiting apoptosis (Figure 3C and D). We analyzed the effect of KPL on apoptosis, and TUNEL staining showed that the half-maximal concentration of H2O2 for inducing apoptosis of PC12 cells in 96-well plates after treatment for 24 h was 200 μM. When 10 μM KPL was added, the number of apoptotic cells was significantly reduced (Figure 6A and B; P < .0001). To confirm the above findings, flow cytometry analysis following double staining of PC12 cells with Annexin V-FITC/PI was performed. Compared with untreated cells treatment, 200 μM KPL-treated cells exhibited a significant reduction in apoptosis following treatment with 4 mM H2O2 for 24 h in 96-well plates. These results showed that treatment with KPL significantly inhibited H2O2-induced apoptosis (Figure 6C to E).

KPL alleviates H2O2-induced apoptosis. (A) Detection of apoptotic PC12 cells in 96-well plate after treatment with 10 μM KPL and 200 μM H2O2 for 24 h by TUNEL staining (×100). (B) Quantification of TUNEL-positive cells. (C) Analysis of PC12 cell apoptosis in a 6-well plate by flow cytometry. (D) Analysis of PC12 cell apoptosis after treatment with 200 μM KPL and 4 mM H2O2 for 24 h (E) or after treatment with 200 μM KPL and 4 mM H2O2 for 24 h by flow cytometry. ****P < .0001.
KPL Inhibits H2O2-Induced Mitochondrial Apoptosis by Inducing Akt1 Phosphorylation
A previous study suggested that Akt1 phosphorylated at Ser-473 is neuroprotective 24 and that overexpression of Akt1 effectively inhibits apoptosis. 25 Akt1 was implicated in the effects of KPL after SCI through network pharmacology and molecular docking studies (Figure 2B, 4A to C). The activation of the mitochondrial apoptotic pathway requires the oligomerization of bcl-2 family-related proteins, their insertion into the mitochondrial membrane, the release of cytochrome C (Cyt C), and further activation of the downstream effector caspase-3. 26 MK2206 is a potent Akt1 inhibitor.27,28 The data showed that 25 μM MK2206 had no significant effect on cell viability (P > .05), but 40 μM MK2206 significantly decreased cell survival (P < .0001; Figure 5C). Next, Western blot analysis was performed to measure the expression levels of proteins involved in the mitochondrial apoptotic pathway. We found that KPL significantly increased pAkt1 (Ser-473) levels and that this effect was significantly inhibited when MK2206 was added (P < .0001; Figures 5D, 7A and B). In our study, KPL significantly inhibited the expression of cleaved caspase-3, Cyt C, and Bax and increased the expression of Bcl-2. When MK2206 was added, the expression of Bcl-2 was inhibited, and the expression levels of cleaved caspase-3, Cyt C, and Bax were significantly increased (Figure 7C to F). Our data suggested that KPL played a critical role in the inhibition of mitochondrial apoptosis in PC12 cells exposed to H2O2 by inducing Akt1 phosphorylation.
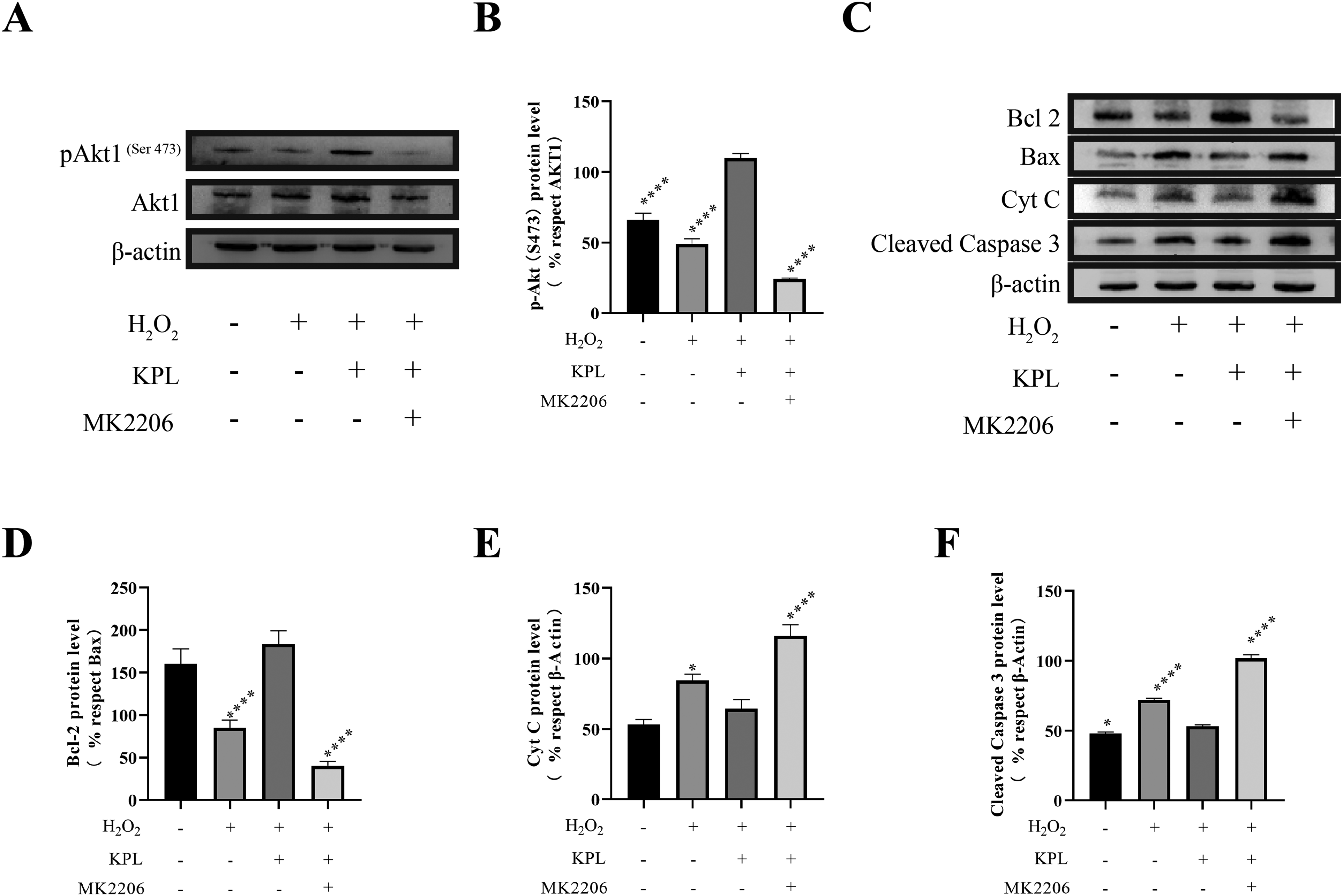
KPL inhibits H2O2-induced apoptosis by inducing Akt1 phosphorylation. (A) Protein levels of pAkt1 (Ser 475) and Akt1 were analyzed after treatment with 20 mM H2O2, 1 mM KPL, 2.5 mM MK2206, or a combination of drugs for 24 h. (B) The pAkt1 (Ser 475)/Akt1 ratio was quantified. (C) Protein levels of Bcl-2, Bax, Cyt C, and cleaved caspase-3 were analyzed after treatment with 20 mM H2O2, 1 mM KPL, 2.5 mM MK2206, or a combination of drugs for 24 h. The Bcl-2/Bax ratio (D), the Cyt C/β-actin ratio (E), and the cleaved caspase-3/β-actin ratio (F) were quantified. The results (mean ± SEM) are expressed as the percentage relative to control PC12 cells. *P < .05, ****P < .0001 compared to the KPL group.
Discussion
KPL can significantly protect neural cells from damage and reduce protease cleavage and nuclear apoptosis by inhibiting the oxidative stress response. 29 A previous study revealed that the pathophysiology of SCI mainly includes acute and chronic phases and is accompanied by a cascade of destructive events, such as oxidative stress and apoptotic pathways.30–32 However, there are no studies either predicting or validating the protective mechanism of KPL after SCI.
Network pharmacology is used to identify relevant biological networks and analyze the links between drugs, targets, and diseases in these networks 33 ; there are 2 distinguishing features of network pharmacology: its systematic nature and its predictive nature. 34 Network pharmacology can be used to identify herbal components that act on the network associated with specific diseases and comprehensively screen all possible targets to clarify the efficiency of drug therapy.35,36 In our study, we constructed a KPL-SCI gene set that consisted of 46 target genes and found 11 hub targets, including AKT1, TNF, JUN, CASP3, PTGS2, and PPARG. GO enrichment and KEGG pathway analyses showed that the effect of KPL after SCI was related to the regulation of the apoptotic signaling pathway.
Molecular docking has an important role in drug discovery, as it can predict and confirm the action of drugs on specific targets. 37 The success of molecular docking experiments is based on selecting the correct docking mode and predicting binding affinities. 38 Akt1 was the gene of interest that was most significantly associated with apoptosis, and molecular docking was performed to validate the interaction between the ligand KPL and the receptor Akt1 (binding energy: −5.72 kcal/mol; Figure 4).
PC12 cells exposed to H2O2 are often used as a neurological injury model.39–41 Therefore, we used the PC12 cell line to mimic neural cell damage after SCI and to further explore the mechanism of KPL in the treatment of SCI. The CCK-8 assay was used to determine the half-maximal inhibitory concentration of H2O2 and to assess the effect of KPL or both KPL and H2O2 on cell viability. In our study, we observed that 200 μM H2O2 resulted in ∼50% cell viability. We chose 10 μM KPL as the optimal concentration for the subsequent experiment, as it did not reduce cell viability. Our data showed that KPL significantly increased the viability of PC12 cells exposed to H2O2 (P < .0001; Figure 5D). These findings are in agreement with those from a previous study. 42
Inhibition of apoptosis is the main mechanism underlying the protective effect of a variety of traditional Chinese medicinal compounds on PC12 cells.40,43 Previous studies have shown that KPL treats a variety of diseases, such as osteoporosis, 44 knee osteoarthritis, 45 and ischemia-reperfusion injury,13,46 by inhibiting apoptosis. Research indicates that there are 3 main apoptotic pathways: the mitochondrial pathway, the endoplasmic reticulum pathway, and the death receptors pathway.47,48 TUNEL staining was used to detect apoptotic cells, as it labels free 3′-hydroxyl ends. 49 Herein, we used TUNEL staining and annexin V-FITC/PI staining to evaluate apoptosis. The SCI group (200 μM H2O2) exhibited elevation of apoptosis in PC12 cells in 96-well plates, which was significantly inhibited when 10 μM KPL was added (P < .0001). Subsequently, we verified the above results with annexin V-FITC/PI staining (cells treated with 200 μM KPL and 4 mM H2O2 for 24 h in a 6-well plate).
In vertebrates, mitochondria are thought to be a key for apoptosis regulation, and death signals cause apoptosis via activation of the BH3-domain-containing Bcl-2 family members (eg, Bax), their insertion into the mitochondrial membrane, the release of Cyt C, and finally activation of the caspase cascade and the downstream molecule caspase-3. 26 After SCI, neurons can be protected by alleviating mitochondrial-dependent apoptosis. 50 Consistent with the findings of previous studies, by examining the expression of key proteins and apoptotic factors, we found that the phosphorylation of Akt1 at S473 was induced by KPL, Bax, and Cyt C, and cleaved caspase-3 (Asp 175) expression was significantly increased in the SCI group, and Bcl-2 was significantly increased in the KPL group. When MK2206, 27 an Akt1 inhibitor, was added, the phosphorylation of Akt1 at S473 was inhibited, and the expression of apoptosis-related factors (Bax, Cyt C, and cleaved caspase-3) was markedly increased. Our data strongly suggested that KPL could inhibit mitochondrial apoptosis by activating Akt1 in PC12 cells.
Conclusion
In summary, our research systematically investigated the mechanisms of KPL after SCI. We found that KPL could inhibit apoptosis in PC12 cells exposed to H2O2 in vitro by inducing the phosphorylation of Akt1, which suggested that KPL is a very promising candidate for the treatment of SCI.
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was financially supported by the National Natural Science Foundation of China (No. 82174166) and the Innovation Capability Support Program of Shaanxi (No. 2021TD-45).
