Abstract
Introduction
Incidence of gastric cancer (GC), a very common cancer worldwide, is second highest compared to all other malignancies in China. 1 Unfortunately, most patients with GC are diagnosed in the progressive stage.2,3 Although, surgery and chemotherapy play a crucial role in GC therapy, the overall 5-year rate remains low. 4 Therefore, the exploration of new therapeutic strategies and key drug molecules is necessary to improve the treatment efficacy for GC.
Aloin (ALO) is a bioactive component obtained from Aloe species. 5 Recent studies have reported that ALO exhibits antitumor properties, and it has been observed to be pro-apoptosis, antiproliferation, and antimigration, thereby helping control metastasis.5–8 It has been reported that natural agents, such as sinomenine and quercetin can regulate the expression of many microRNAs (miRNAs), thereby promoting tumor cell apoptosis.9,10 ALO has been reported to decelerate the progression of hepatocellular carcinoma through the circ_0011385/miR-149-5p/WT1 axis, 5 and suppress osteoclastogenesis via inhibition of miR-21 expression. 11 However, it is not known whether ALO can induce GC cell apoptosis through regulation of miRNA-mediated target protein expression.
The high mobility group box 1 (HMGB1), a nonhistone DNA-binding protein, has been reported to regulate cancer cell apoptosis, proliferation, and migration.6,12–14 In addition, several miRNAs play an important role in cancer pathogenesis by regulating HMGB1 expression.15–17 Bioinformatics analysis has revealed that miR-5683 and HMGB1 are abnormally expressed in GC tissue. Moreover, TargetScan database has predicted that HMGB1 is the target gene for miR-5683.
In this study, we have investigated the ALO-induced effects of the miR-5683/HMGB1 axis on GC cell apoptosis. Specifically, a direct interaction has been observed between miR-5683 and the 3′ untranslated region (UTRs) of HMGB1. Our data suggest that ALO can promote GC cell apoptosis via the miR-5683/HMGB1 axis. This study has successfully provided a novel molecular mechanism by which ALO regulates GC cell apoptosis.
Materials and Methods
Reagents and Antibodies
ALO was purchased from Selleck Chemicals (Houston, Texas, USA) and dissolved in dimethyl sulfoxide; miR-5683 mimics and inhibitors from General Bio (Chuzhou, China); Lipofectamine® 3000 transfection reagent and 4′,6-diamidino-2-phenylindole (DAPI) from Thermo Fisher Scientific, Inc. (Waltham, MA, USA); CCK-8 (cell counting kit) and Annexin-FITC/PI (fluorescein isothiocyanate/propidium iodide) apoptotic kit from Keygen Biotech Co., Ltd (Nanjing, China); primary antibodies of PARP (poly ADP ribose polymerase), pro-caspase 3, cleaved caspase 7, and bcl-2 (B cell lymphoma 2) from Cell Signaling Technology (Beverly, MA, USA); HMGB1 from Thermo Fisher Scientific, Inc.; and horseradish peroxidase (HRP)-conjugated antimouse and antirabbit IgG secondary antibodies from Cell Signaling Technology (Beverly, MA, USA).
Cell Lines and Cultures
The MGC-803 cell line, a human gastric adenocarcinoma cell line, purchased from Zhiwei Biotechnology Co., Ltd (Hefei, China), was authenticated by short tandem repeat profiling. The GES-1 cell line, a normal gastric mucosal epithelial cell line, was obtained from Cellcook Biotech Co., Ltd (Guangzhou, China). The above cells were cultured in Roswell Park Memorial Institute (RPMI)-1640 medium (Gibco) supplemented with 10% fetal bovine serum (FBS) (Lonsera), 100 mg/mL streptomycin, and 100 U/mL penicillin at 37°C in 5% CO2.
CCK-8 Assay
Cell viability was detected using the CCK-8 assay. Briefly, MGC-803 cells were seeded in 96 well cell culture plates at a density of 1 × 104/well. After treatment with different doses of ALO, and incubating for 24 h, the cell culture medium was discarded, and the cells in the wells were cultured with new medium added with 10 μL/well of CCK-8 work fluid, and absorbance was measured at 450 nm after 2 h incubation, using a Multiskan™GO plate reader (Thermo Fisher Scientific, Inc.). Each treatment was performed in triplicate. The data are shown as mean ± SD (standard deviation).
Cell Apoptosis Detection
DAPI Staining
DAPI staining was used to observe ALO-induced morphological changes in the cells. MGC-803 cells were stimulated with different doses of ALO, and incubated for 24 h, following which the medium was discarded. Cells were then stained with DAPI work fluid for 5 min at room temperature in the dark. An inverted fluorescence microscope (Olympus, Japan) was used to observe the nuclear morphology.
Flow Cytometry Analysis
The apoptosis rate was determined using the Annexin V-FITC/PI double staining assay. In brief, MGC-803 cells were harvested by trypsin digestion after stimulation with different concentrations of ALO for 24 h. Binding buffer was used to re-suspend the cells, and then Annexin V and PI were added. The apoptosis rate was measured by BD flow cytometry (FCM) (Biosciences, Franklin Lakes, NJ, USA). The data were analyzed using Flowjo 7.6 software.
Protein Extraction and Western Blotting
Total protein was extracted from MGC-803 cells using Radio Immunoprecipitation Assay (RIPA) buffer containing phenyl methyl sulfonyl fluoride protein inhibitors (Beyotime). Lysates were collected by centrifugation at 4°C, at the speed of 12 000 rpm for 15 min. The total protein was separated by 10% sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE), and then transferred to negative control (NC) (naturally hydrophilic) membranes (Pall Corporation, Port Washington, NY, USA). The membranes were blocked in 5% fat-free milk powder for 1 h at room temperature, and then incubated with the indicated primary antibodies overnight at 4 °C. After washing with TBST (tris buffered saline with Tween 20) 3 times, HRP-conjugated secondary antibodies were incubated with the membranes for another 2 h. The proteins were visualized using an enhanced chemiluminescence (ECL) high-sensitivity kit (Abbkine). Finally, the protein images were captured using a chemiluminescence imaging system (Clinx, Shanghai, China) and semiquantified by Image J version 1.52 software.
Reverse Transcription Quantitative Real-Time Polymerase Chain Reaction
Total RNA was extracted using a GeneJET RNA Purification Kit (Thermo) and reversed into complementary DNA (cDNA) using RevertAid Master Mix (Thermo), notably, the reverse transcription of miR-5683 using a specific stem-loop primer (GENERAL BIOL). The cDNA amplification was performed using MonApTM ChemoHS qPCR Mix (Monad, China) according to the protocol provided by the manufacturer. Glyceraldehyde 3-phosphate dehydrogenase (GAPDH) and U6 were used as internal controls. The primers are shown below:
HMGB1 forward: 5′-CAGAGCGGAGAGAGTGAGGA-3′ HMGB1 reverse: 5′-TTGCCTCTCGGCTTCTTAGG-3′ GAPDH forward: 5′-AAAGCCTGCCGGTGACTAA-3′ GADPH reverse: 5′-AGAGTTAAAAGCAGCCCTGG-3′ miR-5683 forward: 5′-CTACAGATGCAGATTCTCTGACTTC-3′ miR-5683 reverse: 5′-ATCCAGTGCAGGGTCCGAGG-3′ U6 forward: 5′-CTCGCTTCGGCAGCACA-3′ U6 reverse: 5′-AACGCTTCACGAATTTGCGT-3′ Predenaturation: 95°C for 5 min Denaturation: 30 cycles at 95°C for 30 s Annealing: 55°C for 30 s Extension: 72°C for 1 min Extension: 72°C for 10 min
The steps in the thermal cycle reaction were as follows:
The relative gene expression levels were calculated using the 2–ΔΔCt method.
Cell Transfection
miR-5683 mimics, inhibitor, and NC were designed and synthesized by General Biology Co., Ltd (Chuzhou, Anhui, China), with the miRNA regulated sequence as follows.
miR-5683 mimic NC: UCACAACCUCCUAGAAAGAGUAGA miR-5683 mimics: UACAGAUGCAGAUUCUCUGACUUC miR-5683 inhibitor NC: UCUACUCUUUCUAGGAGGUUGUGA miR-5683 inhibitor: GAAGUCAGAGAAUCUGCAUCUGUA. Cell transfection was performed using Lipo3000, following the manufacturers’ protocols.
Dual-Luciferase Reporter Assay
The psicheck 2.0 reporter vector carrying either wild-type (WT) or mutant (MUT) HMGB1 was co-transfected with either the miR-5683 mimics or miR-NC into HEK-293T cells. After transfection for 48 h, luciferase activities were measured with a double luciferase reporter gene detection kit (Beyotime, China) using a multimode microplate reader (cat. No.:Synergy HT, BioTek, USA).
Bioinformatic Analysis
The differently expressed miRNAs in GC tissue and adjacent normal tissue were analyzed using TCGA (cancer genome atlas) and TargetScan databases. The TargetScan database was used to predict the putative target genes of miR-5683.
Statistical Analysis
Data are shown as mean ± SD. Statistical Package for the Social Sciences (SPSS) 170.0 software was used for the statistical analysis. The differences between multiple groups were analyzed by using one-way analysis of variance with Tukey's post hoc test. Student's t-test was used to analyze the differences between the 2 groups; P < .05 indicated statistical difference.
Results
ALO-Induced GC MGC-803 Cell Apoptosis
MGC-803 cells were stimulated with different doses of ALO for 24 h, and CCK-8 assay was used to evaluate the cell viability. We found that ALO reduced cell viability at concentrations above 100 μg/mL in a dose-dependent manner (Figure 1A). In view of the above results, we selected 100, 200, and 400 μg/mL of ALO for the subsequent experiments. DAPI staining and FCM were used to determine the apoptotic effects of ALO on MGC-803 cells. As shown in Figure 1B, the nuclei of the control cells were stained light blue, nuclear morphology was round, and the sizes were uniform. However, in the case of cells treated with ALO, the cell nuclei showed condensation and fragmentation, some nuclear cracking, and occasional multiple apoptotic bodies were noticed. The apoptotic rate in the control group was 7.99%, and ALO stimulation at the prementioned concentrations of 100, 200, and 400 μg/mL increased the apoptotic rate to 11.69%, 16.35%, and 22.28%, respectively (Figure 1C). Western blotting was used to detect expression levels of apoptosis-related proteins. The expression levels of cleaved PARP and cleaved-caspase 7 increased in a dose-dependent manner after ALO treatment for 24 h, while the levels of pro-caspase 3 and bcl-2 reduced in a dose-dependent manner, as expected (Figure 1D).
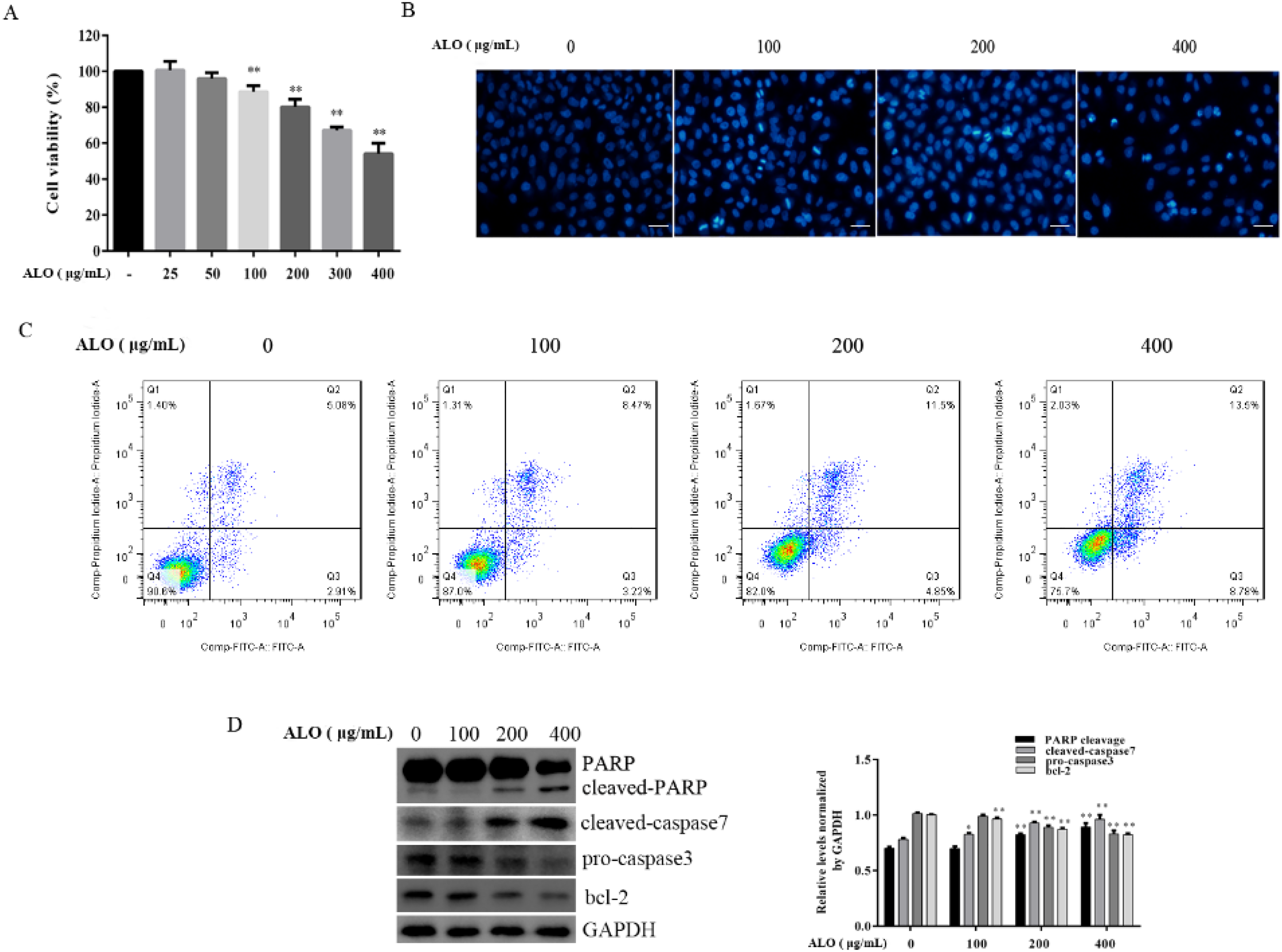
ALO-induced GC MGC-803 cell apoptosis. GC MGC-803 cells were treated with different doses of ALO for 24 h before CCK-8 assay was used to detect cell viability (A); DAPI staining was used to observe the nuclear morphological change (B). Magnification, 100 ×; scale bar:100 μm. (C) Cell apoptotic rates were detected using FCM; (D) the levels of apoptosis-related proteins were measured by WB. Data are shown as mean ± SD. *P < .05 and **P < .01 versus the control group. Abbreviations: ALO, aloin; CCK-8, cell counting kit 8; DAPI, 4′,6-diamidino-2-phenylindole; FCM, flow cytometry; GC, gastric cancer; SD, standard deviation; WB, western blotting.
ALO Decreased the Expression Level of HMGB1 at mRNA and Protein Levels
Our previous study demonstrated that ALO can regulate GC HGC-27 cell apoptosis by targeting HMGB1. However, the potential molecular mechanism of ALO on HMGB1 expression in GC cells has not been determined. We used different doses of ALO to treat MGC-803 cells for 24 h. The total protein was extracted, and western blotting was used to measure the levels of expressed HMGB1 protein. Figure 2A shows that ALO reduces expression levels of HMGB1 protein, and the inhibitory effect was seen to be dose dependent. In another experiment, MGC-803 cells were stimulated with 400 μg/mL of ALO for different time intervals. Total RNA was extracted and the expression level of HMGB1 mRNA was measured using reverse transcription quantitative real-time polymerase chain reaction (RT-qPCR). As expected, the data demonstrated that ALO attenuated HMGB1 mRNA levels after treatment for 6 to 24 h (Figure 2B).

ALO attenuated the expression of HMGB1 in MGC-803 cells. MGC-803 cells were stimulated with different concentrations of ALO for 24 h before total protein was extracted, and the levels of HMGB1 were detected using WB (A); MGC-803 cells were treated with ALO (400 μg/mL) for different times before total RNA was extracted, and the level of HMGB1 mRNA was determined by RT-qPCR (B). Data are expressed as mean ± SD. **P < .01 versus the control group. Abbreviations: ALO, aloin; HMGB1, high mobility group box 1; RT-qPCR, reverse transcription quantitative real-time polymerase chain reaction; SD, standard deviation; WB, western blotting.
ALO Upregulated the Expression Level of miR-5683
In order to investigate the underlying molecular mechanism by which ALO downregulates HMGB1 expression, we screened the differently expressed miRNAs using TCGA and TargetScan databases. We then predicted the possible miRNAs that target the HMGB1 gene, using the TargetScan database. Through bioinformatics analysis, we found that 6 miRNAs targeting the HMGB1 gene; miR-1258, miR-4770, miR-802, miR-5680, miR-944, and miR-5683 were downregulated in GC tissue (Table 1). Three miRNAs with the most significantly differential expression (miR-5680, miR-944, and miR-5683) were selected for verification at the cellular level. Total RNA in normal gastric mucosal epithelial cells GES-1 and GC MGC-803 cells was extracted, and the concentration and purity were determined using NanoDrop-1000 (Thermo). RT-qPCR was used to detect the expression levels of miR-5683, miR-5680, and miR-944. As evident from Figure 3A, their levels were downregulated in MGC-803 cells compared with those in GES-1 cells.

ALO enhanced the expression level of miR-5683 in MGC-803 cells. Total RNA in normal gastric mucosal epithelial cells GES-1 and GC MGC-803 cells were extracted, and RT-qPCR was used to detect the levels of miR-944, miR-5683, and miR-5680 (A); MGC-803 cells were treated with ALO (400 μg/mL) for different times before total RNA was extracted, and the levels of miR-944 (B), miR-5680 (C), and miR-5683 (D) were determined using RT-qPCR. Data are expressed as mean ± SD. **P < .01 versus the control group. Abbreviations: ALO, aloin; GC, gastric cancer; RT-qPCR, reverse transcription quantitative real-time polymerase chain reaction; SD, standard deviation.
Differently Expressed miRNAs That Target HMGB1 Gene in GC and Adjacent Tissues.
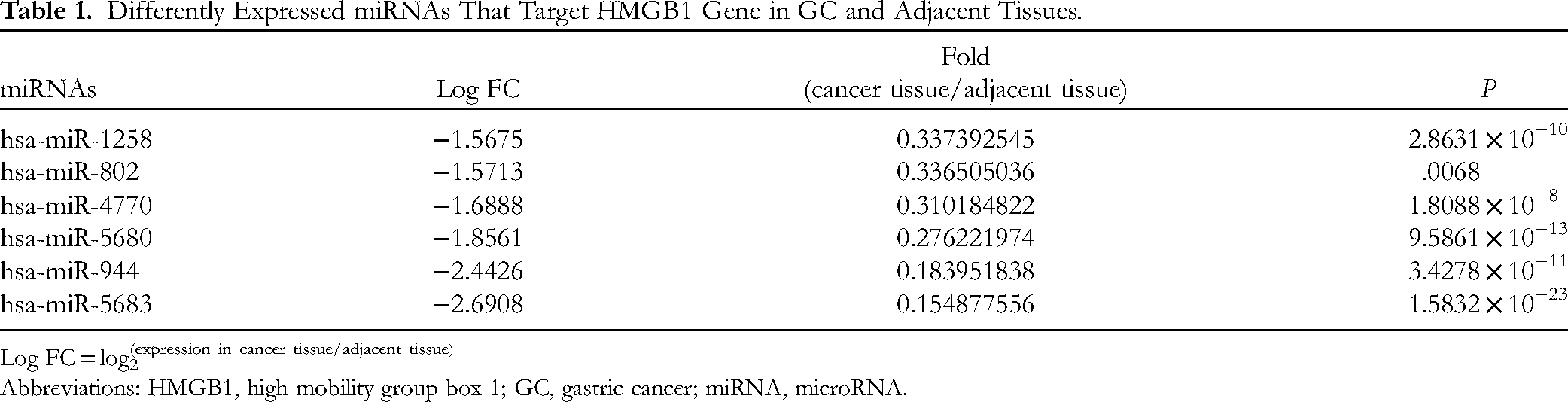

Log FC = log2(expression in cancer tissue/adjacent tissue)
Abbreviations: HMGB1, high mobility group box 1; GC, gastric cancer; miRNA, microRNA.
To explore the effects of ALO on these miRNAs, MGC-803 cells were treated with 400 μg/mL of ALO for different time intervals (3, 6, 12, and 24 h). Total RNA was extracted and the expression levels of miRNAs were determined using RT-qPCR. As shown in Figure 3B, ALO attenuated miR-5680 expression, and considerably enhanced the miR-5683 expression level at 6 h, but had no effect on the expression of miR-944.
miR-5683 Downregulated the Expression Level of HMGB1 at the Post-Transcription Level
The levels of HMGB1 in GES-1 and MGC-803 cells were measured, and the results indicated that HMGB1 was highly expressed in MGC-803 cells (Figure 4A). Regarding the mutually opposing expressions of HMGB1 and miR-5683 in GC tissue and cells, and considering the fact that miRNA can inhibit gene expression by directly binding with target mRNAs, we suspected that miR-5683 might be responsible for the inhibitory effect of ALO on HMGB1. To prove this hypothesis, MGC-803 cells were transfected with miR-5683 mimics and an NC for 24 h. Following this, the transfection efficiency was verified by RT-qPCR (Figure 4B); the levels of HMGB1 were measured by western blotting. It was observed that miR-5683 mimics distinctly inhibited HMGB1 expression, as evident from Figure 4C. To further confirm that miR-5683 influenced ALO-induced HMGB1 expression, we transfected MGC-803 cells with miR-5683 mimics and an NC, and then carried out stimulation with ALO for 24 h. HMGB1 levels were then determined using western blotting. As shown in Figure 4D, miR-5683 mimics were seen to enhance the ALO-induced downregulation of HMGB1. On the contrary, when we transfected MGC-803 cells with miR-5683 inhibitor for 24 h, the result was exactly opposite and HMGB1 levels were enhanced (Figure 4E-F).

ALO inhibited HMGB1 expression via upregulating miR-5683 in MGC-803 cells.Total protein and RNA in normal gastric mucosal epithelial cells GES-1 and GC MGC-803 cells were extracted, and the HMGB1 level was measured using WB (A); miR-5683 expression was detected by RT-qPCR (B), **P < .01 versus GES-1. MiR-5683 mimics and NC were transfected in MGC-803 cells, and after transfection for 24 h, total protein was extracted. WB was used to detect the expression level of HMGB1 (C); miR-5683 mimics and NC were transfected in MGC-803 cells for 24 h, and then the cells were stimulated either with or without 400 μg/mL of ALO for another 24 h, after which total protein was collected, and the expression of HMGB1 was measured using WB (D); miR-5683 inhibitor and NC were transfected in MGC-803 cells, and after transfection for 24 h, total protein was extracted, WB was used to detect the expression level of HMGB1 (E); miR-5683 inhibitor and NC were transfected in MGC-803 cells for 24 h, and then the cells were stimulated either with or without 400 μg/mL of ALO for another 24 h. Total protein was collected, and the expression of HMGB1 was measured using WB (F). Data are shown as mean ± SD. **P < .01 versus miR-5683 NC; ##P < .01 versus miR-5683 NC + ALO. Abbreviations: ALO, aloin; GC, gastric cancer; HMGB1, high mobility group box 1; NC, negative control; RT-qPCR, reverse transcription quantitative real-time polymerase chain reaction; SD, standard deviation; WB, western blotting.
miR-5683 Directly Binds to the 3′-UTRs of HMGB1 mRNA
TargetScan database was used to predict the interaction between miR-5683 and HMGB1, and the binding site between them were located (Figure 5A). To further validate the interaction between miR-5683 and HMGB1, and to determine whether miR-5683 directly binds to the 3′-UTRs of HMGB1 mRNA, the dual-luciferase reporter assay was performed. It was observed that miR-5683 mimics significantly attenuated luciferase activity, which is evident from the fact that luciferase activity in HMGB1-3′ UTRs-WT (wild type) group decreased by 24% compared with that in the control group (Figure 5B). However, no such effect was observed in the mutant type. After a mutation in the HMGB1-3′-UTR region, the luciferase activity of the mutant type increased by 27.6% compared with HMGB1-3′-UTRs-WT. From the results, it is evident that miR-5683 can be combined with the predicted target sequence of the HMGB1 gene.

miR-5683 directly binds to the 3′-UTR of HMGB1 mRNA. The binding site between miR-5683 and the 3ʹ-UTRs of HMGB1 was predicted using the TargetScan database (A). The interactions of miR-5683 with either wild-type HMGB1 or mutated HMGB1 were verified by a dual luciferase reporter assay (B). **P < .01vs miR-NC or miR-5683 + HMGB1 WT. Abbreviations: HMGB1, high mobility group box 1; mRNA, messenger RNA; UTR, untranslated region; WT, wild type.
miR-5683-Mediated ALO-Induced GC Cell Apoptosis
To investigate further the effect of miR-5683 on ALO-induced apoptosis, we transfected miR-5683 mimics, inhibitor, and an NC in MGC-803 cells. DAPI staining and FCM were used to verify the apoptotic effects. DAPI staining revealed that in the cells transfected by miR-5683 mimics, the presence of apoptotic bodies, nuclear condensation and nuclear fragmentation was obvious, compared to cells transfected with the NC. In addition, miR-5683 mimics were seen to enhance significantly the ALO-induced nuclear condensation and fragmentation (Figure 6A). From the FCM results, it was evident that miR-5683 mimics not only promoted MGC-803 cell apoptosis but also enhanced ALO-induced apoptosis. The apoptosis rate of miR-5683 mimic-transfected cells was 25.9%, which was significantly higher than that of the NC; moreover, miR-5683 mimics significantly enhanced ALO-induced apoptosis, and the apoptotic rate increased from 28.1% to 39% (Figure 6B). However, transfection with miR-5683 inhibitor reversed the apoptotic effects of ALO on MGC-803 cells (Figure 6C-D).
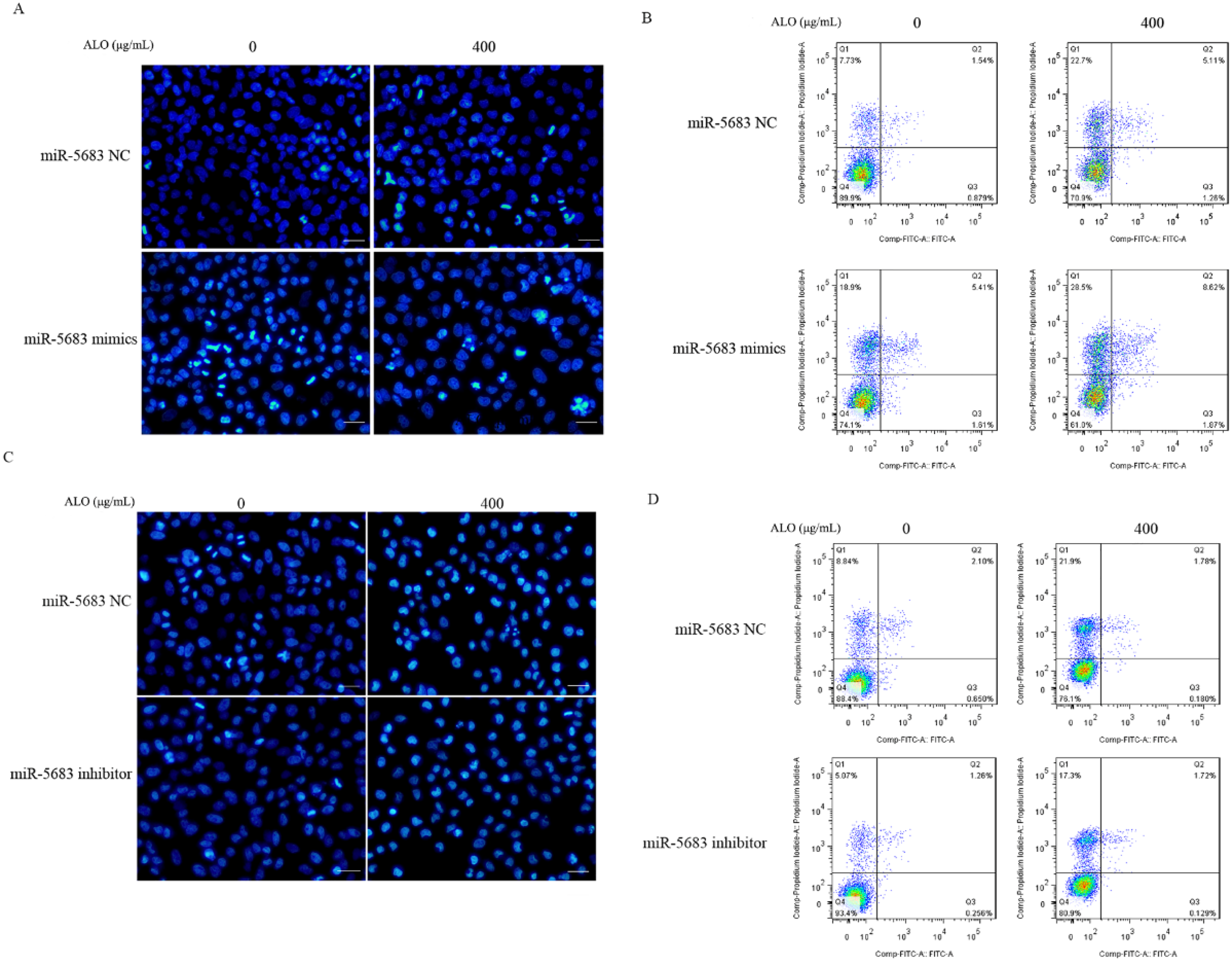
miR-5683 regulated ALO-induced MGC-803 cell apoptosis. MGC-803 cells were transfected with either miR-5683 mimics or negative control for 24 h, and then stimulated with ALO for 24 h. DAPI staining was used to observe the nuclear morphology change (A); FCM was used to measure the apoptotic rate (B). mIR-5683 inhibitor and negative control were transfected with MGC-803 cells and then ALO treated for 24 h, DAPI staining was used to observe the nuclear morphology change (C); FCM was used to measure the apoptotic rate (D). Magnification, 100 ×; scale bar:100 μm. Abbreviations: ALO, aloin; DAPI, 4′,6-diamidino-2-phenylindole; FCM, flow cytometry.
Discussion
Although our previous study demonstrated that ALO could induce apoptosis of GC HGC-27 cells via HMGB1 downregulation, 6 the underlying mechanism of this needs to be clarified.
In the present study, we first went on to prove the apoptotic effect of ALO on MGC-803 cells. The results and data obtained from the CCK-8, DAPI, as well as FCM studies have revealed that ALO clearly promotes apoptosis of MGC-803 cells (Figure 1A-C). These results are consistent with the previously reported findings.18,19 PARP cleavage, cleaved caspase, and bcl-2 are known to play an important role in cancer cell apoptosis.20,21 Our data demonstrated that ALO enhanced the expression of cleaved PARP and cleaved caspase 7 and reduced the levels of pro-caspase 3 and bcl-2 in a dose-dependent manner (Figure 1D).
Subsequently, we explored the mechanism of ALO-induced MGC-803 cell apoptosis. HMGB1 is known to be associated with the proliferation, migration, and apoptosis of cancer cells.22,23 Our study indicated that ALO inhibits HMGB1 expression at the mRNA and protein levels (Figure 2). How does ALO downregulate the expression of HMGB1? miRNAs have been reported to play crucial roles in modulating the progression of tumors through regulation of the target gene expression, by binding to 3′-UTRs of the target genes.24,25 For example, miR-129-5p and miR-1179 attenuated tumor proliferation via targeting HMGB1 in GC cells26,27 miR-29b induced prostate cancer apoptosis by increasing Bim expression, 28 and miR-107 enhanced the radiosensitivity of nonsmall cell lung cancer by downregulating expression of HMGB1. 29
Extrapolating the results of the above research, we first analyzed the distribution of miRNAs in GC and adjacent normal tissues using the TCGA and TargetScan database, and then screened the differentially expressed miRNAs. We then predicted the identity of the miRNAs that possibly bind to the 3′-UTRs of the HMGB1 gene using the TargetScan database (Table 1). Among these miRNAs, we selected 3, which were miR-5683, miR-5680, and miR-944, and then performed RT-qPCR to validate their differential expression at the cellular level (Figure 3A). The results were consistent with the results of the bioinformatics analysis. The presence of HMGB1 in GES-1 and MGC-803 cells was detected by western blotting. The data showed that HMGB1 was highly expressed in MGC-803 cells. The expression levels of HMGB1 in the presence of the 3 abovementioned miRNAs were reduced. This suggests that these miRNAs possibly mediate the effect of ALO on HMGB1.
To confirm the hypothesis that the 3 miRNAs, miR-5683, miR-5680, and miR-944, could mediate the inhibitory effect of ALO on HMGB1, we treated MGC-803 cells with ALO for various time intervals, and then performed RT-qPCR to measure the expression of miRNAs. We found that ALO significantly upregulated miR-5683 and downregulated miR-5680. However, it did not affect the expression of miR-944 (Figure 3B-D). Thus, we concluded that miR-5683 mediated the effect of ALO on HMGB1. To prove our hypothesis, we transfected MGC-803 cells with miR-5683 mimics and miR-5683 inhibitor, and then detected the level of HMGB1. We found that miR-5683 mimics not only inhibited the expression of HMGB1, but also enhanced the inhibitory effect of ALO on HMGB1. Moreover, transfection with the miR-5683 inhibitor showed the expected opposite effect on HMGB1 by enhancing its expression (Figure 4).
To verify that miR-5683 binds to HMGB1, we performed a bioinformatics analysis on the TargetScan database. The data indicated that miR-5683 could indeed bind with the 3ʹ-UTR of HMGB1 (Figure 5A). Although the lack of RNA-RNA binding or RNA-protein assays to confirm this association is a limitation to the present study, luciferase reporter assay results revealed that the luciferase activity of WT (HMGB1-WT) was reduced by miR-5683 mimics. However, the miR-5683 mimics had no effect on HMGB1-3ʹ-UTR-MUT (Figure 5B). Combination of all the above results indicates that ALO can attenuate HMGB1 expression via miR-5683.
Our previous study has shown that ALO can induce GC cell apoptosis through downregulation of HMGB1 expression. 6 However, the role of miR-5683 in ALO-induced GC apoptosis is still unclear. Hence, we proceeded to observe the apoptosis resulting from the transfection of MGC-803 cells with miR-5683 mimics and miR-5683 inhibitor with and without ALO treatment using FCM. The data revealed that miR-5683 mimics clearly enhanced ALO-induced GC cell apoptosis. However, the miR-5683 inhibitor exerted the reverse effect and attenuated apoptosis (Figure 6).
The results of this study (Figure 3D) indicated that ALO upregulated miR-5683 expression at 6 h, but sharply downregulated it at 12 and 24 h. Future studies are required to elucidate these results and the associated mechanisms. miRNAs are known to associate with Argonaute (AGO) proteins for directing widespread post-transcriptional gene repression. Although this association with AGO typically protects miRNAs from nucleases, extensive pairing with unusual target RNAs can trigger miRNA degradation. It is possible that ZSWIM8 ubiquitin ligase recognizes the conformational changes caused by extensive pairing with the miRNA 3′ region, which leads to polyubiquitination of AGO, followed by degradation by the 26S proteasome, thereby exposing the miRNA to cytoplasmic nucleases. 30 Dicer is an enzyme required for the processing of precursor miRNA (pre-miRNA) into mature miRNA. It has been reported that different stress conditions, such as apoptosis and hypoxia, could regulate the expression of Dicer.31,32 In addition, Dicer protein undergoes post-translational modifications such as phosphorylation and SUMOylation (SUMO: small ubiquitin-related modifier). These modifications may affect Dicer abundance and/or activity. 33
In view of the above research, we postulate that the possible reasons for the decline of miR-5683 after ALO treatment for 12 and 24 h are as follows: (1) ALO downregulates the expression of Dicer at transcription and post-translation levels; (2) ALO inhibits the activity of Dicer; and (3) ALO affects the polyubiquitination of AGO and its degradation. Exploring the molecular mechanism of ALO downregulating miR-5683 expression after 12 h is the focus of our next work.
From the present study, we can conclude that ALO promotes apoptosis of GC MGC-803 cells through regulation of the miR-5683/HMGB1 axis. This study provided a new insight into the antitumor effect of ALO on GC cells.
Footnotes
Authors’ Contributions
XC was involved in investigation, methodology, software, and original draft preparation. XW was involved in investigation, methodology, and software; FG in investigation and methodology; ZC in software, formal analysis; JC in methodology and software; and ZQ in supervision, funding acquisition, and writing—review & editing. All authors have read and approved the final manuscript.
Availability of Data and Materials
The data that support the findings of this study are available from the corresponding author on reasonable request.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Natural Science Foundation of China (grant nos. 81601380 and 81872371); Natural Science Research Project of Anhui Provincial Colleges and Universities (grant no. KJ2020ZD54); Top academic talents in discipline (specialty) Funded project (gxbjzd2021060); Active Biological Macromolecules Research Provincial Key Laboratory Project (grant no.1306C083008). College Students’ innovation and Entrepreneurship training program project (grant nos. 202010368052 and S202110368109); and Wannan Medical College Scientific Research Grant Program for university students (grant nos. WK2021XS07 and WK2021XS64).
