Abstract
Introduction
The Food and Agriculture Organization (FAO) estimates that 20–40% of annual global crop losses are due to plant pathogenic infections, both directly and indirectly. 1 This translates to an annual cost of approximately 40 billion dollars to the global economy. 2 In Ethiopia, crops like bananas (Musa acuminate), enset (Ensete ventricosum), coffee (Coffea Arabica), and ginger (Zingiber officinale) are seriously threatened by plant microbial diseases such as blight and wilt, which significantly affect their economic viability. 3 The widespread use of pesticides worldwide, including fungicides, 4 nematicides, 5 and antibiotics for bacterial plant pathogens, 6 has indeed led to significant increases in crop yields. However, this extensive pesticide use brings several challenges, such as the development of pathogen resistance, changes in plant pathogen growth patterns, and concerns about toxicity to both humans and the environment.7–9 Hence, there is a pressing need to search for new, safe, and effective drugs, including natural products, capable of efficiently preventing and controlling plant pathogens while also overcoming the limitations of current drugs, such as the development of resistance and toxicity. 10
Aloe pirottae A. Berger, a member of the Asphodelaceae family, is indigenous to Ethiopia growing at altitudes ranging from 1300 to 1820 m. 11 In Ethiopian traditional medicine, the roots, leaves, and the leaf latex of A. pirottae have been used to treat various ailments, including tropical ulcer, malaria, snake bites, gallstones, hemorrhoids, wounds, eye infections, gonorrhea, parasites, muscular pain, colon cleaner, constipation, diabetes mellitus, gastritis, and social uses such as weaning children from breastfeeding, mosquito and insect repellent effect.8,12,13 Previous studies have also investigated the antibacterial, 14 anticancer, 15 and antimalarial 16 activities of different parts of A. pirottae. Hence, this study aimed to isolate, characterize, and perform molecular docking of compound(s) isolated from the leaf latex of A. pirottae. The goal was to gain further insights into the enzyme targets of these compounds and to understand how they bind to the active sites of the enzymes.
Results and Discussion
Characterization of the Isolated Compound (1 )
Phytochemical analysis of the leaf latex of A. pirottae was conducted using column chromatography, resulting in the isolation of a yellow amorphous compound (
In the negative-mode HRESI-MS (Fig. S1), compound

Chemical structure of aloin A/B.
Antibacterial Activity of the Leaf Latex and Aloin A/B
Table 1 summarizes the antibacterial effects of both the leaf latex and aloin A/B against three bacterial strains: Pseudomonas syringae pv. gaarcae, Ralstonia solanacearum and Xanthomonas vasicola pv. musacearum. In the disk diffusion method, X. vasicola showed the highest susceptibility to both the leaf latex and aloin A/B at a concentration of 25 mg/mL, with the inhibition zone diameters (IZD) of 9.68 mm and 14.47 mm, respectively. Among the strains tested, R. solanacearum exhibited the lowest sensitivity to the leaf latex (IZD = 9.91 mm; 50 mg/mL), while P. syringae showed the least activity against aloin A/B, with IZD of 11.54 mm at the highest concentration tested (25 mg/mL).
Inhibition Zone Diameter (IZD) and minimum Inhibitory Concentration (MIC) of Leaf Latex and Aloin A/B.
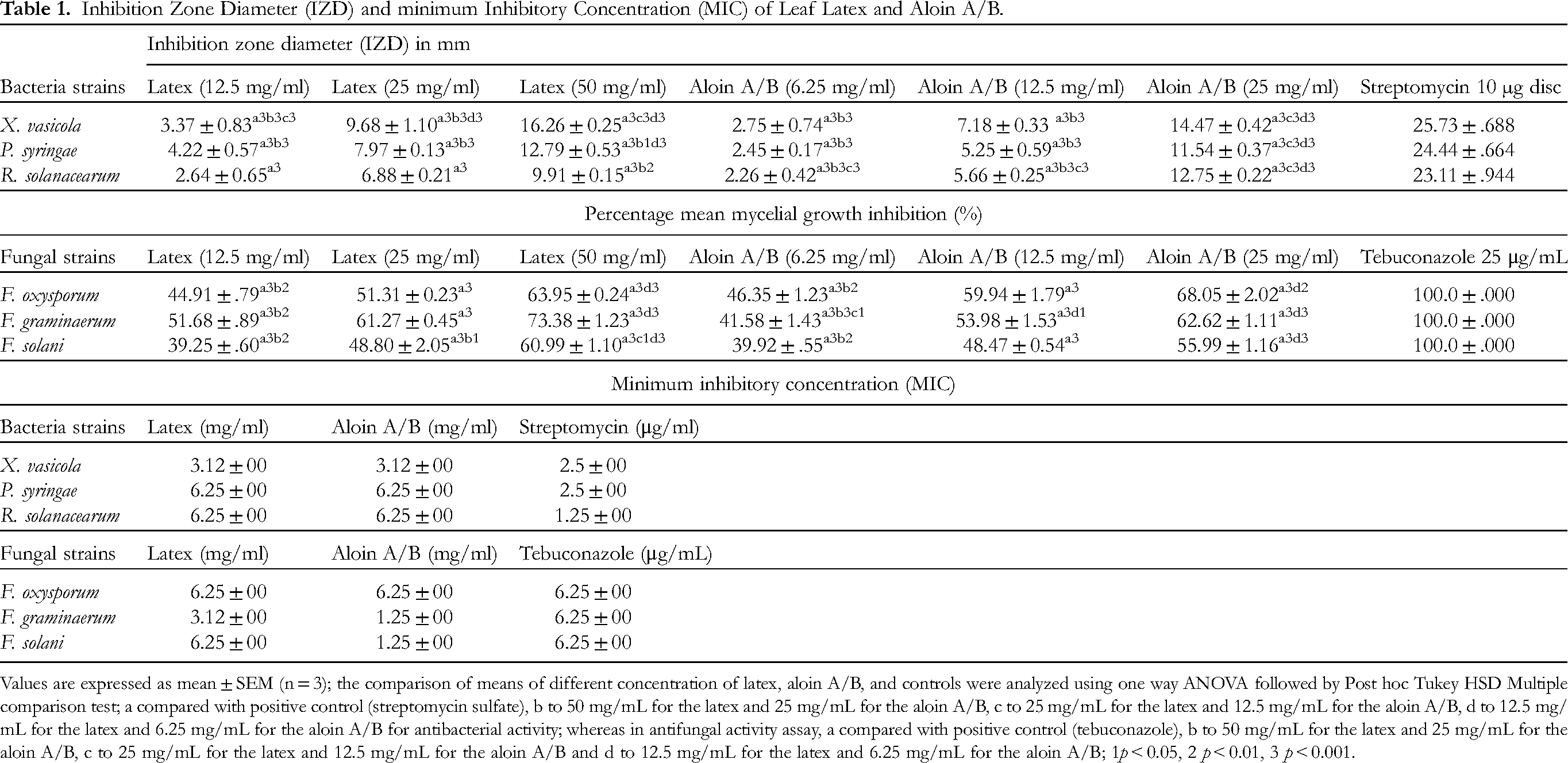

Values are expressed as mean ± SEM (n = 3); the comparison of means of different concentration of latex, aloin A/B, and controls were analyzed using one way ANOVA followed by Post hoc Tukey HSD Multiple comparison test; a compared with positive control (streptomycin sulfate), b to 50 mg/mL for the latex and 25 mg/mL for the aloin A/B, c to 25 mg/mL for the latex and 12.5 mg/mL for the aloin A/B, d to 12.5 mg/mL for the latex and 6.25 mg/mL for the aloin A/B for antibacterial activity; whereas in antifungal activity assay, a compared with positive control (tebuconazole), b to 50 mg/mL for the latex and 25 mg/mL for the aloin A/B, c to 25 mg/mL for the latex and 12.5 mg/mL for the aloin A/B and d to 12.5 mg/mL for the latex and 6.25 mg/mL for the aloin A/B; 1p < 0.05, 2 p < 0.01, 3 p < 0.001.
Consistent with the results from the disk diffusion method, both the leaf latex and aloin A/B (Table 1) exhibited the highest antibacterial activity against X. vasicola, with a minimum inhibition concentration (MIC) value of 3.12 mg/mL for each. Furthermore, the pathogen R. solanacearum was found to be more susceptible to A. pirottae leaf latex (MIC = 6.25 mg/mL) compared to a previous study using Aloe vera extract (MIC = 25 mg/mL). 18 It worth mentioning that a product formulated from Aloe vera has been utilized to combat bacterial, fungal, and viral plant pathogens. 19
Antifungal Activity of the Leaf Latex and Aloin A/B
The antifungal activity of the leaf latex and aloin A/B was evaluated against three fungal strains, namely Fusarium graminarum, Fusaium oxysporum, and Fusarium solani using the poisoned food technique.20,21 The results presented in Table 1 showed that F. graminarum was most susceptible to the leaf latex, with a maximum inhibition rate of 73.38% at a concentration of 50 mg/mL. Meanwhile, aloin A/B exhibited a maximum inhibition rate of 68.05% against F. oxysporum at a concentration of 25 mg/mL. Additionally, the standard drug (tebuconazole; 25 μg/mL) completely suppressed the mycelial growth of all tested fungal strains.
The mean diameter of inhibition zones produced by both the latex and aloin A/B at all dose levels against the test bacterial and fungal strains (p < 0.05) exhibited statistically significant differences compared to those produced by their respective positive controls. In a previous study conducted by Kankamol et al (2021) 18 on Aloe vera extract, a mycelial growth inhibition rate of 54.70% against F. oxysporum was observed at a concentration of 25 mg/mL. Interestingly, this inhibition rate was comparable to the mycelial growth inhibition rate of A. pirottae leaf latex (51.32%) against F. oxysporum at the same concentration level (25 mg/mL), as revealed in the current study. This similarity may be attributed to common phytochemical constituents among the plants. Moreover, when the broth dilution method was employed, leaf latex of A. pirottae demonstrated the strongest activity against F. graminaerum, with an MIC value of 3.12 mg/mL. On the other hand, F. oxysporum exhibited the highest susceptibility to aloin A/B, with MIC value of 6.25 mg/mL.
Molecular Docking Analysis
Both topoisomerase II and MurE are well-established targets in antimicrobial research. The selection of these enzymes for molecular docking studies in Fusarium oxysporum and Xanthomonas vasicola was based on prior in silico and experimental evidence. Studies have shown that naturally derived anthrones inhibit topoisomerase II22–26 and MurE (UDP-N-acetylmuramoyl-L-alanyl-D-glutamate-2,6-diaminopimelate ligase).27,28
Topoisomerase II is essential for DNA replication, transcription, recombination, and repair, making it a valuable target for antifungal and antibacterial agents.29,30 MurE, crucial in peptidoglycan biosynthesis, plays a key role in bacterial cell wall formation. Inhibiting MurE can weaken the cell wall, leading to cell lysis and death, which makes it a promising target for antibacterial drug development.31,32
Homology Modeling and Validations of Modeled Proteins
The crystal structures of the target enzymes were unavailable in the Protein Data Bank (PDB). Therefore, homology models were created using SWISS-MODEL and a template library generated from a BLAST search of UniProt, utilizing the closest crystal structures: murE from Xanthomonas vasicola (PDB: 8F5D) and DNA topoisomerase II from Fusarium oxysporum (PDB: 4GFH).
The homology model of murE of Xanthomonas vasicola was built with a 50.76% sequence identity, a GMQE score of 0.73, and a QMEAN score of −2.35, indicating good quality and reliability. Similarly, the homology model of DNA topoisomerase II of Fusarium oxysporum was built with a 53.70% sequence identity, a GMQE score of 0.60, and QMEAN of −1.72, also indicating good quality and reliability. The QMEAN score of below −4.0 depicts low quality of the predicted structure, while the GMQE score ranges between 0 and 1, and a higher score corresponds to higher reliability.33,34 The Ramachandran plot (Figure 2), which assesses the precision and dependability of the models, showed favorable results for both proteins. The Ramachandran plot demonstrated that murE of Xanthomonas vasicola protein showed a 95.00% Ramachandran favored, and 1.52% Ramachandran outliers. The DNA topoisomerase II of Fusarium oxysporum protein showed 95.03% Ramachandran favored and 0.66% Ramachandran outliers. Consequently, these results indicate that the proposed homology model is reliable and acceptable.

3D model structure of protein from murE (A) and DNA topoisomerase II DNA topoisomerase II (B) ramachandran plot analysis of the homology modeled residues murE (C) and DNA topoisomerase II DNA topoisomerase II (D).
Docking Analyses
The molecular docking studies revealed a favorable binding affinity between Aloin A/B and the modeled protein structures from the species that showed the highest antimicrobial activity in the experimental study.
Aloin A/B achieved the highest docking score of −10.060 kcal/mol against DNA topoisomerase II of Fusarium oxysporum. The molecular docking analysis revealed important interactions, including hydrogen bonds, polar interactions, hydrophobic interactions, and charged interactions between aloin A/B and the amino acid residues of the enzyme. In Figure 3, aloin A/B exhibited five hydrogen bond interactions with DNA topoisomerase II of Fusarium oxysporum, specifically involving amino acid residues Ser 308, Asn 279, Asn 250, Asn 254, and Gly 283. Additionally, hydrophobic interactions were observed with Ile 286, Ile 300, Ala 251, and Pro 285. Charged interactions were also noted with Lys 316, Arg 257, Lys 282, and Asp 258.

Molecular docking interactions between Aloin A/B and the binding sites of FUSOX: (A) 2D model of the interactions; (B) 3D model of the interactions.
In Figure 4, the best conformation of Aloin A/B showed a docking score of −8.861 kcal/mol against murE of Xanthomonas vasicola. Aloin A/B formed five hydrogen bond interactions with polar amino acid residues Thr 153, Ser 180, His 206, and Asn 409. Charged interactions were also observed with Glu 463, and a pi-pi interaction occurred with Arg 384. Additionally, hydrophobic interactions were also noted with Phe 152, Tyr 352, and Tyr 465.

Molecular docking interactions between Aloin A/B and the binding sites of murE: (A) 2D model of the interactions; (B) 3D model of the interactions.
As shown in Figures 2 and 3, the 2D model representations of molecular docking interactions between aloin A/B and the binding sites of murE and DNA topoisomerase 2 indicate that the highest binding affinities for both enzymes occur with the S-configuration at C10, corresponding to aloin B.
Different parts of A. pirottae have been previously investigated for their effects on various human ailments. However, this is the first time it has been studied against plant pathogens. In the present study, a mixture of the two diastereomeric anthrones, aloin A and aloin B, was isolated, characterized, and evaluated against plant pathogenic microbes. Anthrones are among the predominant secondary metabolites present in Aloe species. 35 Aloin A/B is commonly found in the leaf latex of several Ethiopian Aloe species, including Aloe sinana, 36 Aloe harlana, 37 Aloe trichosantha, 38 and Aloe trigonantha. 39
In this study, aloin A/B demonstrated antimicrobial activity against plant pathogenic microbes. This antimicrobial activity was further supported by its high binding affinity to DNA topoisomerase II of Fusarium oxysporum, evidenced by a docking score of −10.060 kcal/mol. DNA topoisomerase II, also known as DNA gyrase in bacteria, is essential for maintaining the structural stability of the bacterial chromosome, facilitating critical DNA processes, and ensuring proper cellular functions. 40 The information from key interactions (hydrogen bonds, hydrophobic interactions, π-π stacking) that contribute to binding affinity can be used to modify the lead compound to enhance these interactions. For example, functional groups can be added or substituted to strengthen interactions with the target enzyme or to improve pharmacokinetic properties.
Anthrones, such as aloin A/B, have been reported to disrupt the integrity of microbial cell walls, contributing to their antimicrobial activity. 36 Similarly, anthraquinones, which are structurally similar to anthrones and isolated from Aloe barbadensis, have been shown to inhibit bacterial protein synthesis by binding to the ribosomal A site. 41 Additionally, both anthraquinone and 1,8-dihydroxyanthraquinone inhibit the growth of human and plant pathogenic microbes by targeting the cell membrane and crucial enzymes involved in their metabolic processes.42–44 Several studies also support the anthraquinones inhibitory activity on various cell lines through binding on topoisomerase II enzyme. 45 Anthraquinones prevent metabolic transformation of fungal aflatoxins to toxic derivatives or promoting their conversion to nontoxic products. 46 Anthraquinone inhibits DNA topoisomerase II and promotes DNA double-strand breaks through the stabilization of the topoisomerase II–DNA cleavage complex ultimately inhibiting ATP hydrolysis.29,47
Aloin A/B shows promise in combating plant pathogens, but several limitations still need to be addressed. The effectiveness of aloin A/B in real-world agricultural settings has yet to be thoroughly investigated. Its spectrum of activity against various pathogens is unclear, and the potential for resistance development over time is not addressed. Additionally, the potential toxicity to non-target organisms and the environmental impact of large-scale use are not considered. Addressing these limitations through further research is essential for validating aloin A/B as a viable agricultural antimicrobial agent.
In the end, given the high concentration of aloin A/B in the leaf latex of A. pirottae and its presence in several other Ethiopian Aloe species, aloin A/B holds promising potential for combatting plant pathogens in Ethiopia.
Conclusion
This study demonstrated that the leaf latex of A. pirottae has antibacterial and antifungal properties against plant pathogenic bacteria and fungi in an in vitro model. The results from the assays on plant pathogenic bacteria and fungi indicate that the antimicrobial activity of A. pirottae is, at least in part, due to the presence of aloin A/B. The antimicrobial activity of aloin A/B was evident from molecular docking studies, which showed a strong binding affinity of aloin A/B towards topoisomerase II in Fusarium oxysporum and UDP-N-acetylmuramoyl-L-alanyl-D-glutamate-2,6-diaminopimelate ligase (murE) in Xanthomonas vasicola. However, the greater antimicrobial activity of the leaf latex compared to aloin A/B against both plant pathogenic bacteria and fungi could be due to the higher availability of phytochemicals in the leaf latex, along with their diverse mechanisms or synergistic activities. Finally, this study provides insights into potential candidates for the proposed molecular targets, based on the binding affinities of aloin A/B. However, experimental characterization is crucial for further validating these protein targets. In the end, the present findings highlight aloin A/B as promising natural antimicrobial agents and lead compounds for new plant pathogen treatments. Further research is recommended to explore their activity against a wider range of plant pathogens.
Materials and Methods
General
The 1H-NMR (400 MHz), 13C-NMR (100 MHz), DEPT-90, and DEPT-135 were run on a Bruker Advance DMX400 FT-NMR spectrometer instrument using tetramethylsilane (TMS) as internal standard. Electrospray Ionization mass spectrum (ESI-MS) was generated using Ultimate 3000 LC-MS instrument, Ultraviolet Spectrophotometer (UV) (Cammag, Switherland), Normal phase column chromatography, pre-coated plates with silica gel 60 F254 (Merck KGaA, Darmstandt, Germany) for analytical thin layer chromatography (TLC).
Plant Material
The leaf latex of A. pirottae A. Berger was collected in November 2018 from Erer valley of Babile Wereda, East Hararge zone, Oromia region, Ethiopia. The plant was authenticated by Dr Anteneh Desta at the National Herbarium and given a voucher herbarium specimen of AP01.
Extraction and Isolation
The latex from the leaves of A. pirottae was collected using the methods described in our earlier study.
48
The yellow sap was collected by cutting the fresh leaves of A. pirottae at the base. The collected sap was then left to stand for a week to allow water to evaporate and the sap to dry, resulting in the formation of the leaf latex. Silica gel column chromatography was employed to isolate compounds from the leaf latex of A. pirottae. The column was initially packed with slurry consisting a mixture of silica gel and chloroform. The methanol soluble portion of the leaf latex (2 g) was then adsorbed on silica gel and loaded onto the column. A total of 100 fractions, each 10 mL, were collected using a gradient elution of chloroform and methanol. The first fraction was eluted with 100% chloroform, and continued with 5% increments of methanol in chloroform. The purity of the fractions was monitored using TLC with CHCl3/MeOH (4:1 v/v) as a solvent system. Fractions 89–95, eluted with a CHCl3 and MeOH (9:1) mixture, yielded a yellowish amorphous solid substance (185 mg) and coded as compound
Compound
Antimicrobial Activity
Bacterial and Fungal Strains
Bacterial strains (Pseudomonas syringae pv. gaarcae, Xanthomonas vasicola pv. musacearum, and Ralstonia solanacearum) and fungal strains (Fusarium graminarum, Fusarium oxysporum, and Fusarium solani) were sourced from the Ambo Agricultural Research Institute in Ambo, Ethiopia.
Antibacterial Activity Test
The antibacterial activity of the test samples were evaluated using the agar well diffusion method. 49 Sterile nutrient agar plates were prepared and the bacteria were spread over the plates using a sterile swab. Four wells each of 6 mm diameter were punched into the plates using separate sterile borer, and the test samples (100 μL) with three concentrations (50, 25 and 12.5 mg/mL) in 2% dimethyl sulfoxide (DMSO) were added to their respective wells using sterile micropipette. A standard drug streptomycin sulfate (10 µg) disc carefully placed onto the middle part of the media50,51 and 2% DMSO (100 μL) was added as a negative control. All the plates were then incubated for 24 h at 30 °C, and the resulting zone of inhibitions were measured by digital caliper in mm. All assays were conducted in triplicate.
Antifungal Activity Test
The antifungal activity of the test samples was assessed by the poisoned food technique. The test samples (500 μL) of three concentrations (50, 25 and 12.5 mg/mL) were spread evenly onto three different previously prepared and solidified sterile potato dextrose agar (PDA) plates using a sterile glass spreader. A mycelial disc of 0.5 cm in diameter cut out from the 7-day old fungal culture aseptically placed on the center of different PDA plates containing test samples, negative (DMSO and PDA) and positive control (tebuconazole; 25 μg/mL), respectively. The inoculated plates were incubated at 26 ± 1 °C for 7 days and the mycelial growth inhibition rate was calculated on the seventh day.
52
Rate of mycelial growth inhibition was calculated as follows: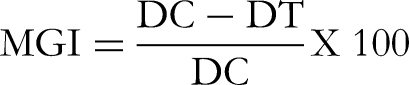
Determination of Minimum Inhibitory Concentrations (MICs)
MICs of the test samples were determined using the Broth dilution method. 53 To prepare the test samples, the latex and isolated compound were dissolved in 2%DMSO solution. The concentrations tested ranged from 50 mg/mL to 0.098 mg/mL for the latex and 25 mg/mL to 0.049 mg/mL for the isolated compound.54,55 Streptomycin sulfate (10 μg/mL) and 2%DMSO both in Mueller Hinton Broth (MHB) were used as a positive and negative control respectively for antibacterial assay. 56 Tebuconazole (25 μg/mL) and 2%DMSO both in potato dextrose broth (PDB) were used as a positive and negative control respectively for antifungal assay. 17 The plates were covered with sterile lead and incubated at 30 °C for 24 h for bacteria and 25 °C for 48 h for fungi.18,57 After the designated incubation periods, ie 24 h for (bacteria) and 48 h (fungi), the MICs were assessed by adding 40 μL of a 2% solution of iodonitrotetrazolium (INT) to each well58,59 followed by a 25-min incubation at 30 oC for bacteria and 25 oC for fungi. The metabolically active bacteria and fungi would reduce the colorless test sample to a pink color, and the MIC was considered to be the lowest sample concentration that prevented this change. 18 The assay was conducted in triplicate.
Molecular Docking Studies
Homology Models Building and Validation
The crystal structures of the target enzymes were not available in the PDB. Consequently, homology models were created using SWISS-MODEL and a template library generated from a BLAST search of UniProt, as previously described. The closest crystal structures used as templates were murE of Xanthomonas vasicola (PDB: 8F5D) and DNA topoisomerase II of Fusarium oxysporum (PDB: 4GFH). The amino acid sequence of the DNA topoisomerase II and DNA gyrase subunit B proteins for Fusarium oxysporum and Xanthomonas vasicola was obtained in FASTA format from the Uniprot database UniProtKB (Universal Protein Resource Knowledgebase; http://www.uniprot.org) using the primary accession codes A0A8B5YVD4 and W9KBZ2, respectively. 60 The identification of suitable templates for homology modeling was conducted using the Basic Local Alignment Search Tool (BLAST) through SWISS-MODEL online workspace (https://www.swiss model.expasy.org). The reliability of the modeled 3D structure was assessed using the Qualitative Model Energy Analysis (QMEAN) and Global Model Quality Estimation (GMQE) scores. A QMEAN score below −4.0 indicated low quality of the predicted structure, while the GMQE score ranges between 0 and 1, with a higher score indicating higher reliability.33,34
Docking Study
For protein and ligand preparation, the homology modeled target proteins were downloaded in pdb format from the SWISS-MODEL workspace and prepared using the Protein Preparation Wizard module of Schrödinger suite. 61 The protein was prepared by adding hydrogens, fixing any errors such as missing side chains, assigning correct bond orders, adjusting ionization and tautomeric states (via Epik), and deleting water molecules beyond 5 Å. Optimization and subsequent minimization were performed on all atoms using OPLS3 force fields, with the convergence of heavy atoms set to an RMSD of 0.3 Å (default).
For ligand preparation, the compounds drawn in SD format using ChemDraw were imported and prepared using the LigPrep module, which added hydrogens, performed a 2D to 3D conversion, generated ionization and tautomeric states (via Epik) at physiological pH 7.0 ± 2.0, and determined ring conformations using default settings. The docking of the isolated compounds was conducted using the Glide program with the previously prepared receptor grid and ligand molecules. The favorable interactions between the ligand molecules and the receptor were scored using Glide, and all docking calculations were performed in extra precision (XP) mode and using the OPLS-2005 force field.
Statistical Analysis
Data analysis was carried out using IBM SPSS (Statistical Package for Social Sciences) Statistics for Windows, Version 20.0. Results were reported as mean ± standard error of mean (M ± SEM). The statistical significance was determined by one-way ANOVA followed by Tukey post hoc test to compare inhibition activity of the treatment and control groups. p < 0.05 was considered significant.
Supplemental Material
sj-docx-1-npx-10.1177_1934578X241281497 - Supplemental material for Antimicrobial and Molecular Docking Studies of Anthrones from the Leaf Latex of Aloe pirottae A. Berger against Some Plant Pathogenic Microbes
Supplemental material, sj-docx-1-npx-10.1177_1934578X241281497 for Antimicrobial and Molecular Docking Studies of Anthrones from the Leaf Latex of Aloe pirottae A. Berger against Some Plant Pathogenic Microbes by Shewaneh Ayele, Misgana Aragaw, Denberu Kebede, Bayoush Birke and Daniel Bisrat in Natural Product Communications
Footnotes
Acknowledgements
The authors are thankful to Ambo Agriculture and Research Center for their cooperation and the facilities provided for conducting the biological experiment.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval
Ethical approval is not applicable for this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The present study was financially supported by the School of Graduate Studies of Addis Ababa University research grant (AAU/SGS/Pharm-0022).
School of Graduate Studies of Addis Ababa University, (grant number AAU/SGS/Pharm-0022).
Statement of Informed Consent
There are no human subjects in this article and informed consent is not applicable.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
