Abstract
Introduction
Myocardial ischemia (MI) is a diseased myocardial state caused by an imbalance between muscular oxygen demand and coronary artery blood supply, 1 and it is one of the primary sources of human morbidity and mortality worldwide. 2 The cardiac is a high-energy consuming organ and the metabolism of energy substrates is critical to sustaining normal heart structure and function.3,4 Conversely, MI leads to disturbances in cardiac metabolism and intestinal flora that can affect the progression of the disease.
The intestinal flora is engaged in adjusting diverse functions in the body, such as supplying metabolic substances to the organism, being involved in growth and immune regulation, clearing pathogenic microbes, and sustaining the integrity of the gut barrier and internal homeostasis. 5 The intestinal microbiota is associated with the progression of different human diseases. Human and animal studies have shown that alterations in the composition and function of the intestinal flora, thought to be dysbiosis of the gut flora ecology, can accelerate the progression of cardiovascular disease. 6 The intestinal microbiota exerts its biological effects through the generation of certain metabolites that work on the gut barrier or pass into the blood to mediate the target organs.7,8 However, the mechanism of action by which gut flora affects MI is unclear.
Herbal/herbal preparations can be used as probiotic bacteria to regulate the intestinal flora composition and metabolic phenotype of the host, and as a new potential source of drug precursors to manage intestinal flora target diseases.9-11 Fuling-Guizhi (FLGZ) is a well-known pair of herbs with good anti-inflammatory and antioxidant properties used to treat different cardiovascular diseases, such as MI and arrhythmias. 12 Studies have shown that Fuling can add diversity to the intestinal flora, mainly by adding the helpful bacteria Lactobacillus, Bifidobacterium and reducing the Desulfovibrio and the inflammation-related bacteria Mucospora, and Staphylococcus. 13 The active ingredient of Guizhi has good anti-inflammatory and immunomodulatory properties, which can change the dysbiosis of intestinal microbial ecology. 14
However, the interaction between MI and gut flora, and the mechanism of FLGZ modulation of gut flora in MI is not clear. Therefore, this study combined blood metabolomics and 16s rRNA gene sequencing analysis to further investigate the alteration of intestinal flora in MI rats comprehensively and the effect of FLGZ on gut flora in MI.
Materials and Methods
Extraction of FLGZ Solution
FLGZ decoction was made by the Ministry of Education Key Laboratory of Traditional Chinese Medicine Resources and Traditional Chinese Medicine Compounding, Hubei University of Chinese Medicine. The filtrate was mixed by 2 decoctions of 2 kinds of tablets, Fuling and Guizhi, in a mass ratio of 1:1, and concentrated into a concentrate containing 0.15 g of raw herbs per mL, which was stored in a refrigerator at −20 °C. To ensure the comparability of the experimental results, the same batch of herbs was used for the experiments of the same sample.
Animal
Male Sprague-Dawley (SD) rats, 200 to 240 g (6-8 weeks), acquired from Hubei Animal Research Center (Certificate of Conformity No. SCXK (E) 2015–0018). All rats were placed in the Specific Pathogen Free (SPF) experimental animal room at the Hubei University of Chinese Medicine with a room temperature of 22 ± 2 °C, the humidity of 50 ± 10%, and a light/dark cycle of 12 h. Rats were randomly divided into four groups: control check (CK), model (MX), propranolol (XDAZ), and FLGZ groups, with eight rats in each group. Except for the CK group, the rats in all groups were injected subcutaneously with ISO (0.04 mg.kg−1.d−1) for six days. 15 Equal amounts of saline were administered to the CK and MX groups, propranolol (20 mg.kg−1.d−1) to the XDAZ group, and FLGZ (1.5 g.kg−1.d−1, converted by body surface area) to the FLGZ group, and all rats were administered once daily for 14 days. After the final gavage administration, rats were anesthetized by intraperitoneal injection of 2% sodium pentobarbital, blood was taken from the abdominal aorta, blood was centrifuged to separate the serum, and the serum was stored frozen at −20 °C for metabolomic analysis.
Histopathological Analysis
Cardiac tissues were fixed in 4% paraformaldehyde and stored in a 4 °C refrigerator. Processing was performed by dehydration, embedding, and sectioning with staining steps. The morphological structure of every sample was inspected with an optical microscope (BH-2, Olympus).
16s rRNA Gene Sequencing
DNA Extraction
We used the OMEGA Soil DNA Kit (D5625-01) (Omega Bio-Tek) kit to extract nucleic acids. The quantity and quality of the extracted DNA were evaluated by Bio Spectrometer UV spectrophotometer (Eppendorf) and 0.8% agarose gel electrophoresis, respectively.
16S rRNA Gene Amplicon Sequencing
PCR amplification of the V3-V4 region of the bacterial 16S rRNA gene was carried out using forward primer 338F (5ʹ-ACTCCTACGGGGAGGGCAGCA-3ʹ) and reverse primer 806R (5ʹ-GGACTACHVGGGTWTCTAAT-3ʹ). Sample-specific 7-bp barcodes were merged into the primers for multiplex sequencing. PCR products were quantified using the Quant-iT PicoGreen dsDNA Assay Kit on a Microplate reader (Bio-Tek, FLx800), and then mixed according to the amount of data required for each sample. Sequencing libraries were built using Illumina's TruSeq Nano DNA LT Library Prep Kit, and the NovaSeq 6000 SP Reagent Kit (500 cycles) was used on an Illumina NovaSeq machine for 2 × 250 bp double-end sequencing.
Sequence Analysis
Microbiome bioinformatics was performed using QIIME2 2019.4, 16 with raw sequence data demultiplexed using the demux plugin and then primer cleavage was performed using the cutadapt plugin. The sequences were quality filtered using the DADA2 plugin. 17 Nonmonoclonal amplicon sequence variants (ASVs) were removed using mafft for comparison and constructed phylogenetically using fasttree2.
The alpha-diversity of the samples was characterized by abundance (Chao1) diversity (Shannon, Simpson), and coverage (Good's coverage). Beta-diversity analysis (PCoA, UPGMA) was carried out with the Bray-Curtis distance algorithm. Taxa rich in between-group differences were detected by linear discriminant analysis of effect sizes (LEfSe). 18 In addition, PICRUSt2 19 predicted microbial functions on the basis of the Kyoto Encyclopedia of Genes and Genomes (KEGG) database and visualized the pathways of microbial metabolism by R packages.
Metabolomics Analysis
Sample Handling
Serum samples stored at −20 °C were thawed at room temperature, 100 μL was aspirated and placed in a 1.5 mL centrifuge tube, 1 mL of methanol was added, and mixed well. Aspirate 600 μL of serum homogenate into a centrifuge tube, vortex for 1 min, sonicated in an ice bath for 15 min, spin-off (13 000 r/min) for 10 min, then pipette 200 μL of supernatant into a sample bottle and concentrate by centrifugation in a fast centrifuge concentrator and desiccator. Add 10 μL of D27-myristate (0.75 mg/mL) followed by 40 μL of methoxy pyridine solution (40 mg/mL) and react for 90 min at 30 °C on a shaker. Add 80 μL MSTFA + 1% TMCS derivatization reagent and react for 60 min. After cooling to room temperature, 20 μL of hexane was added to terminate the derivatization reaction, centrifuged, and the supernatant was withdrawn into a microsampling tube for gas chromatography-mass spectrometry (GC-MS).
Metabolic Data Analysis
Raw data acquisition for GC-MS was carried out with Agilent Masshunter software (Agilent Technologies, Inc.). The spectra were deconvoluted using the Automated Mass Spectrometry Resolution and Identification System (AMDIS) tool from the National Institute of Standards and Technology (NIST) library. Metabolites with P values < .05 and VIP > 1 were regarded as differential metabolites. Principal component analysis (PCA), partial least squares-discriminant analysis (PLS-DA), relative intensity, and pathway analysis were analyzed using MetaboAnalyst 5.0 website (https://www.metaboanalyst.ca/). R2 indicates the degree of model fit and Q2 indicates the percentage of the model prediction. The closer the values of R2 and Q2 are to 1, the more well the model is built. The differential metabolite heat map was constructed using the Genescloud platform (https://www.genescloud.cn).
Correlation Analysis Between Serum Differential Metabolites and Microorganisms
Pearson correlation analysis is a valid method to assess the relationship between differential metabolites and variable genera. According to the correlation coefficient (|r| > .5), strong correlations were displayed as heat maps (*P < .05, **P < .01).
Statistical Analysis
GraphPad Prism 8.0 statistical software was used to analyze and process the data, and the results were expressed as the mean ± SD. Independent samples t-test was applied for inter-group comparison, one-way ANOVA was also used for multi-group comparisons, and differences were deemed statistically meaningful at P < .05. The flowchart for the integration of metabolomics and gut flora studies is shown in Figure 1.

Flowchart of integration of metabolomics and gut flora studies.
Results
Evaluation of MI Models and Efficacy of FLGZ
The criteria for MI were ST-segment elevation or depression. The electrocardiogram (ECG) in Figure 2A shows that the ST segment of the CK group remained at the horizontal level, and the mean voltages of the MX group were shifted higher, approaching 0.2 mV, indicating that the MI model was constructed successfully.20,21 Compared with the MX group, the ST segment voltage of propranolol and FLGZ group was close to that of the CK group, indicating the efficacy of FLGZ on MI.

(A) FLGZ on MI electrocardiogram in all groups of rats. (B) Representative H&E stained images of paraffin sections of each group of hearts.
H&E stained sections of myocardial tissue in the CK group showed regular cell arrangement, clear demarcation, and no obvious abnormalities in morphology and structure; compared with the CK group, myocardial cells in the MX group disappeared in several places and were replaced by proliferating connective tissue with a small amount of neovascularization within the connective tissue, associated with a few lymphocytic infiltrates and minor localized hemorrhage; compared with the MX group, The XDAZ group showed that the localized cardiomyocytes in the tissue disappeared and were replaced by a few proliferating connective tissues with a small infiltration of lymphocytes, and no other significant abnormalities were seen; after FLGZ treatment, tissue cardiomyocytes were regularly arranged, clearly delineated, some cardiomyocytes had vacuolar degeneration, tiny round vacuoles were seen in the cytoplasm, no other obvious abnormalities were seen. The above pathological changes were significantly reduced after FLGZ treatment, indicating that FLGZ has certain efficacy on MI (Figure 2B).
The FLGZ composition (Supplemental Table S1) was studied in our previous work. 22
Alpha Diversity Analysis
To validate whether the quantity sequenced in our research was sufficient to reflect the raw microbial diversity, we performed an Alpha diversity analysis on the 16S rRNA sequencing results. A bigger Chao1 index indicates a higher species richness. Higher Shannon and Simpson indices indicate higher species diversity. Coverage is applied to gauge OTU coverage and indicates if the sequencing outcome represents the real status of microorganisms in the sample. Supplemental Figure S1 shows that the rarefaction curve gradually smooths out with increasing sampling volume in a certain range, suggesting that the sequencing volume per sample is sufficient. Supplemental Figure S2 Chao1 index shows that the FLGZ group had the highest bacterial flora abundance, followed by the CK and MX groups; Shannon and Simpson indices showed that species diversity was higher in the FLGZ group than in the CK and MX groups. This also suggests that FLGZ influences the abundance and diversity of intestinal flora in MI rats. The coverage index indicated that all sample libraries were > 99, suggesting that the scale of the libraries in this research was adequate to reflect the majority of bacteria.
Species Annotation Analysis
Figure 3A and B displays the results of species annotation of the CK, MX, and FLGZ groups at the phylum and genus levels obtained from 16S rRNA sequencing analysis. The figure shows only the 10 species of bacteria with the highest abundance at the phylum and genus level, and combines the remaining species as “others.” At the phylum level, Firmicutes and Bacteroidetes were the major phyla in every group of rat intestinal bacteria. In Figure 3A, in contrast to the CK group, the MX group displayed a decline in Firmicutes and an increase in the Bacteroidetes, Proteobacteria, and Actinobacteria. In contrast, compared to the MX group, the FLGZ group displayed an increase in Firmicutes and a decline in Proteobacteria and Actinobacteria.

Histogram of species distribution of CK, MX, and FLGZ groups at the phylum (A) and genus (B) levels (colors indicate various phylum or genus levels of bacteria). PCA between CK, MX, and FLGZ groups (C). CK, MX, and FLGZ groups hierarchical clustering analysis (D).
Beta Diversity Analysis
As shown in Figure 3C, a significant separation of intestinal flora was observed between the CK, MX, and FLGZ groups. Hierarchical cluster analysis using the Unweighted pair-group method with arithmetic means (UPGMA) showed significant clustering of the CK, MX, and FLGZ groups in the respective groups (Figure 3D). The closer the samples and the shorter the branches, the more similar the species composition of the two samples.
Screening of Significantly Altered microbiota
We performed a LEfSe with LDA collapse = 4. The LDA score plot (Figure 4A) showed that Phascolarctobacterium was the dominant genus in the CK group; Allobaculum and Ruminococcus were the dominant genus in the MX group compared to the CK group; Oscillospira was the dominant genus in the FLGZ group. The species composition heat map (Figure 4B) shows the clustering of the top 20 genera at the genus level, showing that the vast majority of genera returned to the normal group level, also indicating the efficacy of FLGZ on the intestinal flora of MI rats.

Linear discriminant analysis of effect sizes (LEfSe) for differences in microbiota composition between the three groups, visualized by cladogram (A), and species composition heat map (B).
Differential Microbial Functional Analysis
KEGG functional analysis was conducted on the obtained differential flora. The findings revealed that these microorganisms are primarily associated with the metabolism of amino acids, carbohydrates, energy, lipid, cofactors and vitamins, other amino acids, terpenoids, and polyketides (Figure 5).

Predictive analysis of Kyoto Encyclopedia of Genes and Genomes (KEGG) function in microorganisms with genus-level differences.
Serum Metabolomics Analysis
We performed a PLS-DA analysis of the GC-MS data obtained (Figure 6A) to determine metabolite differences between the three groups. Serum metabolic profiles showed good separation between the CK, MX, and FLGZ groups. In addition, further PLS-DA analysis also showed a clear separation between the three groups (Figure 6B and C). Cross-validation in the PLS-DA analysis indicated that the model was reliable (Figure 6D). According to VIP > 1 and P < .05, 14 different metabolites were identified by two comparisons between CK-MX and MX-FLGZ groups (Table 1), while differential metabolites were also significantly different in the ion flow diagrams of the three groups (Figure 7).

PCA plots for CK, MX, and FLGZ groups (A). PLS-DA score plots for CK with MX and MX with FLGZ groups (B, C). R2 and Q2 of PLS-DA (D).
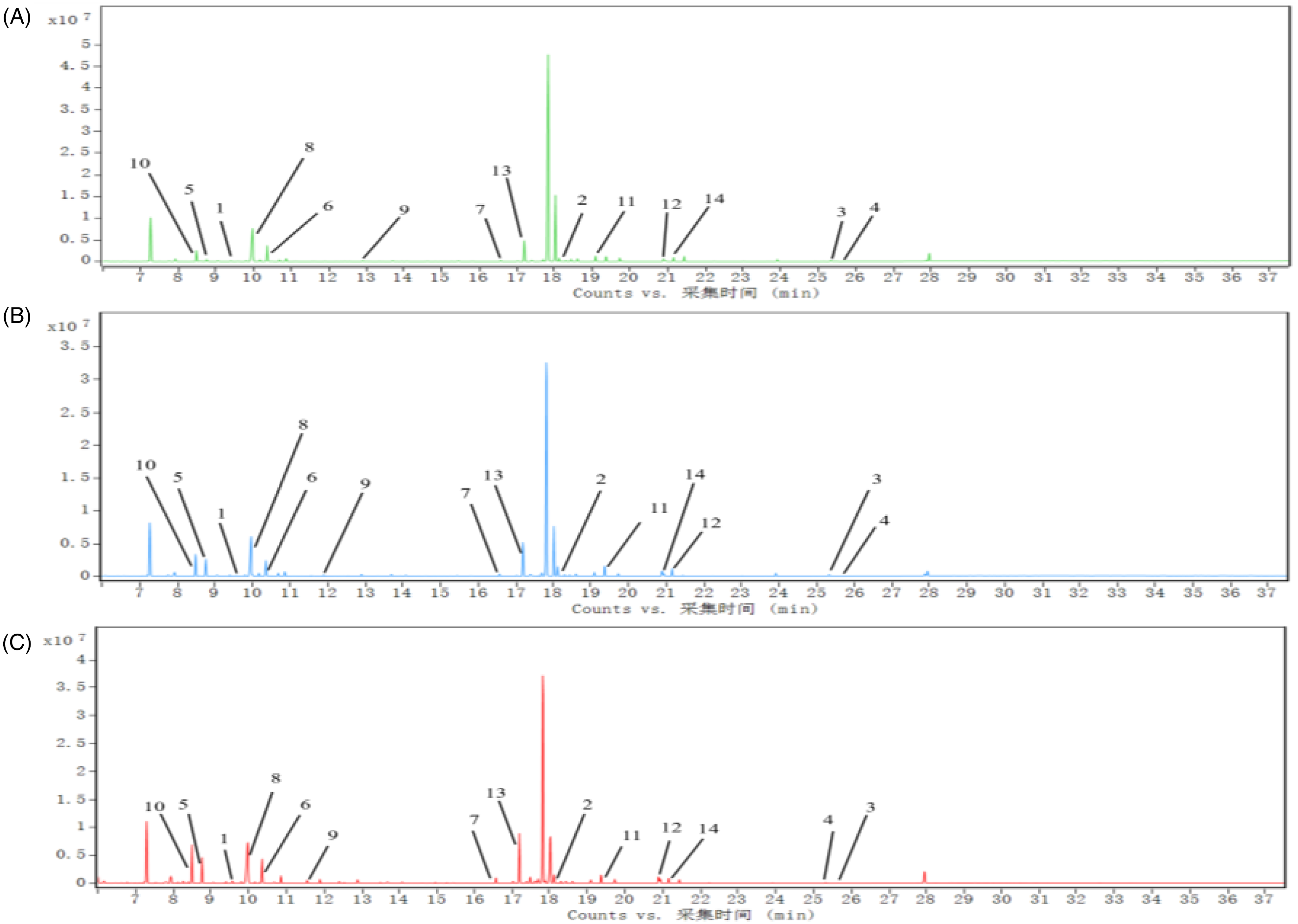
Plots of total ion flow of metabolites obtained from rat sera of CK (A), MX (B), and FLGZ groups (C). The x-axis indicates retention time and the y-axis indicates peak abundance.
Identification of Potential Biomarkers.

Abbreviations: FLGZ, Fuling-Guizhi; CK, control check; MX, model; VIP, variable importance in projection.
Metabolic Pathway Analysis
The results of metabolic pathway analysis revealed that the main metabolic pathways of FLGZ-treated MI (P < .05) were unsaturated fatty acid biosynthesis, linoleic acid metabolism, nitrogen metabolism, and aminoacyl tRNA biosynthesis (Figure 8A, Supplemental Table S2). The color and size of the circles in the figure are based on the P-value and the impact value. The heat map of the differential metabolite interaction origin showed that the FLGZ group's differential metabolite levels were consistent with the CK group (Figure 8B).
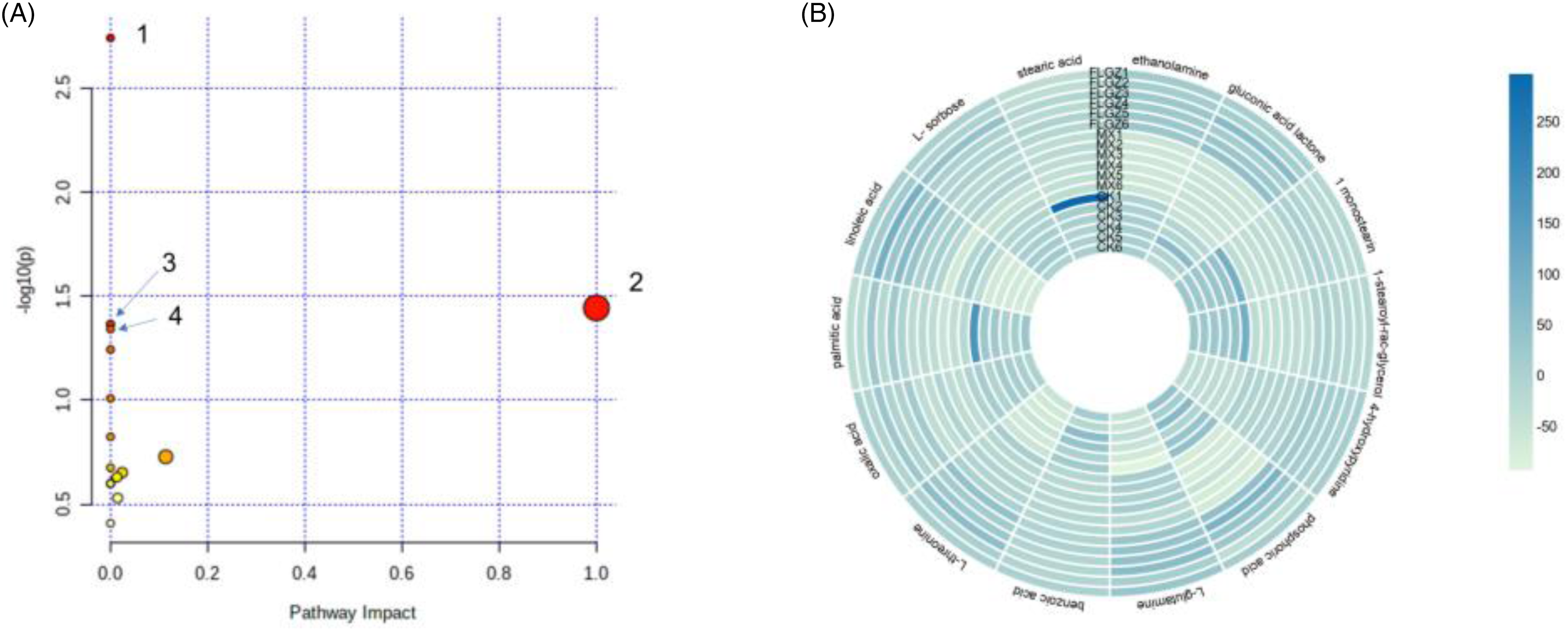
Pathway analysis of differential metabolites using MetaboAnalyst (A). Key pathways include 1—biosynthesis of unsaturated fatty acids; 2—linoleic acid metabolism; 3—nitrogen metabolism; and 4—aminoacyl-tRNA biosynthesis. Heat map of differential metabolite interactions (B).
Correlation Analysis of Differential Metabolites and Microbes
Pearson correlation analysis was performed to further reveal the correlation between serum differential metabolites and gut microbes. The correlation coefficient between differential metabolites and gut microbiota is shown in Figure 9A. Phascolarctobacterium is positively correlated with linoleic acid, L-glutamine, L-threonine, and 4-hydroxypyridine. It was negatively correlated with 1-monostearin and 1-stearoyl-rac-glycerol. Oscillospira was negatively correlated with ethanolamine, phosphoric acid, and oxalic acid. Allobaculum, Rminococcus, ethanolamine, phosphoric acid, and oxalic acid were positively correlated (Figure 9B).

Correlation between genus-level gut microbiota and serum differential metabolites. Pearson correlation heat map: positive correlation (red), negative correlation (blue), *P < .05, **P < .01 (A). Network diagram of gut microbiota–metabolites based on the correlation analysis, with red (blue) lines indicating the positive (negative) correlation between gut microbiota and metabolites (B).
Discussion
Metabolomic results showed that the differential metabolites were mainly associated with disorders of amino acid, carbohydrate, lipid, and energy metabolism (Supplemental Table S3).
Amino Acid Metabolism
We identified two amino acids (L-glutamine and L-threonine) from the serum differential metabolites. Studies have shown that glutamine is also required for glucose metabolism to the hexosamine biosynthetic pathway (HBP) for glycosylation of nuclear and cytoplasmic proteins. 23 The protection of L-glutamine against myocardial ischemia is mediated by an enhanced flux of HBP, leading to increased levels of O-linked N-acetylglucosaminoglucose (O-GlcNAc) on nuclear-cytoplasmic proteins, and increased levels of O-GlcNAc reduce ischemia-reperfusion-induced cell death. 24 Amino acids have the potential for non-oxidative metabolism, and by supplementing cells with glutamate and glutamine, cellular function during hypoxia can be prolonged and post-hypoxic recovery improved because these specific amino acids can be readily converted to α-ketoglutarate, which not only maintains levels of Krebs cycle intermediates but also provides cellular energy through substrate level phosphorylation during conversion to succinate. 25 Thus, cardiac ischemia and cellular injury were predicted to be followed by changes in amino acid fluxes during and after the ischemic episode.
Carbohydrate Metabolism
The metabolite associated with carbohydrate metabolism is oxalic acid. Clinical studies have shown that oxalate is a potential trigger for systemic inflammation, and cardiovascular complications and that patients with higher serum oxalate concentrations are more likely to experience cardiovascular death than patients with lower serum oxalate levels. 26
Lipid Metabolism
Metabolites associated with lipid metabolism are ethanolamine, linoleic acid, ethanolamine, palmitic acid, and stearic acid were downregulated in the MX group and upregulated in the FLGZ group. Ethanolamine is a bioamine that exists natively in vivo as a portion of membrane lipids. Signal transducer and activator of transcription 3 (STAT-3) is a critical promoter of MI, and ethanolamine has a new cardioprotective effect against MI injury by activating STAT-3. 27 MI is related to the formation and action of lipid mediators derived from polyunsaturated fatty acids, and the linoleic acid-derived mediator epoxyoctadecanoic acid (EpOME) may have an essential effect in MI. 28 The dynamic changes in metabolites suggest that the development of myocardial ischemia and disturbances in lipid metabolism are closely related.
Energy Metabolism
Among the differential metabolites related to energy metabolism are mainly phosphoric acid, which is involved in the oxidative phosphorylation pathway (OXPHOS), and oxygen-depleted mitochondrial OXPHOS is the physiological tool for ATP generation in cardiac. 29 The heart has a very high energy requirement and must continuously produce large amounts of ATP to maintain its contractile function, while MI causes the heart to lose its metabolic flexibility and become energy deficient due to its reduced ability to produce ATP.30,31
In the species annotation analysis, both PCA and compositional structure showed significant differences between the three groups (Figure 3A to C). Research has shown that the onset of cardiovascular disease can cause dysbiosis in the ecology of the intestinal flora. 32 Proteobacteria and Actinobacteria are markers of cardiovascular disease risk, with increased proportions of Proteobacteria and Actinobacteria found in cardiac events such as stroke, coronary artery disease, and myocardial infarction, 33 which also occurred in the MX group.
By combining gut microbiota and differential metabolites, the results indicate that gut microbes and host metabolites are closely related in myocardial ischemic rats. As shown in Figure 4A, the microbially related genera predicted from each differential metabolite were Phascolarctobacterium, Oscillospira, Allobaculum, and Ruminococcus. It has been shown that Phascolarctobacterium and TMAO levels are positively correlated, 34 however trimethylamine N-oxide (TMAO) is involved in the mechanisms by which intestinal flora affect cardiovascular disease,35-37 TMAO is a metabolite of dietary choline and other trimethylamine-containing intestinal microorganisms that accelerate the process of LV dilation, cardiac fibrosis, and LV remodeling. 38 In our study, six metabolites were statistically significant with Phascolarctobacterium, including 1-monostearin, 1-stearoyl-rac-glycero, 4-hydroxypyridine, L-glutamine, L-threonine, and linoleic acid, many of which were associated with MI.
Related studies have shown that patients with MI have increased concentrations of pro-inflammatory biomarkers, such as C-reactive protein (CRP), interleukin 6 (IL-6), and white blood cells (WBC), 39 while Allobaculum improves the damaged gut, reduces lipopolysaccharide production and circulation, and decreases IL-6 and interleukin 12 (IL-12) expression in the myocardium. 40 Previous studies have found an association between Ruminococcus and inflammatory markers; in addition, Ruminococcus abundance was significantly associated with TMAO levels and was higher in patients with atherosclerotic cardiovascular disease (ACVD) than in controls. 41 Oscillospira may be a next-generation probiotic candidate with weight loss, lipid-lowering, bariatric, and metabolic syndrome-relieving effects, and has great potential for health applications. 42 It is hypothesized that Oscillospira may produce the short-chain fatty acid butyrate, 43 which is capable of reducing I/R-induced inflammation, oxidative stress, and apoptosis, 44 and numerous clinical research demonstrated a negative correlation between the abundance of australoplasma and inflammation.45-48 As shown in Figure 10, the abundance of Phascolarctobacterium, Allobaculum, and Ruminococcus were all higher in the model group than in the FLGZ group, and all of these microorganisms were associated with inflammation, which also suggests that the ISO-induced MI rat model was related to the inflammatory factor release. In contrast, Oscillospira richness was higher in the FLGZ group than in the MX group, indicating that inflammatory factors were downregulated by FLGZ intervention, which is consistent with the previously described result that Oscillospira is negatively associated with inflammation. The main metabolic pathways associated with these microorganisms are the metabolism of amino acids, carbohydrates, energy, lipids, cofactors and vitamins, other amino acids, terpenoids, and polyketides, consistent with metabolite association pathways.

Box line diagram of the difference test for intestinal flora.
Conclusions
In this study, 16S rRNA gene sequencing was combined with metabolomics to evaluate the effects of FLGZ on intestinal flora and serum metabolites. The results showed that FLGZ has an effect on the composition of intestinal microorganisms (Phascolarctobacterium, Oscillospira, Allobaculum, and Ruminococcus) as well as on metabolic pathways (amino acid, carbohydrate, lipid, and energy metabolism). Therefore, FLGZ may affect amino acid, carbohydrate, lipid, and energy metabolism through Phascolarctobacterium, Oscillospira, Allobaculum, and Ruminococcus, thereby attenuating inflammatory responses, oxidative stress, and apoptosis, and further studies on key microorganisms and metabolites will be conducted later.
Footnotes
Abbreviations
Authors’ Contributions
Fang Huang and Bailu Duan conceived and designed the study. Bailu Duan, Yan Ye, Xiangfa Zeng, Zhenxiang Zhou, and Lintao Han analyzed the data. Yan Ye and Jianghua Liu wrote the manuscript. Bailu Duan, Yan Ye, Ping Huang, Fengyun Zhang, Jianghua Liu, Dan Zheng, Wusheng Wang, and Xuan Cheng conducted the experiments.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The project was supported by the Guiding Project of the Scientific Research Program of the Hubei Provincial Education Department (B2021117).
Ethical Approval
Ethical approval is not applicable to this article.
Statement of Human and Animal Rights
All experiments were conducted under the guidelines established by the Animal Ethics Committee of the Hubei University of Chinese Medicine (HUCMS-201903001).
Statement of Informed Consent
There are no human subjects in this article and informed consent is not applicable.
Supplemental Material
Supplemental material for this article is available online.
