Abstract
Alizarin (1,2-dihydroxyanthraquinone) is a natural red dye extracted from the roots of
Introduction
Alizarin (1,2-dihydroxyanthraquinone), a natural red dye extracted from the roots of
The aryl hydrocarbon receptor (AhR) is the transcription factor which belongs to the basic-loop-helix/Per-ARNT-Sim family. AhR is activated by aromatic hydrocarbons, including benzo[

The structural formulas of alizarin and other AhR ligands, including BaP, TCDD, ITE.
Recently, AhR has been reported to have multiple functional roles in the development of the organ system, immunity, and reproduction. Impaired production of AhR ligand is associated with glucose metabolic syndrome and hepatic steatosis; this highlights that restoration of AhR signaling by the AhR ligand induces beneficial effects on the metabolic syndrome. AhR signals induce interleukin (IL)-22 production in AhR-expressing innate and adaptive T cells. 19 The AhR protein and transcripts are present in ovarian cell types of several species, indicating its role in ovarian function. 20 Evidence supports the hypothesis that AhR is involved in the regulation of the oocyte apoptosis rate in germ cell nests during embryonic development and oocyte survival in the fetal and neonatal ovary. 21 Therefore, identification of the AhR ligand or agonist is important in AhR-linked physiological pathways. Here, we have studied the mechanism and pathway of AhR induced by alizarin via the expression of CYP1A1 nuclear translocation of AhR, activated AhR signaling by monitoring the induction of the AhR target gene, and the CYP1A1 expression in hepatocellular carcinoma cells, including Hepa-1c1c7 and HepG2 cells.
Materials and Methods
Materials
Alizarin, BaP, 1-methyl-N-[2-methyl-4-[2-(2-methylphenyl) diazenyl]phenyl-1H-pyrazole-5-carboxamide (CH-223191), Hoechst 33342, antibody (Ab) against β-actin, and other chemical reagents were obtained from Sigma-Aldrich (St. Louis, MO, USA). Fetal bovine serum (FBS), minimum essential medium α (MEM), Dulbecco's modified eagle medium-nutrient mixture F12 (DMEM-F12), phosphate-buffered saline (PBS), antibiotics, and other culture media were obtained from Invitrogen (Grand Island, NY, USA). Antibodies that include CYP1A1, glyceraldehyde 3-phosphate dehydrogenase (GAPDH), and AhR were purchased from Santa Cruz Biotechnology (Santa Cruz, CA, USA). Horseradish peroxidase (HRP)-conjugated secondary antibodies were obtained from Abcam (CA, USA). All chemicals were dissolved in dimethylsulfoxide (DMSO), with the final concentration less than 0.5% (v/v).
Cell Culture
The human hepatoma cell line, HepG2 cells, and the murine hepatoma cell line, Hepa-1c1c7 cells, were obtained from the ATCC (Rockville, MD). The C4 (B13NBii1) cell line, derived from Hepa-1c1c7 cells, is the dysfunction of AhR nuclear translocator protein (ARNT) due to a point mutation (Gly-326 to Asp) in the
Quantitative Reverse Transcription Polymerase Chain Reaction (qRT-PCR)
After the cells were incubated with alizarin at the indicated concentration and duration, total RNA was isolated with NucleoZOL reagents. Reverse transcription was performed to convert RNA to complementary DNA using the Magic RT cDNA synthesis kit with oligo-(dT)18 and random hexamer. The levels of mouse CYP1A1, human CYP1A1, human CYP1A1, mouse β-actin, and human GAPDH mRNA were evaluated by qRT-PCR performed using IQ2 SYBR Green Fast qPCR System Master Mix on a CFX96 Real-Time PCR Detection System (Bio-Rad), as described previously.23,24 The relative expression level of target genes was determined using the ΔΔCT method.
Western Blotting
After treatment, the cell protein was extracted using radioimmunoprecipitation assay buffer. The levels of proteins were determined using a BCA Protein Assay kit. Equal protein loaded to the gel was separated by electrophoresis and transferred onto polyvinylidene difluoride membranes. The membranes were blocked by 5% skim milk and then incubated with primary antibodies overnight. The membranes were washed and incubated in secondary antibodies. After washing, the membranes were visualized with chemiluminescence reagents. 25 ImageJ software was used to generate densitometric images for quantitative image analysis.
Immunofluorescent Staining
After reaction, the cells were fixed with 4% formaldehyde in PBS for 15 min, as described in detail previously . 21 After washing with PBS, the slices were incubated with primary antibodies and then probed by fluorescence binding secondary antibodies, and Hoechst 33342. All images were captured with a fluorescence microscope. ImageJ software was used to calculate the fluorescent intensity in the nucleus and cytoplasm.
Plasmid Construction and Reporter Activity Assay
The pAhRDtkLuc3 reporter plasmid comprises the Cyp1a1 AhRD enhancer in the pGL3-basic vector, as described previously.22,23 The Rous sarcoma virus (RSV)-
Molecular Docking
Molecular docking could simulate the interaction of protein targets and small molecule compounds. The molecular structures of AhR stimulators and agonists of structured field introducer format were obtained from the PubChem database (https://pubchem.ncbi.nlm.nih.gov/). The molecular structures of hAhR (PDB: 6px0) and mAhR (PDB: 4m4x) in PDB format were obtained from the US Research Collaboratory for Structural Bioinformatics Protein Data Bank (RCSB PDB) database (https://www.rcsb.org/). Then, AutoDock, a program that can automate the docking of unknown compounds with biomolecular complexes, was used to predict the potential interactions, and was generally applied in molecular design. AutoDock was used to simulate the interaction of AhR with B(a)P, TCDD, alizarin, and ITE.
Statistical Analysis
GraphPad Prism 5.0 software was used to analyze the data. The values from all experiment were expressed as mean ± standard deviation (SD). Comparisons of several test series were analyzed using one-way analysis of variance (ANOVA tests). The results were considered statistically significant when
Results
Alizarin Treatment Induces CYP1A1 mRNA Expression
To determine whether alizarin can activate AhR signaling, we analyzed the target gene
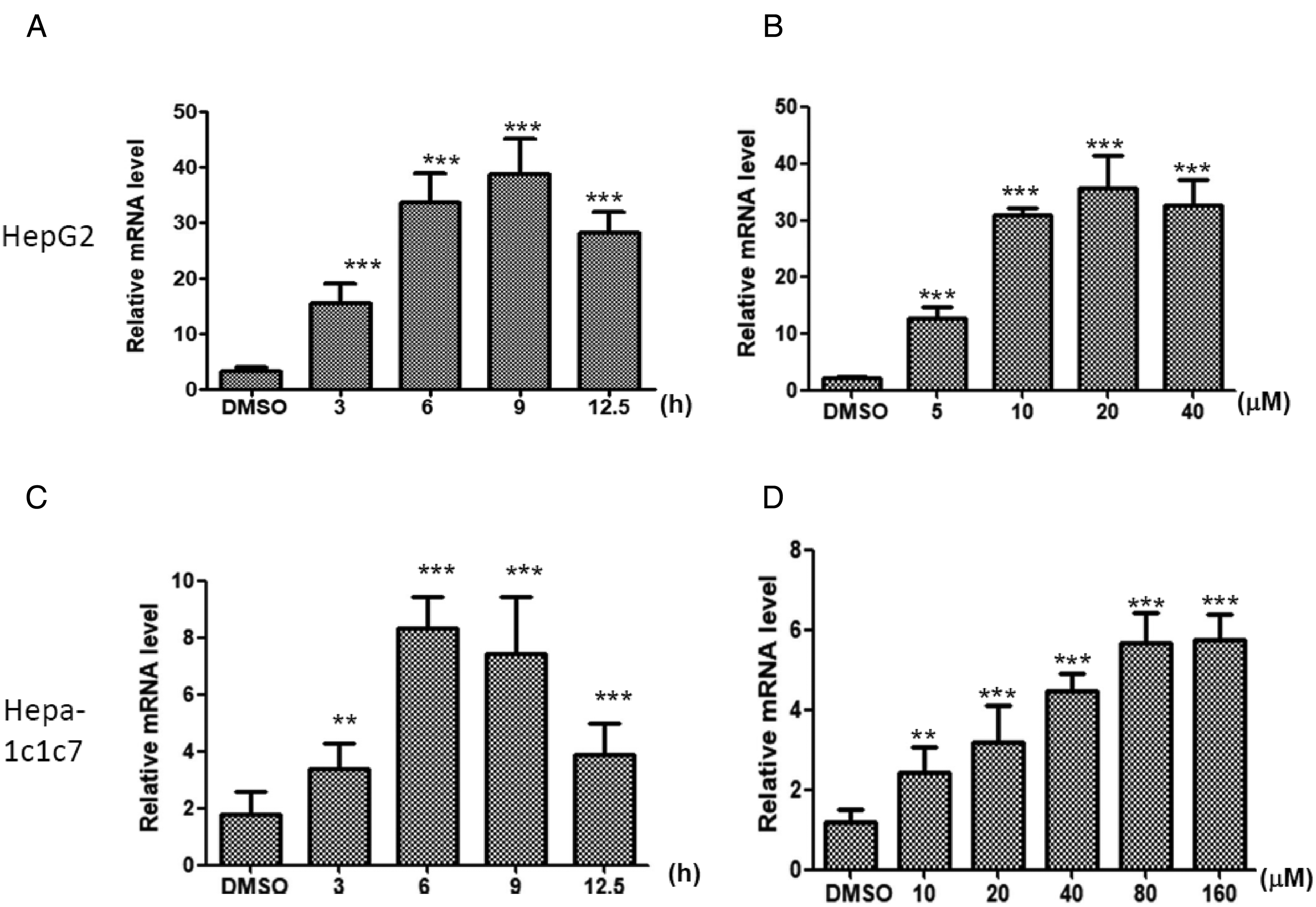
Alizarin treatment increases cytochrome P450 1A1 (CYP1A1) mRNA expression in human and mouse hepatoma cell lines. CYP1A1 mRNA expression was analyzed after treatment with 20 μM of alizarin at (A) various time points or (B) different concentrations of alizarin treatment in HepG2 cells. CYP1A1 mRNA expression were analyzed after 80 μM of alizarin treatment at (C) various time points or (D) different concentrations of alizarin in Hepa-1c1c7 cells.
Hepa-1c1c7, a murine hepatoma cell line expressing AhR and CYP1A1, is suitable for studying AhR and CYP1A1. We treated Hepa-1c1c7 cells with different concentrations of alizarin at various time points. The induction of CYP1A1 mRNA expression increased approximately three-fold at 3 h and reached maximum levels at 6 h (Figure 2C). After 3 h treatment of Hepa-1c1c7 cells with different concentrations of alizarin, the expression of the CYP1A1 mRNA level reached a maximum with 80 μM of alizarin treatment (Figure 2D). This indicates that alizarin induced CYP1A1 mRNA expression in Hepa-1c1c7 cells in a concentration- and time-dependent manner. The results showed that alizarin treatment induced CYP1A1 mRNA expression in both human and murine hepatoma cell lines.
Alizarin Treatment Increased CYP1A1 Protein Expression
After confirmation that alizarin treatment induced CYP1A1 mRNA expression in a concentration- and time-dependent manner, we then tested whether the protein expression of CYP1A1 was increased. Alizarin was applied to HepG2 and Hep-1c1c7 cells at different time points, and the results of Western blot showed that the protein expression of CYP1A1 increased with longer exposure of these two cell lines to alizarin (Figure 3A and B). The protein expression of CYP1A1 also showed a concentration-dependent manner with various concentrations of alizarin treatment (Figure 3C and D). This indicated that alizarin treatment induced CYP1A1 expression in human and murine hepatoma cells. To determine whether other cells showed a similar response after alizarin treatment, we investigated the protein expression of CYP1A1 in HO23, an immortalized human ovary granulosa cell line, after alizarin treatment. These data indicated that alizarin induced the protein expression of CYP1A1 in HepG2 and Hepa-1c1c7 hepatoma cells.

Alizarin treatment increases cytochrome P450 1A1 (CYP1A1) protein expression in human and mouse hepatoma cell lines. CYP1A1 protein expression was analyzed after treatment with 20 μM of alizarin at various time points in (A) HepG2 and (B) Hepa-1c1c7 cells. CYP1A1 protein expression was analyzed after the indicated concentration of alizarin treatment for 12 h in (C) HepG2 and (D) Hepa-1c1c7 cells.
Alizarin Treatment Activates AhR Translocation into the Nucleus
We showed that alizarin treatment induced CYP1A1, a major AhR target gene, mRNA, and protein expression. Upon activation, AhR will translocate to the nucleus. If alizarin induced CYP1A1 expression through AhR activation, AhR should be translocated into the nucleus after alizarin treatment. Hepa-1c1c7 cells were treated with different concentrations of alizarin for 1 and 2 h and with nuclear localization of activated AhR by fluorescent dye staining (Hoechst 33342). The green fluorescence, a representative of the AhR protein, was shifted from the cytoplasm to the nucleus after alizarin treatment. The nuclear translocation of the quantity of green fluorescence increased as the alizarin concentration increased to 20 μM (Figure 4). Nuclear translocation of AhR protein was evident as early as 1 h after alizarin treatment, but a slight decrease in nuclear translocation was found after 2 h of alizarin treatment.

Alizarin treatment induced aryl hydrocarbon receptor (AhR) translocation in the nucleus of Hepa-1c1c7 cells. Hepa-1c1c7 cells were treated with alizarin (10 and 20 μM) for 1 and 2 h. Cells were fixed and stained with AhR antibody. Hoechst 33342 was used for nuclei staining. Fluorescence images were examined under a fluorescence microscope. The relative nuclear/cytoplasmic fluorescent intensity was calculated by ImageJ software and is listed under the image of AhR.
Alizarin Transactivates AHRE Activity
After activation, AhR translocates to the nucleus, interacts with the AHRE of its target genes, and induces gene expression.17,18 Therefore, we analyzed the AHRE reporter activity after alizarin treatment. The pAhRDtkLuc3 reporter plasmid containing three AHRE motifs was transfected into HepG2 and Hepa-1c1c7 cells. The reporter plasmid RSV-
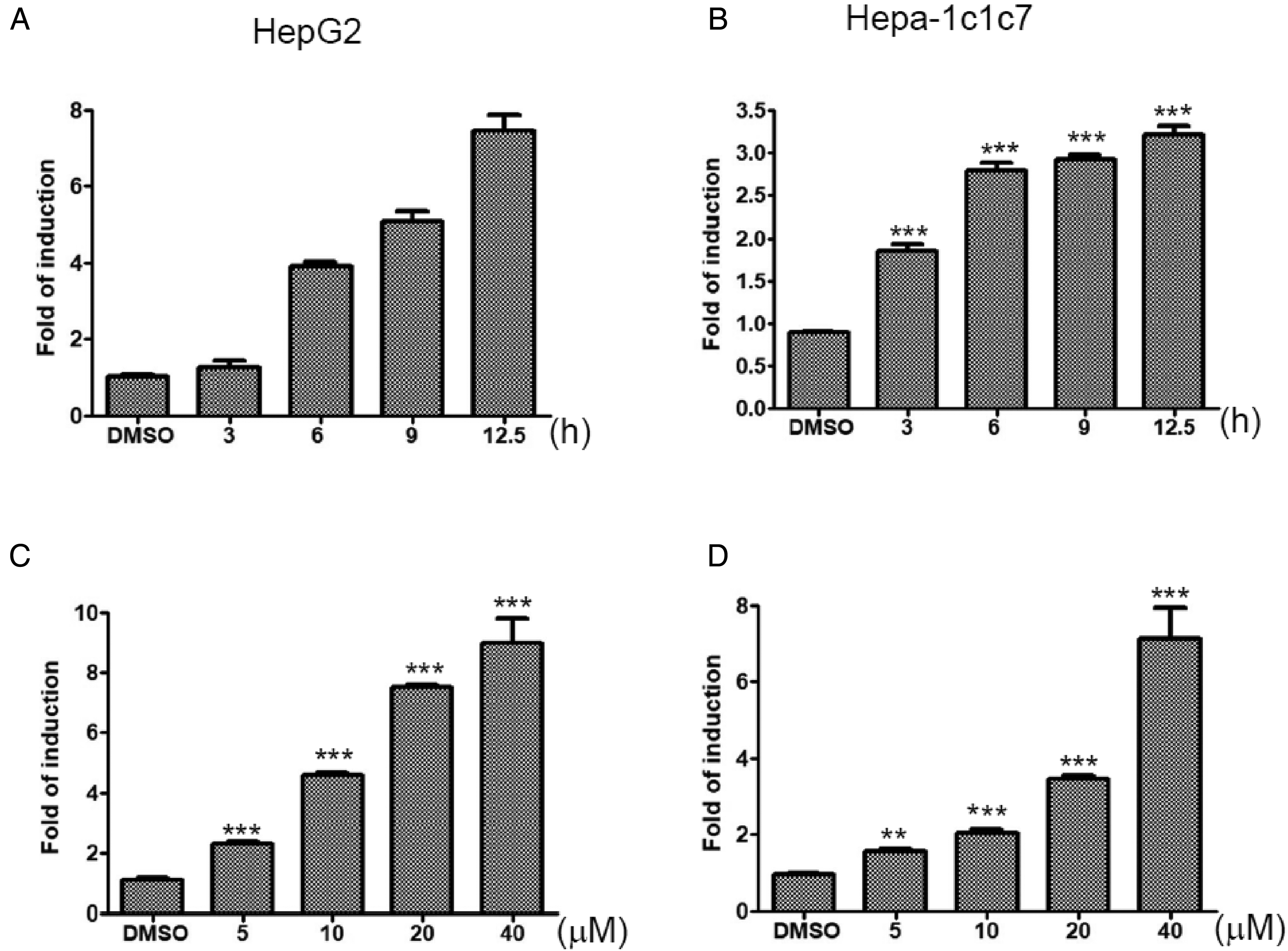
Effects of alizarin treatment on the transactivation activity of the aryl hydrocarbon response element. (A) HepG2 and (B) Hepa-1c1c7 cells were treated with alizarin (20 μM) for 3 to 12 h. (C) HepG2 and (D) Hepa-1c1c7 cells were treated with the indicated concentration of alizarin for 12 h. At completion of incubation with alizarin, cells were harvested, and cell lysates were collected for an activity assay of luciferase and β-galactosidase, as described in the “Materials and Methods” section.
AhR Antagonist Inhibits CYP1A1 Expression Induced by Alizarin Treatment
To determine further the specificity of CYP1A1 expression after alizarin treatment through AhR activation, we applied AhR antagonists, CH-223191
24
and salicylamide
25
and detected CYP1A1 expression in cells treated with alizarin. CYP1A1 expression was detected in alizarin-treated human and murine hepatoma cells, HepG2 and Hepa-1c1c7 cells. When cells were pretreated with the AhR antagonist, CYP1A1, expression was suppressed, indicating that alizarin activated AhR and induced CYP1A1 expression (Figure 6A and B). Both C4 and C12 cells were derived from parental Hepa-1c1c7 cells. The C4 cells lack functional ARNT due to a point mutation (Gly-326 to Asp) in the

CYP1A1 protein expression induced by alizarin treatment was blocked by AhR antagonist. (A) Hepa-1c1c7 cells were pretreated with AhR antagonist CH-223191 (CH) for 1 h, followed by alizarin (20 μM) treatment for 12 h. (B) Hepa-1c1c7 cells were pretreated with CH and salicylamide (SAL) for 1 h, followed by alizarin (20 μM) for 12 h. (C) Hepa-1c1c7, C4 (B13NBii1), and C12 (B15ECiii2) cells were treated with alizarin (20 μM) for 6 h. Treated cells were harvested and subjected to Western blotting analysis to determine CYP1A1 protein expression. Glyceraldehyde 3-phosphate dehydrogenase (GAPDH) and β-actin of their cell lysates were analyzed by Western blotting. The relative protein expression level was calculated by ImageJ software and is presented under the Western blotting subsection. Cells without alizarin treatment were used as control for comparison.
Alizarin Bound to hAhR and mAhR
The lower the binding energy, a combination of the two was better. Afterwards, AhR was docked with B(a)P, TCDD, alizarin, and ITE. The results showed that the affinity of hAhR with B(a)P, TCDD, alizarin, and ITE were −6.94, −6.63, −5.73, and −5.44, respectively (Figure 7A). On the other hand, the affinity of mAhR with compounds was −9.0, −7.91, −7.0, and −6.91, respectively. Both showed that AhR stimulators, including B(a)P and TCDD, always had a higher binding energy than agonists, including alizarin and ITE. It is worth noting that alizarin had better affinity than ITE whether in hAhR or mAhR. The AutoDock visualizing data indicated that B(a)P, TCDD, alizarin, and ITE had nearby binding sites in hAhR and mAhR. According to the results, alizarin and ITE were binding strongly to AhR and were potentially able to antagonize B(a)P and TCDD (Figure 7B to I).

Molecular docking model of B(a)P, TCDD, alizarin, and ITE with AhR. (A) Affinity of B(a)P, TCDD, Alizarin, and ITE with hAhR and mAhR, respectively. (B) B(a)P and hAhR, affinity = −6.9 kcal/mol. (C) TCDD and hAhR, affinity = −6.63 kcal/mol. (D) Alizarin and hAhR, affinity = −5.73 kcal/mol. (E) ITE and hAhR, affinity = −5.4 kcal/mol. (F) B(a)P and mAhR, affinity = −9.0 kcal/mol. (G) TCDD and mAhR, affinity = −7.91 kcal/mol. (H) Alizarin and mAhR, affinity = −7.0 kcal/mol. (I) ITE and mAhR, affinity = −6.91 kcal/mol.
Discussion
Alizarin was reported recently to have uncovered biological activities. In this study, AhR and its downstream target CYP1A1 were activated by alizarin. Moreover, alizarin treatment induced CYP1A1 mRNA and protein expression in human and murine hepatoma cells. Alizarin induced AhR translocation into the nucleus and activated the AHRE reporter activity. Alizarin did not induce CYP1A1 protein expression in cells pretreated with AhR antagonists. Therefore, alizarin activates AhR signaling and its downstream target CYP1A1 expression.
Overexpression of AhR has been reported in various cancers. 26 Binding of BaP to AhR accelerates cell proliferation, 27 suggesting a role for AhR in tumor development. On the contrary, TCDD inhibits cell proliferation in some AhR-dependent cancer cells.28,29 These results indicated that AhR can trigger contradictory physiological functions in cells, depending on which ligand is bound to AhR, emphasizing that the binding ligand is a key regulator in controlling AhR function.
Alizarin is a constituent of
CYP1A1 is a major enzyme in the cytochrome P450 superfamily and is responsible for the metabolization of xenobiotics. CYP1A1 is supposed to be an AhR-sensitive target and is often used as an indicator of AhR activation. 16 However, recent evidence indicates that the induction of CYP1A1 activity does not reflect the activation of AhR signaling. To avoid this possible discrepancy, we further examined nuclear translocation of AhR, AhR promoter reporter activity, specific application of the AhR antagonist, and AhR-deficient cell lines after alizarin treatment. The results showed that alizarin induced CYP1A1 mRNA and protein expression in a concentration- and time-dependent manner. Both C4 and C12 cell lines represent Hepa-1c1c7 cell derivatives with respective deficiencies in ARNT and AhR. Alizarin induced CYP1A1 protein expression in Hepa-1c1c7 cells, but was unable to induce it in C4 and C12 cells. In addition, CH-223191 and SAL, which are AhR antagonists, suppressed alizarin induced CYP1A1 protein expression, indicating that AhR mediated the action of alizarin to induce the expression of CYP1A1.
In summary, our results indicate that alizarin activated AhR and induced expression of the AhR-targeted gene CYP1A1 (Figure 8). The structure of alizarin is distinctly different from that of AhR ligand prototypes, such as BaP and TCDD, and this information provides a clue for studies of AhR ligand structures. Our results also demonstrate that alizarin toxicity decreased in cells with activated AhR signal, indicating that alizarin was metabolized by AhR-induced metabolic enzymes into metabolites with lower toxicity. This report provides valuable information for further studies of the biological application of alizarin.

Scheme of alizarin as the activator of the AhR signaling pathway.
Footnotes
Acknowledgments
We are very grateful to Chiung-Yao Fang Associate Researcher (Medical research, Ditmanson Medical Foundation Chia-Yi Christian Hospital) for her great help in the structure of the entire article and in editing the scientific methods. We thank Dr Alvaro Puga (University of Cincinnati Medical Center, Cincinnati, OH, USA) for the generous gift of the pAhRDtkLuc3 plasmids. We thank Dr Abraham Amsterdam (The Weizmann Institute of Science, Rehovot, Israel) for the generous gift of HO23 cells.
Author Contributions
Conceptualization: Yin-Che Lu and Yu-Hsiang Kuan. Data curation: Yin-Che Lu, Chih-Yi Lin, and Shan-Chun Chen. Formal analysis: Yu-Ting Chou, Guan-Lun Gao, and Chiang Wei Hsu. Investigation: Yin-Che Lu, Yu-Ting Chou, Guan-Lun Gao, and Jyan-Gwo Joseph Su. Methodology: Chih-Yi Lin, Yu-Ting Chou, Shan-Chun Chen, Guan-Lun Gao, and Chiang Wei Hsu. Project administration: Jyan-Gwo Joseph Su. Supervision: Jyan-Gwo Joseph Su. Validation: Yin-Che Lu and Chih-Yi Lin. Writing—original draft: Yin-Che Lu, Yu-Hsiang Kuan, Shan-Chun Chen, and Chiang Wei Hsu. Writing—review & editing: Yin-Che Lu, Yu-Hsiang Kuan, and Jyan-Gwo Joseph Su.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by a grant (R101-029) from Ditmanson Medical Foundation Chia-Yi Christian Hospital Research Program. This study was supported by research grants from the Ministry of Science and Technology of Taiwan (MOST 109-2320-B-040-MY3.
Correction (January 2023):
Funding statement is updated with an additional information: This study was supported by research grants from the Ministry of Science and Technology of Taiwan (MOST 109-2320-B-040-MY3.
