Abstract
Introduction
Lung cancer (LC) is the most commonly diagnosed cancer globally (accounting for 11.4% of all cases) and the leading cause of cancer death (accounting for 18.0% of all cancer deaths). 1 The incidence of LC has gradually increased in recent years, especially in developing countries. Like in China, it is estimated that 870982 patients will be newly diagnosed with LC in 2022, accounting for 18.1% of all newly diagnosed cancers. And there will be 766898 new LC deaths, accounting for 23.9% of all cancer deaths. 2 Lung squamous cell carcinoma (LUSC) accounts for about 30% of all cases diagnosed with LC.
LC treatment has changed in the twenty-first century, and the prognosis has been continually improving. The driver gene can be found in more than 60% of LC cases, 3 including the epidermal growth factor receptor (EGFR), echinoderm microtubule-associated protein-like 4/Ana lipoma kinase (ALK), C-ROS oncogene 1 receptor tyrosinase (ROS 1) gene fusion, etc. The targeted drugs for the mutations mentioned above are constantly innovating, improving the survival of LC patients, however, for LUSC, which accounts for 25% to 30% of all LCs, 4 the incidence of common driver genes such as EGFR mutation and ALK gene rearrangement is very low, at about 2.7% and 1.5% to 2.5%, respectively. Only a few patients with LUSC have the opportunity to be treated with EGFR-tyrosinase inhibitor (EGFR-TKI) or ALK inhibitor, and this treatment is also full of challenges. It is therefore imperative to develop new drugs.
There are approximately 80 kinds of antitumor drugs commonly used, some of which come from plants or traditional Chinese medicine (TCM), including paclitaxel, camptothecin, vinblastine, resveratrol, podophyllotoxin, artemisinin, ginseng, etc. 5 They have a definite curative effect and few adverse reactions in tumor treatment, and have come to play a leading role among antitumor drugs. Screening antitumor drugs from plants or TCM is therefore currently the focus of antitumor drug research, and expected to be so in the future.
Phytoestrogen is a natural estrogen receptor (ER) regulator that is similar in structure and function to endogenous mammalian estrogen. 6 Flavonoids, stilbenes, coumarins, and lignans are phytoestrogens with apparent affinity and selectivity to ERs. Studies have shown that flavonoids, stilbenes, and lignans have antitumor effects. 7
Bioinformatics analysis has proved to be an efficient method for identifying hub genes for tumor progression. 8 Network pharmacology is a new discipline that can design multitarget molecule drugs through selected target genes. Using bioinformatics to screen antitumor medicines can improve the efficiency of drug screening. Some scholars previously used bioinformatics methods to filter the key biomarkers and therapeutic targets for LUSC, 9 however, most experiments have not been further verified.
In this study, based on bioinformatics, we screened hub genes related to the occurrence and development of LUSC. We screened the effective components of TCM, targeting hub genes according to network pharmacology. Our results are further verified by in vitro experiments.
The workflow for the bioinformatics strategy of LUSC is illustrated in Figure 1a.

(a) Flow chart of the study. Volcano plots of DEGs in 6 GEO datasets and TCGA datasets. Volcano plots of GSE19188 (b), GSE21933 (c), GSE33479 (d), GSE40275 (e), GSE62113 (f), GSE74706 (g), and TCGA (h). Red represents genes with differences, and black represents genes without differences. (i) Veen diagram 7 datasets. (j) The dataset–disease–gene networks. Abbreviations: DEG, differentially expressed genes; LUSC, lung squamous cell carcinoma; GEO, Gene Expression Omnibus; TCGA, the Cancer Genome Atlas.
Materials and Methods
Data Collection
We downloaded datasets from the Gene Expression Omnibus (GEO) database (https://www.ncbi.nlm.nih.gov/) according to the following criteria: (1) the gene expression data series contained LUSC tissue and normal tissue; (2) the species of the samples was Homo sapiens; and (3) the information of each sample was consistent and without missing values. Finally, 6 gene expression profiles, GSE19188, 10 GSE21933, 11 GSE33479, 12 GSE40275, 13 GSE62113, 14 and GSE74706 15 were collected. Details of these microarray studies are shown in Table 1. We downloaded the Cancer Genome Atlas (TCGA)-LUSC RNA-seq data from the University of California - Santa Cruz (UCSC) database (https://xenabrowser.net/hub/), which included 501 LUSC samples and 49 normal samples. The information above was freely available online.
Details of Associated Microarray Datasets from the GEO Database in the Study.
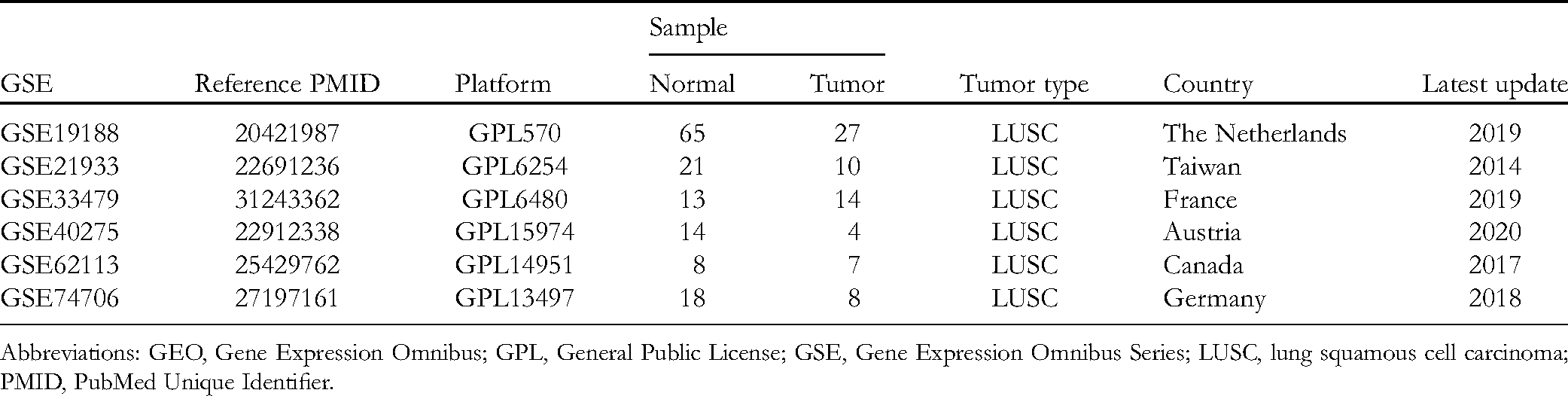
Abbreviations: GEO, Gene Expression Omnibus; GPL, General Public License; GSE, Gene Expression Omnibus Series; LUSC, lung squamous cell carcinoma; PMID, PubMed Unique Identifier.
Identification of Differentially Expressed Genes
GEO2R (http://www.ncbi.nlm.nih.gov/geo/geo2r) is a free online analysis tool to identify differentially expressed genes (DEGs) in the GEO datasets. We screened DEGs according to the following criteria: |log2FC| > 2 and adj. P value < .05. In the TCGA dataset, we selected 49 para-cancer normal samples and 49 LUSC samples after matching clinical information. We then used the “limma” package (|Log2FC| > 2 and adj. P value < .05) to identify DEGs.
Screening Hub Genes
Search Tool for the Retrieval of Interacting Genes/Protein (STRING) (https://cn.string-db.org/) is a database that can analyze the interaction between proteins. We imported the DEGs of each dataset into the STRING database, where the reliability is 0.40, and the species is H sapiens. 16
The PPI network generated by the STRING database was visualized using Cytoscape v3.8.2. 17 The 30 candidate genes in the PPI network of each dataset were screened using the maximal clique centrality (MCC) methods of cytoHubba, a plug-in of Cytoscape software. 18 We used the “Veen” software package to analyze the intersection genes of 7 candidate gene sets, and finally obtained 7 intersection genes as the hub genes of LUSC.
Verification of Hub Genes
The human protein Atlas database (HAP: https://www.proteinatlas.org/) contains the distribution information of 24 000 kinds of human protein in tissues and cells, and can be queried for free. We therefore analyzed the protein expression of the hub genes in tumor and normal tissues in the HAP database.
The gene chip and RNA-seq data of the Kaplan–Meier Plotter database (http://kmplot.com/analysis) are mainly from public databases, such as GEO and TCGA. The Kaplan–Meier Plotter database can evaluate the role of 54 000 genes in 21 cancer types, including breast, ovarian, lung, and stomach cancer. 19 We used the Kaplan–Meier Plotter database to analyze the effect of hub genes on the survival rate for LUSC. We selected overall survival (OS) for 5 years as the observation index and included 377 cases. According to the auto-select best cutoff, the cohorts were divided into high expression and low expression groups. The p <0 .05 was considered statistically significant.
Construction of Hub Gene–Drug Interaction Network
The HERB database (http://herb.ac.cn/), a database based on high-throughput experiments and reference guides of TCM, relates 12 933 targets and 28 212 diseases to 7263 kinds of TCM and 49 258 ingredients and provides the pairwise relationship between them. We used the HERB database to screen herbal medicines affecting hub genes. The practical components of herbal medicines were then selected for cell experiments in vitro. The network of disease-hub genes-TCM was drawn in Cytoscape.
Cell Lines and Reagents
The H226 cell line was purchased from the Shanghai Enzyme Research Biotechnology Co. Ltd (sourced from American type culture collection (ATCC) Co. Ltd), and Roswell Park Memorial Institute (RPMI)-1640 was bought from the Shanghai Yuanpei Biotechnology Co. Ltd South American fetal bovine serum (FBS) was acquired from PAN BIOTECH. Streptomycin was gained from Roche. The coumestrol (CM; purity: high-performance liquid chromatography ≥98%) was a product of Chengdu Manster Biotechnology Co. Ltd, prepared with 20 mmol/L in dimethyl sulfoxide (DMSO) and stored at −20 °C. Before the experiment, CM was diluted with the RMPI-1640 medium to the working concentration. Beijing Qingke Biotechnology Co. Ltd, synthesized the primers used for reverse transcription–polymerase chain reaction (RT-PCR).
The H226 cells were cultured in an RMPI-1640 medium containing 10% FBS, 100 U/mL penicillin, and 100 mg/mL streptomycin, cultured in an incubator at 37 °C and 5% CO2 saturation humidity, and subcultured. Logarithmic growth cells were taken for experiments.
Observation of Cell Morphology Under a Light Microscope
The H226 cells in the logarithmic growth phase were digested with 0.25% trypsin and then inoculated in a culture dish. Cells were divided into the CM group (CM concentration was 20 and 40 μmol/L) and the control group (DMSO). The final DMSO concentration was 0.25% to reduce error, which was consistent with 40 μmol/L CM concentration). After 48 to 72 h of culture, the morphological changes of cells were observed directly under the light microscope.
RT-PCR Amplification of Hub Genes
Cells were divided into a CM group (0, 10, 20, and 40 μmol/L) and control group (0.25% DMSO). The H226 cells were treated for 72 h, and the total RNA was extracted using the Trizol method. The reverse transcription of total RNA was then performed with an iScript cDNA synthesis kit.
The primer sequence was as follows. The primer sequences for CCNB2 were 5′-GGCTGGTACAAGTCCACTCC-3′ (sense) and 5′-GAAGCCAAGAGCAGAGCAGT-3′ (antisense). The primer sequences for CENPF were 5′-TGGGATTTAGAGGAAAAACTGC-3′ (sense) and 5′-TCTCCATCTGTAGGTTTTGTCG-3′ (antisense). The primer sequences for KIF11 were 5′-TGTTTGATGATCCCCGTAACAAG-3′ and 5′-CTGAGTGGGAACGACTAGAGT-3′ (antisense). The primer sequences for MELK were 5′-TATTCACCTCGATGATGATTGCG-3′ and 5′-AGAAAGCCTTAAACGAACTGGTT-3′ (antisense). The primer sequences for nucleolar and spindle-associated protein 1 (NUSAP1) were 5′-TGGCCCTGCAAGTCAGAGTA-3′ (sense) and 5′-TCTGGCTGGAGTCTTGGTCA-3′ (antisense). The primer sequences for PBK were 5′-CCAAACATTGTTGGTTATCGTGC-3′ (sense) and 5′-GGCTGGCTTTATATCGTTCTTCT-3′ (antisense). The primer sequences for RRM2 were 5′-GTGGAGCGATTTAGCCAAGAA-3′ (sense) and 5′-CACAAGGCATCGTTTCAATGG-3′ (antisense). The primer sequences for Glyceraldehyde-3-Phosphate Dehydrogenase (GAPDH) were 5′-GTCTCCTCTGACTTCAACAGCG-3′ (sense) and 5′-ACCACCCTGTTGCTGTAGCCAA-3′ (antisense).
The relative messenger RNA (mRNA) expression of the target gene was calculated using the 2-ΔΔCt method and calibrated with the internal reference GAPDH.
Statistical Analysis
Statistical analysis was performed using R version 4.1.0 (https://www.R-project.org). The P < .05 was considered statistically significant.
Results
Screening of DEGs
A volcano map was used to visualize the DEGs of each dataset. There are 825 DEGs in the GSE19188 dataset, including 494 downregulated and 331 upregulated genes (Figure 1b). In total, 1303 DEGs were generated in the GSE21933 dataset, including 732 downregulated and 571 upregulated genes (Figure 1c). A total of 980 DEGs were generated in the GSE33479 dataset, including 531 downregulated genes and 449 upregulated genes (Figure 1d). In total, 1359 DEGs were generated in the GSE40275 dataset, including 586 downregulated and 773 upregulated genes (Figure 1e). A total of 864 DEGs were generated in the GSE62113 dataset, including 246 downregulated and 618 upregulated genes (Figure 1f). In total, 1868 DEGs were generated in the GSE74706 dataset, including 1115 downregulated and 753 upregulated genes (Figure 1g). A total of 2306 DEGs were generated in the TCGA-LUSC dataset, including 1224 downregulated and 1082 upregulated genes (Figure 1h).
Select Hub Genes of LUSC
We imported the DEGs of each dataset into the STRING database to generate interactive network data. Cytoscape visualizes the protein-protein interaction (PPI) network (Supplemental Figures S1-S7). The first 30 genes scored using the MCC method in CytoHubba were selected as candidate genes, and the candidate genes of 7 datasets are shown in Supplemental Table S1. A Veen diagram shows the relationship between the candidate genes of the 7 datasets (Figure 1i). The dataset–disease–gene network is shown in Figure 1j. The intersection genes from 7 datasets were selected as hub genes, which were RRM2, KIF11, PBK, CENPF, NUSAP1, MELK, and CCNB2 (Table 2).
Details of 7 Hub Genes.

Abbreviations: BUB1, Budding Uninhibited by Benzimidazoles 1; PDZ; Post-synaptic density protein-95, Discs large, Zonula occludens 1; UP, up.
Verification of the Hub Genes
We explored the protein expression level of the hub genes in LUSC tissues and normal tissues in the HPA database. We found that CCNB2, CENPF, NUSAP1, PBK, and RRM2 were not detected in normal lung tissues but could be detected in tumor tissues (Figure 2a, b, and e–g). KIF11 was weakly expressed in normal tissues and strongly expressed in LUSC tissues (Figure 2c). MELK was strongly expressed in normal lungs and LUSC, while the density of normal tissues was lower than that of LUSC (Figure 2d). The protein translated by the hub genes is therefore highly expressed in LUSC. More details of the immunohistochemistry are shown in Supplemental Table S2.

Immunohistochemistry validation for hub genes. (a) CCNB2, (b) CENPF, (c) KIF11, (d) MELK, (e) NUSAP1, (f) PBK, and (g) RRM2.
Survival analysis showed that CCNB2, CENPF, KIF11, and MELK were significantly correlated with a prognosis of LUSC (Figure 3a–d), while NUSAP1, PBK, and RRM2 were not associated with a prognosis of LUSC. The results are shown in Figure 3e–g.

OS analysis of hub genes in LUSC patients. (a) CCNB2, (b) CENPF, (c) KIF11, (d) MELK, (e) NUSAP1, (f) PBK, and (g) RRM2. (h) The disease–gene–drug network. Abbreviation: LUSC, lung squamous cell carcinoma.
Establish a Hub Gene–Drug Interaction Network
The HERB website was used to study the interaction between hub genes and drugs from TCM. We found that many drugs can act on these 7 hub genes (Figure 3h), among which strawberry clover can act on all 7 hub genes. As the main component of strawberry clover is a phytoestrogen CM, we chose CM for the follow-up study (see Supplemental Table S3 for more details).
Observation of Cell Morphology
Under the light microscope, the H226 cells in the control group were more significant in size, spindle-shaped or irregular triangular, with less debris and a faster growth rate, however, H226 treated by CM showed apparent cell damage, gradual shedding, a rounded or irregular shape, increased cell debris, and an unclear boundary (Figure 4). The effect of the 72h group was more evident than the 48 h group, so we studied the expression of 7 hub genes further after 72 h of CM treatment.

Morphological changes of H226 cells treated by coumestrol (CM).
Effect of CM on the Expression of Hub Genes
All 7 genes were highly expressed in H226 cells (Figure 5a). Compared with the control, a low concentration of DMSO does not affect the expression of 7 hub genes. The expressions of CCNB2, KIF11, and NUSAP1 decreased after 72 h of treatment at 10, 20, and 40 µM. Compared with the control, the difference was statistically significant. The expression of CENPH, MELK, PBK, and RRM2 was not affected, and there was no significant difference compared with the control (Figure 5b).

(a) Expression of 7 hub genes in H226 cells. (b) The expression of 7 hub genes after CM treatment. ※ Compared with the control of NO DMSO, P < .05. Abbreviations: CM, coumestrol; DMSO, dimethyl sulfoxide; NO, no.
Discussion
LUSC is a unique subtype of LC characterized by poor prognosis and limited effective treatments. Insufficient effective targeted therapy options mean that conventional chemotherapy and radiation are the primary systemic therapeutic strategies. 20 The rapid development of next-generation sequencing in GEO and integrated multiomics measurements in the TCGA database enabled the research to reveal the significant molecular heterogeneity of LCSC. The analysis of multiple GEO datasets makes the research on the relationship between cancers and genes more reliable. Liao et al 21 discovered 8 hub genes related to small-cell LC through several GEO databases, including Budding Uninhibited by Benzimidazoles 1 Homolog Beta (BUB1B), Centromere Protein E (CENPE), and Kinesin Family Member 2C (KIF2C). Bioinformatics analyses can thus identify specific molecular targets for LUSC. 22
In this study, we found that CCNB2, CENPF, KIF11, MELK, NUSAP1, PBK, and RRM2 may play a vital role in the occurrence and development of LUSC. There are many studies on the relationship between hub genes and LUSC, and some scholars have used these genes as tumor therapeutic targets, such as CENPF and MELK.23, 24
The occurrence of cancer is a long-term, multifactor, and multistage process. It results from the comprehensive action of protooncogene, tumor suppressor gene, and carcinogen (including physical, chemical, and biological). Gene regulation is a very complex network system. Long noncoding RNA (lncRNA) and microRNA (miRNA) do not have the protein-coding function but regulate protein synthesis to regulate human diseases. Therefore, lncRNA and miRNAs may be potential targets for tumor therapy. For example, Homeobox Cluster Antisense RNA 3 (HOXC-AS3) can inhibit the early activation of MiR-96 in non-small-cell LC (NSCLC) cells and increase the growth and invasion of NSCLC cells. Therefore, regulating the expression of HOXC-AS3 may be a potential target for treating NSCLC. 25
Genes can directly affect the occurrence and development of cancer and can also affect cancer by acting on the immune network. 26 Indoleamine-Pyrrole 2,3-Dioxygenase (IDO-1) can mediate the function and accumulation of myeloid-derived suppressor cells (MDSC), which leads to anti-Programmed Cell Death 1(PD-1) drug resistance in NSCLC patients. Therefore, treatment with an IDO-1 inhibitor (INCB023843) can produce an anti-PD-1 effect.26,27 Signal Transducer and Activator of Transcription 3 (STAT3) can activate many immunosuppressive functions through MDSC and other mechanisms that may contribute to tumor escape and growth. Therefore, the STAT3 pathway may be a potential target for LC treatment. 28
Many studies have found that the reproductive hormone was associated with the risk of LC. ER expression was significantly related to a poor prognosis in LUSC, especially for male patients after radical resection. 29 Studies have also found that exposure to menstrual and reproductive factors may play a role in the occurrence of LC for females, but the mechanism remains unclear. The type and extent of the immune response are affected by gender and age, so the immune state is different between males and females. Sex hormones can regulate the anatomical differences between males and females, and directly regulate the immune response, resulting in different risks of immune diseases. Overall, women are at greater risk for autoimmune diseases such as systemic sclerosis and systemic lupus erythematosus, while men are more likely to suffer from infectious and parasitic diseases. 30 Men are also at a higher risk for nonreproductive cancer. 31 Some studies, however, have shown that estrogen may have a pro-inflammatory property and that androgens may have anti-inflammatory properties. 32
Most phytoestrogens have antiestrogen properties so phytoestrogens may become effective antitumor drugs. Phytoestrogen can prevent many diseases, such as osteoporosis, cardiovascular diseases, nervous system diseases, hormone-dependent breast cancer, ovarian cancer, prostate cancer, and climacteric symptoms, by competitively binding Estrogen Receptor 1 (ESR1)/ESR2 or G Protein-Coupled Estrogen Receptor 1 (GPER) in cytoplasm/nucleus. 33 Studies have found that phytoestrogens can inhibit the cell cycle and have anti-inflammatory, antiangiogenesis, and antimetastasis effects on cancer cells. 34
CM is a naturally occurring dietary phytoestrogen. CM possesses many biological activities, such as anticancer, anti-inflammation, antiosteoporosis, lung protection, cardiovascular protection, neuroprotection, antiobesity, anti-SLE, anti-autoimmune thyroiditis, antioxidation, skin protection, and biological activity of estrogenic. 35 CM’s potential health effects are reported to include estrogen and nonestrogen effects. 36
In food replacement therapy, foods containing CM may reverse fatty liver degeneration, but the mechanism is unclear. 37 A dietary intake of isoflavones is associated with an increased risk of prostate cancer, but CM is not related to the risk of prostate cancer. 38 It has also been found that in countries with a low incidence of breast cancer and prostate cancer, the dietary intake of phytoestrogens, including CM, is significantly higher, which indicates that they may have an effect as chemopreventive agents. 39 Interestingly, studies have shown that high-dose CM intake increases thyroid cancer. 40 The contradictory effect on tumors may be related to the different binding forces of CM and other phytoestrogens with varying ERs. 41 Coumarin phytoestrogens, including CM, have higher affinity and selectivity with ERβ than ERα. 42
CM is found in Strawberry Clover, Szechwan Chiberry Fruit, the root of lobed kudzuvine, and Officinal Dandelion. CM is also found in alfalfa, beans, Brussels sprouts, spinach, clover, and soybeans. Epidemiological and experimental evidence shows that phytoestrogens can prevent or reduce the incidence of estrogen-dependent cancers, so people are very interested in phytoestrogens in their diet. CM can inhibit the proliferation of ovarian cancer cells by blocking the phosphorylation and activation of inositol phosphate 3-kinase (Human Phosphoinositide-3 kinase (PI3K))/Serine-Threonine Protein Kinase (AKT) and Rat Sarcoma (RAS)/Mitogen-Activated Protein Kinase (MAPK) signal pathways. 43 CM can inhibit the proliferation and migration of prostate cancer cells by regulating the signal transduction cascade of AKT, Extracellular Regulated Protein Kinases 1 (ERK1)/2, and Jun Oncogene (JNK)/MAPK. 44 Research on the antitumor effect of CM mainly focuses on the protein level, however, and little on the gene level.
We found that phytoestrogen CM may act on the hub genes of LUSC through network pharmacology. The inhibitory effect of CM on LUSC was confirmed in cell experiments, and RT-PCR showed that CM inhibited the growth of LUSC cells by inhibiting the expression of CCNB2, KIF11, and NUSAP1.
We found that CM can inhibit the growth of H226 by inhibiting the expression of CCNB2, KIF11, and NUSAP1. Cyclin B2 is a member of the cyclin family and an important part of the cell cycle regulation mechanism. High expression of CCNB2 is associated with a poor prognosis of NSCLC. 45 KIF11 has been found to play an important role in the G2-M phase transition of tumor cells, thus affecting the survival of tumor cells. KIF11 is an important anticancer target. 46 NUSAP1 regulates tumor cell proliferation, metastasis, and drug resistance. NUSAP1 knockdown can reduce the expression levels of PI3K and phosphorylated serine/threonine kinase (p-AKT); inhibit the proliferation, migration, and invasion of tumor cells through the AKT/Mechanistic Target of Rapamycin Kinase (mTOR) signaling pathway; and promote the apoptosis of tumor cells.47,48
In conclusion, our bioinformatics analysis confirmed that RRM2, KIF11, PBK, CENPF, NUSAP1, MELK, and CCNB2 were the hub genes in our research. CM was selected as an anticancer drug through network pharmacology. In cell experiments, CM may inhibit the growth of LUSC cells by inhibiting the expression of CCNB2, KIF11, and NUSAP1 genes. CM may become a potential drug for treating LUSC.
Supplemental Material
sj-docx-1-npx-10.1177_1934578X221127960 - Supplemental material for The Effect of Coumestrol on Hub Genes in Lung Squamous Cell Carcinoma Based on Bioinformatic Strategy
Supplemental material, sj-docx-1-npx-10.1177_1934578X221127960 for The Effect of Coumestrol on Hub Genes in Lung Squamous Cell Carcinoma Based on Bioinformatic Strategy by Qingqing Shan, Yifan Zhang, Xu Zhang, Wei Wang and Zongan Liang in Natural Product Communications
Supplemental Material
sj-docx-2-npx-10.1177_1934578X221127960 - Supplemental material for The Effect of Coumestrol on Hub Genes in Lung Squamous Cell Carcinoma Based on Bioinformatic Strategy
Supplemental material, sj-docx-2-npx-10.1177_1934578X221127960 for The Effect of Coumestrol on Hub Genes in Lung Squamous Cell Carcinoma Based on Bioinformatic Strategy by Qingqing Shan, Yifan Zhang, Xu Zhang, Wei Wang and Zongan Liang in Natural Product Communications
Footnotes
Author Contributions
QS analyzed and interpreted the data and contributed to writing the manuscript. XZ and WW checked spelling and grammar. YZ reviewed the manuscript, and all the authors read and approved the final manuscript.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval
Ethical approval is not applicable for this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Department of Science and Technology of Sichuan Province, Chengdu Science and Technology Bureau (grant number 2021YFS0272, 2019-YF05-00496-SN).
Statement of Human and Animal Rights
The data of the bioinformatics analysis can be downloaded from the public website as written in the manuscripts. Laboratory experiment does not contain any studies with human or animal subjects.
Statement of Informed Consent
There are no human subjects in this article and informed consent is not applicable.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
