Abstract
Astragali Radix (AR) and Schisandrae chinensis Fructus (SCF) have been used individually and in traditional Chinese medicine (TCM) formulas for treating non-small cell lung carcinoma (NSCLC). Qi Wei Wan (QWW), a 2-herb TCM formula composed of AR and SCF, is used to treat blood deficiency, fatigue, and metabolic abnormalities. We speculate that QWW may be more effective in treating NSCLC than AR or SCF alone. We identified 28 bioactive compounds in QWW and 322 targets of these compounds from databases. Network pharmacology analysis was used to identify 248 putative NSCLC-related gene targets of the bioactive compounds in QWW. Common target genes were analyzed to build protein–protein interaction networks. Implicated biological functions and pathways (p53, PI3K-Akt, etc) were identified by Kyoto Encyclopedia of Genes and Genomes and Gene Ontology analyses. Molecular docking of core target proteins with the key active compounds was also performed. This study identified the potential gene targets and mechanisms involved in the anti-NSCLC effects of QWW.
Introduction
Cancer accounted for 10 million deaths in 2020. 1 Lung cancer is the third most frequently diagnosed form of cancer (age-standardized incidence rate: 22.4/100 000). 1 It is the most commonly diagnosed cancer in China, accounting for 787 000 cases in 2015. 2 In the United States, 236 740 new cases are estimated in 2022. 3 Lung cancer is classified as small-cell lung cancer (SCLC) and non-small cell lung cancer (NSCLC). 4 NSCLC can be subcategorized into lung squamous cell carcinoma, lung adenocarcinoma, and large cell carcinoma. 5 NSCLC accounts for more than 85% of lung cancer cases, with wheezing, cough, hemoptysis, chest pain, trouble breathing, and fever being the main clinical symptoms with diagnosis usually being made at late stages. 6 Treatment methods include surgery, chemotherapy, radiotherapy, and targeted therapy. Although overall cancer survival rates have increased with improvements in technology, survival rates remain relatively low.3,7 In China, lung cancer's 5-year survival rates (2012-2015) in men and women were 16.8% and 25.1%, respectively. 8 Worldwide 5-year survival rates ranged from 4% to 17%, depending on stage and region. 9
Traditional Chinese medicine (TCM) has been practiced for centuries and has been established as a primary medical system in China.10,11 In the recent past, the consumption of TCM has also gained recognition as complementary and alternative medicine, particularly for the treatment of cancer. 12 Astragali Radix (AR) and Schisandrae chinensis Fructus (SCF) have been used individually or as key components in formulas for thousands of years in China. The functions of AR and SCF have been documented in “Ben Cao Feng Yuan” (The Medicinal Herbs in the Wild, Zhang, 1695). The functions of AR include enriching and nourishing the lung and the spleen system, while SCF strengthens the lung, resolves systemic infections, and invigorates the kidney system. Furthermore, it has been reported that SCF encourages oxygen exchange and reduces coughing and pulmonary inflammation in guinea pigs. 13 Some ingredients of AR and SCF have been reported to have functions on NSCLC.14–16 The functions of the Qi Wei Wan (QWW) formula (composed of AR and SCF) have been documented in “Pu Ji Fang” (Prescriptions of Universal Relief, 1406). It is described as a supplement for the treatment of blood deficiency syndrome. However, the mechanisms and pathways involved in the functions of QWW remain elusive and warrant further research. In this study, the potential of QWW as an anticancer medicine for the treatment of NSCLC was investigated using a network pharmacology-based approach.
Network pharmacology involves systems biology, network analysis, protein–protein and protein–compound interactions, redundancy, and pleiotropy. 17 It is a primary data-driven approach, which potentiates drug discovery by incorporating multiple parameters, such as compound bioactivity, application, assimilation, and clinical efficacy to identify drug targets, toxicity, and side effects. 17 Network pharmacology has successfully been used to predict the potential functions of natural products.18,19 For example, the probability of the compound Matrine being a macropinocytosis inducer was predicted by network pharmacology and then verified by biochemical experiments. 18 Similarly, Danhong Injection, an injectable Chinese compound medicine, was also predicted to specifically target anti-inflammation pathways. 19 Biochemical analysis revealed that Danhong Injection specifically inhibited the NF-kB pathway. 19 Collectively, such research provides explicit evidence for the practical application of network pharmacology.
The QWW formula is a multicomponent and multitarget therapeutic agent that has the potential to comprehensively treat complex disorders including NSCLC. Thus, network pharmacology analysis of QWW can predict target pathways and targets by the bioactive components of QWW, with a high probability that the results can subsequently be verified using biochemical techniques.
A recent study has demonstrated that the combination of platinum-based chemotherapeutic agents with Kangai Injection (a Chinese patent medicine, composed of 3 herbs including AR) for the treatment of stage III/IV NSCLC significantly improved treatment outcome and had reduced toxicity compared to single treatment with conventional chemotherapeutic agents. 20 Another study has demonstrated that the combination of chemotherapeutic agents with Shengmai Injection (a Chinese patent medicine, composed of 3 herbs including SCF) for the treatment of chemotherapy-induced cardiotoxicity significantly improved the efficacy of the treatment and had significantly less toxicity compared to solitary treatment with chemotherapeutic agents. 21 Furthermore, a patent TCM named Huangqi Shengmai Yin or Jiawei Shengmai San (adding AR to Shengmai Yin which contains SCF) has also been shown to protect against radiation-induced cardiac fibrosis injury in the process of cancer treatment. 22
Therefore, as QWW is composed of AR and SCF, it can be hypothesized that using this formula may be effective in treating NSCLC. At present, the existing literature does not address this issue. In this paper, the potential gene targets, pathways, and mechanisms of QWW in the treatment of NSCLC were investigated through cross-database network pharmacology analysis.
Results
Overall Strategy
Our initial aim was to construct an interaction network of the compounds in QWW and NSCLC-related genes. The names of the compounds in QWW and their targets were collected from the Traditional Chinese Medicine Systems Pharmacology (TCMSP) database. The NSCLC-related genes were collected from the GeneCards and DisGeNet databases. The 2 groups of data were imported into the UniProt KB database to unify target names and identify common targets (genes shared between the 2 groups). A protein–protein interaction (PPI) network of the common targets was then generated, and the core target genes were identified based on the abundance of the interactions. Common targets were also analyzed by Kyoto Encyclopedia of Genes and Genomes (KEGG) and Gene Ontology (GO) processes to support the prominence of the core targets involved in NSCLC. Finally, molecular docking was performed to further support the interactions of the QWW key compounds with core NSCLC-related targets.
Bioactive Compounds in QWW and Their Targets
The TCMSP database was used to identify all reported compounds in AR and SCF. Some 217 compounds, 87 from AR and 130 from SCF, were found. Among these compounds, key bioactive compounds were identified using 2 thresholds (Drug-likeness [DL] ≥ 0.18 and oral bioavailability [OB] ≥ 30). A total of 28 key bioactive compounds in QWW satisfying the threshold criteria were identified (AR: 20 compounds, 23.0%; SCF: 8 compounds 6.2%). The key bioactive compounds in QWW are shown in Table 1. A total of 322 proteins as the potential targets of the 28 compounds were also collected from the TCMSP database.
Detailed Information of Active Compounds (OB ≥ 30% and DL ≥ 0.18) in QWW.a

Abbreviations: QWW, Qi Wei Wan; AR, Astragali Radix; SCF, Schisandrae chinensis Fructus; OB, oral bioavailability; DL, drug-likeness.
QWW, A 2-herb formula composed of AR and SCF.
Non-Small Cell Lung Carcinoma–Related Genes and Analysis of the Drug-Compound-Target Network
To identify proteins involved in NSCLC that are potential targets of the bioactive compounds in QWW, we searched the GeneCards and DisGeNet databases (keywords: “non-small-cell lung carcinoma” and “NSCLC”) and identified 6223 verified genes that are involved in NSCLC. To analyze the interactions between NSCLC-related proteins and the key bioactive compounds in QWW, the protein targets of the compounds in QWW were compared to the NSCLC-related genes using the UniProt KB database. Among the 6223 NSCLC-related genes and 322 targets of the key bioactive compounds in QWW, 248 genes, referred to as “the common targets” that are both the targets of the QWW compounds and NSCLC-related genes, were identified as the putative NSCLC targets of the 28 key bioactive compounds in QWW (Figure 1).

Venn diagram of the potential anti-lung cancer targets. Some 6223 NSCLC-related genes (purple) and 322 gene targets of QWW compounds (yellow) were identified. Common gene targets (248) shared between the targets of QWW compounds and NSCLC-related genes were identified and used for PPI network analysis. The remaining 74 targets of QWW are not related to NSCLC. NSCLC, non-small cell lung carcinoma; QWW, Qi Wei Wan; PPI, protein–protein interaction.
A drug-compound-target network was then constructed to further analyze the 248 NSCLC-related genes targeted by the key bioactive compounds in QWW (Figure 2). In the network, most compounds have many overlapping gene targets. The average number of interactions that each compound has with a gene (degree value) is 20.5. A total of 8 compounds in QWW have a degree value greater than 20.5, namely, with the degree value in parentheses, quercetin (124), (3R)-7,2′-dihydroxy-3′,4′-dimethoxyisoflavan (recorded as (3R)-3-(2-hydroxy-3,4-dimethoxyphenyl)chroman-7-ol in the TCMSP database; CAS#64474-51-7) (85), isoflavanone with 4 hydroxyl groups naturally substituted (6,7-dimethoxyl-3-(2,4-dihydroxyl)phenol-benzopyran-4-one, recorded as isoflavanone in the TCMSP database) (80), kaempferol (51), formononetin (33), 7-O-methylisomucronulatol (31), isorhamnetin (30), and calycosin (21). The high-degree values suggest that these compounds may have important roles in NSCLC treatment owing to the higher number of interactions with NSCLC-related genes.

Compound-target network analysis. The network of 28 QWW key compounds (OB ≥ 30% and DL ≥ 0.18; red circles) and 248 common genes (yellow squares) in NSCLC targeted by QWW compounds. Larger circles mean more interactions in the network. OB, oral bioavailability; DL, drug-likeness; NSCLC, non-small cell lung carcinoma; QWW, Qi Wei Wan.
Protein–Protein Interaction and Network Analysis
Of the 248 common genes that were identified (Figure 2), a PPI network of the gene products was generated (17 proteins without interactions were removed) (Figure 3). A screening threshold, that is, P < .05 and minimum interaction score of 0.700 (“high confidence”), was applied. The network was analyzed using Cytoscape software (231 dots and 3762 edges). Major topological features were also identified (“degree,” “betweenness,” and “closeness”). Nodes were selected that had a median topological feature value (for individual proteins) greater than the threshold (“degree” 12, “betweenness” 0.001681, and “closeness” 0.4054). From this screen, 82 “core targets” were identified, and they were considered to be highly important in the application of QWW for the treatment NSCLC. The names of core targets are shown in Table 2, and the network of the core targets and the QWW bioactive compounds is shown in Figure 4. Targets with the highest degree values were TP53, AKT1, SRC, HSP90AA1, and JUN.
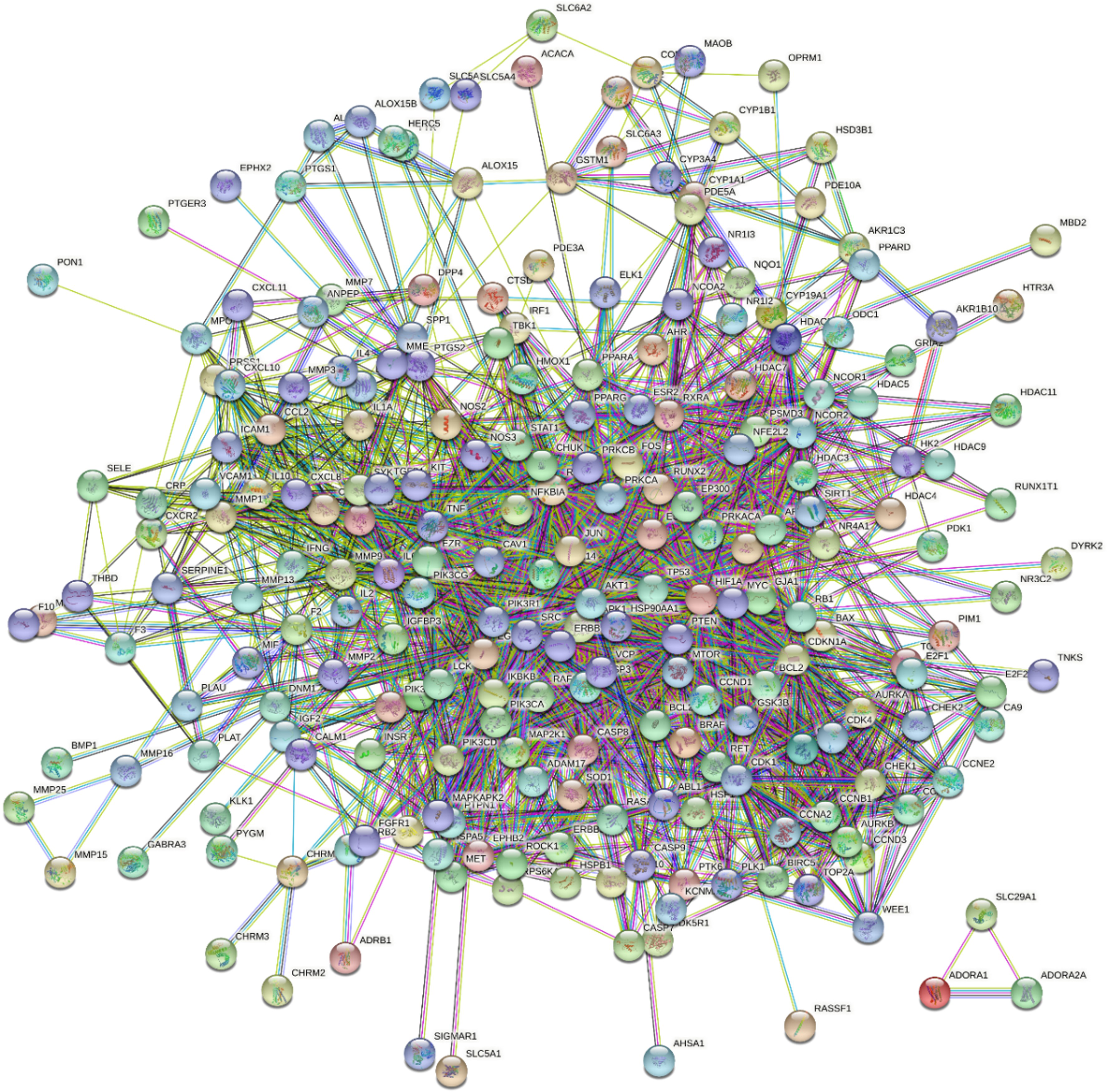
PPI network analysis. Analysis of the interactions of 248 common genes in NSCLC targeted by the QWW compounds, with a screening threshold of P < .05 and “high confidence (0.700)” from STRING. P < .05 represents the results collected have strong evidence against the null hypothesis. After removing disconnected genes, the remaining 231 common genes have interactions with other genes. NSCLC, non-small cell lung carcinoma; QWW, Qi Wei Wan; PPI, protein–protein interaction.

Core compound-core target network analysis. The network of 82 core target genes (yellow squares) in NSCLC targeted by QWW compounds (red squares). Core target genes are defined as having a median topological feature value (for individual proteins) greater than the threshold (“degree” 12, “betweenness” 0.001681, and “closeness” 0.4054). NSCLC, non-small cell lung carcinoma; QWW, Qi Wei Wan.
Details of 82 Targets in the Core Network.

Gene Ontology and KEGG Enrichment Analysis
GO and KEGG pathway analyses were carried out to identify the key potential pathways that are targeted by QWW in the treatment of NSCLC, using Metascape software. The 248 NSCLC-related genes targeted by QWW were screened in the GO analysis process (P < .05). The pathways that had the highest degree values (representing the top 10 biological processes, cellular components, and molecular functions) are listed (Figure 5A). KEGG pathway analysis was carried out to further refine the search. The top 20 enrichment pathways are shown in Figure 5B.

Gene Ontology (GO) analysis and KEGG Pathway Enrichment Analyses. (A) GO analysis showing the top 10 biological processes, cellular components, and molecular functions. The Y-axis shows the count of the targets (enrichment score), and the X-axis shows highly enriched GO categories of the target genes. (B) KEGG Pathway Enrichment Analysis. The Y-axis shows the top 20 significantly enriched KEGG pathways of the target genes, and the X-axis is the fold enrichment. The fold enrichment represents the ratio of the number of target genes in a pathway to the total number of annotated genes in the pathway. The size of dots represents the number of genes in the pathway. The colors of the dots correspond to the different p value ranges. KEGG, Kyoto Encyclopedia of Genes and Genomes.
Interestingly, several pathways that are known to be involved in carcinogenesis were identified, such as the p53 signaling pathway, thyroid cancer, IL17 signaling pathway, and mitogen-activated protein kinase (MAPK) signaling pathway. Consolidation of the KEGG analysis and PPI network analysis identified the IL17, MAPK/ERK, PI3K-Akt, and p53 signaling pathways as being high-degree value targets. In addition, platinum drug resistance pathways were also identified as one of the top 20 KEGG enrichment pathways, suggesting that QWW treatment may reduce the resistance to platinum-based drugs in NSCLC treatment.
Molecular Docking Analysis
Molecular docking was carried out using the AutoDockTool application and processed with PyMol and BIOVIA Discovery Studio software. The proteins identified with the highest degree values (Figure 3) and the 8 compounds in QWW that have a degree value greater than 20.5 (Figure 2) were used to perform molecular docking (Table 3). Complexes with the lowest affinity energy were considered to be high probability structures. Three-dimensional diagrams of the interactions were generated (Figure 6). The binding energy of the top 3 active ingredients with each target protein was mostly less than −7 kcal/mol, indicating relatively high affinity. 23 Especially, the interactions of Akt1 and Src with quercetin and (3R)-7,2′-Dihydroxy-3′,4′-dimethoxyisoflavan have the lowest binding energy, at lower than −8 kcal/mol.

Molecular docking of the core compounds with core targets. Docking results showing the structures and affinity of p53, Akt1, Src, and HSP90AA1 with (3R)-7,2′-Dihydroxy-3′,4′-dimethoxyisoflavan; Src with quercetin; and c-Jun with calycosin.
Binding Energy Between the Core Targets and Core Compounds.
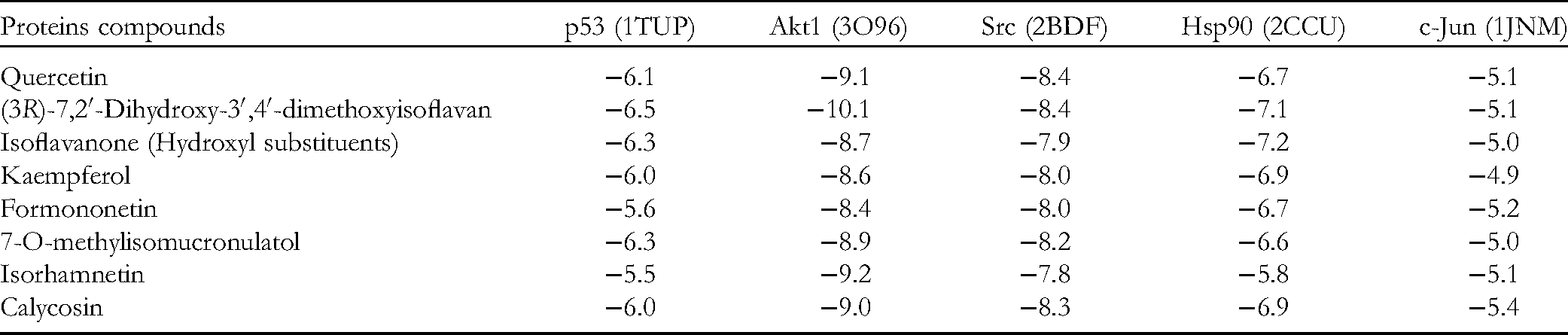
Figure S2 shows putative protein-ligand interactions and the specific amino acids that bind to the compounds. The result for the p53 interaction with (3R)-7,2′-Dihydroxy-3′,4′-dimethoxyisoflavan (Figure S2A) indicated that the interactions occur at the 98P, 156R, 158R, and 258E residues. The docking result for the Akt1 interaction with (3R)-7,2′-Dihydroxy-3′,4′-dimethoxyisoflavan (Figure S2B) shows that the interaction occurs at the 79Q, 80W, 268K, and 270V residues. The Src interactions with (3R)-7,2′-Dihydroxy-3′,4′-dimethoxyisoflavan (Figure S2C) occur at the 273L, 281V, 293A, 295K, 323V, 348D, 393L, and 403A residues. Src interacts with quercetin through 273L, 281V, 293A, 295K, and 393L (Figure S2D). HSP90AA1 and (3R)-7,2′-Dihydroxy-3′,4′-dimethoxyisoflavan interact mostly by van der Waals forces but also with binding sites, which are 98M and 102D (Figure S2E). Calycosin and c-Jun interact through 300M, 301L, 305V, 306A, and 309K (Figure S2F). The molecular docking results support the protein-ligand interactions of the core targets with the core compounds.
Discussion
Network pharmacology is a powerful approach that has been used to identify potential biological targets of bioactive compounds in TCM, within the context of specific diseases. 24 In this study, network pharmacology analysis identified quercetin, (3R)-7,2′-Dihydroxy-3′,4′-dimethoxyisoflavan, isoflavanone, kaempferol, and formononetin as leading compounds (with high-degree values) that interact with NSCLC-related proteins. Some compounds that were identified in this study with lower degree values (relative to the leading key compounds), such as isorhamnetin, 7-O-methylisomucronulatol, and calycosin, have also been reported to act on NSCLC-related proteins.25,26 Molecular docking results support the structural interactions between the leading key compounds and the major hub genes (Figure 6).
Important functions of the key compounds analyzed by network pharmacology in this study have been shown in previous studies. (1) Quercetin: It has been reported that quercetin treatment inactivates Akt-1 and alters the expression of proteins in the Bcl-2 family, leading to apoptosis in A549 lung carcinoma cells. 27 The consumption of quercetin-rich foods has been associated with decreased risk of developing lung carcinoma. 28 (2) Isoflavanone: AR is rich in isoflavanone and isoflavone, as are many other food sources such as soy, cereals, and meat. Research has shown that isoflavone inhibits NF-kB and Akt signaling, to abrogate cancer cell growth. 29 (3) Kaempferol: Kaempferol is known to inhibit cell progression in many cancer cell types, such as breast, liver, and lung. 30 Studies have shown that kaempferol treatment suppresses the PI3K/AKT and ERK pathways, leading to cell death in NSCLC.31,32 (4) Formononetin: Formononetin treatment has been shown to activate p53, leading to cell cycle arrest and apoptosis in NSCLC. 33 (5) Isorhamnetin: Isorhamnetin treatment has also been reported to inhibit proliferation and induce apoptosis in lung cancer cells. 34 (6) Calycosin: Calycosin treatment inhibits the PKC-α/ERK1/2 pathway, impairing NSCLC proliferation and metastasis. 26 Moreover, treatment with (3R)-7,2′-Dihydroxy-3′,4′-dimethoxyisoflavan has been proved to inhibit tumor growth in preclinical studies. 35 Although most functional compounds in the QWW formula are attributed to the AR herb, a bioactive compound in SCF, Gomisin R, has been shown to inhibit the development of preneoplastic lesions in rat livers. 36 Furthermore, Gomisin R treatment has been reported to activate AMPK and p38 to induce G0/G1-phase arrest in colorectal cancer cells and prevent the formation of metastatic tumors in the lung. 37
Aside from key compounds, hub genes identified by network pharmacology in this study have been shown to play important roles in cancer treatment. The PPI network analysis (Figure 3) identified TP53, AKT1, SRC, HSP90AA1, and JUN as major hub genes essential in the application of QWW for the treatment of NSCLC. PPI network analysis identified TP53 as having the highest degree value. TP53 is a tumor suppressor gene whose mutations play important roles in the tumorigenesis of lung epithelial cells. 38 The inactivation of TP53 may lead to increased cancer malignancy, poor patient survival, and resistance to treatment. 39 In NSCLC patients, somatic TP53 mutations and increased TP53 expression were 23% and 65%, respectively. 40 Moreover, TP53 mutations occur at relatively early stages of lung cancer development and are retained during tumor progression and metastasis. 41
Our study identified AKT as having the second-highest degree value and lowest affinity energy. AKT is an essential kinase in the PI3k-AKT pathway activated by PI3K to regulate the proliferation, growth, and apoptosis of cells. Akt/PKB activity is constitutive in NSCLC cells and PI3K dependent, and it promotes NSCLC cell survival. 42 The activation of Akt/PKB can also protect NSCLC cells from chemotherapy and radiation-induced apoptosis. 42 In addition, research has shown that AKT1 silencing decreases MEK/ERK1/2 activity and IκB-α expression, leading to NSCLC cell death. 43 AKT1 is altered in 0.91% of NSCLC patients with AKT1 mutation present in 0.63% of all NSCLC patients. 44 Therefore, AKT is considered to be an attractive anticancer drug target. Another study has reported that AKT1 inactivation is associated with increased NSCLC cell metastatic potential, while activation inhibits migration and invasion. 45
Src, a nonreceptor tyrosine kinase protein involved in processes such as proliferation, differentiation, and adhesion, was identified as having the second-lowest binding energy with the core bioactive compounds in our study. The Src-family of kinase is activated in NSCLC and has been reported to promote survival in various NSCLC cell lines. 46 In addition, inhibition of Src-kinases induces apoptosis in NSCLC cell lines. 46 The critical role of Src in tumor growth, metastasis, and angiogenesis makes it a promising therapeutic target for the treatment of solid tumors. 47
Heat shock protein 90 (Hsp90)-alpha, also known as heat-associated protein 90, is also considered to be an attractive cancer drug target as it can simultaneously inhibit multiple signaling pathways. 48 Clinical studies have shown that average plasma levels of Hsp90α in lung cancer patients are significantly high, potentiating its use as a diagnostic marker for lung cancer. 49 The JUN gene encodes the c-Jun protein. The transcription of JUN can be regulated by c-Jun. 50 Extracellular stimuli, such as peptide growth factors and UV radiation, induce c-Jun protein synthesis. 51 Also, the c-Jun protein is reported to be overexpressed in aggressive, invasive, and metastatic cancers. 52
The target genes identified in this study are implicated in the development of various cancers. Therefore, it is rational to conclude that these genes are likely involved in the anti-NSCLC properties of the QWW formula. KEGG pathway enrichment analysis further strengthens this hypothesis (Figure 5B). Thyroid cancer and IL17 signaling pathways had the highest enrichment. Moreover, other highly enriched pathways, such as the regulation of lipolysis in adipocytes and MAPK signaling pathways, are also known to be involved in carcinogenesis.53,54 Consolidation of the KEGG analysis and PPI network analysis identified the IL17, MAPK/ERK, PI3K-Akt, and p53 signaling pathways as being high-degree value targets. Platinum drug resistance pathways were identified as one of the top 20 KEGG enrichment pathways. In addition, Kangai Injection (which contains SCF) has been proved to reduce adverse effects and regulate tumor immune function, 20 suggesting that QWW treatment may reduce the resistance and the associated toxicity of platinum-based drugs in NSCLC treatment. Moreover, our molecular docking models indicate the possibility of interactions between the QWW compounds and NSCLC-related proteins. Further biochemical experiments are warranted to fully elucidate the targets and mechanisms of the bioactive compounds of QWW in the treatment of NSCLC.
Conclusions
QWW may play the role in inhibiting NSCLC by targeting some cancer-related proteins such as p53, Akt, and MEK/ERK The therapeutic effects of QWW involve the IL17, MAPK/ERK, PI3K-Akt, and p53 signaling pathways. The results of this study lay a foundation for future studies to examine the clinical application of QWW in NSCLC.
Materials and Methods
Data Construction of Active Compounds and Prediction of Potential Targets of QWW
The data of the chemical ingredients in the 2 herbs in QWW were collected from TCMSP database and platform (https://tcmsp-e.com/, Ver.2.3). TCMSP contains 499 herbs registered in the Chinese pharmacopoeia (2010), with a total of 12 144 chemicals. 55 The chemicals were screened according to criteria that the OB is equal to or greater than 30%, and DL is equal to or greater than 0.18. To predict potential targets of the bioactive compounds of QWW, target information was obtained from TCMSP and transformed into UniProt gene identifiers based on the UniProt KB database (https://www.uniprot.org/). If the target information of certain components was not available in the TCMSP database, the 2D structure of the components was downloaded from the PubChem database (https://pubchem.ncbi.nlm.nih.gov/) and submitted to the Swiss Target Prediction database (http://www.swisstargetprediction.ch/) to retrieve the target information. After collecting target protein information from TCMSP and Swiss Target Prediction databases, data were compared with NSCLC-related genes using the common gene name database, UniProt KB. Finally, the targets (322 targets of the QWW compounds and 6223 targets of NSCLC) obtained from the databases (TCMSP, Genecards, and DisGeNet) were imported into Venny 2.1 (https://bioinfogp.cnb.csic.es/tools/venny/) and combined after removing duplicates.
Construction of Drug-Compound-Target Network
To analyze the network of all known compounds of QWW and their targets, we used Cytoscape 3.8.2 software for diagram construction. The information was imported into Cytoscape, and the degrees, edges, and nodes were integrated.
Screening of Potential Targets of QWW Compounds in NSCLC
NSCLC-related protein targets were identified from Genecards (https://www.genecards.org/) and DisGeNET (https://www.disgenet.org/) databases. The Genecards database integrates the genomic, proteomic, and transcriptomic information of human genes. 56 DisGeNET is a discovery platform that contains one of the most comprehensive public collections of genes and variants associated with human diseases based on data integration from expert-curated repositories, GWAS catalogs, animal experiments, and scientific literature. 57
To analyze the interactions between NSCLC-related proteins and the key bioactive compounds in QWW, the protein targets of the compounds in QWW were compared to the NSCLC-related genes using the UniProt KB database. The key word, “Non-Small Cell Lung Carcinoma,” and relevant target proteins were standardized according to the Uniprot.
Construction of PPI Network
STRING (https://www.string-db.org/) database was used to predict the interactions among the common target genes of the QWW compounds in NSCLC. The “organism” was limited to “Homo sapiens,” and the minimum required score was “high confidence (0.700),” and disconnected genes were removed.
Gene Ontology and KEGG Pathway Enrichment Analysis and Network Construction of Targets Pathways
Gene ontology function enrichment and KEGG pathway enrichment analysis of common targets were implemented in the Metascape (http://metascape.org). Histogram and bubble chart were generated from the data.
Molecular Docking
A structure analysis software, AutoDockTools, was used for molecular docking based on the Vina docking algorithm. 58 The structures of proteins were collected from RSCBPDB (https://www.rcsb.org/), which contains tens of thousands of protein structures. PyMol and BIOVIA Discovery Studio graphic tools were used to lay out the detailed connections and bonds of the interaction between proteins and ligands.23,59
Candidate genes identified by the PPI network analysis were imported into the RSCBPDB database to download the specific proteins in PDB format. The structures of the bioactive compounds from AR and SCF were downloaded from the PubChem database (https://pubchem.ncbi.nlm.nih.gov/). Proteins and chemicals were imported into AutoDockTools to obtain the best docking results. The results were further analyzed by PyMol and BIOVIA Discovery Studio software to render the 3D and 2D structures of the related proteins residues and binding bonds.
Supplemental Material
sj-xlsx-1-npx-10.1177_1934578X221120215 - Supplemental material for Network Pharmacology Analysis of Bioactive Components and Mechanisms of Action of Qi Wei Wan Formula for Treating Non-Small Cell Lung Carcinoma
Supplemental material, sj-xlsx-1-npx-10.1177_1934578X221120215 for Network Pharmacology Analysis of Bioactive Components and Mechanisms of Action of Qi Wei Wan Formula for Treating Non-Small Cell Lung Carcinoma by Minghe Zhang, Ye Wang, Aftab Amin, Muhammad Ajmal Khan, Zhiling Yu and Chun Liang in Natural Product Communications
Supplemental Material
sj-xlsx-2-npx-10.1177_1934578X221120215 - Supplemental material for Network Pharmacology Analysis of Bioactive Components and Mechanisms of Action of Qi Wei Wan Formula for Treating Non-Small Cell Lung Carcinoma
Supplemental material, sj-xlsx-2-npx-10.1177_1934578X221120215 for Network Pharmacology Analysis of Bioactive Components and Mechanisms of Action of Qi Wei Wan Formula for Treating Non-Small Cell Lung Carcinoma by Minghe Zhang, Ye Wang, Aftab Amin, Muhammad Ajmal Khan, Zhiling Yu and Chun Liang in Natural Product Communications
Supplemental Material
sj-xlsx-3-npx-10.1177_1934578X221120215 - Supplemental material for Network Pharmacology Analysis of Bioactive Components and Mechanisms of Action of Qi Wei Wan Formula for Treating Non-Small Cell Lung Carcinoma
Supplemental material, sj-xlsx-3-npx-10.1177_1934578X221120215 for Network Pharmacology Analysis of Bioactive Components and Mechanisms of Action of Qi Wei Wan Formula for Treating Non-Small Cell Lung Carcinoma by Minghe Zhang, Ye Wang, Aftab Amin, Muhammad Ajmal Khan, Zhiling Yu and Chun Liang in Natural Product Communications
Supplemental Material
sj-docx-4-npx-10.1177_1934578X221120215 - Supplemental material for Network Pharmacology Analysis of Bioactive Components and Mechanisms of Action of Qi Wei Wan Formula for Treating Non-Small Cell Lung Carcinoma
Supplemental material, sj-docx-4-npx-10.1177_1934578X221120215 for Network Pharmacology Analysis of Bioactive Components and Mechanisms of Action of Qi Wei Wan Formula for Treating Non-Small Cell Lung Carcinoma by Minghe Zhang, Ye Wang, Aftab Amin, Muhammad Ajmal Khan, Zhiling Yu and Chun Liang in Natural Product Communications
Footnotes
Authors’ Note
Minghe Zhang and Ye Wang contributed equally. Data are contained within the article.
Acknowledgements
This work was supported by the Guangzhou Innovation and Entrepreneurship Leading Team Project [202009020005] and the Science and Technology Innovation Bureau of Guangzhou Development District [CY2019-005].
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Science and Technology Innovation Bureau of Guangzhou Development District, the Guangzhou Innovation and Entrepreneurship Leading Team Project (grant number CY2019-005, 202009020005).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
