Abstract
Yiqi Sanjie formula (YQSJF) is mainly applied clinically for the treatment of lung neoplasms. The purpose of this study was to explore the pharmacodynamics of the active components of YQSJF and the mechanism of therapeutic effects in the treatment of lung neoplasm diseases based on network pharmacology. The network of component-target, target-pathway, and pathway-disease of YQSJF was constructed by using Cytoscape software. According to the screening result, 37 key components, 57 important targets, and 866 candidate pathways were obtained. The enrichment analysis results indicated that YQSJF might play a therapeutic role in lung cancer by regulating several signaling pathways, such as the PI3K-AKT, non-small cell lung cancer, small cell lung cancer, and apoptosis pathways. There were 53 intersection genes between YQSJF and the lung cancer gene, 52 common genes, and 11 key targets, including CASP8, CASP9, AR, ESR1, PTGS2, NOS3, PGR, TGFB1, PPARG, RELA, and NOS2, screened by using Protein-Protein Interaction (PPI) analysis. These could be the potential therapeutic targets of YQSJF against lung cancer. Enrichment analysis of the intersection gene pathways revealed 10 major functional pathways, including the VEGF, apoptosis, and IL-17 signaling pathways. The molecular docking results showed the potential regulating activity of kaempferol against AR, pelargonidin against PGR, and baicalein against both PTGS2 and AR. In conclusion, combinational network pharmacology analysis results indicated that YQSJF might present its efficacy of alleviating lung neoplasm symptoms through multiple targets in a synergetic way.
Lung cancer is one of the most common malignant tumors in the world. According to the survey released by the international agency for research on cancer (IARC) of the World Health Organization in 2019 1 World Health Organization in 2019, the number of cases of lung cancer worldwide was 7 74 323 in 2018. It accounts for 18.1% of new cancer cases in the world, ranking the first among malignant tumors. Current treatment guidelines for lung cancer recommend concomitant platinum-based chemo-radiotherapy plus prophylactic cranial irradiation, based on the premise that it disseminates early, and is chemo-sensitive. 2 However, this treatment has a high recurrence rate and a relatively low cure rate. 3 Studies have shown the unique advantages of traditional Chinese medicine (TCM) in combinational treatment for preventing cancer development, recurrence and metastasis owing to its characteristics of multiple pathways, multiple targets, and multiple efficacies. 4
YQSJF is a TCM prescription clinically applied for lung cancer treatment with good clinical efficacy. It consists of 16 TCM medicinal materials: Panacis Quinquefolii Radix (Xiyangshen), Codonopsis Radix (Dangshen), Poria (Fuling), Atractylodis Macrocephalae Rhizoma (Baizhu), Pinelliae Rhizoma Praeparatum (Fabanxia), Fritilariae Thunbergii Bulbus (Zhebeimu), Persicae Semen (Taoren), Armeniacae Semen Amarum (Kuxingren), Trichosanthis pericarpium (Gualoupi), Houttuyniae Herba (Yuxingcao), Gekko swinhonis Guenther (Bihu), Cremastrae Pseudobulbus Pleiones Pseudobulbus (Shancigu), Polistes mandarinus Saussure (Lufengfang), Solanum nigrum L. (Longkui), Sarcandrae Herba (Zhongjiefeng), and Ranunculi Ternati Radix (Maozhaocao). YQSJF in combination with chemotherapy can effectively treat Lewis lung cancer in mice. 5 YQSJF recipe has a certain inhibitory effect on A549 lung cancer in mice and its possible mechanism is relevant to the increase of expression of caspase-4 and DNA-PK protein. 6 Because of the complexities of the pathogenesis and the pathological factors of lung cancer, together with the multifarious YQSJF components and unidentified active components, it is difficult to determine the material basis and mechanism of action of YQSJF in treating lung cancer. A comprehensive analysis of YQSJF based on systems pharmacology in cooperation with virtual docking turned to be an alternative strategy applied for decoding the pharmaceutical rules implied in a TCM formula.
The evaluation of high-throughput biological data and network pharmacology has been proven to be a suitable method for exploring synergistic effects in complex diseases and in the mechanisms of TCMs. 7,8 Based on existing biological data, networks of component-targets, target-pathways, and pathway-diseases interactions can be established to identify potential drug targets and clarify the main mechanism of a drug. 9 This study used network pharmacology and molecular docking to predict the core components and core targets and signal pathways of YQSJF in the treatment of lung tumors, and explore the latent mechanisms of YQSJF in their treatment. This laid the potential foundation for subsequent molecular biology and pharmacological experiments, and also provided some referential ideas for the development of new drugs for the treatment of lung cancer.
Materials and Methods
Establishment of the Chemical Composition Database of YQSJF
Based on traditional Chinese medicine systems pharmacology database and analysis platform (TCMSP, https://tcmspw.com/tcmsp.php), 10 traditional Chinese medicine integrative database (TCMID, http://www.megabionet.org/tcmid/), BATMAN-TCM online analysis tool (http://bionet.ncpsb.org/batman-tcm/), 11 and CNKI (https://www.cnki.net/), the chemical components of all medicinal materials in YQSJF were retrieved. According to the selection criteria of oral bioavailability (OB) ≥30% and drug-likeness (DL) ≥0.18%, 12 the biological information set of YQSJF could be obtained.
Collection of Candidate Targets, Pathways, and Diseases
DrugBank (https://www.drugbank.ca/) was applied for candidate compound targets screening and the Uniprot (https://www.uniprot.org/) database was then used to obtain gene names of target proteins. Then the KEGG PATHWAY database (https://www.kegg.jp/kegg/pathway.html) was used to identify target corresponding pathways and indicate related diseases based on pathway information.
Establishment of Component-Target, Target-Pathway, and Pathway-Disease Networks
A component-target, target-pathway and pathway-disease network was established using the biological informative software, Cytoscape3.6.1 13 (https://cytoscape.org/). Because of the complexity of the network and the purpose of highlighting the key network information, the degree greater than the average node was set as the screening criterion.
GO and Pathway Annotation
The Database for Annotation, Visualization and Integrated Discovery (DAVID) v6.8 (https://david.ncifcrf.gov/) 14 was put into used to analyze pathway enrichment and biological process, and the criteria of P < 0.05 and FDR < 0.05 were set, indicating statistical significance.
The Biological Information Set of Lung Neoplasms
The search for related genes of lung cancer was carried out using Comparative Toxicogenomics Database (CTD, http://ctdbase.org/). 15 The retrieval was restricted to the disease. The key words were inputted into Lung Neoplasms to obtain the corresponding gene target set.
YQSJF—Lung Neoplasms Gene Data Set
The data set of key genes and lung neoplasm genes that had been screened in YQSJF was imported into Draw Venn Diagram (http:// bioinformatics. psb. ugent. Be/webtools/Venn/) for intersection analysis. The results were imported into String 11.0 to explore the protein-protein interaction (PPI) of the intersection of the YQSJF gene and lung neoplasms gene data, and the composite score of >0.4 was used as the screening threshold. 16 Cytoscape software was used to construct the visualized PPI relationship diagram, and the plug-in MCODE was used to analyze the important modules in the PPI network relationship. The screening criteria were degree cutoff equal to 2, node score cutoff equal to 0.2, k-score equal to 2, Max deapth equal to 100, and P < 0.05 (indicating statistical significance). 17
Enrichment Pathway Through ClueGO Plug-In
By using the gene enrichment analysis plug-in ClueGO in Cytoscape 3.6.1, the selected YQSJF-lung neoplasms gene data set was used for signaling pathway enrichment. The ClueGO plug-in tool annotates the gene clusters in the network, analyzes the relationship between the gene clusters and signal pathways in the biological network, and discusses the mechanism of YQSJF in the treatment of lung tumors.
Validation of compound-target interaction via docking simulation
The experimental X-ray structure complexes of AR, PTGS2 and PGR proteins and their ligands were retrieved from the PDB protein database (http://www.rcsb.org/) with the corresponding PDB IDs 5JJM, 18 3INO, 19 and 1A28. 20 Chain A or C of each protein was selected as the docking receptor, and the ligands binding with chain B of each protein (B-DHT in 5JJM, B-52B in 3IN0, and B-STR in 1A28) were isolated as reference compounds for docking mode confirmation and binding energy comparison. Three dimensional structures of all the listed compounds in Table 3 were downloaded from the TCMSP website, energy minimized and used for the virtual screening study. The AutoDock Vina 21 (http://vina.scripps.edu/) module of PyRX software 22 (https://pyrx.sourceforge.io/) was used to implement virtual screening according to the standard protocol linkage, step by step multiple ligand molecular docking using PyRx (Vina), in the FAQ section of the website. The lower the binding energy, the more stable the complex. Then the most stable complexes of the screened compounds were uploaded into the Protein-Ligand Interaction Profiler (https://projects. biotec. tu-dresden.de/plip-web/plip/index) website, 23 the ligand protein ID inputted for analysis, and then the interaction analysis results between small molecule ligand and protein were obtained as 3D interaction diagrams for manual inspection. The cartoons of ligand-protein interaction were then generated using PyMOL software. 24
Results and Discussion
Active Components Screening, Corresponding Targets Identification, and Component-Target Network Establishment of YQSJF
Through TCMSP, TCMID, and BATMAN-TCM online analysis tools and CNKI retrieval and screen, 153 compounds and 1773 corresponding targets were obtained. Because of the complexity of components and targets, node degree 8.7 was set as the screening sub-threshold value; 38 key components and 57 corresponding targets were obtained (Table 1, see Supplemental Table S1 in the Supplementary Material for the detailed information of all the compounds and targets in YQSJF). The higher the degree value, the more relationships the node has and the more profound its meaning. 25 According to the topology of the network, the nodes with a large degree were selected for analysis. These nodes that connect more compounds or targets play a pivotal role in the whole network, and could be the key compounds or targets of YQSJF (Figure 1(A)).
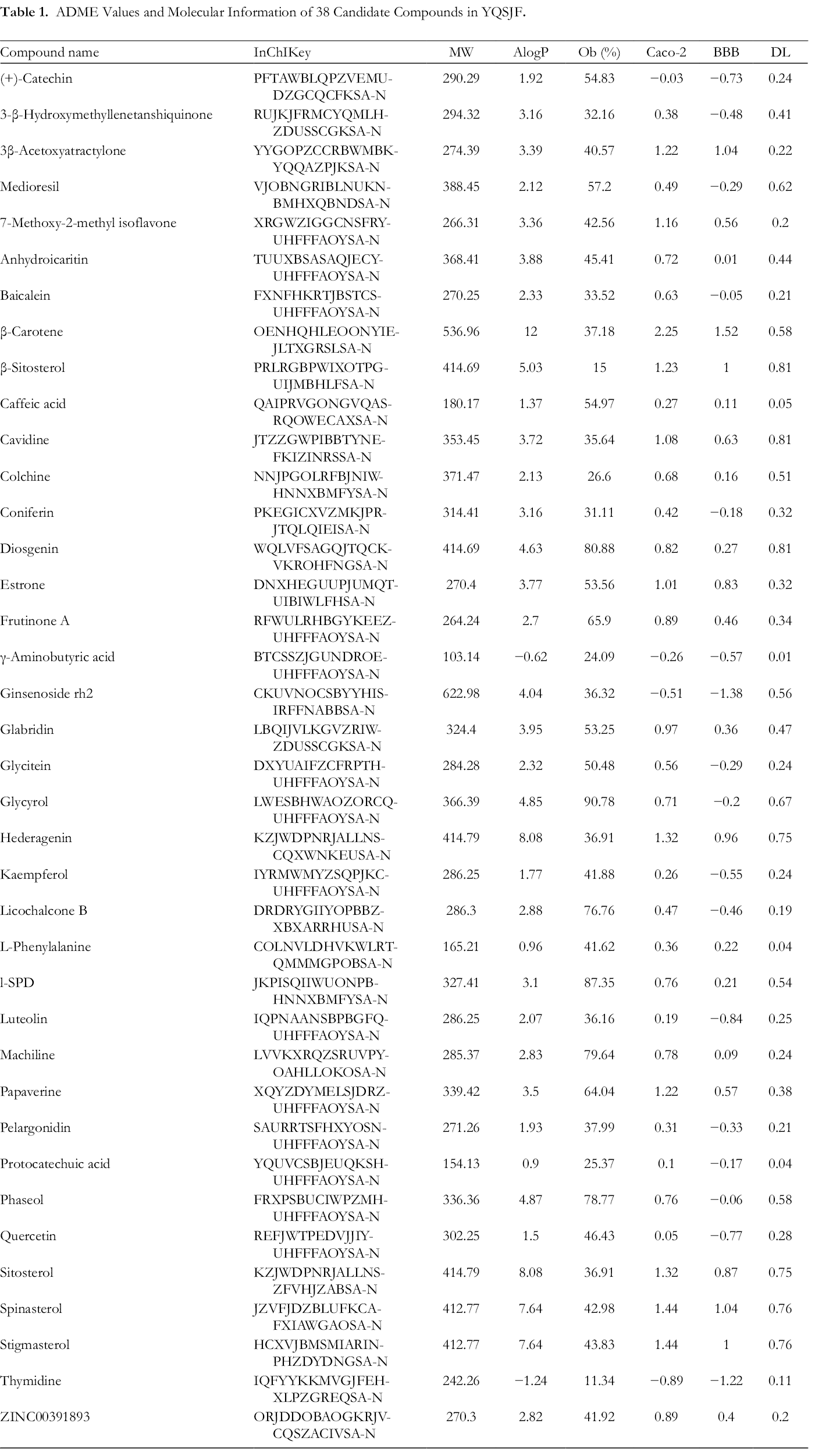
ADME Values and Molecular Information of 38 Candidate Compounds in YQSJF.

A visual network diagram demonstrating the relationship between compounds, targets and pathways in YQSJF. (A) Compound-target network analysis. Green nodes represent target protein, blue nodes represent anti-tumor compounds, orange nodes represent invigorating striking and replenishing qi functional compounds, and pink to divide phlegmy functional compounds. If a compound in the network acts on a target protein, the compound and the target protein are connected by a line. The size of the node is proportional to the number of connected lines. When there are more lines connected, the larger the node in this network. 38 candidate compounds were linked with 57 candidate targets. (B) Target-pathway network analysis. The green node is the target protein and the blue node is the pathway. The target protein is connected to its corresponding pathway. The size of the nodes is proportional to their degree values. 27 targets were connected to 37 pathways. (C) Pathway-disease network analysis. Red nodes are pathways, and blue nodes are diseases. The node size is proportional to the value of degree. There are 27 pathways connected to 164 diseases.
Among them, the compounds with the largest degree value were β-sitosterol and quercetin, which were interacting with 190 and 462 targets, respectively. Quercetin is contained by Sarcandrae Herba, Solanum nigrum and Houttuyniae Herba. β-Sitosterol is a common compound of Sarcandrae Herba, Radix Panacis Quinquefolii, Pinelliae Rhizoma Praeparatum, Fritilariae Thunbergii Bulbus, and Persicae Semen. Studies have confirmed the role of β-sitosterol and quercetin in the treatment of lung cancer. β-Sitosterol can induce the apoptotic cell death and G1 arrest of human lung cancer A549 cells. 26 Animal studies of the anti-cancer effects of β-sitosterol have shown that a diet containing 2% phytosterol compounds can inhibit the growth of cancer cells, reduce tumor size, and reduce the incidence of colon, breast, and liver cancers. 27 -30 Quercetin strongly inhibited cell proliferation, and increased sub-G1 and apoptotic cell populations. 31 Quercetin has potent inhibitory activity on non-small cell lung cancer by regulating the ratio of Bax/Bcl-2. 32 Quercetin is a competitive inhibitor of MMP-9 and can reduce the expression of MMP-9 and TGF-β1, which play an important role in A549 apoptosis. 33
Additionally, the compositions of YQSJF are very distinctive. In order to fully explore mechanisms of YQSJF in the treatment of lung cancer, we combined the results of Table 1 to conduct an inductive analysis of the components of the characteristic Chinese herbal medicine Pinelliae Rhizoma Praeparatum, Polistes mandarinus Saussure, and Solanum nigrum. Pinelliae Rhizoma Praeparatum contains stigmasterol and baicalein. Stigmasterol has anti-tumor and lowering of blood lipid concentration effects, and can prevent cardiovascular diseases. 34 Baicalein has the potential to enhance the antitumor efficacy of docetaxel in a β-catenin-dependent manner and guarantee safety, as well in non-small cell lung cancer (NSCLC). 35 Polistes mandarinus contains protocatechuic acid. Protocatechuic acid can decline FAK, mitogen-activated protein kinase, and NF-κB activation, which subsequently decrease the production of cytokines and growth factors, and consequently inhibit proliferation on NSCLC cells. 36 Solanum nigrum contains diosgenin. Studies have shown that diosgenin can inhibit telomerase activity by down-regulation of hTERT gene expression in the A549 lung cancer cell line. 37
The key targets are prostaglandin G/H synthase 2 (PTGS2), prostaglandin G/H synthase 1 (PTGS1), and nuclear receptor co-activator 2 (NCOA2), which interact with 68, 43, and 54 compounds, respectively. Conversion of arachidonate to prostaglandin H2 is mediated by 2 different isozymes, the constitutive PTGS1 and the inducible PTGS2. PGHS1 is expressed constitutively and generally produces prostanoids acutely in response to hormonal stimuli to fine-tune physiological processes requiring instantaneous, continuous regulation (eg, hemostasis). PGHS2 is inducible and typically produces prostanoids that mediate responses to physiological stresses such as infection and inflammation. With the development of research methods and tools, the role of arachidonate and its metabolites in tumor cell proliferation and metastasis, apoptosis, angiogenesis, and inflammation, as well as its treatment and prognosis has been gradually discovered. 38 PTGS1 and PTGS2 may be potential targets for early detection and treatment of cancer. NCOA2 is a transcriptional co-activator of steroid receptors and nuclear receptors. It is a critical regulator of glucose metabolism regulation, acts as RORA coactivator to specifically modulate G6PC expression. It is also involved in the positive regulation of the transcriptional activity of the glucocorticoid receptor NR3C1 by sumoylation enhancer RWDD3. Glucocorticoids are good anti-inflammatory drugs, which have a good inhibitory effect on chronic inflammation of the respiratory tract. Chronic inflammation is the common pathogenesis of chronic obstructive pulmonary disease and lung cancer. 39 Therefore, it can be speculated that NCOA2 may have an effect on lung cancer by up-regulating glucocorticoid expression.
Analysis of Target-Pathway and Pathway-Disease Networks
For the purpose of systematically deciphering the multiple underlying mechanisms of YQSJF, all of the pathways interacting with candidate targets were extracted from the KEGG Pathways database using DAVID, and then the target-pathway network was constructed (Figure 1(B), see Supplemental Table S2 in the Supplementary Material for presenting the degree value information of candidate active compounds screened from YQSJF).
After calculating the value of degree for each node in the target-pathway network, the average value was 7.Therefore, degree ≥7 was set as the screening standard, and 27 targets and 37 corresponding pathways were obtained. These pathways included pathways in cancer, apoptosis, PI3K-AkT signaling pathway, hepatitis B, cholinergic synapse, and small cell lung cancer. Signaling pathways have been regarded as one of the most important parts of systems pharmacology. 40 For all the pathways, 21 of them belonged to human diseases. Of these pathways for human disease, 4 are cancer pathways: viral carcinogenesis, small cell lung cancer, pathways in cancer, and colorectal cancer. Nine of the pathways are organic systems. Among them, 3 belong to nervous systems: cholinergic synapse, dopaminergic synapse, and serotonergic synapse. Four belong to endocrine systems: thyroid hormone signaling pathway, relaxin signaling pathway, oxytocin signaling pathway, and estrogen signaling pathway. Five of the pathways are signal transduction: Calcium signaling, cAMP signaling, cGMP-PKG, MAPK signaling, and PI3K-AkT signaling.
In Figure 1(C), there are 191 nodes of which the 37 red circle nodes correspond to candidate pathways and the remaining 154 blue circle nodes represent diseases. The node size represents the size of degree. The higher the degree value, the greater is the significance. Therefore, the 3 most important nodes were the MAPK, PI3K-AkT and calcium signaling pathways. They correspond to 47, 27, and 25 different diseases, respectively. All the corresponding diseases were classified by KEGG; nervous system related diseases account for the largest proportion, followed by cancer, infectious disease, cardiovascular disease, endocrine disease, musculoskeletal disease, and congenital malformation.
Analysis of GO Enrichment and KEGG Enrichment
The GO and PATHWAY enrichment analysis functions of online database platform DAVID 6.8 were used to further explore the 57 selected targets. Gene ontology enrichment analysis consisted of 3 parts, biological process (BP), cellular component (CC), and molecular function (MF). A total of 330 results were obtained by the GO enrichment function. In order to make the results statistically significant, 41 P < 0.05 and FDR < 0.05 were selected as the screening criteria and displayed in the order of significance degree (Table 2). It can predict that the biological process of YQSJF is significantly enriched in response to drug, aging, and response to estradiol and response to lipopolysaccharide. Plasma membrane, cell junction, and postsynaptic membrane were ranked as the top 3 cellular components. The main molecular functions induced by YQSJF are enzyme binding, protein homo-dimerization activity, drug binding, and G-protein coupled acetylcholine receptor activity. Through the analysis of enrichment results of biological process, molecular function, and cell components, the therapeutic effect of YQSJF may be closely related to steroid hormone, protein binding, signal transduction, and g-protein-coupled acetylcholine receptor.
KEGG Pathway Enrichment and GO Enrichment Information of Major Targets in YQSJF.

Many studies have reported a significant association between steroid hormones and the treatment of lung diseases. On the 1 hoursand, steroid hormones can accelerate the apoptosis of eosinophils by inhibiting the survival of IL-3, IL-5, GM-CSF, and at the same time reduce the expression level of intercellular adhesion molecule (ICAM-1). Steroid hormone receptors can also induce apoptosis by increasing the ratio of pro-apoptotic factor Bax to anti-apoptotic factor Bcl-2, 42 in order to reduce the damage of inflammatory cells of lung tissue and maintain the integrity of lung epithelial cells and vascular endothelial cells. On the other hand, steroid hormones can regulate the apoptosis of fiber repair cells in the later stage of lung injury, prevent excessive fibrosis of lung tissue, promote the recovery of respiratory function, and play a protective role in lung tissue. 43
A total of 85 results were obtained by the KEGG Pathway enrichment function. In order to make the results statistically significant, P < 0.05 and FDR < 0.05 were selected as the screening criteria and displayed in order of significance degree (Table 2). Among them, the most enriched pathways are cancer, neuro-active ligand-receptor interaction, calcium signaling, and hepatitis B. It can be seen from the table that small cell lung cancer is also one of the pathways with higher enrichment.
Therefore, the GO enrichment and pathway enrichment results confirmed that YQSJF is mainly for use for the treatment of lung neoplasms.
Analysis of YQSJF-Lung Neoplasms-Gene Data Set and its PPI Network
Fifty-seven key gene targets in YQSJF were compared with 33 364 gene targets related to lung neoplasms, and 53 genes with the drug-disease interactions obtained are shown in Figure 2(D), including acetylcholinesterase (AchE), transcription factor p65 (RELA), transcription factor AP-1, nitric oxide synthase, inducible (NOS2), carbonic anhydrase II (CA2), caspase-3 (CASP3), muscarinic acetylcholine receptor M1 (CHRM1), and dipeptidyl peptidase IV (DPP4).

Sub-network of core genes. (D) Venn diagram of YQSJF and lung neoplasms-related targets. The red area is the target protein where the 2 intersect. The green area is the key target protein screened in YQSJF. The blue area is the lung neoplasms-related target protein. (E) Interaction module of YQSJF-lung neoplasms-gene based on MCODE. (F) The PPI network diagram of key YQSJF targets and lung neoplasms-related targets.
A PPI network was established by using the common gene target of YQSJF-lung neoplasms (Figure 2(F)). The pathway of YQSJF in the treatment of lung cancer should be a complex regulatory network, and there will be a certain degree of signal transduction between different signal pathways and targets. Therefore, we believe that the therapeutic effect of YQSJF is not only the direct combination of components and targets, but also the combination of direct regulation of targets and indirect regulation of other targets. Therefore, the construction and mining of protein-protein interaction (PPI) data provides a possibility for the direct and indirect regulation of target discovery. 44 The PPI network of YQSJF-Lung Neoplasms-Gene contained 52 nodes and 234 interactions. The stronger the interaction between proteins, the thicker the connecting lines. To obtain the PPI network module with significant interaction, Cytoscape 3.6.1 plug-in MCODE was used for analysis. As shown in Figure 2(E) and 11 key gene targets were obtained: CASP8, CASP9, AR, ESR1, PTGS2, NOS3, PGR, TGFB1, PPARG, RELA, and NOS2.
Caspases (CASP) are a very important family of peptidases in the process of apoptosis signal transduction and execution. 45 In the exogenous apoptosis pathway, CASP8 can act on Bid and cause it to be cut into tBid, which has a strong apoptotic activity and can be transferred to mitochondria to promote the release of cytochrome C and cause apoptosis. In the endogenous apoptosis pathway, the combination of CASP9 and CASP3 promotes the apoptosis of lung cancer cells. The intervention targeting CASP9 may be one of the strategies for the treatment of lung cancer in patients with advanced lung cancer. 46 The positive expression of AR may be correlated with lung cancer progression and lymph node metastasis. 47 Estrogen can promote the proliferation of NSCLC cells, and estrogen receptor is closely related to the expression and prognosis of NSCLC. 48 Studies have shown the importance of PR through PPD interaction in EGFR-mediated NSCLC proliferation and signal transduction. 49 Therefore, endocrine regulation is predicted to be helpful in the treatment of lung cancer. PGHS2 can be induced to produce prostaglandins, which mediate responses to physiological stress such as infection and inflammation. The expression of NOS, especially NOS1 and NOS3, can inhibit the progression of lung cancer. 50 TGFB1 may suppress tumor initiation by blocking cell proliferation. 51 The overexpression of PPARG protein, as well as its induction using the agonist, rosiglitazone was able to stimulate radiation-induced cell death in an otherwise radio resistant NSCLC A549 cell line. 52 Combined expression of RELA and ACTN4 promotes apoptosis of lung cancer cells. 53 In summary, YQSJF may induce apoptosis of lung cancer cells by regulating the expression of the YQSJF- Lung Neoplasms gene, thus providing help for the treatment of lung cancer.
Molecular Docking Assay
The mechanisms of YQSJF are reflected by the interaction between compounds and targets. Therefore, molecular docking was applied for the simulation of their interactions. Table 3 lists the 3 potential targets with the highest degree values involved in the cancer pathway and the correlated 8 compounds. The results of the docking simulation show that the identified small-molecular ligands could be docked to the binding sites of their corresponding targets, as shown in Figure 3. The binding energy of receptor AR docking with positive control ligand B-DHT was −10.6 kcal/mol, and all the correlated compounds from YQSJF could also form stable complexes with AR, and achieve a certain activity effect (the binding energy ranked from −8.5 to −4.0). The binding energy of receptor PTGS2 docking with positive control ligand B-52B was −8.8 kcal/mol, and that of PGR docking control ligand A-STR was −10.9 kcal/mol. Except for hederagenin with PTGS2 (+0.4 kcal/mol), and diosgenin with PGR (+7.9 kcal/mol), other compounds formed stable complexes with corresponding receptors and should be able to induce certain activities. According to the results of the binding energy of the complex, in the cancer pathway, the binding energy of kaempferol with AR, pelargonidin with PGR, and baicalein with PTGS2 and AR were close to the positive control ligands selected in this experimental process, indicating that these compounds may be potential inhibitors of the corresponded targets. As shown in Figure 3, the ligand and protein interactions of the above 4 complexes were demonstrated in detail.
Binding Affinity Prediction Results of Key Compounds With Corresponded Target Proteins Based on Virtual Docking.

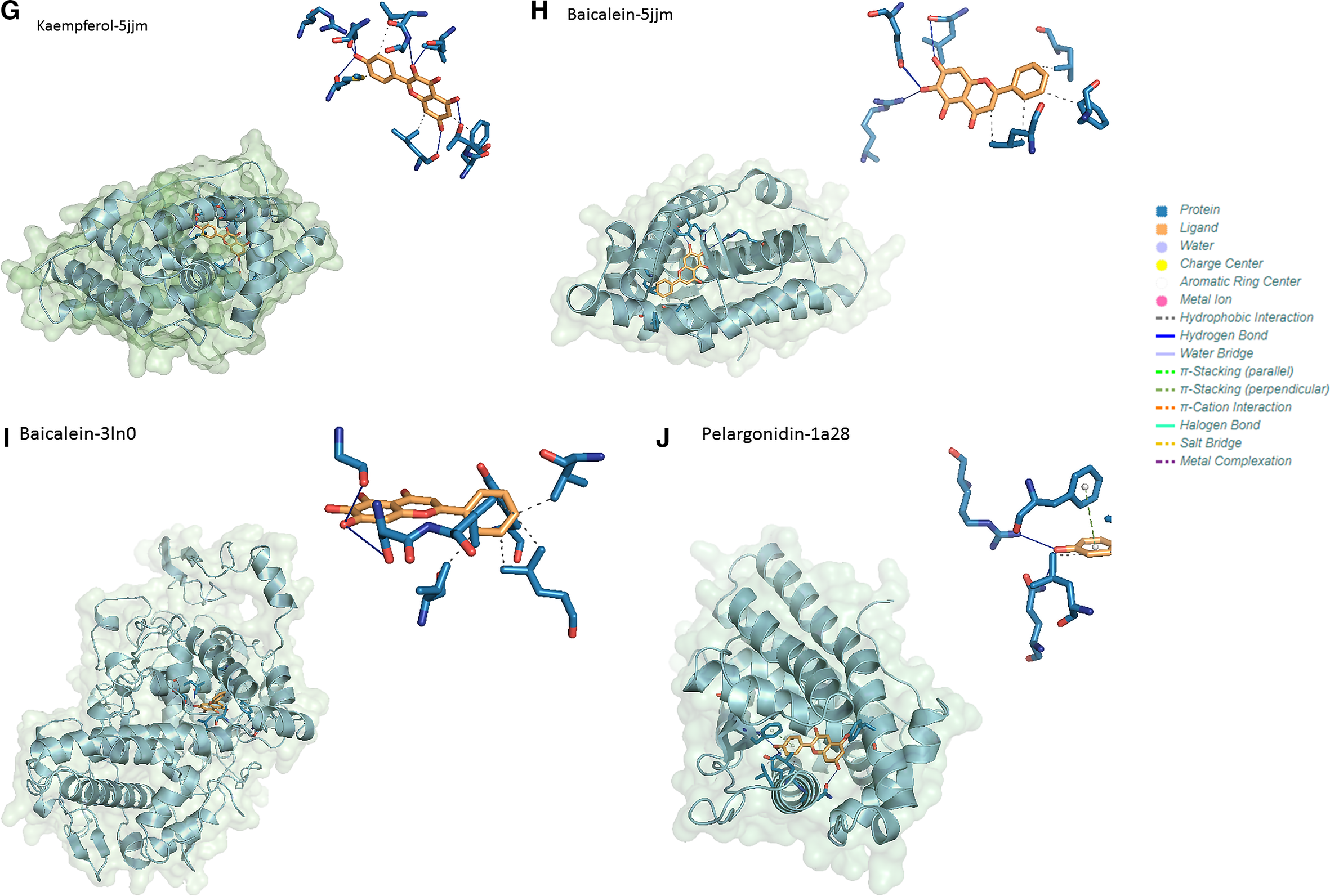
Optimal binding modes of compounds and target proteins. Next to each complex is a diagram of the interaction between the optimal ligand and protein. Different types of lines in the figure represent different forces. For example, the solid blue line represents hydrogen bonding, the gray dashed line represents hydrophobic interaction, and the green dashed line represents Π bond folding interaction. (G) Kaempferol and AR complex. (H) Baicalein and AR complex. (I) Baicalein and PTGS2 complex. (J) Pelargonidin and PGR complex.
Respectively, the main interactions between the complexes formed by the target protein AR with kaempferol and baicalein were hydrophobic interactions and hydrogen bonding forces. Their common amino acid residues that produce hydrophobic forces are LEU and PHE. The common hydrogen bond interactions were mainly formed by phenolic hydroxyl groups on a benzene ring and amino acid residues GLN and ARG. The interactions between the complexes formed by PTGS2 with baicalein were also hydrophobic interactions and hydrogen bonding forces. The amino acids residues that produced hydrophobic forces were VAL, TYR, and LEU. The hydrogen bond interactions were mainly formed by phenolic hydroxyl groups on a benzene ring and amino acid residues SER and GLY. The interactions between the complexes formed by PGR with pelargonidin were hydrophobic interactions, hydrogen bonding forces and Π-stacking. The amino acid residues that produce hydrophobic forces were LEU and TYR. The hydrogen bond interactions were mainly formed by phenolic hydroxyl groups on a benzene ring and amino acid residues GLN and ARG. The Π-Stacking interaction between protein and ligand was mainly generated by PHE. Different compounds were able to form complexes with the same protein receptor, and the binding sites where the forces were generated were basically the same. The results of the molecular docking simulation above showed that AR, PGR, and PTGS2 tended to be the potential target involved in cancer pathways, and kaempferol, pelargonidin, and baicalein could be active compounds to alleviate the symptoms of lung neoplasms. The detailed results of the interaction force are presented in supplementary material Supplemental Tables S3–S5.
Analysis of the Mechanism of YQSJF in the Treatment of Lung Neoplasms
Through gene enrichment analysis plug-in ClueGO, 54 signal pathway enrichment analysis was conducted on 53 key intersection targets of YQSJF and lung neoplasms, and the main signal pathways involved in the key targets were analyzed to study the possible mechanism of YQSJF in the treatment of lung cancer. Among them, nodes of the same color represent the same type of signal pathway, and the size of the node represents the significance of the signal pathway. The larger the node, the higher the significance of the signal pathway. 55 Ten major functional pathways were obtained by pathway enrichment analysis of intersection genes, such as non-small cell lung cancer, cocaine addiction, VEGF signaling pathway, human cytomegalovirus infection, morphine addiction, and salivary secretion, as shown in Figure 4 (Supplemental Table S6 in the Supplementary Material for the detailed information of pathway enrichment by ClueGO). Among them, some pathways have been confirmed by studies to be related to the treatment of lung cancer. For example, PVT1 promotes angiogenesis through targeting the miR-29c/VEGF signaling pathway in non-small-cell lung cancer. 56 IL-17 up-regulates VEGF-C expression to promote lymphangiogenesis, leading to transfer of non-small cell lung cancer. 57 IL-17 antibody has been used to reduce lung inflammation in mice exposed to long-term exposure to cigarette smoke. 58

Signal pathway enrichment analysis chart of YQSJF treatment for lung neoplasms. The signal pathway diagram is divided into 10 clusters by color code.
Conclusion
This network pharmacology method integrates ADME screening, establishes component-target-pathway-disease network visualization, PPI and biological function enrichment analysis, molecular docking, ClueGO plug-in enrichment pathway, and systematically analyzes the potential molecular mechanisms and biologically active ingredients of QYSJF in the treatment of lung cancer. Experimental studies have shown that YQSJF could effectively increase the biological processes in response to drugs, aging, and estrogen, regulate the release of neurotransmitters, and promote the combination of enzymes and drugs. In this study, 38 active ingredients and 57 corresponding core targets were identified from YQSJF, of which 53 targets are related to lung tumors. Kaempferol, β-sitosterol, quercetin, luteolin, hederagenin, stigmasterol, pelargonidin, 7-methoxy-2-methyl isoflavone, and baicalein acted on multiple targets, with far-reaching significance. The results of molecular docking indicated that almost all identified core components had strong affinities with AR, PGR, and PTGS2 proteins. Among them, AR docking with kaempferol, baicalein, PTGS2 docking with baicalein, and PGR docking with pelargonidin obtained the lowest binding energies, respectively, compared to other screened compounds. This suggests that kaempferol, baicalein, and pelargonidin might be important components of YQSJF in the treatment of lung cancer. Studies have shown that kaempferol inhibits the growth of A549 lung cancer cells and induces apoptosis by activating MEK-MAPK. 59 Baicalin improves the safety and effectiveness of drugs in the treatment of non-small cell lung cancer (NSCLC). Pelargonidin presents its anti-tumor efficacies by inducing autophagy, eliminating mitochondrial membrane potential, blocking the G2/M cell cycle, and down-regulating the PI3K/AKT signaling pathway. 60 All in all, many studies have confirmed that kaempferol, baicalin, and pelargonidin have the effects of assisting organisms to fight against lung neoplasms. By constructing a PPI network, it was indicated that the close effects of YQSJF between the various targets of lung neoplasms treatment were mainly related to the 11 target proteins CASP8, CASP9, AR, ESR1, PTGS2, NOS3, PGR, TGFB1, PPARG, RELA, and NOS2. The enrichment of biological processes pointed out that the active compounds of YQSJF participate in the activation of G-protein coupled acetylcholine receptor activity, steroid hormones, and other biological functions through core targets, and regulate the hepatitis b pathway, morphine addiction pathway and VEGF signaling pathway, so as to play roles in the treatment of lung tumors. VEGF coordinates the molecular mechanism of neurovascular homeostasis, plays a central role in the pathogenesis of various cancers, and is a key node in tumor immunotherapy. 61 This study elucidated that the active compounds of YQSJF might also be effective in treating lung tumors by regulating the relevant signal pathways of vascular diseases, which provide a potential direction for the next step in the research.
In general, this study confirmed that the treatment of lung cancer by YQSJF was a multi-compound, multi-target and multi-pathway synergistic effect, and predicted the possible material basis of efficacy and potential of the anticancer activity mechanisms of YQSJF by using network pharmacology and molecular virtual docking, providing an alternative strategy for the clinical treatment of lung neoplasms.
Supplemental Material
Supplementary Material 1 - Supplemental material for Identification of Targets and Active Components of Yiqi SanJie Formula Against Lung Neoplasms Based on Network Pharmacology Analysis and Molecular Docking
Supplemental material, Supplementary Material 1, for Identification of Targets and Active Components of Yiqi SanJie Formula Against Lung Neoplasms Based on Network Pharmacology Analysis and Molecular Docking by Tian-jiao Zhou, Jun-feng Liu, Ping Wang, An-na Hu, Lin-lin Chen and Jun-feng Zan in Natural Product Communications
Supplemental Material
Supplementary Material 2 - Supplemental material for Identification of Targets and Active Components of Yiqi SanJie Formula Against Lung Neoplasms Based on Network Pharmacology Analysis and Molecular Docking
Supplemental material, Supplementary Material 2, for Identification of Targets and Active Components of Yiqi SanJie Formula Against Lung Neoplasms Based on Network Pharmacology Analysis and Molecular Docking by Tian-jiao Zhou, Jun-feng Liu, Ping Wang, An-na Hu, Lin-lin Chen and Jun-feng Zan in Natural Product Communications
Supplemental Material
Supplementary Material 3 - Supplemental material for Identification of Targets and Active Components of Yiqi SanJie Formula Against Lung Neoplasms Based on Network Pharmacology Analysis and Molecular Docking
Supplemental material, Supplementary Material 3, for Identification of Targets and Active Components of Yiqi SanJie Formula Against Lung Neoplasms Based on Network Pharmacology Analysis and Molecular Docking by Tian-jiao Zhou, Jun-feng Liu, Ping Wang, An-na Hu, Lin-lin Chen and Jun-feng Zan in Natural Product Communications
Supplemental Material
Supplementary Material 4 - Supplemental material for Identification of Targets and Active Components of Yiqi SanJie Formula Against Lung Neoplasms Based on Network Pharmacology Analysis and Molecular Docking
Supplemental material, Supplementary Material 4, for Identification of Targets and Active Components of Yiqi SanJie Formula Against Lung Neoplasms Based on Network Pharmacology Analysis and Molecular Docking by Tian-jiao Zhou, Jun-feng Liu, Ping Wang, An-na Hu, Lin-lin Chen and Jun-feng Zan in Natural Product Communications
Supplemental Material
Supplementary Material 5 - Supplemental material for Identification of Targets and Active Components of Yiqi SanJie Formula Against Lung Neoplasms Based on Network Pharmacology Analysis and Molecular Docking
Supplemental material, Supplementary Material 5, for Identification of Targets and Active Components of Yiqi SanJie Formula Against Lung Neoplasms Based on Network Pharmacology Analysis and Molecular Docking by Tian-jiao Zhou, Jun-feng Liu, Ping Wang, An-na Hu, Lin-lin Chen and Jun-feng Zan in Natural Product Communications
Supplemental Material
Supplementary Material 6 - Supplemental material for Identification of Targets and Active Components of Yiqi SanJie Formula Against Lung Neoplasms Based on Network Pharmacology Analysis and Molecular Docking
Supplemental material, Supplementary Material 6, for Identification of Targets and Active Components of Yiqi SanJie Formula Against Lung Neoplasms Based on Network Pharmacology Analysis and Molecular Docking by Tian-jiao Zhou, Jun-feng Liu, Ping Wang, An-na Hu, Lin-lin Chen and Jun-feng Zan in Natural Product Communications
Footnotes
Acknowledgments
The authors would like to thank all the colleagues and students who contributed to this study.
Author note
Conceptualization, Junfeng Zan and Junfeng Liu; methodology, Junfeng Liu; software, Anna Hu; validation, Tianjiao Zhou and Junfeng Zan; formal analysis, Junfeng Liu; investigation, Ping Wang; resources, Tianjiao Zhou; data curation, Linlin Chen; writing—original draft preparation, Tianjiao Zhou; writing—review and editing, Junfeng Liu; visualization, Tianjiao Zhou; supervision, Junfeng Zan; project administration, Junfeng Zan; funding acquisition, Junfeng Liu. All authors have read and agreed to the published version of the manuscript.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was funded by Key projects of Hubei Provincial Department of Education, Study on the basis of antitumor active substance of phlegm-reducing agents and its mechanism of action, grant number D20142003.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
