Abstract
Two new xanthone derivatives, spinosusones A (
Introduction
In recent years, marine-derived fungi have attracted much attention from natural medicinal chemists, and many secondary metabolites possessing intriguing structures and various activities have been reported.
1
Aspergillus is a chemically brilliant fungal genus with immense potential to produce a wide range of unique secondary metabolites including polyketides,2–4 alkaloids,5–7 peptides8,9 and terpenoids.10,11 Some of these metabolites exhibit intriguing biological properties, including antimicrobial,5,7,11 pro-angiogenesis,
6
and cytotoxic,2,4,10 as well as enzyme-inhibiting activities.
8
In a current study, Aspergillus sp. WHUF03110 was fermented through a solid fermentation methodology and the ethyl acetate extract of the strain showed antibacterial activity against Staphylococcus aureus and Escherichia coli. Continuing our efforts on the discovery of the chemical diversity and biological activities of natural products, chemical investigation was performed. Three new compounds, spinosusone A (

Structures of compounds (1-12) isolated from Aspergillus sp. WHUF03110.
Results and Discussion
Compound

The key HMBC correlations of 1-3.
1H and 13C NMR Data for Compounds 1 and 2 (600/150 MHz, in CDCl3, δ ppm).
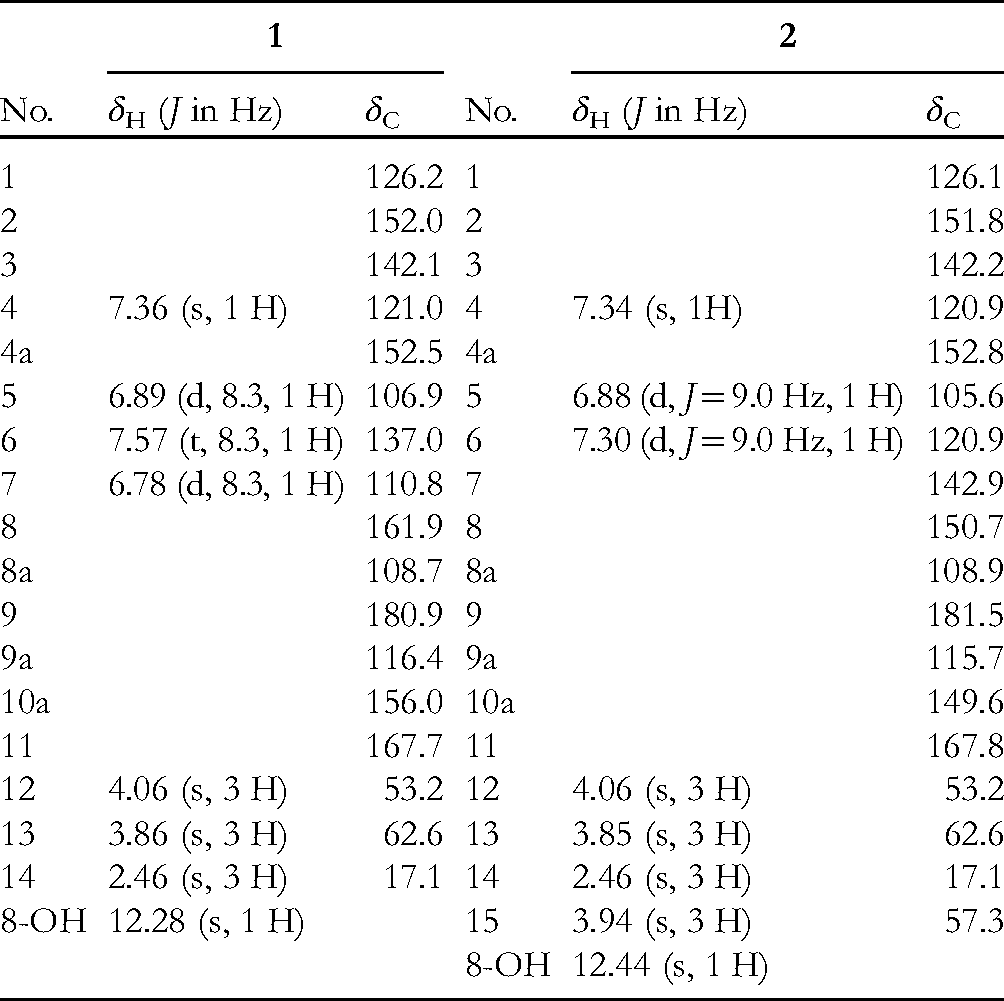
Compound
Compound
1H and 13C NMR Data for Compound 3 (600/150 MHz, in CDCl3, δ ppm).

Most of the compounds were evaluated for their antimicrobial activities against ten Gram-negative bacteria (H. pylori 26695, H. pylori G27, H. pylori 159, H. pylori 129, E. coli MG1655, P. aeruginosa PAO1, A. baumannii ATCC19606, K. neumoniae ATCC35657, S. typhimurium 14028 s, S. dysenteriae), seven Gram-positive bacteria (E. faecalis FA2-2, E. faecium ATCC19434, B. subtilis 168, S. aureus ATCC25923, S. aureus NEWMAN, S. aureus USA300, S. aureus NRS 271), M. smegmatis ATCC 607, C. albicans ATCC SC5314 and C. albicans YY-1-4. The results are shown in Table S1. Compound
Experimental
General
Optical rotations were measured using a JASCO P-1020 polarimeter (Jasco Tokyo Japan), and the CD spectrum on a JASCO J-815 instrument. The HRESIMS data were obtained on an Agilent 6210 TOF MS system (Agilent Technologies, Santa Clara, CA, USA), UV spectra, recorded in MeOH, on a Shimadzu UV spectrometer-1800 (Shimadzu Corp., Kyoto, Japan), and IR spectra (KBr) on a Nicolet 6700 FT-IR spectrometer (Thermo electric nicoli, United States). 1H and 13C NMR spectra were obtained on a Bruker AVANCE III 600 MHZ spectrometer (600 MHz for 1H NMR and 150 MHz for 13C NMR) with TMS as internal standard (Bruker company, Switzerland). Semi-preparative reversed-phase HPLC (PR-HPLC) was performed on an Agilent ZORBAC SB-C18 column (5 μm, 250 × 9.4 mm id) with Agilent 1260 separation. Silica gel GF254 for TLC, supplied by the Yantai Zhifu Huangwu Silicone Factory, Yantai, China, and Sephadex LH-20 gel (GE Healthcare, Uppsala, Sweden) were used. Compounds were detected on TLC either under UV light or by heating after spraying with a solution of 40 mL H2SO4 and 10 g Vanillin in 500 mL H2O. The microorganism test strains were: Gram-negative Bacteria (Helicobacter pylor 26695, H. pylor G27, H. pylor 159, H. pylor 129, Escherichia coli MG1655, Pseudomonas aeruginosa PAO1, Acinetobacter baumannii ATCC19606, Klebsiella pneumoniae ATCC35657, Serovar typhimurium 14028 s, Shigella dysenteriae); Gram-positive bacteria (Enterococcus faecalis FA2-2, Enterococcus faecium ATCC19434, Bacillus subtilis 168, Staphylococcus aureus ATCC25923, S. aureus NEWMAN, S. aureus USA300, S. aureus NRS 271); Mycobacterium smegmatis ATCC 607, Candida albicans ATCC SC5314 and C. albicans YY-1 to 4. All were provided by the Department of Pathogen Biology & Jiangsu Key Laboratory of Pathogen Biology, Nanjing Medical University, Nanjing, China.
Fungal Material
Strain WHUF03110 was isolated on GPY medium (glucose 20 g, peptone 10 g, yeast extract 10 g, sea salts 15 g, agar 20 g, ddH2O 1 L, pH 7.5) from a mangrove soil sample collected from Yalong Bay, at Sanya, Hainan, China, in Dec, 2018. The strain was first selected by the strong antibacterial activities against Staphylococcus aureus and Escherichia coli of its crude 80% methanol extract of its agar plate culture on PDA (Potato Dextrose Water 25 g, agar 20 g, sea-salt 15 g, ddH2O, 1 L, pH 6.0 ± 0.2). The fungus was identified as Aspergillus sp. based on its morphological characteristics and the internal transcribed spacer (ITS) sequence (626 bp), which has 99.65% similarity to Aspergillus spinosus NRRL 5034 (NCBI Reference Sequence: NR_137489.1); the sequenced data derived from the fungal strain have been deposited in GenBank (accession no. MZ661122). The strain is stored at −80°C in the School of Pharmaceutical Sciences, Wuhan University, China.
Fermentation
The fungus was cultured on seed A medium-agar plates at 28°C for 3 days. The seed A medium (soluble starch 15 g, glucose 5 g, peptone 5 g, yeast extract 5 g, [NH4]2SO4 0.5 g, K2HPO4 0.5 g, NaCl 0.5 g, MgSO4 0.5 g, CaCO3 1 g, made up to 1 L with water, pH 7.5) was inoculated with strain WHUF03110 and incubated at 28 °C for 72 h on a rotating shaker (180 rpm). The mass scale fermentation of strain WHUF03110 was carried out using solid rice medium in 1000 mL flasks (rice 200 g, distilled water 200 mL) inoculated with 15 mL of seed solution. Flasks were statically fermented for 30 days at 26 °C in a solid rice medium.
Extraction and Isolation
The entire fermented cultures from 80 flasks were harvested and ultrasonically extracted with EtOAc, three times for 30 min each time. The EtOAc extract (57.0 g) was fractionated by silica gel (200-300 mesh) column chromatography, eluting with a light petroleum-EtOAc gradient system (10:1, 4:1, 1:1, v/v), and a dichloromethane-MeOH gradient system (10:1, 0:1, v/v) to afford nine fractions A-I. Fraction D (3.6 g) was separated on a silica gel column, eluting with light petroleum-EtOAc (15:1, 8:1, 2:1, v/v), to yield fractions D1-D8. Fraction D2 (1.8 g) was subjected to silica gel column chromatography eluting with light petroleum-EtOAc (15:1, 12:1,8:1, v/v) to give 6 subfractions (Fr. D2A-D2F). Fr. D2C (650.0 mg) was purified by HPLC eluting with MeOH-H2O (65:35, v/v) to afford
Spinosusone A (
Spinosusone B (
Asperdiazapinone G (
Antimicrobial Assays
The antibacterial and antifungal assays were conducted in 96-well plates using a broth micro dilution method. 25 The specific experiment was as follows: the initial cultures for H. pylori strains were maintained on Brain Heart Infusion (BHI) medium, and other pathogenic strains were maintained on Luria Broth (LB) agar. A single bacterial colony was picked, and suspended in LB/BHI broth to approximately 106 CFU/mL. Each test compound was dissolved in MeOH and serially diluted to 9 different concentrations (16, 8, 4, 2, 1, 0.5, 0.25, 0.125, 0.0625 μg/mL) using LB broth on a 96-well plate. An aliquot (10 μL) of bacterial suspension was then added to each well. Sterile water was used as the negative control. Amoxicillin, vancomycin, and amphotericin B were used as the positive control for H. pylori, S. aureus, C. albicans strains, respectively. Ampicillin was used as the positive control for the other strains. The plate was incubated at 37°C aerobically for 24 h. MIC values were defined as the minimum concentration of compound that inhibited visible bacterial growth.
Conclusions
In conclusion, twelve compounds, including three new compounds (
Footnotes
Acknowledgments
This research was funded by grants from National Key Research and Development Program of China. (Grant Numbers: 2018YFC0311000) and National Natural Science Foundation of China. (Grant Numbers: 81603255). We thank Jun-Nan Shen and De-Sheng Li for their contribution on strain isolation and crude extracts preparation.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Key Research and Development Program of China (grant number 2018YFC0311000, 81603255).
Ethical Approval
Our institution does not require ethical approval for reporting individual cases or case series.
Statement of Human and Animal Rights
This article does not contain any studies with human or animal subjects.
Statement of Informed Consent
There are no human subjects in this article and informed consent is not applicable.
Trial Registration
Not applicable, because this article does not contain any clinical trials.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
