Abstract
Endolichenic fungi (ELF) serve as a novel source of natural products with distinctive bioactivities. A total of 58 ELF isolated from 29 lichens collected from mangrove and mangrove-associated plants of Negombo lagoon, Sri Lanka were identified using morphological and DNA barcoding techniques. Ethyl acetate extracts of 18 such identified ELF isolates were subjected to
Introduction
The discovery of biologically active compounds has opened a diverse “chemical space” in natural product chemistry. 1 Microorganisms have played a major role in the discovery of novel bioactive small molecules, contributing to the identification of more than 22 500 bioactive metabolites. 2 Among them, lichens and endolichenic fungi (ELF) are emerging as a treasure trove for novel bioactive metabolites.
Lichens are a symbiosis between a fungal component, which makes up the bulk of the thalli while providing access to water and minerals, and a cyanobacterium and/or alga, which provides the photosynthate. Interestingly, this is a physiological combination of microorganisms belonging to 3 different Kingdoms. Moreover, these lichens provide grounds for the exploration of more microorganisms that live on them asymptomatically.3,4 ELF are one such group, which live inside the thallus of a lichen, analogous to endophytic fungi residing in plant tissues. 5
Since the first report of heptaketides from ELF published in 2007,
5
many molecules have been identified with potent bioactivities around the world.
6
Such potent metabolites have been discovered during previous studies conducted in Sri Lanka as well, which include polyketides from ELF
Our previous research was conducted using the lichens in the mangrove ecosystem of Puttalam lagoon in the dry zone of Sri Lanka 11 while the current study was undertaken in the mangroves of the Negombo lagoon in the wet zone of the country. Constant exposure to high salinity, extreme tides and temperature changes in this area make it a challenging environment not only for lichens and ELF but also for any organism to survive. In the current study, ELF from the mangrove ecosystem were identified using DNA barcoding of the internal transcribed spacer (ITS) region and their crude extracts were screened for antioxidant, anti-inflammatory, antibacterial and tyrosinase inhibitory potentials. Liquid chromatography–mass spectrometry (LC–MS) dereplication, an integrated approach of combining traditional fraction collection, sample preparation, and multicomponent analysis into a single step, was performed for the understanding of metabolite profiles of each representative isolate obtained in the study. This is the first such study focusing on metabolic profiles of the mangrove associated ELF in Sri Lanka.
Results and Discussion
Identification of Lichens
It is reported that nearly 1200 lichen species are known from Sri Lanka.
12
Host plant and lichen species of the 29 lichen samples collected are shown in Table 1. The majority of the samples belonged to the crustose type, which included

Selected lichens collected from mangrove and mangrove-associated plants in Negombo lagoon, Sri Lanka. (A)
Endolichenic Fungal Strains Isolated From the Lichens of Mangrove and Mangrove-Associated Plants in Negombo Lagoon, Sri Lanka.
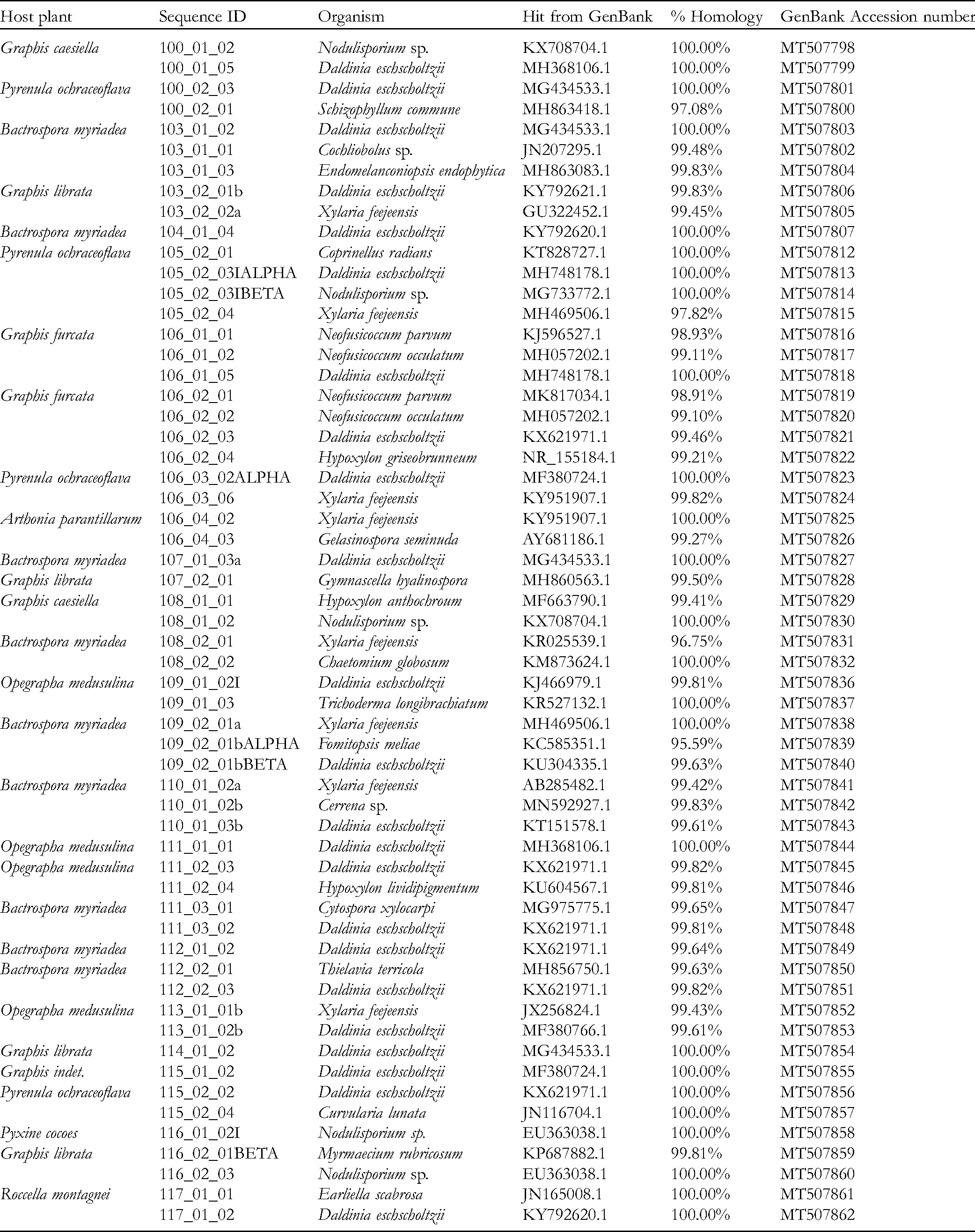
Isolation and Identification of ELF
A total of 58 fungal isolates were isolated into pure cultures from 29 lichen samples. Species confirmation was achieved by comparing sequences of ITS-rDNA region with the established (voucher specimens or published) records with the highest % similarity of the GenBank accessions and are shown in Table 1. Isolates belonged to 23 different species, out of which 18 belonged to phylum Ascomycota and only 5 to Basidiomycota. Out of the Ascomycetes, 9 genera belonged to the class Sordariomycetes, 5 to Dothideomycetes and only 1 to Eurotiomycetes. All genera found under the phylum Basidiomycota belonged to the class Agaricomycetes. The highest number of isolates (23) belonged to the species
Molecular phylogenetic analysis was performed using ITS sequences. Multiple sequence alignment (MSA) was made using ∼500 to 600 length of ITS sequences. The maximum likelihood tree with the highest log likelihood (−813.33) is shown in Figure 2. The tree was drawn to scale in combination with the already available sequences for each species in GenBank. The branch lengths were measured by the number of substitutions per site. In the analysis, isolates were clearly grouped according to families and genera. Fungal species clustered together regardless of their lichen host demonstrating that in most cases, ELF diversity is not determined by the lichen host.
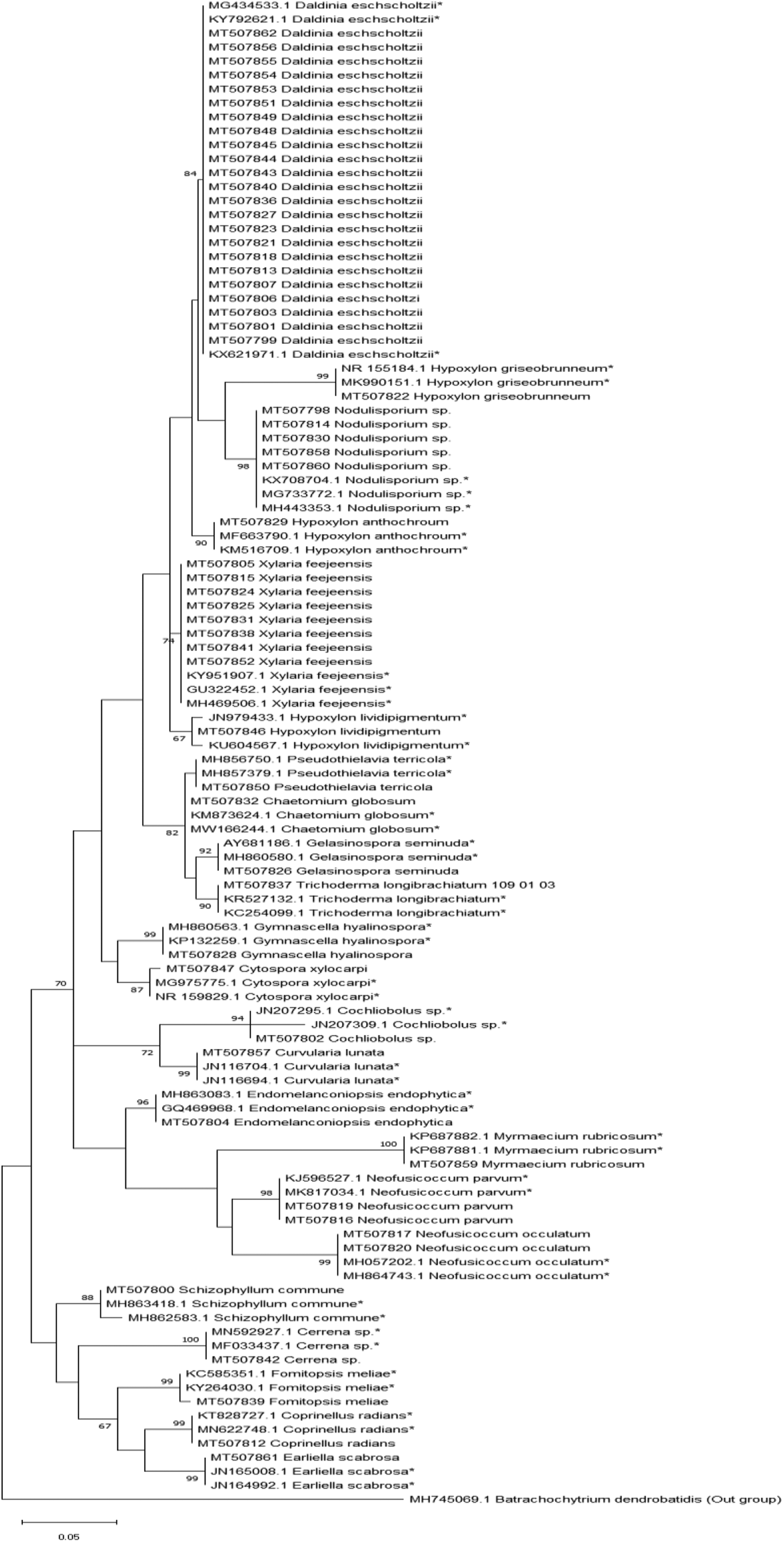
Maximum likelihood tree with the highest log likelihood showing phylogenetic placement of endolichenic fungi isolated from lichens in the mangrove and mangrove-associated plants in Negombo lagoon, Sri Lanka. *Indicates the reference sequences from the GenBank.
Qualitative Analysis of Metabolites in Extracts of ELF
Qualitative analysis of crude extracts performed using thin-layer chromatography (TLC) indicated that the chemical constituents of the same fungal species isolated from different lichens were similar to each other. Based on their close phylogenetic relationships, only one isolate per selected species was used in bioassays.
Bioassays and LC–MS dereplication were conducted for 18 selected isolates due to the sterility of the remaining 5 strains which obstructed further culturing.
Screening of Radical Scavenging Activity (DPPH Assay)
The extreme reactivity of free radicals can cause damage to macromolecules inside our body, making antioxidants important in protecting tissues from many noncommunicable diseases. 14 2, 2-diphenyl-1-picrylhydrazyl (DPPH) consists of a delocalized electron and produces a characteristic purple color in a solution of methanol, which changes to a residual pale yellow in the presence of an antioxidant. This absorbance can be measured at 517 nm. 15
The lowest half-maximal scavenging concentration (SC50) values were shown by the crude extracts of
Moreover, a considerable amount of activity was observed for
Activity displayed by the extracts of
Anti-inflammatory Assay (HRBC Stabilization Method):
The regulation of an inflammatory response can be achieved by stabilizing the lysosomal membrane which otherwise would release the constituents to trigger inflammation. Since the lysosomal membrane is analogous to the erythrocytes’ membrane in humans, this assay provides the technique to assess anti-inflammatory potency of the extracts using the heat-induced hemolysis principle. 16
The extract of
The extracts of
The IC50 values shown by
Tyrosinase Inhibitory Assay
While giving protection from ultraviolet (UV) rays, overproduction of melanin can cause skin pigment disorders and therefore the cosmetic industry is also interested in the reduction of melanin production. This reduction can be achieved by inhibiting the enzyme tyrosinase, which will prevent a major step in the production of melanin inside melanocytes. 21
In contrast to other bioassays, the tyrosinase inhibitory activity shown by extracts was slightly less. However, the extract of
The extract of
Antibacterial Assay (Agar Well Diffusion Method)
Antibiotic resistance is of major global concern which has highlighted the importance of the discovery of clinically proven new antibiotics during the past few decades.
25
Hence this assay was carried out to assess activity against common aerobic bacterial species
Considering the inhibition zone values, extract of
The extract of
The extracts of
Inhibition against
Only
Activity comparisons in bioassays are presented graphically in Figure 3.

The activity comparison of bioassays. (A) SC50 comparison for DPPH (BHT as the positive control); (B) IC50 comparison for HRBC stabilization assay (aspirin as the positive control); (C) IC50 comparison for tyrosinase inhibitory assay (kojic acid as the positive control); (D) antibacterial assay inhibition zone comparison for highest dose (azithromycin for
LC–MS Profile of Crude Extracts
LC–MS dereplication has been found to be an efficient method to scrutinize the chemical diversity of organisms in the field of natural product chemistry. It provides an opportunity to differentiate previously studied secondary metabolites from novel active compounds.
36
LC–HRMS analysis was performed and the accurate mass data were used along with the database Dictionary of Natural Products (DNP) to find the reported close hit for the particular peak. Multiple filters were applied in this process such as the inclusion of genus, species and subsequently fungi as a broad category. All the suggested structures have shown similar UV profiles along with the accurate mass defect error below 10 PPM, which is well within the acceptable range. Accordingly, from the filter of species, 4 molecules were dereplicated from the fungus
Suggested structures in LC–MS dereplication in species and genus level search

Abbreviations: LC–MS, liquid chromatography–mass spectrometry; UV, ultraviolet; ELF, endolichenic fungi
Furthermore, TLC visualization conducted using visibility under UV at 254 and 366 nm proved the existence of aromatic and highly conjugated compounds in the extracts. The derivatization conducted using anisaldehyde–sulfuric acid reagent showed blue, violet, red, and yellow spots, which were indicative of phenols and terpenes being present. 44 The TLC profiles of the crude extracts in the respective optimized solvent systems are available in the supplemental material Supplemental Material (File S3).
Materials and Methods
Ethical Statement
A permit to collect lichen samples from the mangrove forest in Negombo lagoon was obtained from the Forest Department of Sri Lanka. The samples were collected from the trees in the area of “Kadolkele” under the observation of a representative of the department, without causing any disturbance to the surroundings.
Collection of Lichen Samples
Collection of 29 lichen specimens was carried out on March 31, 2018 at the mangrove forest of Negombo lagoon, Sri Lanka. The study site was “Kadolkele” area situated at 7.195528 N 79.84389 E GPS coordinates belonging to the lagoon area. The lichens were collected from mangrove and mangrove associated plants belonging to 6 different species positioned mainly at the water’s edge and a few towards the land. Samples were collected from root, stem and bark with an approximated elevation of 1 m from the sea level. Collected specimens were placed in acid-free paper bags, labeled appropriately and transported to the Department of Chemistry, University of Kelaniya. Samples were stored at 4 °C to be processed for the isolation of ELF within 2 weeks of sampling.
Identification of Lichens
Portions of lichens of 2 × 2 cm were refrigerated for 2 weeks and air dried. The identification of these 29 specimens (Figure 2) was carried out using morphological and molecular techniques at the Natural History Museum, London and photographs were taken using Olympus stereomicroscopes and Olympus compound microscopes with interference contrast, connected to a Nikon Coolpix digital camera. Tap water was utilized for section mounting for all measurements. Duplicates of each specimen are deposited in the Department of Chemistry, University of Kelaniya.
Isolation of Endolichenic Fungi
The lichen thalli were washed in running tap water to remove unwanted particles and then surface sterilized using consecutive immersion in 96% ethanol for 10 s, 0.5% sodium hypochlorite for 2 min, and 70% ethanol for 2 min and dried using sterile filter paper. Small segments of the lichen thallus (∼3 × 3 mm) were then cut and placed on 2% malt extract agar supplemented with 0.01% streptomycin. Sealed plates were then incubated under ambient conditions (room temperature 30 °C) for 14 days and observed regularly. Hyphal tips from each morphologically distinct colony were placed on new potato dextrose agar (PDA) plates and subjected to several sub-cultures to obtain a pure culture.
Extraction of Genomic DNA and Polymerase Chain Reaction
The isolates were inoculated in potato dextrose broth and incubated at room temperature (30 ± 32 °C) for 1-week. Mycelia were separated from the broth, washed with sterilized water, excess water was removed, and DNA extracted using a slightly modified method of the CTAB DNA extraction protocol. 45 The quality of extracted DNA samples was tested using 1% agarose gel electrophoresis and samples were stored at −20 °C for further utilization. The nuclear ribosomal DNA ITS region (rDNA-ITS) was amplified using polymerase chain reaction (PCR); the fungal specific ITS1/ITS5 forward and ITS4 reverse universal primers were used. 46 The PCR reaction mixture was comprised of 2 μL fungal DNA, 1 × PCR buffer, 1.5 mM MgCl2, 0.2 mM each dNTP, 1 μM each forward and reverse primer, and 1.25 units of Taq DNA polymerase (Promega). PCR protocol consisted of an initial denaturation step at 94 °C for 5 min, followed by 30 cycles of 94 °C for 30 s, 55 °C for 30 s, and 72 °C for 30 s, with a final extension step of 72 °C for 10 min and was performed in a Veriti™ Thermal Cycler (Applied Biosystems). The amplification of the ITS region was confirmed by separating the amplified product using 1% agarose gel electrophoresis. The PCR products were subjected to Sanger dideoxy sequencing bi-directionally at Genentech Institute, Colombo, Sri Lanka. DNA sequences were manually edited using BioEdit sequence Alignment Editor (Version 7.2.5) and compared with the sequences available in the GenBank using Basic Local Alignment Search Tool (BLAST) to assess homology. DNA sequences of the identified ELF species were deposited at the National Center for Biotechnology Information (NCBI) database and accession numbers were obtained. ELFs identified in this study are shown in Table 1 with the percent similarity to the published/authenticated sequences at the GenBank and the accession numbers of the deposited sequences.
Phylogenetic Analysis of ELF
DNA sequences were aligned by using an MSA algorithm, Multiple Sequence Comparison by Log-Expectation (MUSCLE),
47
and poorly aligned positions and divergent regions of the alignment were removed/edited manually. All characters were equally weighted and the gaps were treated as missing data. A sequence of the ITS region of
Extraction of Secondary Metabolites
Each endolichenic fungus was cultured on 5 PDA 150 × 25 mm plates and incubated at room temperature for 2 weeks. The mycelium of each fungus with the medium was cut into small pieces, extracted with ethyl acetate (6 × 500 mL) and the solvent evaporated under reduced pressure using a Rotary Evaporator (IKA RV 10B S96). The evaporated ethyl acetate extracts were then transferred to glass vials separately and N2 was passed through the samples to remove any remaining solvent. The resulting semisolid extracts were stored at 0 °C for further analysis. 7
Radical Scavenging Ability by DPPH Method
The radical scavenging ability assay was carried out in a flat-bottom 96-well microtiter plate, according to the colorimetric method described by Chatatikunand Chiabchalard,
49
with slight modifications. A volume of 160 µL with a series of doses of each test sample and standard antioxidant, BHT (12.50, 25.00, 50.00, 100.00, 200.00, 400.00, 800.00 μg/mL) was added to 40 μL of 0.25 mM methanolic DPPH solution in a 96-well plate. All reagents were mixed and incubated for 15 min at room temperature protected from light. The absorbance of each well was measured at 517 nm with a Microplate Reader (Biotek). The percentage inhibition was calculated using Equation (1). The SC50 values of all the crude extracts were calculated using GraphPad Prism 7 Statistical Software. The experiment was carried out in triplicates.
Anti-inflammatory Assay (HRBC Stabilization Method)
The anti-inflammatory assay was conducted using the human red blood cells (HRBCs) stabilization method, which uses the heat-induced hemolysis principle. 50 Test samples and the positive control Aspirin (1 mg of each) were dissolved in 0.5 mL of dimethyl sulfoxide (DMSO) and diluted up to 5 mL by addition of normal saline. A half dilution series was prepared with a control for each sample and the reaction mixture was prepared with 2.5 mL of the solution and 0.25 mL of 10% v/v RBC suspension. All the centrifuge tubes containing reaction mixtures were incubated in a water bath at 56 °C for 30 min after which the tubes were cooled under running tap water. The reaction mixture was centrifuged at 3000 rpm for 10 min to obtain a clear liquid phase and the absorbance of the supernatants was measured at 560 nm using Microplate Reader (Biotek) by using aliquots of 200 µL from respective mixtures for each well in a 96 well-plate. The test was performed in triplicates and the percentage stability was calculated using Equation (1). IC50 values were calculated using the statistical software GraphPad Prism 7..
Tyrosinase Inhibitory Assay
Tyrosinase inhibitory assay was carried out in a flat-bottom 96-well microtiter plate according to a modified version of the method described by Souza et al
21
Crude extracts and positive control kojic acid were dissolved in 10% DMSO in phosphate buffer (pH 6.5) and a half dilution series of doses was prepared for each (0.0, 10.41, 20.83, 41.66, 83.33, 166.60, 333.30 μg/mL). An aliquot of 70 μL of each extract dilution was combined with 30 μL of tyrosinase (333 U/mL). Following incubation at room temperature for 5 min, 110 μL of substrate (2 mM L-3,4-dihydroxyphenylalanine [L−DOPA]) was added to each well. Incubation was continued for 30 min at room temperature under dark conditions and absorbance was measured at 490 nm with a Microplate Reader (Biotek). The percentage inhibition was calculated using the Equation (2).
Antibacterial Assay (agar Well Diffusion Method)
Antibacterial assays against aerobic bacterial species
In the case of the anaerobic species, an aliquot of overnight cultures in trypticase soy broth (supplemented with blood with lysed RBC and grown under anaerobic conditions in a jar containing anaerobic sachets at 37 °C) was used in the same manner as previously described on trypticase soy agar (supplemented with lysed RBC) plates. Well and extract preparation and filling were also conducted in a similar manner with similar concentrations and volumes. Chlorhexidine (0.2%) was used as the positive control. After an incubation of 24 h at 37 °C under anaerobic conditions, the diameters of observable inhibition zones were measured in cm.
Analyzing LC–MS Profile of Crude Extracts
LC–MS dereplication was carried out for some extracts to discern interesting active ingredients and novel molecules present in them. Chromatographic separation of crude extracts was performed on an Agilent 1290 Infinity 2 utra high performance liquid chromatography (UHPLC) system (with photodiode-array (PDA) detector, electrospray ionization (ESI) source, quadrupole time-of-flight (Q-TOF) analyzer, 5 µm and 4.6 × 250 mm C18 column) using a 1 mL/min solvent gradient. Samples with a concentration of 5 mg/mL were prepared in LC–MS graded methanol. The solutions were filtered through a 0.22 µM durapore filter and transferred to clean and dry vials. In total, 5 µL was injected into the column from each extract. The mobile phase system consisted of 0.1% formic acid (A) and methanol (B), and the gradient was set up as 0 to 2 min 30% A/70% B, a linear gradient from 2 to 8 min to 5% A/95% B and a linear gradient from 12 to 15 min to again 30% A/70% B. The obtained masses were thoroughly searched in DNP and SciFinder, using genus and species level specificity for rationalization of the obtained profiles. TLC visualization was also carried out to aid the validation of the presence of different metabolites in the extract. The visualizing was conducted under UV wavelengths 254 and 366 nm and derivatization was conducted using anisaldehyde–sulfuric acid reagent (0.5 mL anisaldehyde, 10 mL glacial acetic acid, 5 mL concentrated sulfuric acid and 85 mL methanol).
Conclusions
ELF has become a promising source for providing novel bioactive compounds and this study adds more evidence to that already proven statement. In the present study, 58 ELF strains, isolated from 29 lichens collected from mangrove and mangrove associated plants of Negombo lagoon, Sri Lanka, were identified using molecular and morphological techniques. In order to determine antioxidant, anti-inflammatory, tyrosinase inhibitory, and antibacterial potency, ethyl acetate extracts of ELF belonging to 18 species were utilized.
The LC–MS dereplication study conducted for the same 18 extracts revealed that they are comprised of interesting metabolites reported from the literature while containing some unknown major compounds as well. Many major peaks observed in the extracts of ELF
Supplemental Material
sj-docx-1-npx-10.1177_1934578X211048652 - Supplemental material for Bioactive Properties and Metabolite Profiles of Endolichenic Fungi in Mangrove Ecosystem of Negombo Lagoon, Sri Lanka
Supplemental material, sj-docx-1-npx-10.1177_1934578X211048652 for Bioactive Properties and Metabolite Profiles of Endolichenic Fungi in Mangrove Ecosystem of Negombo Lagoon, Sri Lanka by Ramani H. Weerasinghe, Kasun Maduranga, Renuka N. Attanayake, Chaitrali Shevkar, Abhijeet S. Kate, Gothamie Weerakoon, Kiran Kalia and Priyani A. Paranagama in Natural Product Communications
Supplemental Material
sj-docx-2-npx-10.1177_1934578X211048652 - Supplemental material for Bioactive Properties and Metabolite Profiles of Endolichenic Fungi in Mangrove Ecosystem of Negombo Lagoon, Sri Lanka
Supplemental material, sj-docx-2-npx-10.1177_1934578X211048652 for Bioactive Properties and Metabolite Profiles of Endolichenic Fungi in Mangrove Ecosystem of Negombo Lagoon, Sri Lanka by Ramani H. Weerasinghe, Kasun Maduranga, Renuka N. Attanayake, Chaitrali Shevkar, Abhijeet S. Kate, Gothamie Weerakoon, Kiran Kalia and Priyani A. Paranagama in Natural Product Communications
Supplemental Material
sj-docx-3-npx-10.1177_1934578X211048652 - Supplemental material for Bioactive Properties and Metabolite Profiles of Endolichenic Fungi in Mangrove Ecosystem of Negombo Lagoon, Sri Lanka
Supplemental material, sj-docx-3-npx-10.1177_1934578X211048652 for Bioactive Properties and Metabolite Profiles of Endolichenic Fungi in Mangrove Ecosystem of Negombo Lagoon, Sri Lanka by Ramani H. Weerasinghe, Kasun Maduranga, Renuka N. Attanayake, Chaitrali Shevkar, Abhijeet S. Kate, Gothamie Weerakoon, Kiran Kalia and Priyani A. Paranagama in Natural Product Communications
Footnotes
Acknowledgments
Authors thank Ms Patricia Wolseley, Algae, Fungi and Plants Division, Department of Life Sciences, The Natural History Museum, London for the support rendered in lichen identification and reviewing of the draft and Prof. Mala Amarasinghe, Department of Plant and Molecular Biology, University of Kelaniya, for helping in the identification of mangrove host plants.
Author Contributions
Experimentation: Collection, isolation, screening and data analysis: Ramani Weerasinghe; Collection assistance and phylogenetic analysis: Kasun Maduranga; LCMS analysis: Chaitrali Shevkar; Fungal molecular identification and analysis: Renuka Attanayake; Lichen identification: Gothamie Weerakoon; LCMS analysis and dereplication: Abhijeet Kate, Kiran Kalia. Writing-original draft: Ramani Weerasinghe, Kasun Maduranga, Chaitrali Shevkar. Writing-review and editing: Renuka Attanayake, Gothamie Weerakoon, Abhijeet Kate, Kiran Kalia, Priyani Paranagama. Funding acquisition: Priyani Paranagama. Overall supervision: Priyani Paranagama.
Data Availability Statement
The data used to support the findings of this study are available from the corresponding author upon request.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Ministry of Science Technology and Research, Sri Lanka (grant number MSTR/TRD/AGR/03/02/07).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
