Abstract
Introduction
Lichens, a fascinating symbiotic association between fungi (mycobiont) and photosynthetic partners (photobiont), are a treasure trove for natural product discovery.
1
These unique organisms are known to produce a diverse array of structurally complex and biologically active secondary metabolites.
2
Recent advancements in culturing lichen mycobionts independently of the photobiont have opened exciting new avenues for exploring their biosynthetic potential.
3
Mycobionts derived from lichens have produced a diverse range of secondary metabolites, some of which exhibit uniqueness and notable divergence from compounds obtained from the original lichens.2,4,5 The chemical structures of these metabolites show homology to fungal secondary metabolites.6–8

Chemical structures of
Materials and Methods
General Experimental Procedures
The HRESI mass spectrum was obtained using a MicrOTOF–Q mass spectrometer coupled with an LC-Agilent 1100 LC-MSD Trap spectrometer. NMR spectra were recorded on a Bruker Avance III spectrometer (500 MHz for 1H NMR and 125 MHz for 13C NMR). Experimental ECD data were recorded using a JASCO J-815 Spectropolarimeter. Gravity column chromatography was carried out on silica gel 60 (0.040-0.063 mm, Himedia). Thin-layer chromatography (TLC) was conducted on precoated silica gel 60 F254 or silica gel 60 RP–18 F254S (Merck), and spots were visualized by spraying with a 10% H2SO4 solution followed by heating.
Lichen Material
The lichen sample used in this research was obtained from the bark of tree located in Quang Ngai city (15.104 N; 108.806 E), Vietnam (approx. 9 m alt.), during March 2021. Identification of the lichen specimen as
Mycobiont Culture
This research applied the mycobiont cultivation method outlined by Do et al (2022).
5
In brief, tiny fragments of mycobiont colonies were aseptically transferred onto the surface of MY10 medium (composed of malt extract 1%, yeast extract 0.4%, sucrose 10%, and agar 1.5%,
Extraction and Isolation
Following the method established by Nguyen et al (2023),
12
the mycobiont colonies obtained (58 g dry weight) were ground into a fine powder, followed by maceration with EtOAc (in 10 portions of 300 mL each) at room temperature. The combined extracts were concentrated using a rotary evaporator to yield a crude extract (340 mg). The resulting crude extract was fractionated by silica gel column chromatography (CC), eluted with a gradient system of
Hydroxydiorygmone A (1 )
Colorless oil; [
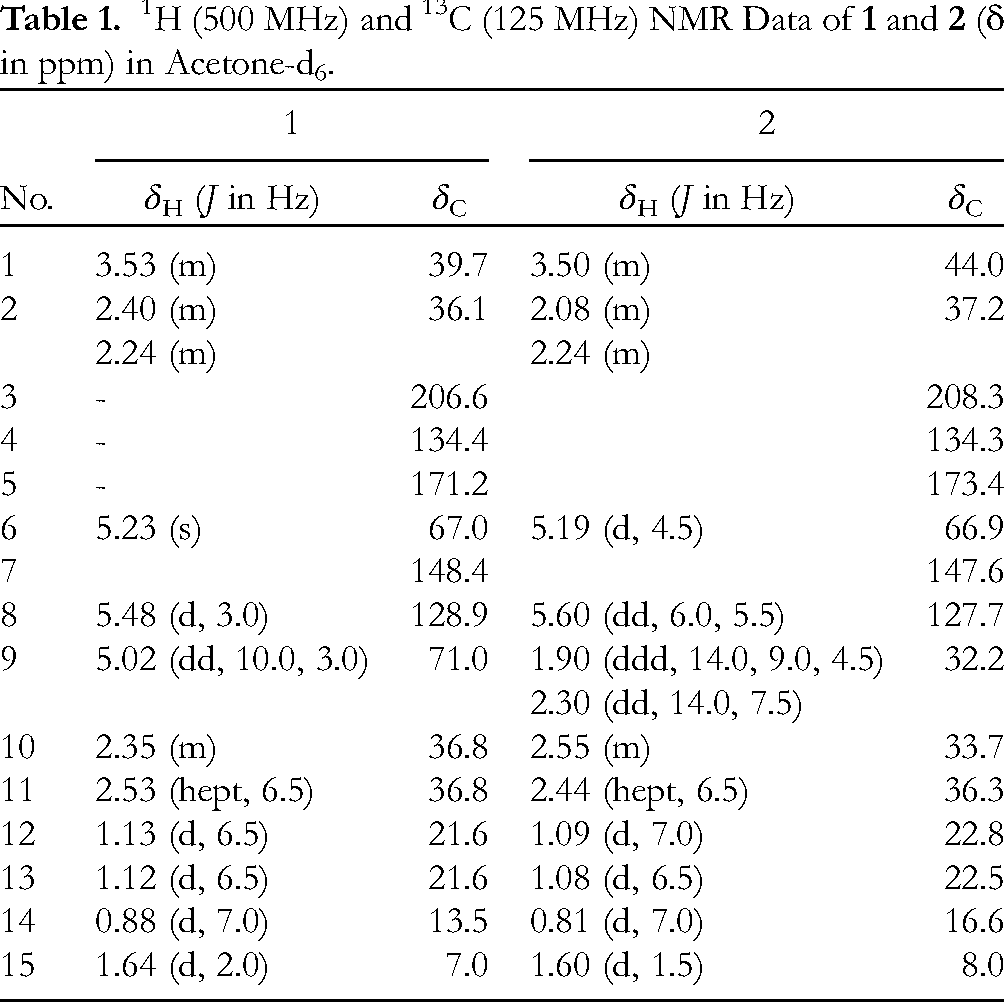
1H (500 MHz) and 13C (125 MHz) NMR Data of
Cytotoxicity Assay
The method used followed that reported by Nguyen et al (2022).
12
Briefly, HepG2 and RAW-264.7 cells in a complete medium containing DMEM (HyClone, Cytiva, USA) and 10% fetal bovine serum (FBS) were seeded into a 96-well plate at 5 × 104 cells per well. The cells were then incubated at 37 °C with 5% CO2. After culturing for 24 h, the old medium was replaced by diluted compounds (in dimethyl sulfoxide (DMSO, Sigma-Aldrich, Germany) for HepG2 cells or DMSO supplemented with lipopolysaccharide (LPS, Sigma Aldrich, Germany) for RAW-264.7 cells), positive control Doxorubicin (DOX, Fesenius Kabi, Germany), and negative control DMSO or DMSO+LPS with the corresponding concentrations as nitric oxide production assay. After 72 h of treatment, 3-(4,5-Dimethyl-2-thiazolyl)-2,5-diphenyl-2H-tetrazolium bromide (MTT) was added into each well of the test plate. The plate was incubated at 37 °C with 5% CO2 for another 3 h. DMSO was used to dissolve the crystals from the interaction between MTT and viable cells. The absorbance of samples was read at 570 nm by the ELISA Reader (BioTek, USA). Cell death (% inhibition) was estimated by the following formula:
α-Glucosidase Inhibition Assay
A previously established protocol by Duong et al (2020),
4
with minor adjustments, was employed to investigate the inhibitory effects of various compounds on
Measurement of nitric oxide production assay
Determination of nitric oxide (NO) production and cell viability assay were performed using the same method described in our previous reports.13,14 The sample was analyzed in triplicate at five different concentrations around the IC50 values, and the mean values were retained. N(G)-monomethyl-L-arginine (L-NMMA) was used as a positive control.
Molecular Docking
The
Statistical Analysis
Data are expressed as the mean (±) with the standard error of the mean (SEM). Statistical significance was set at
Results and Discussion
Structure Elucidation
Compound

Selected HMBC and NOESY correlations of
The relative configuration of

ECD spectra of
Biological Activity
The inhibitory effect of

Cytotoxicity of

Cytotoxicity of
Molecular Docking
XP docking analysis revealed that both hydroxydiorygmone A (

The binding modes of two ligands with Human PPAR-gamma protein compared to native ligand.
The XP Docking scores, the MMGBSA and number of H-bonds of Two Ligands in Human PPAR-Gamma Active Site.

XP docking analyses confirmed the binding potential of

The binding modes of two ligands with
The XP Docking scores, the MMGBSA and number of H-bonds of Two Ligands in α-Glucosidase Active Site.

*(in parentheses): The Glide GScore value, after state penalty compensation.
Based on the aforementioned
Supplemental Material
sj-docx-1-npx-10.1177_1934578X241297992 - Supplemental material for Hydroxydiorygmone A, a New Guaiane-Sesquiterpene from the Cultured Lichen Mycobiont of Diorygma sp
Supplemental material, sj-docx-1-npx-10.1177_1934578X241297992 for Hydroxydiorygmone A, a New Guaiane-Sesquiterpene from the Cultured Lichen Mycobiont of
Footnotes
Acknowledgements
This work was supported by Ton Duc Thang University.
Authors’ Contributions
Conceptualization, T.P.N, N.H.N ; methodology, T.H.D, N.H.N, H.T.L; software, G.H.D, H.T.N, T.H.T.N; formal analysis, T.H.D, N.H.N, H.T.N; investigation, T.H.D, N.H.N, T.M.D.T; resources, T.P.G.V, G.H.D, N.H.N.P, T.H.T.N; data curation, T.P.G.V, N.H.N.P, T.H.T.N; writing—original draft preparation, T.M.D.T, N.H.N, T.P.N; writing—review and editing, all authors. All authors have read and agreed to the published version of the manuscript.
Availability of Data and Materials
Datasets generated in this study are available from the corresponding author on request.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect tothe research, authorship, and/or publication of this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval
Ethical Approval is not applicable for this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Statement of Human and Animal Rights
This article does not contain any studies with human or animal subjects.
Statement of Informed Consent
There are no human subjects in this article and informed consent is not applicable.
Supporting Information
Supporting information accompanies this paper was found online.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
