Abstract
The profound interconnectedness of the sciences and technologies embodied in the Fourth Industrial Revolution is discussed in terms of the global role of natural products, and how that interplays with the development of sustainable and climate-conscious practices of cyberecoethnopharmacolomics within the Quintuple Helix for the promotion of a healthier planet and society.
Keywords
Introduction and the Fourth Industrial Revolution
Products derived from the natural environment are an essential aspect of the life of every human being, whether in terms of shelter, food, clothing, writing materials, gums, waxes, fragrances, or drugs. However, the mitigating factor of population, currently (February, 2021) at 7.85 bn, and the anthropogenic impacts from the generation of resources to sustain that population have brought humanity to a crisis point where the UN, government groups, international organizations, and many scientific groups are demanding action to minimize the changes induced through climate modulation and environmental degradation. Over time, particularly in the past three centuries, the Earth has witnessed two unprecedented events, a dramatic increase in population and the resource needs that has generated, and stunning changes in day-to-day activities as innovation after innovation has spurred the global interweaving of societal activities. Perhaps the most notable outcome, emphasized during the current pandemic crisis, is real-time connectedness with almost anyone and anything, including all knowledge, on the planet. To reach this edge to the future most societies in the world have passed through a series of industrial revolutions, and are in the throes of the next, the Fourth Industrial Revolution (known colloquially as 4IR or Industry 4.0). As fundamental as natural products are to daily global life, it is pertinent to ask, what will be the role of natural products in the era ahead? Who will be creating that role, and where? This review will attempt to respond to aspects of those questions, while examining the functioning of an evolving global society under the Quintuple Helix, within which the integrated natural products sciences and technologies operate.
An early suggestion of a forthcoming Industrial Revolution was made in 1984 by Maugh in reference to the impact of nanotechnology and its products. 1 The formal, societal-wide, multi-faceted concept of a Fourth Industrial Revolution was introduced by Schwab at the World Economic Forum in Davos, Switzerland in 2016. 2 It reflects the increased blurring of the boundaries that formerly existed between the physical, biological, and digital realms through constructive interaction. It is about automation, interconnectedness, and smart devices; the evolution of cyber physical systems, at home, at work, and at play. Some contemporary examples include automated movement technologies, drones, artificial intelligence, chemical and biological sensors, nanosystems, and the Internet of Things (IoT). 3 Twelve emerging technologies were described as the essential core of the 4IR. As applied to the future development of natural products they converge as: artificial intelligence and robotics, ubiquitous linked sensors, new computing technologies, 3D printing (additive manufacturing), advanced materials and nanomaterials, biotechnologies, and blockchain and distributed ledger technologies. Aspects of how these areas of development relate to the evolution of bioactive natural products will be discussed in this commentary. So how did we get here, and what are these new opportunities, as they apply to natural products in the future?

The First Industrial Revolution is regarded as the period from about 1760 to 1820, and reflects the advances made through the introduction of mechanical production equipment and steam powered engines. The Second Industrial Revolution, from 1871 to 1914, embraces the movement to machine-made from hand-made, the advent and distribution of electrical power and telegraphic messaging, railroads, and mass production for a burgeoning marketplace. The Third Industrial Revolution (3IR) originated around 1947, and grew rapidly after 1969. It includes the mass availability of electronics, desk- and home-based computers, data systems, globalization of goods and electronic services, rapid transportation systems, miniaturization of data storage capacity, smartphones, biotechnology and genomics, and automated production for the global market.
The natural product sciences have integrated well the steady advances in science and technology during the periods of 1IR, 2IR, and 3IR, and are currently leading initiatives particularly in the areas of biotechnology and genomics. Early efforts at the isolation of pure natural products began in 1IR and were based on traditional medicine. The structure determination of natural products and their associated chemistry and synthesis began in 2IR and continued ponderously in the period until the early 1940s. The relationships between structure and biological activity, improved separation and detection methods, enzyme-facilitated reactions, an expanded reagent pool, enhanced synthetic methods, the tools for structure determination, the development of automated cell- and enzyme-based testing systems, and an understanding of the formation of natural products at the enzymatic, and subsequently the gene, level, have empowered natural products research in the 3IR from 1969 to the present.
What does the future hold and what might we imagine will be the impactful areas of 4IR on the natural product sciences, and how will those outcomes enhance society? Let us consider aspects of the abovementioned areas. In 2017, a compiled a list of sixty challenges for the field of natural products for 2030 was presented 4 ; some are relevant to this discussion and will be described. Subsequently, the concept of “cyberecoethnopharmacolomics” was promulgated as a synergistic and holistic approach to the scientific, technological, and societal integration of natural products activities. 5,6 As we delve into some of the emerging technologies of 4IR and their application to natural products, it is also important to indicate two global factors which will be highly impactful as 4IR continues to evolve, sustainability and climate change. 7
Drug and Natural Product Discovery and the 4IR
Artificial Intelligence (AI) and Robotics
The evolution of artificial intelligence and robotics technology is important for the improvement of future research in the natural product sciences. 8,9 Over the last few decades, laboratories incorporated more complementary systems, including the tandem use of HPLC and mass-spectrometry analysis, as well as the application of more advanced NMR spectroscopy technologies. 10 In the era of the 4IR, the application of chemical robotics may offer more rapid solutions to research problems. As an early example, in 1996, Whitten et al. reported a new method for lead optimization of corticotropin-releasing Factor1 receptor antagonist. In this study, the use of robotics-driven synthesis, notably rapid microscale synthesis, resulted in the identification of several potent antagonists in a shorter time than employing traditional methods. 11 Additionally, in 2018, Carmelli et al. 12 reported the use of an affordable robotic system to explore azo-coupling reactions. In traditional natural product research for isolation and structure elucidation, a crude matrix of metabolites is extracted from the organic material, and a series of liquid chromatography steps (flash, Sephadex, HPLC) results in the isolation of relatively pure samples, which may then be subjected to final stage purification using a variety of acceptable methods. This is followed by the typical protocols for compound identification and structure elucidation. 10 What if an instrument system existed which included the sequential use of HPLC technology, mass spectrometry, IR, and bench-top NMR, which could identify “known” compounds, as well as fractionate other compounds based on the usual principles? Could we see the universal use of soft robotics technology in such a sequential manner? In addition to creating a more robust research atmosphere, the development and increased use of such soft robotics systems will afford more centralized laboratory data storage, allowing for more rapid and efficient data processing and human review.
Automated Plant Identification
The concept of positively identifying a plant in the field through a photograph (now a digital image) is not new 13 ; making it real for a large and complex genus like Ficus is more challenging. 14 The need for such capacity, particularly in a hand-held device, is manifold. Whether it is a commercial collector of medicinal plants, a farmer assuring crop identification, a botanist seeking new species or examining areas for diversity from a conservation perspective, or a range farmer identifying plants that bear animal toxins, rapid and accurate identification on site is a long-sought goal.
There are about 393,900 vascular plant species known 15 making it impossible for any one individual taxonomist to assign a positive identification of more than a few thousand species. While ca. 2000 new species are identified each year, taxonomic specialists are declining in number, and ecological monitoring and biodiversity monitoring is becoming a continuous process, the need for on-site, high accuracy identification becomes even more essential to assign conservation value.
Early efforts in this regard 16 -20 achieved modest success. With the advent of convolutional neural networks (CNNs) visual object classification accuracy, even for plants, has been enhanced. 21,22 This success and future prospects were reviewed. 23 Using different architectures and pretrained datasets, a 99.5% accuracy was obtained on a sample of 44 species, 24,25 while a different system provided 99.65% accuracy on a 32-species dataset. 26 Numerous factors impact success, including the nature of the images used for training and testing (eg, scan vs structured photo vs in-field photo), the growth status of the plant and the comprehensive nature of the training set of images for a species based on the number of plant parts included. 27,28 Apps are also available, for example, Leafscan and Pl@ntnet. The former can give a 96.8% recognition rate for 18 tree species, 29 while the latter could provide 69% accuracy from 2,200 species based on a single photo. 30 As these datasets expand, including through crowdsourcing, accuracy should increase.
Significant advances in image recognition are anticipated in the future as machine learning architectures evolve. 23 SqueezeNet 31 for example may be applicable in the future for hand-held devices in the field. Enhancing the size of the datasets with fully authenticated images is a major challenge, especially if there is a need to assess a plant at different stages of the growth cycle. Thus, transparent networks of diverse data sets are the likely pathway to address the holistic challenge. Other attributes of the image, date, topographic location, etc. should also be used in plant identification based on prior distribution data. This information, as well as images of the estimated 350 million plant samples in the 3,000 herbaria in the world, 32 become extremely important input for a comprehensive assessment of global biodiversity, though they will be selectively suitable for AI training purposes due to shape and color changes on drying. The key to the future success of high accuracy, in-field identification, is a strong collaboration between taxonomists and computer and imaging specialists. At some point one can envisage that a transition to the video-based, 3D-identification of plants in the field will also occur. Examples of the application of artificial intelligence in natural products research are illustrated in Figure 1.

Applications of artificial intelligence in natural product research.
Chemical Synthesis
Natural product chemistry is frequently concerned with the identification of known, and the structure elucidation of new, metabolites. It also embraces the semi-synthesis and total synthesis of compounds, and the targeted (in silico driven) and untargeted modification of bioactive compounds to potentiate their effectiveness in particular assays. 33 However, despite advances made in organic synthesis, there remain failed synthetic transformation attempts and those which yield new products from unexpected rearrangements. This issue was recognized from a computational perspective many years ago, 34 and remains a major consideration in the success of retrosynthetic design.
There is a long-standing desire to put organic synthesis into a more highly predictive state, where failure is minimized, and resources optimized, thereby following green chemistry initiatives regarding solvent and reagent use, as well as atom efficiency considerations. Predictability in synthetic outcome is also a core aspect of automated synthesis and for the processing of designed structural motifs. 35 For complex transformational sequences other approaches were developed and will undoubtedly be extended through AI and automation.
In the late 1960s, Corey and Wipke 36 advanced computer-driven retrosynthetic analysis for total and correlative partial synthesis. Strategies for selective, sequential bond formations were proposed based on computer-derived analyses, such as the evolving algorithm Logic and Heuristics Applied to Synthetic Analysis (LHASA). 37 -39 The program was one of the earliest tools to have a graphical input and output for structures, and (in part) led to the award of the Nobel Prize in Chemistry for Corey in 1990. 40
Numerous programs for retrosynthetic analysis followed, including, SYNCHEM 41 and SYNLMA, 42 and later commercial 43,44 and open source programs 45,46 became available. These systems rely on a paradigm of systematic bond breaking of synthesizable bonds based on known chemical reactions until simple, available starting materials are identified. Alternative machine learning methods often fail to identify a pathway when viewed by practicing chemists. 47 A data-driven retrosynthetic analysis method (DDRAM) was recently described and uses millions of known synthetic procedures, including those from laboratory notebooks, 45,47,48 and interfaces that with accepted synthetic reaction ontology. It brings together information on reactants, precedents, and scalability which can be pursued through automated synthesis. 47 Output indicated that in-house corporate synthetic procedures for existing drugs might be improved in 40% of the single step cases, and that limiting the millions of reactions to a few thousand alternatives allowed up to ten plausible routes to be developed for a target molecule in seconds. Coupled with holistic initiatives to increase drug discovery throughput and the targeting of derivatives of hit compounds, 49 together with automated chemical synthesis, productivity in identifying lead molecules was anticipated to be enhanced. 47 Improvements in predictive retrosynthetic pathway outcomes have also been claimed through the use of deep neural networks, 50 and for a framework based on graph convolutional networks. 51 A template-free approach gave a 99.6% molecular validity for single step transformations, and was extended to examine published strategies for successful multistep syntheses. 52 A robotic system which reads a synthetic method from the literature and automatically translates that into a reaction has been described. 53 In addition, a combined expert (with reaction rules) and machine learning (literature data) approach showed that the processes can be synergistic in improving overall synthetic accuracy. 54 Collaboration between machine learning scientists and bench chemists will be important for further advances in predictable, high-yield pathway outcomes.
A major omission in many of the accumulated datasets is the recording of negative reaction results; where a reaction which did not give the anticipated product outcome, was unexpectedly slow, or did not work at all, is not reported. 35 Standardization across more public datasets, and the interoperability for datasets in the public domain, will be important for chemists in emerging countries to improve their efficiency. Sustainability and green chemistry considerations with respect to reagents and waste products will need to become more important considerations in the design of synthetic pathways, including those using nanoparticular approaches. 55 In addition, AI should also focus on reaction discovery, the development of new reaction possibilities, and must consider spatial approaches based on integration with modeling data from in silico enzyme interactions. Procedures in the future will likely be based on a paradigm of chemist-AI-robot-AI-chemist-robot to generate a final compound of higher value. These strategies are independent of compound origin, and a greater emphasis on underutilized, available, diverse natural product scaffolds for in silico modeling may lead to new precursor molecules which can be sustainably sourced. 56
Biotechnologies
The use of enzymes from a variety of natural sources to conduct chemical reactions, regiospecifically and enantioselectively, is well-established. 57 These systems were typically discovered empirically, and a particularly useful attribute is the serendipitous ability to activate carbons which would otherwise be deemed inactive to canonical organic reagents. One early classic example was the C-11 α-hydroxylation of progesterone by Rhizopus arrhius or Aspergillus niger, now a commercial process. 58 Since then, it has been demonstrated that numerous reactions, such as the stereoselective aldol reaction, 59 can be performed, and, from the perspective of flexibility, that enzyme reactivity can occur in nonaqueous systems. 60 The ability to make “reagent” enzymes available on a larger scale through the polymerase chain reaction (PCR) transformed the implications of their use, 61,62 and made them commercially available. 63 However, to fulfil the obvious potential in terms of green chemistry and the use of sustainable, renewable reagents, one question was whether more targeted approaches are possible?
The structural diversity of almost 200,000 natural metabolites provides a wide range of biosynthetic transformations, and the enzymes which are responsible for some of those pathway steps are being isolated and characterized. 64,65 In particular, those enzymes which have been cloned and expressed from bacterial metabolite pathways through extensive efforts of genome mining, cloning heterologous expression, and functional analysis, are of significant interest for selective synthetic reaction studies. One need at present is for an available database of sources and chemo-activities, and the commercialization of more functionally diverse, stable, and substrate promiscuous, enzymes.
Indeed, historically, one of the profound challenges in this arena is substrate specificity, which frequently limits the chemical diversity of an accepted reactant. Other issues which in the past limited enthusiasm for biocatalysis include enzyme stability, feedback inhibition, and lack of regioselectivity. 57 Directed evolution, a cyclic process of gene mutagenesis, expression, functionality screening, has over the past 25 years, partially addressed these issues. Much more needs to be uncovered about the precise catalytic sites for function. Collation of the reactivities can then be used to develop a computational and systematic approach for expanding the site specificity for the spatial acceptance of a broader range of substrates. 66 This will also allow for the more robust inclusion in AI systems of highly predictable, biocatalytic steps in retrosynthetic analyses, 67 thereby improving the greening of natural product synthesis and transformation. The notion of multistep synthetic pathways of non-natural products based on biocatalysis remains a worthwhile goal, 68 in the form of either cell-based or solid support systems.
The synthetically desirable biosynthetic reactions include reductions, oxidations, esterifications, hydrolyses, and cyclization reactions, among others. 57 Enzymes for some of these processes have been well-studied including Diels-Alderases 69 and Pictet-Spenglerases. 70 Important challenges for the future are the demonstration that metabolic pathways can be completed from acknowledged precursors ex situ and the enzymes utilized to enhance structural diversity for bioassessment. An early example of the former possibility derives from the work of Scott using a pool of 12 enzymes of vitamin B12 biosynthesis, 71 and examples of total synthesis have been described. 72,73
Is it necessary to process the plant or microbial source for the isolated enzyme to effect some of these transformations? Possibly not. Studies on the use of whole plant systems, including vegetables such as manihot, carrots, coconut juice, and sugar cane, where no enzyme isolation occurs, as chemical reagents for chiral carbonyl reduction reactions are well established. 74 Unlike a synthetic heavy metal or chiral hydride catalyst used for the same purpose of carbonyl reduction, these reagents can be used repeatedly without loss of activity, and significant further exploration of cheap and abundant resources is warranted.
More explorations are anticipated which can diversify the synthetic capacity for a variety of transformations as further plants, microbes, and substrate explorations are conducted. There is an industrial need to seek greener reactions for a variety of stereoselective reductions and regioselective oxidations. 57,75,76 From a green chemistry, economic, and sustainability perspective, and considering the biotechnology concepts for the 4IR, the importance of these pioneering synthetic transformations should not be underestimated, especially for those reactions in which heavy metals are used as chiral reagents or catalysts.
Future of Biosynthesis
The global needs for new drugs and agrochemicals are well established. Old diseases, new diseases, well-honed ones, and neglected ones are all demanding biological agents. This is particularly the case for a range of diseases (cancer, microbial and parasitic infections) and plant pathogenic organisms where resistance has developed. 77 The innovative strategies for a synthetic chemical approach to this global crisis have been discussed earlier. What is the future role of natural products in meeting these needs? The screening approach for plant and antimicrobial extracts presented earlier remains an option, with caveats of clean-up, stability, and the dereplication of known actives and interfering compounds. 78 However, with the dramatic advances in microbial genomics, coupled with bioinformatics, and systems biology, there are numerous avenues evolving for natural products to meet these societal needs with highly creative approaches in rethinking the way that bioactive metabolites are identified. 79 -83
“Nature does not produce what it cannot produce”. Since the early characterizations of natural products there has been a growing curiosity as to how the almost 200,000 natural products from marine and terrestrial sources are produced. What are the building blocks, the mechanisms, the sequences, the rearrangements? Over the years, the methods used for studying biosynthetic pathways have evolved dramatically from radio- and then stable isotope analyses to the isolation of enzymes, the identification, cloning, and expression of gene clusters, to various forms of point mutations looking at substrate specificity in non-ribosomal peptide synthesis (NRPS) and polyketide synthesis (PKS) pathways. 84 Assembly sequences for NRPS and NRPS/PKS-derived products are now routinely deduced, and that the sequences can be specifically modified at the functional level reveals the available options for reconstruction. In the process, the sequences have become bespoke assembly lines exclusively for the creation of new metabolites. 83
One of the new learnings that occurred when whole genomes of bacteria and fungi were examined through next generation sequencing 85,86 and bioinformatics 87 was that there are typically many biosynthetic gene clusters (BGCs) present in the genome whose encoded enzymatic products are likely unknown and possibly novel. 88 -90 For example, the genome of the organism Streptomyces coelicolor A3(2) revealed 16 unknown pathways, 91 which were suggested through bioinformatics to include new scaffolds. 89,91 As noted, more accurate estimates of the number of biosynthetic gene clusters are derived from the finished genome sequences rather than the draft one. 92 Identifying the gene cluster does not predicate a natural product structure. Although several algorithms are available to assist, 93 heterologous expression, isolation, and structure elucidation are still required at the present time. As more gene clusters are characterized and the substrate specificities recognized, higher levels of predictive accuracy should be expected, particularly if non-functional modules within a gene product in an assembly sequence can be identified.
Culturing the organism to promote the expression of new pathways is one of the great challenges in biosynthesis for seeking new metabolites and potentially new biological agents. 94,95 Techniques to stimulate the production of metabolites from the cryptic pathways have been summarized, 80 and include changing media, enhancing pathway expression, identifying and mutating regulatory genes, modulating pathway-specific transcription factors, and heterologous expression in a suitable host organism.
Another significant challenge to enhancing structure diversity relates to substrate specificity, and the exploration of what flexibility is inherent, and what can be introduced into assembly line processes. Typically, these are empirical approaches, such as knockout studies, down regulation, or adaptive laboratory evolution. 83 Targeted metabolite biosynthesis requires a higher level of knowledge concerning specific regulators and functional sites within an identified assembly module. The module “parts” derived from assembly line genome sequencing converge in systems biology, where the “parts” can be introduced through combinatorial biosynthetic approaches to specifically modulate a structural outcome 96 ; the extensive work on daptomycin and the related A54145 are examples. 82 The goal is to turn a billion-year evolutionary process to one achieved in a year or less.
AI will be of critical importance as the amino acid sequences and functioning of the modules in assembly systems are established, modified, and the targeted products analyzed in order to develop rational genetic modification strategies, and directed genomic evolution for new metabolites. 82,83 There is an important machine learning process 97 which relates to the conserved protein architecture responsible for NRPS, PKS, and terpenoid synthase systems which can recognize and trace, with high accuracy, the substrate specificity, or flexibility as appropriate, for a new biosynthetic product. 98 The approaches for targeted directed genome evolution have been reviewed. 83 The subsequent developmental steps will be, as in synthetic chemistry, to move, with expressed modules available, to automated systems capable of producing specifically designed target molecules through programmed biosynthesis 99 in a design-build-test-learn paradigm. 100
The sustainability of known, commercially significant compounds are also a target for metabolic engineering and synthetic biology. One of these is shikimic acid, an isolate of star anise seeds [Illicium verum Hook. f. (Schisandraceae)], and an important synthetic precursor for oseltamivir (Tamiflu®), as well as other commercial entities. However, isolation from the plant source is inefficient and costly. Understanding the balance of the precursor levels for the shikimate formation, and the subsequent fate of shikimate at the genetic level from a pathway and regulatory perspective, allowed the development of high-producing strains of E. coli using different metabolic engineering approaches. 101
Synthetic Biology
Closely interwoven with the application of metabolic engineering is synthetic biology, 102 -108 and its’ strong relationship with the United Nations Sustainability Development Goals (SDGs) 109 have been discussed. 104,106 Human needs for the products of nature, such as food and medicine, are growing rapidly for a population reaching 9.7 billion people by 2050, especially with 50% more people in polluted urban settings. 110 Innovation in many areas of agriculture and medicine will be necessary as a part of 4IR to address those needs globally. At the present, synthetic biology is teaching us what is not known about how nature operates, a welcome opportunity for future inspiration and creativity. As the discussion unfolds, challenges will arise, from the small issues to be resolved, and more significantly whether grander solutions that are widely applicable can be developed. Other complex challenges for synthetic biology include public perception, the reality of risk, and monitoring the possible duality of outcome. 107
There are at least five areas for contributions that synthetic biology could (must?) make to the SDGs, 109 namely: (i) natural chemicals to replace harmful synthetics (e.g., pesticides, herbicides); (ii) using microbiological sources to effect environmental clean-up; (iii) replacing synthetic, non-renewable materials with those derived from nature, such as biofuels and bioplastics; (iv) enhancing marine and terrestrial agricultural yields through genetic strategies; and (v) discovering new biosynthetic products for drug resistance and neglected diseases, and enhancing the production of critical metabolites in commerce.
As a result of cheap and accessible DNA sequencing technology, access to genetic information is expanding rapidly, allowing correlations, interpretations, and actions that were previously impossible. Imagine a natural product chemist sitting at home watching a late-night movie, an idea for a modification to a scaffold comes to mind. Through the phone, the chemist asks the drug development and synthesis laboratory for a structure fit analysis into a particular enzyme niche. It’s a positive. A synthetic protocol from an available core molecule is sought. The projection is for a chemo-enzymatic approach in 3 steps with a projected overall yield of 85%. Isolation, purification, structure proof, and automated in vitro testing will be underway by mid-morning and completed by lunch-time. That’s the (bio)synthetic chemistry approach alluded to earlier. Can that be achieved with natural products? Can synthetic biology meet the challenge of microbial “natural” products on demand? A report from the Broad Institute suggests that it can, for a microbial system. 111 However, the development of a synthetic microbe to assemble a defined structure requires, as noted, the genes to encode for the appropriately substrate promiscuous enzyme, and that the available reactions are limited by those that nature performs. Higher predictability, based on more enzymes from more sources, available on demand, and coupled with artificial intelligence to construct a functional plasmid will also be necessary. 112
At the present, the gap for that process being accomplished for a plant metabolite is vast. Because in utilizing the biosynthetic pathways occurring in plants, the challenges are quite different. They focus on where pathway steps occur in the plant and in which parts of the cell, how intermediates are transported, and whether complex, isolated enzyme systems can be assembled and functionalized to produce needed metabolites, mimicking the pathway models from microorganisms. A far deeper understanding of how a plant operates as a metabolite factory is needed. Only then will it be possible to tweek the genetic apparatus and introduce specific genes in a precise location to effect a desired transformation in a specific pathway.
Plants typically produce a broad range of metabolites based on very few precursor molecules. AI should be able to indicate which plants are likely to be the most appropriate for the introduction of either a single gene or a set of pathway genes to achieve a target compound. Can the gene for an epoxidase from a microorganism be introduced into Catharanthus roseus (L.) G.Don (Apocynaceae) to generate the elusive 14,15-epoxide of vindoline? Can a microbial genome pathway be introduced into a fast-growing crop to promote the local development of a bio-available drug for harvest? This would enhance the accessibility of a drug, such as an antibiotic, in an emerging economy.
Progress in metabolomics has indicated that many disease states produce detectable biomarkers. Predictive monitoring through the application of disease-specific chemo- and/or biosensors, such as for hippuric acid and prostate cancer, 113 will likely expand rapidly. In addition, the application of cybergenetics for controlling cellular processes at the gene level through the interfacing of cellular systems and a computer which turns an embedded “genetic switch” on or off 114,115 may enhance drug delivery technology.
There are untold millions of microbial systems, yet only ten are used industrially at the present. 107 As the need to produce small designer molecules expands, so the range of available microorganisms which can be adapted for synthetic biological purposes will also grow. That implies the development of universal expression systems that can operate in diverse hosts, hence avoiding organism-specific constructs. Technologies involving DNA-editing, such as CRISPR/Cas9, should speed this process. Transferring to cell-free systems also has advantages 116 in which enzyme pathways can be tethered to a nanoparticle ameliorating transport and diffusion issues, 117 while controlling accessibility and concentration effects through microfluidics and compartmentalization as automated biosynthesis proceeds. 118
As indicated, 119 there is a fundamental need for large datasets of the parameters of cellular behavior which can be analyzed through AI or machine learning to eliminate the empiricism, 120 and to provide a set of design standards for the processes and the products of synthetic biology which would afford a higher degree of interoperability for the field. 107 International agreements and regulation development will be needed in the future to deal with risk analysis for organism creation, specifically to avoid dual usage of the technology, and engender a higher level of trust in society for the products of synthetic biology. Critical thinking training regarding these and other ethical concerns for the scientists working in these areas has been recommended. 107
From the perspective of 4IR and the SDGs, automated analysis of genetic products from a risk/benefit perspective should be readily achievable and lead to the use of machine learning to derive products that should inherently be safer toxicologically and environmentally; although extensive safety concerns will still need to be ameliorated through bioassay. There is a need to promote the development of research programs in the emerging world to enhance their bio-economic industrial initiatives to meet local food and health care needs. 109 However, this must not occur through the exploitation of indigenous resources, or through ignoring local environments which have had no exposure to such biological materials previously. The migration of new genetic material must be monitored, as well as its effects on specialized crop rhizospheres. 121 This is a particularly sensitive area of science where the practical issues, the ethical and environmental concerns, and regulatory control coalesce. 109
Network Pharmacology
The role of a purified natural product in drug discovery and development can be traced from the isolation of morphine from a traditional medicine, opium. It continued, also for the reduction of pain, with the marketing of the semisynthetic drug aspirin, derived from salicin, in 1899. In retrospect, those events drove two ideas, the first that a single compound could target a single health condition, while subsequently revealing that in fact multiple targets, not just one, were being “hit”. The history of drug discovery over the past 80 years reflects the first concept of single target-oriented, sometimes called “magic bullet”, drug discovery. 122,123 This approach, although successful in bringing hundreds of effective drugs to patients, also resulted in many late-stage clinical failures due to lack of efficacy and unexpected toxicity, 124,125 which contributed significantly to very high research development costs.
The evolution of the idea and demonstration that a single compound does not target a single target gene, that is, that compounds have a plethora of actions at nodes throughout the human genome, and that they can act synergistically or antagonistically, brought a very different focus to the discovery phase. Network pharmacology 126 recognizes and considers for strategic purposes that these multiple sites of action can be identified and mapped, and that the collated information can be used in several different formats. 123,127 Natural products discovery, or mechanism of action studies on a traditional medicine, becomes very complicated at this point. The complex metabolite matrix represents perhaps 100, 1000, or even more compounds. Each one potentially has its own myriad of nodes of action. In silico approaches in systems biology are therefore of prime importance in beginning to resolve and comprehend this issue. 128 -130 Why those approaches often fail is because only about 1.8% of individual natural products have target binding information in the relevant databases. 131 What does that mean in terms of target space?
For the future in 4IR, the existence of significant, and diverse target space represents a knowledge creation opportunity. It provides the challenge of AI-directed, new targets (and possibly new uses) for natural products with well-established safety and toxicity profiles, so-called drug repositioning. 132 It has the potential to identify new synergistic relationships between two or more compounds. It offers the possibility of new activities for known compounds, and also the identification at an early development stage of the possible implications for untoward effects, including toxicity. Finally, it may provide an understanding of the mechanism of action of compounds and extracts. 127,132,133 A significant issue for natural products at the present time is that the algorithms for the pharmacodynamic and pharmacokinetic aspects of adsorption, distribution, metabolism, and excretion have not transitioned. 134
A list of network mapping resources is available, 127 and one of the well-used, open access sites is Connectivity Map (CMap) at the Broad Institute of Harvard and MIT. 135 Some successful uses for identifying new activities for known compounds, which were subsequently supported through in vitro assays, have been reviewed. 127 As an example, the adjunct supportive effects of a TCM were substantiated through global transcriptional profiling. 136 Astragaloside-IV, a dominant component of a TCM cardiovascular treatment, affected 33 different pathways associated with cardiovascular disease. This led to the conclusion that it is effective through several mechanisms, including anti-inflammation, anti-oxidation, immune regulation, calcium blocking, and vasodilation. 137 The activities of the constituents of foods cannot be ignored. The effects of the widely distributed natural products apigenin, chrysin, and luteolin were similar in CMap, and it was proposed that they may act additively in terms of chemoprevention or synergistically with drugs. 138 The development of more comprehensive functional response profiles in the future will expand the concept of a single time slot transcriptomic response. The Library of Integrated Network-based Cellular Signatures (LINCS) system is one approach 139 which includes binding profiles and phenotypic response profiles.
Also requiring more consideration in these algorithms are the pharmacologic effects attributed to interactions with cell wall proteins and other functional structures. In the future, one can envisage a much more targeted approach to both the discovery and application (distributed medicine) phases where integration of clinical data is reflected in gene modulation considerations, and the appropriate, multicomponent, personalized medicine provided to the patient. It will be coming almost full circle for a single or complex traditional medicine, and would have the support of in silico and experimental evidence for the desired mechanism(s) of action. Deconvoluting traditional medicines for their holistic mechanisms of action therefore remains a high priority to identify the various sites of action of the key metabolites. In addition, there is an urgent need to expand the genomic knowledge for a much broader range of natural products that would be available on a sustainable basis.
Human Microbiome
Advances in genomics and bioinformatics, including genome mining, machine learning, and high throughput screening were mentioned as being important in the identification of biosynthetic gene clusters (BGCs) in microbial systems. All mammals, including humans, also host a vast number of microbes, producing a range of largely unknown metabolites. One algorithm has proposed the existence of >14,000 BGCs in the human bacterial genome, 140 for most of which the products are not characterized 141 and their interactions unknown. Associations between disease states and specific BGCs are being made through bioinformatics using metagenomic data from known BGCs, comparing abundance levels as a criterion through machine learning. This approach was used to identify 43 BGCs in the guts of patients with Parkinson’s disease, 142 and over one thousand oral BGCs associated with caries and periodontal disease. 143 Determining which systems are active is another challenge. 144 Possible outcomes include the identification of new targets for drug discovery, and potentially new metabolites if the organisms can be cultured.
Nanomaterials
Over the last decade, there has been tremendous growth and use of nanoscience and technology in a broad array of applications. 145 For example, there has been substantial progress made in the synthesis, assembly, fabrication, and application of these nanomaterials in pharmaceutical and industrial processes and products. 146,147 With respect to the role of these advanced nanomaterials in academic research, how can these advancements in nanoscience be universally applied or exploited by natural product researchers? Can natural products be detected using sensory nanomaterials, and those systems used routinely in the field? What are the research and development opportunities for nanomaterials derived from single or complex natural products? What are the implications for quality control and standardization?
With respect to sustainability, there are many challenges associated with potent natural product drug candidates, including solubility, stability, bioavailability, metabolism, and targetability. Nanotechnology, in particular the use of nano-formulations as a tool to enhance the bioavailability and therapeutic value of natural products, has been discussed elsewhere. 148,149 In addition to improving therapeutic values, the bioactivity of natural products may be enhanced using nanomaterials. With respect to antibiofilm activity, it is established that the use of nanomaterials in the formulation are effective for the eradication of bacterial and fungal biofilms. 150,151 With the formation of biofilms as a primary factor in the reduced antimicrobial susceptibility among microbes, as seen with Pseudomonas aeruginosa, 152 the application of nanoparticles as a delivery mechanism, either via coating, nano-emulsions, or encapsulation, should be evaluated where possible, for comparative purposes, as a new standard, similar to that of SAR analysis during the typical antibiotic assessment.
Another way to potentiate the development of nanotechnology is the expansive use of nanomaterials based on natural products, 149 with the caveat that natural product research be highly targeted towards more environmentally acceptable nanomaterials. Over the years, as the deleterious effects of the 3IR were observed, attempts were made to rectify the negative contribution of chemical research done by promoting more environmentally friendly techniques. As the 4IR is taking shape, this is an opportunity to start with preventative mechanisms which afford the same output, to account for sustainability, and to assess any contribution of the technology and processes to climate change. It is imperative, as a natural product collective, to offer continuous solutions to ensure the development of environmentally responsible mechanisms and outcomes for industrial applications. There can be, with conscious creativity, the widespread application of nanotechnology as an integral aspect of natural product research in the areas of discovery in the field, applications in the laboratory, and product delivery to the patient. Some of these approaches for natural products are presented in Figure 2.
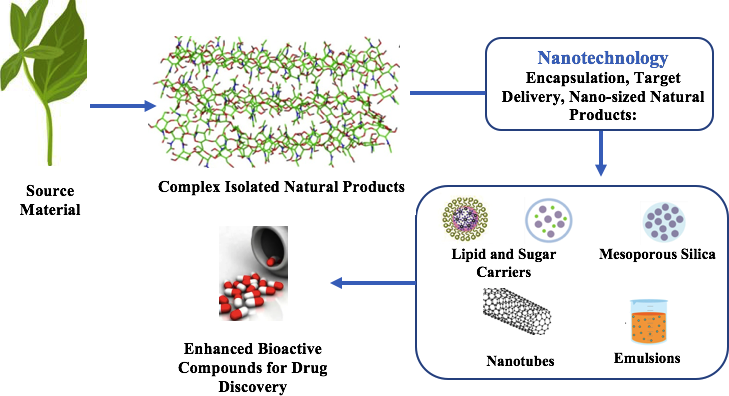
Nanotechnology in natural product drug discovery.
Techniques and Technology in Natural Product Discovery and the 4IR
NMR Assignments and Structure Elucidation
NMR spectroscopy, moving now to the 1.0 and 1.2 GHz range of magnets, remains a fundamental asset in many phases of natural products research, including structure elucidation of small molecules and proteins, and assessing the variable complexity of natural product matrices. In addition to significant advances in reducing the sample size for NMR analysis, through cryogenic cooling of radiofrequency coils and capillary tubes, to the nanomole scale, 153 advances have been made in 2D-NMR analysis for metabolomics, 154 and in quantitative NMR 155 ; applications in these areas are likely to expand significantly. Computational methods for the prediction of proton and carbon-13 NMR shifts provide an important adjunct to the structure elucidation of natural products and synthetic compounds singly or in a matrix. 156 -158 In the future, more algorithms will be developed to automate spectral interpretation in terms of correlative relationships and, as has been achieved with protein structure analysis, to provide a higher level of conformational information for small molecule natural products.
Mass Spectrometry Imaging
The notion that mass spectrometry could be used as a spatial analytical tool was first demonstrated in the 1960s. 159 In the subsequent years, 2D- and 3D-techniques have evolved such that mass spectrometry imaging (MSI) is a very well-established technique. Compounds of interest can be located in diverse spatial environments, including living tissues, plants, single cells, and microbial colonies. 160 -166 Through interfacing with datasets of mass spectra, MSI has evolved as a canonical approach to natural products analysis in real time. A significant advance in that capacity was the development of ambient ion mass spectrometry (AMS). 163,165 -167 This technique examines compounds at the surface of a matrix using soft desorption methods and requires minimal or no sample preparation. AMS methods multiplied rapidly after the pioneering work by Cooks et al. 168 on the desorption electrospray ionization (DESI) technique, and by Cody et al. 169 on direct analysis in real time (DART). A recent review summarized the strengths and weaknesses in spatial application of the currently available vacuum (MALDI, SALDI, SIMS) and ambient (DESI/nanoDESI, LAESI, LESA, LA-DART, IR-MALDESI) techniques. 166 Applications in forensics, cancer and other medical diagnostics, and for drug discovery and development have been described. 163 -165,170 -172 The focus in this brief survey will be on the applications of AMS for natural product metabolomics in plants and microbial systems. A recent review has commented on many of these applications, 165 and only a few illustrative examples will be presented.
The establishment of MSn databases is a critical adjunct to any series of analyses. Studies with cancer biomarkers indicated that interoperability for identifying and grading a particular tumor type in different instrument locations remains a validation issue, 173 although variable tissue density 174 and metabolite heterogeneity 175 indicate the need for multiple biomarkers to be established. One solution may be national or regional repositories where global clinical data can be accumulated, processed, and the diagnostic probability increased in a manner independent of the instrumentation used.
As a surface technology, a crucial aspect of AMS imaging is the quality of the surface itself, bacterial cultures are easier to study than fungi, which are easier than plant leaves or fruit interiors; hard, curved and uneven surfaces, such as nuts, stems, bark, and roots are especially challenging. MSI on bacteria in agar plates could successfully identify 400 metabolites. 176 Fungal strains could be analyzed through metabolite transfer to a secondary surface, such as filter paper, silicon wafers, or porous polytetrafluoroethylene (PTFE) film. 177 A real-time study of 34 living microorganisms (yeast, pathogens, fungi, and marine bacteria) using a liquid microjunction surface sampling probe (LMJ-SSP) with computer-controlled sample orientation, revealed numerous metabolites of biological significance, including virulence factors, surfactins, antibiotics, and signaling molecules, 167 illustrating the potential for novel metabolite discovery with an automated system of culture sampling over time, once dereplication techniques are applied..
For plants, alkaloid distribution in areca nuts [Areca catechu L. (Arecaceae)] was assessed, 178 and the spatial locations of rohitukine and its derivatives in the early growth of Dysoxylum binectariferum (Roxb.) Hook. f. ex Bedd. (Meliaceae) was monitored. 179 MSI studies on potatoes could be used to indicate metabolic changes in the steroidal alkaloids on fungal impact 180 using surface blotting techniques, including specially designed nanofiber mats. 181 A range of monoterpene indole alkaloids was characterized from the leaves of C. roseus indicating a possible role in the quality control of critical medicinal plants. 182 Typically, no additional sample preparation is necessary, although occasionally a rapid solvent wash is included, depending on the polarity of the metabolites to be investigated. 165
Mass spectral imaging has also found intense application in neurological research to explore defects and diseases. 163,183 MALDI-MS has been used for protein analysis down to ~50 µm, while small biomolecules can be detected down to ~100 nm under DESI conditions. Using this technique, murine brain tumors and other disease-specific metabolite modifications could be detected. 183 Multiple classes of lipids (fatty acids, ceramides, sterols, etc.) were identified using a matrix of silver nanoparticles. 184 Metabolomics studies of neurotransmitters, 185,186 various endogenous and exogenous compounds, 183 including 23 metabolites in the purine biosynthesis pathway, have been visualized in mouse brain, 187 illustrating the potential for drug distribution and metabolism studies once matrix interference issues are resolved, The continuing applications of MSI-based metabolomics will promote it as a very important drug discovery and development tool.
These techniques are routine examples of cyber-physical research with numerous practical applications to food and medicine quality control, understanding metabolite accumulation and dissipation in organisms, and drug distribution in living tissue. Historically, quantitation of AMS data has proven to be challenging due to reproducibility, especially the uniform introduction of standards on to tissue surfaces and matrix ion suppression effects, 188,189 although improved techniques have been reported recently. 190,191 For the future, advances in database metabolite identification, the ability to quantify the data, and particularly to view drug distribution and metabolic profile changes over time in a continuous manner, would provide important information regarding catabolic and biosynthetic pathways, as well as the mechanisms and timing of metabolite translocation in plant and animal tissues. Enhancement in spatial resolution requires a corresponding increase in sensitivity. The ability to combine MSI techniques was demonstrated on a multi-organism complex, 192 which may ameliorate deficiencies in the individual techniques, as more diverse biological samples are investigated for their metabolite content.
3D Printing
Additive manufacturing, colloquially referred to as 3D-printing, uses a widening array of materials and methods to make available devices and constructs for deployment in a host of environments for diverse purposes. 193 The ability to conduct such processes locally has the potential to transform the practical and financial aspects of academic and industrial research development, as well as medical device availability, particularly in emerging economies, where laboratory-ware has to be imported. 194,195 One example for chemical laboratories uses low-cost 3D-printing technology involving a robotically controlled syringe which can deposit various printable bioinks at room temperature. 196 Since the gels set through self-healing, reusable reaction-ware can be made on site through this robo-casting technique. 197,198 Computer-aided design adds to the flexibility of the bespoke, highly purpose-driven, finished product. 196,199,200 Functionality, such as catalysts, reagents, and chromatographic processing can be designed into the construct, 201 permitting multistep reactions to be completed, 202 as well as the synthesis of unstable or expensive reagents. 203 Aspects of 3D-printing for synthetic chemistry and chemical engineering have been reviewed. 204 An important consideration from an environmental perspective with 3D-printed materials has been their biodegradability. 205 Poly-lactic acid (PLA) fulfils many requirements for the polymer construction of devices, including mechanical strength, processibility, 206 and complete microbial biodegradability to non-toxic products. 207 -209
Bioprinting is the synchronous positioning of living cells and biomaterials in a defined layer-by-layer stacking process to form a 3D scaffold. 210 A range of applications for 3D-printing occurs within the framework of natural product chemistry and biology. Beyond the aforementioned reaction-ware design and production, there is the generation of targeted chemical and biosensors, bioprinting for specific drug discovery applications, and advanced systems of tissue and selective organ function production for in vitro bioactivity and toxicity testing. One application is to design custom systems for the extraction and processing of natural materials leading to the selective, highly reproducible formation of samples for further processing in an intact system. The development of microfluidic valves 211 allows for both chromatographic separation and selective detection in a designed module, incorporating an enzyme-based biological assessment, 212 which would assist drug discovery and dereplication studies of extracts. 213 Such green systems markedly reduce chromatographic and energy requirements and non-reusable solvent volumes, and for a discovery or biomarker detection initiative, can eliminate the re-isolation of IMPS and PAINS. 214,215
Another important application in drug discovery relates to the earliest possible detection of toxicity in a compound or an extract in the discovery pipeline. Thus, as candidates progress, the simulation of a future application becomes important for unwanted effects. 216 Time to transition from bench to product is a fundamental deterrent to the efficiency of drug discovery because many drugs (up to 40%) fail at the stage of non-clinical pharmacology and toxicology. 217 Enhanced in vitro prediction models are therefore essential, and 3D tissue printing, rather than 2D cellular monolayers, 218 was shown to be an alternative approach to establish a 3D cellular scaffold. 219 Seeding of stacked sheets on a scaffold with cells, or using an extracellular matrix (ECM) before solidification, has been developed. 218 Various other methods have also been described to construct in vitro 3D tissue models, 219 and include bioprinting. 220 -222 Conventional 3D platforms have been used to simulate a variety of specific organs and disease models for assessing bioactivity. 219 However, they pose significant challenges, 223,224 including the application of different cell types in an assay matrix, batch to batch variation in the natural ECM matrices rendering the results inconsistent, the higher cost and time to establish an assay, the introduction of vascularization resulting in limited oxygen transport, and the absence of an ordered structure to mimic the in vivo tissue matrix with its myriad of other metabolites and enzymes in the surrounding space.
Some of the advantages of bioprinting include more anatomically relevant tissue constructs based on magnetic resonance imaging and computer tomography, greater porosity, co-culturing of different cell types, 225 integrated vascularization, 226,227 the controlled introduction of growth factors, genes, and enzymes, 228 and rapid fabrication. As a result, a wide range of tissue scaffolds have been developed 219,221,222 for use in biological evaluation. More exposure to natural products, both extracts and single compounds, in multiple tissue models, is needed to validate the assays and to conduct more tissue-relevant drug discovery and toxicity studies. Note that as synthetic constructs, these bioassays do not fall within the ethical considerations and regulations regarding the in vivo use of animal-based assays.
In the future, one can expect the further miniaturization of physiologically relevant, bioprinted assay scaffolds based on enhanced microfluidics, multiple organ on a chip models, and more diverse detection systems with linked outputs to reflect more holistic modeling. 219,221,222 The focus for natural products should reflect the opportunity for wider testing capabilities for diverse disease states, and the ability to assess potential, site-specific toxicity at an early stage in both a drug discovery and traditional medicine quality control mode. Costs for testing should be reduced in the long-term as assays are stabilized and can be reproduced locally. Regulatory agencies will need clarity of efficacy, reproducibility, and comparability with in vivo assays before such 3D-systems can be included in drug registration applications. Wider disease states will be 3D-modeled, including neglected diseases, which should allow more effective, local discovery programs in situations where resources are limited. Importantly, contributions of the constructs and the data sets to an international repository will advance the bioinformatics aspects of identifying real biomarkers in complex systems, such as a traditional medicine, and identifying in a comparative manner, the most effective testing constructs.
Data in Natural Product Discovery and the 4IR
Molecular Networking
The desire for interlaced natural product databases was discussed earlier in this review and elsewhere. 4 -6,229,230 One of the breakthrough responses to this need is the Global Natural Products Social Molecular Networking (GNPS) construct, a community platform for the storage and analysis of mass spectra. 231 The combined datasets (including MassBank, ReSpect, and NIST) grew rapidly to serve over 100 countries and over 9,200 users, and provides a solid basis for the identification of over 18,000 natural products. The program allows for on-line dereplication, as well as automated molecular networking analysis, and promotes the inclusion of curated new data.
A molecular network represents the interconnectedness of constituents in a metabolomic matrix through the alignment of their mass spectral fragmentation patterns. 232 It thus provides a mass lineage which, if the structures are known, or can be postulated, may also represent biosynthetic pathways, and develop relationships to metabolites of known chemical or biological properties. The molecular network 233 accesses over 272 public data sets to enhance the analytical capacity for an unknown metabolite. The reverse situation of specifically seeking analogs of a compound of interest in a matrix was also presented. 231 A recent use of GNPS revealed three major clusters of metabolites, catechins, flavonoid glycosides, and neolignan glycosides in Huangjinya green tea extract, with the neolignan glycosides showing anticholinesterase activity. 234 Through the addition of Feature-Based Molecular Networking, it became possible to distinguish isomers in metabolomics studies. 235 The use of molecular networking based on MS/MSn analysis is now a mainstay of metabolomics, and as data are refined and expanded will lead to even higher levels of accuracy of identification. The system represents a model of global community collaboration, and hopefully will inspire other areas of spectroscopy, DNA barcoding, and biological datasets to be integrated as a dedicated network response. 229
Big Data
Compartmentalization of data in the natural product sciences no longer fosters the development of effective and efficient research programs. 6 Natural product research is replete with data – big data, on many different phyla, compounds, activities, results, etc. These datasets include: taxonomic data about organisms, genomic data relating to organisms and their biosynthetic gene clusters, chemical data on isolates and their transformation products, methods and conditions of isolation, chromatographic (HPLC and GC) data, infra-red, ultra-violet, NMR (1H- and 13C-), and mass spectral data, and vast volumes of in vitro, in vivo, and clinical data. The evident goal for natural product research must be to have access to all of this information instantaneously, anywhere in the world. 229 It is time for a global effort to develop the threads, the networks, the webs, the lattice which promotes a labyrinth of data interconnectedness for natural product scientists to share. This will achieve an important impact of 4IR for natural products – an accessible large data set. What are the characteristics of “big data”? What constitutes big data? What is the most productive approach to using big data to optimize natural product research goals? Can AI propose new research programs and experiments and then, given resource materials, automatically conduct them? As a chemist, can AI design an improved molecule through an enhanced pathway process? Can it identify a preferred plant or microbe to study for a particular bioactivity? As a biologist, can AI indicate and conduct pertinent and predictive bioassays? What are the purposes of creating integrated and linked big data sets?
The size of the natural product datasets is an obvious characteristic. Another is the volume of data that are generated routinely. Characteristically, there are 3 sources of generated data, i) access to existing, smaller datasets through establishing linkages, ii) new data that appear in the hundreds of journals published each month (often as supplementary materials), and iii) experimental data not appearing in the literature, but directly made available from an investigators’ research, which meets curatorial guidelines and standards. At the present time, most of this natural product data is text (words or pictures); in the future, data will also be in a video format. Laney 236 proposed that three other parameters be used for the qualifications of big data, namely Volume, Variety, and Velocity. Others have proposed three additional attributes, Veracity, Variability, and Value. 237
The Volume of natural product data that already exist around the world is vast. There are 123 databases cited in the literature at the present time which contain various aspects of natural product data, although only 50 of them are open access. 238 A separate review of traditional Chinese medicine (TCM) databases is available, 239 as is a compilation of microbial biotechnology databases, 240 and of natural product structures. 241 The number of locally held (siloed) databases in specific areas of natural products research is not known. Consequently, the real volume of compiled data is completely unknown, although it is probably more than several petabytes. Variety refers to the structural heterogeneity that is inherent in each dataset, be that structured, semi-structured, or unstructured. Velocity indicates the rate at which existing data can be analyzed and correlated with new data to generate a response. One example would be the analysis of HPLC/MSn data for metabolite identification, and its extension to molecular networking (vide infra). 242 Contextualization is a vital component for optimum data usage and interpretation of data, 229 and can lead to more comprehensive metabolite identification with overlays of taxonomic, chemical, and biological data through relationships with other data sources. 243 This approach was used for metabolite identification in the GC/MS analysis of a new human and mammalian metabolite, 244 and the SMART tool was used to analyze >2000 HSQC spectra. 245
Which brings us to Veracity, what is the quality of the information is going into a dataset and thus what is coming out? Curation of data is of prime concern in the interrelationships of datasets, and has to rely on trust from colleagues entering small data sets (NMR or mass spectra or biological results), since rarely can they be verified for accuracy at input. Only subsequently will invalid data be exposed. Veracity when retrieving data also has a strong human component. For example, if an ion peak in a mass spectrum indicates several options for identification, will that be resolved accurately by the dataset, or will options be presented for human researcher interpretation? Variability in data may reflect lack of biological reproducibility, differences in NMR chemical shifts based on concentration and field strength, and different styles and language presentation for information on traditional uses which must be correlated. Value relates how much (the density) of the data in the data set can be utilized and how much is extraneous. That assessment will clearly depend on accessibility and volume of use over time.
What are the analytics that can be applied to such large, integrated datasets? For only then can a value be ascribed to the contained information. How rapidly does analysis and decision making need to occur to have relevance? For a GC-MS profile, ideally, in milliseconds. For an infield plant identification, the complete profile of the biological responses of a particular plant extract, or the isolates from a marine organism, seconds or even minutes, may suffice. Thus, data management, how data sets are assembled and interrelate, and analytics, how insights into the compiled data are obtained, are separate processes. As discussed elsewhere, 4 the ability to query a server verbally for a plant identification and literature background should provoke a response which has accessed, compiled, and integrated various forms of data in a text-based report format based on question, document, and answer processing, which ranks and refines the correlated data. From a texted query, a combination of information extraction from unstructured text, and abstractive summarization from structured datasets would lead to evidence-based decision-making (or the prompting of additional queries). The possibilities here are endless and would provide indescribable opportunities for research groups all over the world far beyond the limited datasets that are presently so carefully siloed and protected. There is a truly desperate need in the field of natural products for a dedicated system of communications between datasets for the processing of known and new information. A discussion of this issue in application to contextualized metabolomics 229 indicated that natural product communities are examining issues of best practice development as a pathway to enhancing sharing and interactivity. 237 Interfacing with systems for automated structure elucidation of natural products 246 also becomes an important aspect of moving such initiatives forward.
Blockchain technologies and their application to natural products are discussed in more detail subsequently. It is worth mentioning here that this technology also generates large interconnected, datasets, and promotes the practice of upfront data integrity which can be examined in an encrypted open-keyed system, thereby improving the likelihood of veracity as each dataset is entered. The fundamental difference is that in a blockchain the data is immutable, whereas in an integrated natural product network, secondary considerations, such as more sophisticated data and human curation, will likely refine datasets over time resulting in a steadily improving accuracy of the output.
The combination of AI and big data is a very powerful tool for natural products discovery research. Among the benefits include the conservation of information over time, and its accessibility and integration to indicate where societally relevant “holes” in data reside, and thus where research is possibly needed. Beyond the identification of neglected diseases and the correlation with ethnomedical uses of plants, interwoven with the in silico structures of known and new metabolites with a “fit” to inhibit an active site at an enzyme, there is an opportunity for drug discovery where resources can be focused on a highly select group of compounds whose probability for biological success is enhanced. 56 Acquisition resources are conserved, in vitro testing is limited, and potential toxicities are constrained; it represents a classical ecopharmacognosy outcome.
Machine Learning
The evolving relationship between large data sets, chemistry, and biology will continue to be enhanced through diverse applications of machine learning (ML). Machine learning is defined as the study and application of algorithms performing pattern recognition-derived predictive tasks. 247 The ability to adjust and “learn” based on additional, perhaps modulated, data input in a feedback loop, and seek new patterns for recognition provides creative insights into the chemical and biological relationships of natural products. 93 Within the natural product sciences, the potential applications are extensive and include recognition of genomic signature elements and predictions about collective outcomes biosynthetically, projections of bioactivity, propositions for (bio)synthetic compound diversity, and disease targeting. 248 Deep learning (DL) is an extension of ML and focuses on layers of neural networks that can assist in predicting protein structures. 249,250 DL has provided next-stage analysis of quantitative structure activity relationship (QSAR) data for mutagenesis, 251 for diverse biological activities using global libraries, 252 and identified the compound halicin as a structurally new, and mechanistically different antibiotic, along with eight additional candidate compounds. 253
An early review of big data analysis in crop plants summarized the potential for analyzing plant breeding characteristics, relationship of yields to climate, impact of pathogens, stress protection, and market needs. 254 It also emphasized the need for global systems that would integrate plant genome databases and provide the bioinformatics tools for analysis. Summaries of machine learning applications in natural products discovery 255 and the identification of priority biological targets 256 have been presented. More specialized datasets have focused on anti-malarial activity in a large dataset of natural products (although cytotoxic compounds were not eliminated, which significantly confounds the structural conclusions). Predictions for compounds to be evaluated were also made. 257 Two active synthetic mimetics were generated through a compound assessment using molecular targets analogous to the Alzheimer’s disease treatment alkaloid galanthamine, and supported by in vitro testing. 258 An analysis of 25,523 natural isolates from fungi and bacteria (NatProdAtlas) revealed new distinctions of evolutionary origin based on their structures and biosynthetic pathways. 259 Using a dataset of 201,791 natural products in comparison with the same number of synthetic products, a machine learning algorithm developed parameters for distinguishing the 2 sets with high accuracy and quantifying natural product-likeness. As expected, the space occupied by the natural products and the overall diversity and complexity (eg, stereocenters) was much larger for the natural products, the number of nitrogen atoms was smaller, and the number of oxygen atoms greater. 241
This is a very rapidly expanding area of natural products research, and promises to advance the discovery of compounds for initial bioscreening in an appropriate assay. The challenge, as in any natural product discovery program, is access to a supply of the pure metabolite to be tested. Thus, while machine learning may provide potential hits, reality dictates that an important refinement of the algorithm must include the accessibility of the originating organism, identification of an alternative source, and sustainability if a “hit” occurs.
Blockchain and Distributed Ledger Technologies
Blockchain technology is a distributed ledgering system, based on cryptographic annotations that allows for the continuous management, verification, and public recording of a sequence of transactions between two or more parties. It provides an immutable and visible record of each point of movement and financial contact in a production chain. 260,261 In previous discussions of the production of traditional medicines and dietary supplements, one of the major concerns from a patient perspective was traceability. 5,230,262 Among these concerns have included questions regarding the point of origin, the age of the plant material, from where, when, and how it was grown and acquired, when it was processed, how it has moved up the value chain from farm (or forest) to store shelf, and how it was distributed to reach the point of sale. Patients have been demanding traceability, transparency, and sustainability certifications in these products for a long time. 263
Blockchain technology can provide significant, transparent answers to those questions for all of the product ingredients. 264 In the process, it markedly reduces the risks associated with potential adulteration, contamination, or degradation, 265 and thus serves to restore the trust that has dissipated over the past years with respect to what is in a traditional medicine or dietary supplement package. From a manufacturer perspective, an enhanced and unequivocal record of the supply chain is generated, thereby fostering profiles for complex ingredient matrices. Several data processing companies operate blockchain technology programs, and some major manufacturers using natural materials, including foodstuffs and cosmetics, are piloting programs for the supply chains of these constituents. 264,265
At the other end of the value chain, the suppliers of the raw materials, who are frequently small-holding farmers in emerging economies, become more assured that they are being paid appropriately for their crops, be that coffee, vanilla, or a medicinal plant. Sustainability concerns can also be addressed, since usage volumes in a particular area, throughput, wastage, and sell-by dates, can all be monitored with mobile phone apps. The demonstrated traceability and transparency in the available information on-line through a lot number also promotes the restoration of trust from the consumer/patient. Blockchain technology is also monitoring the recycling and sustainability aspects of the product packaging chain. Figure 3 summarizes the impact of blockchain technology on the quality control of traditional medicines and phytotherapeuticals.

Blockchain technology in traditional medicine quality control.
A large cosmetic company, in conjunction with a major supplier, has been experimenting with this technology for the production of vanilla from Madagascar, 266 and the sensor-based application for tracing wood from sustainable harvest to the consumer purchase has been described. 267 For the medicinal plants used in dietary supplements and phytotherapy the benefits for the stakeholders in the supply chain, and especially for the patient, blockchain could be highly significant. In addition to the financial perspectives and the traceability benefits, there are aspects of enhancing initial quality, recording the use of pesticides and herbicides, the deployment of organic farming protocols beyond Good Agricultural and Collection Practices, discerning the potential for adulteration and contamination during storage or transportation, and assuring application of the regulatory constraints, including the local implementation of intellectual property rights, as required by the Convention on Biological Diversity and the Nagoya Protocol. 268,269 A pertinent discussion of the impact of blockchain technology in the supply and quality chains of medicinal plants is available. 270 Other applications in biotechnology and pharmacy include the security and distribution of genomic data on plants, microbes, and particularly humans. This will overcome some of the issues currently associated with the silos of data that require delocalization. 271 The basic human right of personal access to one’s genomic data and the importance of that in the future of medicine and the development of personalized treatment regimens remains controversial. 271 Another application in the pharmacy/medicine area which would benefit from blockchain technology is for ongoing diversified clinical trials, where immutable data could be visualized by all approved parties for patient consent and inclusion, assure quality in performance and metrics, and speed up data analysis and conclusions regarding both efficacy and side effects. 272,273
Integration of Data and Technologies in Natural Products Research
Cyberecoethonopharmacolomics
The term “cyberecoethnopharmacolomics” (CEEPO) was proposed as an integration of ecopharmacognosy 230,262 with many disciplines and technologies to promote philosophies and practices which engender a holistic, highly interactive approach to natural product development. For example, opportunities are seen between sustainability and economics, sponsoring commercial development, and aiming to encourage greener processes and products on behalf of the patient, thereby reducing the carbon footprint. The prescribed morphemes are: “cyber” – which embraces all the aspects of big data, data security, AI, robotics, computer-aided synthesis, plant identification, molecular networking, network pharmacology, remote sensing, and more comprehensive access to natural product datasets; “eco” – signifies consideration of the sustainable development aspects, and includes the environmental impact, at each research and development stage in the initiative, and subsequently in production processes; “ethno” – reflects the respect, based on thorough analysis and prioritization, for the global importance of traditional medicine data sets for future health care initiatives in any program focused towards medicines security; “pharmacol” – is the in silico, in vitro, and in vivo determination of toxicity and biological activity in a conserved and sustainable manner; and finally there are several “omics”. There are the taxonomic aspects of the organisms, their genomics, particularly in relation to biosynthesis, the metabolomics of an extract as an identification, drug discovery and quality control tool, and there are the agronomics of sustainable cultivation, marine or terrestrial, for the selected organisms. Finally, the overarching economic aspects must be integrated, which can drive the research and development programs, may develop new product opportunities, and result in an accessible (available and affordable) product for the patient. The holistic term CEEPO fits well with the 4IR, as it embraces many of the twelve emerging technologies. It also is aligned with the strengths of the quintuple helix (vide infra).
Natural products from diverse sources have been, and remain, a vital source of drug leads and drugs 274,275 ; in part due to their exceptional spatial and stereochemical attributes. 248 Libraries of natural products and extracts from biological samples represent a critical resource from a compound spatial perspective for screening as a plethora of biological test systems are developed. 276,277 There are considerations related to conservation and sustainability which must be addressed for natural products discovery programs, 278 and these have dampened, or eliminated natural products, other than select single compounds, from the focused discovery programs of major pharmaceutical companies. As a part of 4IR initiatives towards robotics and automation, big pharma discovery programs are now highly automated with vast libraries of compounds, full robotic processing for high- and medium-throughput screening against a biological target, and prototype automated design-synthesize-test-analyze facilities. 279 These advances favor faster feedback loops for “hits” and failures. Automation can enter at the stage of molecule design based on Lipinski’s “rule of five”, 280 through computational compound profiling, 281,282 or in-house prior design success. 279
Deep learning methods have been applied using neural networks and other approaches for de novo structure design which should enhance green chemistry targets for minimizing derivatives, solvents, reagents, and processing materials for a given set of test materials. The practical challenges of these approaches have been discussed, 279 and besides the reproducibility of the chemical transformation steps and the biological steps, the importance and value of human input in consideration of (false) positive and negative data was emphasized. Next steps may include the miniaturization of automated bench-top devices for synthesis and testing in a single assay, making the technology more widely applicable, and therefore suitable for a wider range of assays for libraries of natural products. 131 Future considerations will focus on the notorious failure rate of “hits” as they proceed through development, as well as considering green synthetic pathways, and algorithms for positive design outcomes that lead to a more predictable biological response. The very limited work that has been conducted on the structure modification of available plant-derived natural products (other than atropine and morphine) makes them ripe for future development, even in big pharma, as “new” skeletons are subjected to computational and in silico binding studies.
In that regard, the Universal Natural Products Database (UNPD) accumulated 197,201 natural product structures and docked them with 332 target proteins related to FDA-approved drugs. 131 Several natural products which could be regarded as likely to interfere in bioassays because they showed many target responses (eg, staurosporine, 283 quercetin, 284 and genistein) were identified, and at the same time new, potential and diverse biological responses were suggested for other natural products. The practical considerations of invalid metabolic panaceas (IMPS) 215 285 and pan assay interference compounds (PAINS) 214 are important. These widespread metabolites (eg, ursolic acid) may be present at such a level in a tested extract to mask the identity of a truly active metabolite, leading to confusion in the discovery decision-making process. 286 Automated dereplication studies based on a chemo-bio-MS assay protocol would markedly assist in identifying these compounds. Similarly, they should not be present in single natural product screening libraries, as they will serve to create false positives. Microfluidics is an important aspect of drug discovery for synthesis, in vitro testing, and mimicking organ tissue for testing (see 3D-printing section). 287 Significant more effort is needed to develop libraries of “clean” extracts from natural sources for bioevaluation which will not confound assessment due to fluorescent or other interfering compounds.
The Triple, Quadruple, and Quintuple Helices
The discussion of cyberecoethnopharmacolomics as a comprehensive, cyclic system of multifaceted and interwoven considerations and actions relating to natural products innovation, 5,6 and its subset ecopharmacognosy, can be illustrated through the development of the quintuple helix model for natural products in society. With the mitigating factors of losses of biodiversity, traditional knowledge, climate change and time, the international response is sustainable development as a series of seventeen goals (SDGs). 109 The relationships between sustainable development and 4IR have been discussed. 288,289 Many of the goals have a direct relationship to the future of natural products in 4IR, including sustainable agriculture, promoting health and well-being, education and life-long learning, sustainable economic growth, industrialized innovation, sustainable consumption, combatting climate change, conserving and managing global terrestrial and marine natural resources, and establishing global partnerships.
While the unattended consequences of the mitigating factors are dire, in fact there is an enormous opportunity for corrective action, as expressed in various statements, agreements, accords, etc. over the past 25 years or so. 290,291 The bottom line, as eloquently indicated in the 12th century by St. Hildegard of Bingen is that “The Earth which sustains humanity must not be destroyed. For without it we cannot survive”. 292 In the early stages of the Anthropocene Era, 293 it is up to humanity to do our utmost to correct (or at least minimize) the damage done, so that future generations may survive in greater harmony with nature. There is no precedent for this moment in human history. There is no prior experience which applies. Corrective action will necessitate innovation, new thinking, and actions, and thus the creation of new knowledge to supplement the wealth of the accumulated knowledge from human historical records. We must examine our lives as a part of a societal sustainable impact assessment; we will all be found “wanting” to a greater or lesser degree. Many countries (eg, US, parts of Europe) will be high on the list of those nations most “wanting” based on resource usage, etc. on a per capita basis, and other countries will be identified on a “volume” basis. 294
The systems for “knowledge” generation and transmission have evolved dramatically in the past 300 years as an aspect of the successive industrial revolutions. For many decades in the 20th century, the basis of actions in society was a triple helix of innovation, in which academia, government, and industry collaborated towards a common, knowledge-generating goal. One example in the area of natural products drug discovery was the National Cooperative Drug Discovery Grant program of the United States National Cancer Institute, 295 a fundamentally linear innovation program.
The challenge for natural products development for society is accompanied by a moral obligation to sustainability through actions based in knowledge. The existing knowledge is available to all and is the fundamental global resource for innovation. Knowledge creation is proceeding each second of each day in all the diverse segments of society, many of which, at their core, are based in nature. How this new knowledge is harnessed and processed, as “big data” in 4IR, for societal benefit is an enormous challenge, one which lies at the very core of the Quintuple Helix model. 296,297
In examining the model of the Quintuple Helix, the first aspect is academia and the traditional role of the university as a center for research where the research and knowledge created was approved by peers before dissemination. The second aspect, classically evolving in World War II with the development of antimalarials and antibiotics, became the role of government in sponsoring, through various mechanisms and levels of investment, research that was needed as societies evolved; the continuing “War on Cancer” is another example. 298 To bring an even greater level of focus and refinement to the knowledge-generating, innovative programs, and to introduce pathways to enhance the transition from bench to bedside, industry became involved in continuing and translating the innovation steps to a practice or a product. Linking basic research with problem-solving resulted in both interdisciplinary thinking and knowledge generation, and successive application, leading to direct societal benefit, such as a new anticancer drug.
The Quadruple Helix added a fourth aspect of the public domain, namely, culture, media, and of a knowledge society in which information is totally accessible. Put another way, there is knowledge democracy on which innovation is based. In the natural products and biological sciences, one form of media transmission is open access journals which are enhancing the information base for scientists all over the world, whose access otherwise would be limited through high-priced journals. This aspect of publishing is likely to continue to undergo rapid change in terms of translation time from bench to public domain. In the covid-19 crisis, communications regarding research on both drugs and vaccines has frequently bypassed the traditional peer-review stage completely, and whether from basic or clinical scientists or big pharma, information has been passed to the media before any peer review process. Other models for publishing research are likely to evolve in the near future as the relationship between the primary literature and database knowledge acquisition and processing becomes more flexible and holistic.
In the Quintuple Helix, the added aspect is the “natural environment” 296 which provides for the considerations of sustainable development and environmental impact (part of the umbrella term “social ecology”). Social ecology examines the relationship between the co-development and the co-evolution of society and nature. It is the relationship between what culture produces physically and through knowledge generation and the dynamic interactions with the natural world and its healthy maintenance for the benefit of society. The European Commission established this transition many years ago as being a major challenge for economies and the development of the societies they support. 299 For natural products and society, this fifth component of socioecological transition builds on the inherent and inseparable relationship between humans and nature. It supports the Gaia hypothesis 300 of Earth as an integrated and fully interactive organism. The ability to generate new knowledge and innovation in this area is consequently the primary force for societal progress in an environmentally conscious era.
The Quintuple Helix model represents a cooperative system of knowledge, experience, and innovation, and promotes a full spectrum of associated inter- and transdisciplinary interactions for the development of the basic and applied natural sciences, as well as the social, economic, and humanitarian sciences. It has been defined as the answer to the question “How do knowledge, innovation, and the environment relate to each other”. 301 The core knowledge system remains the Triple Helix paradigm, 302 expanded with the public system of traditions, values, and experience. It is this system which promotes the knowledge for the society. 303 Adding the fifth helix system, based on the natural environment, is the stimulus for innovation generation, and completes a web of interconnected activities which considers conservation of resources and the development of society as inextricably interlocked activities. These are embodied through the development and application of sustainable and green technologies based on natural resources. 296 The term “eco-innovation” and the business aspects “eco-entrepreneurship” have been suggested to encapsulate these activities 296 ; for natural products it is ecopharmacognosy. 230,262 How these interactions of the 5 subsystems (the helices) operate for natural product development is proposed, based on an existing model. 301
In the education system, the focus is on the development of the “human capital”, comprised of the teachers, the students, scientists, and the academic entrepreneurs who generate, through creative thought and action, the research knowledge of natural products. The economic system is the government component, the business aspect, the companies, the funding sources which underwrite both the original research and the progress of the innovation into diverse products derived from natural sources, with all the machinery and technology that makes that a reality. The natural environment is the ultimate resource for materials for sustainable development, together with the considerations that involve prioritization for research program activities. The media-based and culture-based public is how the 2 systems of social capital (traditions, values, etc.,.) with the canonical media (journals, books, television, etc.,.) interface with the contemporary media (internet, social media) as the so-called capital of information. 301 The political system is about the expressly defined desires of the state with respect to the present and the future, and the boundaries (laws, regulations, etc.,.) within in which the natural product innovations are presented to the users.
These systems are built on an ethos of innovation for the sustainable development of the society. 301 Each system is itself an aspect of information circulation in which there are inputs and outputs, such that an input into a subsystem is knowledge creation in that subsystem. That new knowledge is utilized by the subsystem innovatively to create an output of new knowledge for another system. That may result in further innovation within the same subsystem, or it may become an output to stimulate innovation in other subsystems. In which case, there is a “circulation of knowledge”, which has two outcomes, stimulation for advancing innovation, which eventually creates the “common knowledge” for sustainable development. How can this be applied to a situation of relevant interest, natural product development for medicinal agents?
All research initiatives begin with an analysis of prior knowledge. That knowledge to be analyzed may originate in several of the subsystems. For example, what does society need? What is the previous research that is available? What types and levels of expertise and facilities are needed? What is the most economic and effective approach to conduct the research, and how will that be funded? What are the legal and regulatory constraints with respect to the conduct of the research, and further down the development line to an industrial product, such as a single-agent drug or an evidence-based, standardized, complex traditional medicine? In the process of responding, each question is answered in consideration of the core perspective of sustainable development, and the innovation derived therefrom.
Based on the knowledge analysis of a defined research area identified by the political system (which includes relevant government agencies), an investment stream is elaborated which assures program development over time with respect to new equipment and facilities, new scientist positions, and resources for program function. This investment generates a higher level of knowledge creation (output) from the education system. Supplementary programs for training and new technologies can further enhance output. The investment therefore has a positive, long-term impact on promoting the existence of human capital for future programs. For a traditional medicine program, several high-profile plants might be selected for development based on the sciences within the education system, cognizant of the constraints of the economic system (production costs, market forces) and the political system (defining the regulatory niche), and focused on sustainable access to the raw material, and the greening of the research and production processes.
The result is an input of new knowledge for the economic system which is then challenged to innovate the sustainable development of an appropriate production process, including the marketing and product service aspects, as well as new values for corporate social responsibility such as package disclosures, product consistency, and patient information. Which links to the media-based and culture-based public and the natural environment, since new inputs are generated through a demand for sustainability of the natural resources and the innovation in developing that availability, possibly through cultivation. That output feeds both the education system and the economic system. The innovation may lie in maintaining or restoring the balance of nature (knowledge creation of remediation) which could be applied in other circumstances. An output of the natural environment system is therefore “green know-how”. 297
The “green know-how” innovation for a series of medicinal plants will become input for a greener, media-based and culture-based public. New capital is received regarding both care for the environment and remediation of depleted resource areas. That may incentivize the media-based public to acquire a more expansive appreciation of natural resources in general, and the culture-based public to promote new outputs of needs, wishes, and issues for inputs into the political system. For example, broadening concerns of quality and sustainability to other sectors that rely on natural resources, such as the home construction, the food, or the cosmetics industries. It may stimulate demands for an end to the concept of “buyer beware” in medicines generally when the media-based public is acquiring those information materials through unregulated internet access. That represents a stimulus for program creation and investment in knowledge creation through innovation in a new area to improve the health of the society and the economy. Thus, cyclic distribution of knowledge between the 5 helices promotes sustainable development, an improved knowledge economy, and knowledge democracy, where new knowledge is distributed widely in society (internet access), and is not siloed with the knowledge creators. 301 The model serves to demonstrate the concepts of cyberecoethnopharmacolomics. 5,6
In India, the Digital India initiative is underway to enhance information distribution, productivity, and the quality of life overall based on a knowledge economy in the 156 million rural households. One natural product focus is the development of sustainable ecosystems for small and marginal farmers through the Small Farmers Agribusiness Consortium, including those farmers specializing in growing medicinal plants. 304 Apps for the many rural languages are being developed to shift to a knowledge-based agriculture system for productivity enhancement.
Higher Education
Large datasets are an aspect within big data. Knowledge access can be facilitated. But knowledge does not drive innovation, human (and in the future AI) creativity based on that knowledge and information does. One critical function of academia now, and will be more so in the future, is to expose students to more creative, multidisciplinary opportunities as an integral aspect of “learning” about natural products. There have been several discussions presented regarding the impact of 4IR on higher education, particularly focused on changes in knowledge creation, integrating the impact of emerging technologies and sustainable development, and redesigning education goals to meet the changing skill sets needed for societal development. 305 -307
Based on a survey of 2,350 scientists in South Korea in 2017, it was recommended that the highest government priority be given to improving science and technology education and the research and development system itself, not the development of specific technologies. More precisely, the focus was on increasing creativity as an educational goal in an interdisciplinary manner and streamlining the regulatory and legal aspects of research and technology transfer. 308,309 In terms of the interface with the Triple Helix (academia, industry, and government), there is an inherent core value that technological development is a natural phenomenon empowering people, which can be focused on societal need, and that the result can be value laden. The development of new technologies which relate to drug discovery for neglected or high-need diseases, and the enhanced quality control of traditional medicines are examples for natural products in the future. Such considerations indicate that education systems, particularly universities, need to incorporate more “learning by doing”, or the creativity implicit in “design thinking”, where challenges are identified and open-minded possibilities are analyzed to achieve an inter- and multidisciplinary response. 310 What does that mean for undergraduate student education?
The role of the ivory tower in society will be changing, for that role can no longer only be to provide a well-defined curriculum with a cookie-cutter degree. There is already a societal need for a much stronger and probably life-long educational relationship between universities and students, at all levels, post-graduation, as skills will need to be updated over an extended time frame. Universities are the key societal players in that new dynamic. 311 Rapid change and growth in the respective sciences, and in technology applications, are anticipated core outcomes of 4IR. Consequently, a curriculum, in chemistry, biology, biotechnology, or pharmacy, will require reassessment with unprecedented regularity to reflect those rapid changes.
With all relevant knowledge literally in hand, the competitive “edge” for success will be a creative one, which students, and later, graduates, will be challenged to develop, repeatedly. Faculty development will need to be similarly creative to foster interactions with leaders in the field, and facilitate participation in specialized training opportunities for technology integration and new skills development. Sustainability and ethical issues of impact in society will be highlighted in new learning experiences as the impacts of robotics, the internet of things (IoT), genomics, and biotechnology advances challenge the contemporary boundaries. Understanding and translating those issues into a curriculum and into research practice will be necessary to maintain a creative advantage as an essential aspect of approaching the 2030 goals of sustainable development. Individual and collective (team-processed) creative intellectual activity will be highly valued. Academic units involved in natural product research will be charged with aggressive curriculum reviews for undergraduate and graduate students, as well as greater investment in structured faculty development programs and workshops. Those who have integrated new technologies will be strongly encouraged to share that expertise. The demolition of that old ivory tower is an essential component for a successful 4IR.
Holistic Integration and Deep Work
The world is a busy place. Multitasking is commonplace. All the information we need to live and thrive is, as discussed, in our hand. There is climate change and the need for sustainable development. Technology innovation is rampant. For the future, the differences between the development of science and technology, and therefore economic growth, in various nations will depend, significantly, on knowledge evolution (generation and application), and the innovation and creativity that is generated based on that knowledge, and ultimately the new pathways that are formulated. Creative thinking is sometimes like a light switch, the idea appears, suddenly. At other times, the solution, the new pathway, evolves over time, consciously. Inspiration and creativity often occur only in dedicated time; the time referred to in “Deep Work”, 312 where focused, uninterrupted time is set aside, distraction-free, for concentrated thought. That inspiration for innovation may occur individually or collectively as a team research development activity.
For natural products research in the era of 4IR and the Quintuple Helix there is much to ponder, much to be debated, much to be funded (and some programs unfunded!), and much to be initiated. If you are feeling comfortable with your research programs now, there is something amiss. This is not a time for clinging to the comfort of the status quo. Quite the reverse. It is an exciting time, filled with the opportunities to integrate, innovate, and create action plans. It is a time for the technologically integrated natural product sciences to provide a proactive response, not a reactive one. “Deep work” will be essential to move from the silos of 2IR and 3IR to the exciting possibilities of transforming how and where the natural product sciences are conducted in 4IR, with a focus on sustainability, on the patient, and on the impact of climate change. Research on sustainable practices and climate change on medicinal plants are a reactive response to the effects of the previous three industrial revolutions. With the ability to exploit these new developments in technology, natural product researchers in the 4IR must optimize their research through a proactive approach to sustainable practices and technology integration. That’s a risk, so that as the integration of technologies expands, natural product research will need to actively and consciously participate in ensuring that alternatives and collaborative processes are developed and implemented while in the throes of the full impact of 4IR.
The 2IR and the 3IR focused on reductionism and separation as the sciences evolved and developed. Under 4IR, advancement will be achieved through focused holistic integration. The applications of research must be targeted to societal needs; answering the question “Who in the society is this for?” Wasting resources on dead-end, individualized projects with no societal benefit, or focused on non-sustainable plants or resources, should no longer be supported intellectually or financially; those intellectually luxurious days are over. The proactive and conscious integration of technologies across disciplines, the centralization of access to natural product data, and the application of soft robotic systems are the new universal standards of research in the natural product sciences. They are ready to initiate programs focused on real societal challenges, including medicines security for the burgeoning population.
The covid-19 pandemic crisis has shown us that whatever “normal” was, it can no longer be, and we never will return to, the “old normal”. How we interact with each other through local and international “meetings” has been transformed. How the results of medical science are transmitted, by-passing peer review and appearing as a news headline, is a revelation. The time to approve a new drug (camptothecin took almost 30 years!) or a vaccine, has been summarily slashed to a few months. How does that fit into new models for natural product research, for identifying new classes of drugs, identifying health care gaps for neglected diseases, for new strategies to standardize traditional medicines and phytotherapeuticals, for disclosing new biosynthetic pathways, for systems to conserve and preserve natural resources for our descendants, to improve health care and agriculture, locally and globally, to optimize technology integration into natural product research, and to rewrite fundamentally the legacy of natural products of abuse? In 2017, it was sixty challenges for natural products for 2030. 4 Now, it’s up to natural product researchers to do their own “Deep Work”. Take on a challenge from that list or add to that list with your own global challenges for natural products. As the English philosopher and writer George Bernard Shaw said: “You see things and say Why? I dream things and say Why not?” Acting collectively, cooperatively, and collaborating across many sciences and technologies will make long-term transformational dreams possible and enhance the role of natural products for the benefit of society.
Footnotes
Acknowledgments
The authors appreciate Professor Nor Hadiani Ismail, Atta-ur-Rahman Institute for Natural Products Discovery, Universiti Teknologi Mara, Puncak Alam, Malaysia for initiating the topic of the role of natural products in the Fourth Industrial Revolution.
*
Dedicated to Professor K. Hüsnü Can Başer on the occasion of his 70th birthday.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
