Abstract
Diabetes mellitus (DM) is a chronic disease that is very common and seriously threatens patient health. Gegen Qinlian decoction (GQD) has long been applied clinically, but its mechanism in pharmacology has not been extensively and systematically studied. A GQD protein interaction network and diabetes protein interaction network were constructed based on the methods of system biology. Functional module analysis, Kyoto Encyclopedia of Genes and Genomes pathway analysis, and Gene Ontology (GO) enrichment analysis were carried out on the 2 networks. The hub nodes were filtered by comparative analysis. The topological parameters, interactions, and biological functions of the 2 networks were analyzed in multiple ways. By applying GEO-based external datasets to verify the results of our analysis that the Gene Set Enrichment Analysis (GSEA) displayed metabolic pathways in which hub genes played roles in regulating different expression states. Molecular docking is used to verify the effective components that can be combined with hub nodes. By comparing the 2 networks, 24 hub targets were filtered. There were 7 complex relationships between the networks. The results showed 4 topological parameters of the 24 selected hub targets that were much higher than the median values, suggesting that these hub targets show specific involvement in the network. The hub genes were verified in the GEO database, and these genes were closely related to the biological processes involved in glucose metabolism. Molecular docking results showed that 5,7,2', 6'-tetrahydroxyflavone, magnograndiolide, gancaonin I, isoglycyrol, gancaonin A, worenine, and glyzaglabrin produced the strongest binding effect with 10 hub nodes. This compound–target mode of interaction may be the main mechanism of action of GQD. This study reflected the synergistic characteristics of multiple targets and multiple pathways of traditional Chinese medicine and discussed the mechanism of GQD in the treatment of DM at the molecular pharmacological level.
Keywords
Diabetes mellitus is a collection of metabolic diseases featuring hyperglycemia. 1 Long-term hyperglycemia in diabetes leads to the chronic damage and dysfunction of various tissues, especially the eyes, kidneys, heart, blood vessels, and nerves, which seriously harm the health of patients. 2 Gegen Qinlian decoction (GQD) comprises Pueraria (Gegen), Scutellaria baicalensis (Huangqin), Coptis chinensis (Huanglian), and Glycyrrhiza uralensis (Gancao) at 8:3:3:2. The clinical effect of GQD in the treatment of diabetes has been reported. 3 A large-scale randomized, double-blind, placebo-controlled clinical trial showed that 12 weeks of GQD significantly reduced fasting blood glucose and hemoglobin A1c, increased the number of beneficial bacteria and improved the stability of islet cells in patients with diabetes mellitus. 4 A large number of clinical experiments have shown that GQD has a definite effect on lowering blood sugar and has little side effects after long-term use. 5 -9 The mechanism of GQD is worth further studying. 10
Traditional Chinese medicine (TCM) is a complex system with multiple components, multiaction targets, and multi-action pathways. 11 Network pharmacology is an emerging discipline that reveals the effect of TCM on the body regulation network at the system level and builds a bridge between TCM and modern pharmacology. 12,13 At present, research on the protein interaction network has made great achievements in revealing the correlation between protein functions, the pathogenesis of disease, and the mechanism of drug action. 14,15 Li, Huiying 16 confirmed that a new antidiabetic component, 4-hydroxytoluene, extracted from Puerariae radix, can increase the insulin secretion of Rin-5f cells and improve the insulin resistance of 3T3-L1 fat cells. Liu et al 17 systematically explained the mechanism of TCM in the treatment of polycystic ovary syndrome through the integrated protein interaction network and the disease target network of polycystic ovary syndrome. Based on the difference in gene expression between colon cancer patients and the control group, Yu et al 18 successfully screened the key functional modules and discussed the possible mechanism of colon cancer. Zanzoni et al 19 integrated the proteomic data of different stages of cancer into a functional module of the protein network, reflecting the difference in protein expression at different stages of cancer. Today, this method has been applied broadly in research on drug action mechanisms and disease pathogenesis. 20 In this study, the drug target network and disease target network were integrated, and a GQD protein interaction network and diabetes protein interaction network were constructed to screen hub nodes as potential therapeutic targets for diabetes. This study is designed to illustrate the mechanism of GQD in diabetes mellitus treatment at the molecular network level and to offer a scientific basis for its clinical application.
Materials and Methods
Acquisition of GQD Network Data
The 4 hourserb-related data on GQD were mainly obtained from the pharmacology database of the TCM system (TCMSP, http://ibts.hkbu.edu.hk/LSP/tcmsp.php), 21 SymMap database (SymMap, http://www.symmap.org/), 22 and TCMID databases (TCMID, http://183.129.215.33/tcmid/), 23 the 3 largest data platforms for TCM. These databases contain information on the chemical constituents and pharmacokinetic (ADME) characteristics of TCMs commonly used in the pharmacopoeia of the People’s Republic of China. Oral administration is the most widely used method for Chinese herbal medicines to function in the gastrointestinal tract. 24 Thus, 2 models related to ADME, drug similarity (DL) and oral bioavailability (OB), were chosen to screen the bioactive ingredients of GQD. The screening criteria were as follows: OB >20%, DL >0.18. 25 The target information on the active ingredient was obtained from the TCMSP system pharmacology database and the BATMAN database (http://bionet.ncpsb.org/batman-tcm/index.php), 26 after which we obtained the target corresponding to the active ingredient. The target protein–protein interaction information was obtained from the String 9.1 database (http://string-db.org). 27 All PPIs are rated for confidence in the database. 28 In this paper, PPIs with hub >0.7 were imported into Cytoscape version 3.70 to construct a protein interaction network. 29
Acquisition of Diabetes-Related Targets
Data on diabetes targets were obtained from the human Mendelian genetic online database (OMIM, Online Mendelian Inheritance in Man, http://www.omim.org/), which is an online database of human genetic disease genes. 30 The PPI information on disease targets was obtained from the String database. We selected PPIs with hub >0.7 and imported this information into Cytoscape to construct a protein–protein interaction network.
Functional Analysis of the Network Module
The molecular complex detection (MCODE) algorithm is used to cluster the constructed protein–protein interaction network. 31 A Gene Ontology (GO) enrichment analysis of the identified modules was performed using BinGO plug-ins in Cytoscape. The P value represents the probability that a group of genes belong to the same gene standard. The smaller the P value, the greater the probability that a group of genes belong to the same gene standard. The annotation information is derived from the GO (http://www.geneontology.org/). 32
Hub Gene Screening
To further explore the pharmacological mechanism of GQD in mellitus treatment, the GQD protein interaction network and the diabetes protein interaction network were combined to screen the network hub nodes. Based on the Network Analyser tool in Cytoscape, the topological parameters of each node in the 2 networks were calculated to determine the importance of the hub nodes in the network. 33 The interaction and biological function of the hub nodes were analyzed based on the GeneMANIA plug-in Cytoscape. 34
Hub Target Verification
We verified the accuracy of screened hub genes based on existing literature reports and GEO database gene expression data to determine whether there is differential expression of hub genes between diabetes samples and normal samples. Differentially expressed genes (DEGs) in normal and diabetic samples were screened using the limma package in R. |logFC| > 1, and P <.05 is defined as DEGs.
GSEA Analysis
To illustrate the effect of the high expression or low expression of hub genes on the metabolic pathways regulated by these genes, we used Gene Set Enrichment Analysis (GSEA) version 2.2.2 software to conduct an enrichment analysis of the metabolic pathways related to the hub genes. The expression values of hub genes in the GSE25724 dataset were used as phenotypes. The c2.cp.kegg.v6.1.symbols.gmt dataset in the msigdb database of the GSEA website was selected as the reference gene set. The number of random combinations was set to 1000.
Molecular Docking
Download 3-dimensional (3D) structure of compound molecules based on PubChem database (https://pubchem.ncbi.nlm.nih.gov/) (PubChem: a public information system for analyzing bioactivities of small molecules). The 3D structure of the target is searched and downloaded from the PDB protein database (http://www.rcsb.org/pdb/home/home.do), and the crystal structure with higher resolution is preferably selected, which is compounded with the corresponding biologically active ligand (Inference of macromolecular assemblies from crystalline state). Ten of the protein targets corresponding to the 24 hub nodes have a protein crystal structure. The molecular docking software used in this study is AutoDock Vina, which performs semi-flexible molecular docking calculations (Software news and update AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and Multithreading). Before the docking simulation, all ligands were removed from the protein receptor complex, and polar hydrogen atoms and charges were added. Finally, semi-flexible docking was used to molecularly dock the compounds in astragalus. According to the magnitude of the root mean square deviation (RMSD) of the docked ligand molecule and the original ligand molecule, the rationality of the docking parameter setting is judged. According to the literature, it is generally agreed that the conditions for docking to repeat the ligand–receptor binding mode well are RMSD ≤2.0 nm (Dual anticoagulant/antiplatelet activity of polyphenolic grape seeds extract). At the same time, it can be considered that the docking parameter settings are reasonable.
Results
Data Analysis Flow Chart
The data analysis flow chart of the whole article is shown in Figure 1.
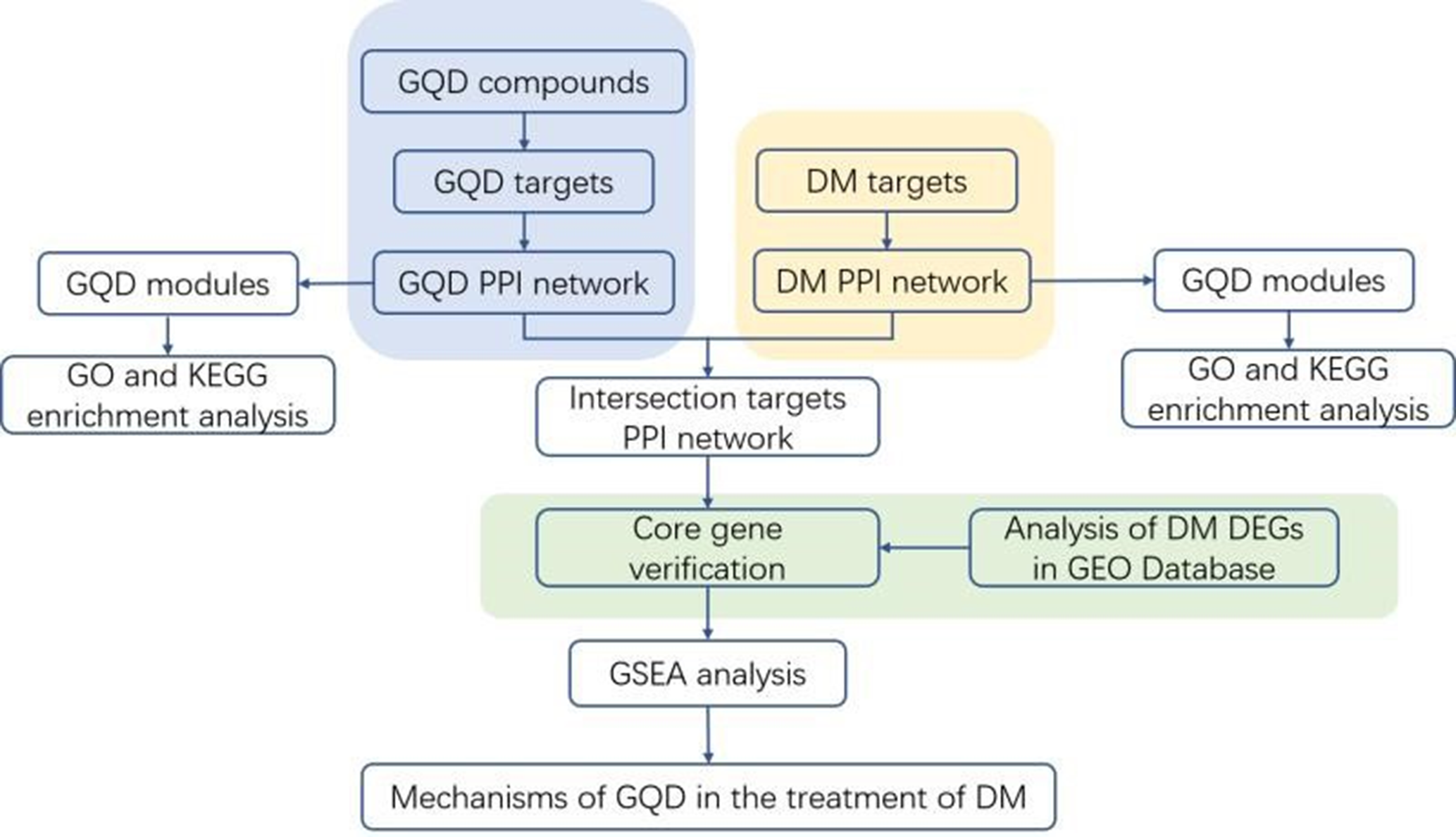
Experimental flow chart. DEG, differentially expressed gene; DM, diabetes mellitus; GO, Gene Ontology; KEGG, Kyoto Encyclopedia of Genes and Genomes.
GQD Compound–Target Network
Four ingredients of TCM in GQD were retrieved from 3 major databases, generating a total of 974 species. After removing the repetition, the study included 43 components shared among the 3 databases, 35 as shown in Figure 2.
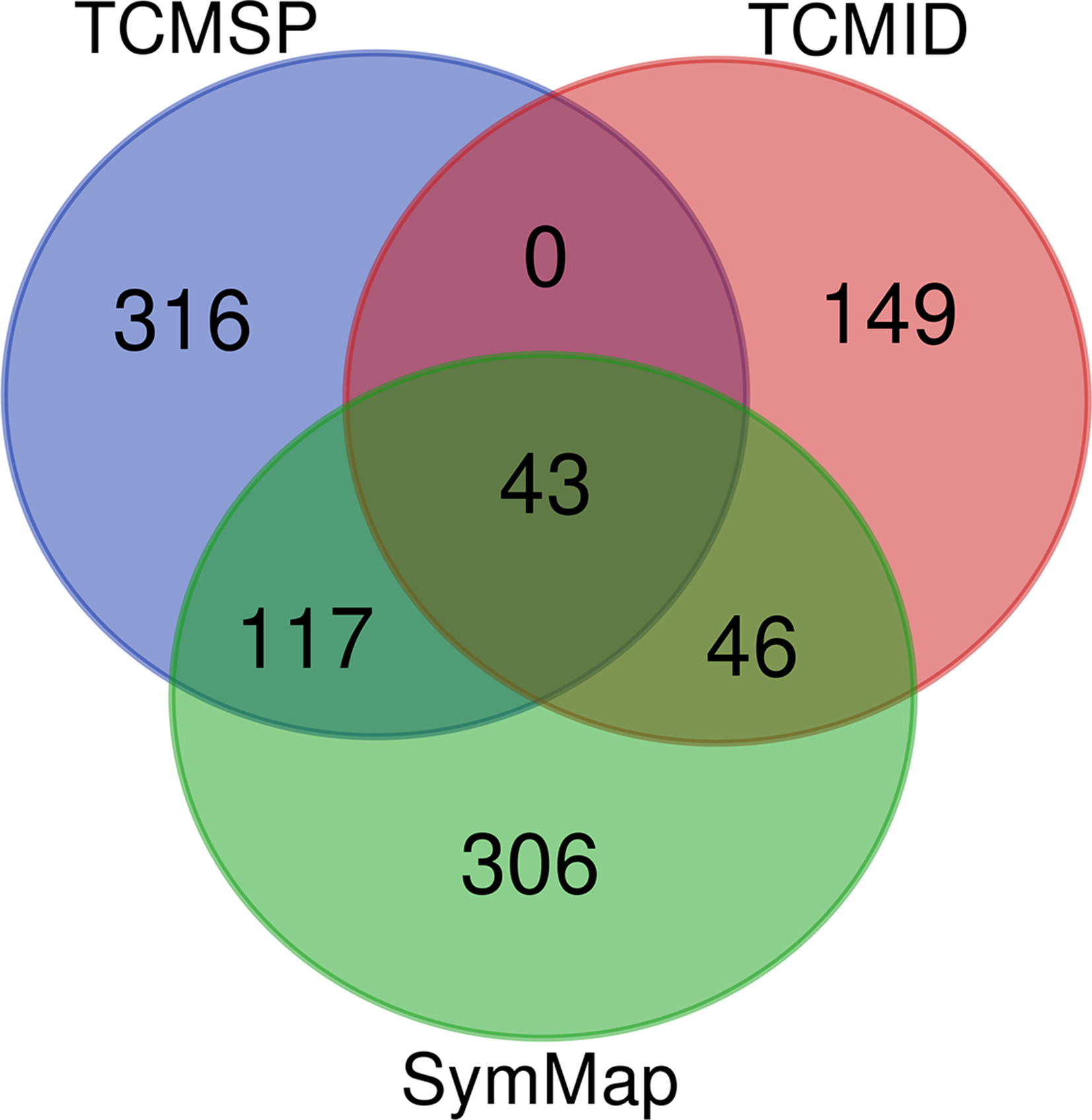
Venn diagram of components in GQD. The blue circle represents components from TCMSP, the red circle represents components from TCMID, and the green circle represents components from SymMap. A total of 974 compounds were obtained from the 3 databases, including 476 from TCMSP, 512 from SymMap, and 238 from TCMID, of which 43 compounds were shared among the 3 databases. GQD, Gegen Qinlian decoction.
Two ADME-related models, OB and DL, were used for screening and grouping, and 25 bioactive components were obtained. The GQD component–target network diagram was constructed using Cytoscape version 3.7.0, which is shown in Figure S1.
The network contains 232 nodes and 571 edges. The 232 nodes contain 25 active ingredients and 207 targets of TCM (Table S1). The degree of the node indicates the number of nodes or edges connected with the node. In this network, we selected the components of the top 5 degree rankings, which are Stigmasterol, 3'-methoxyglabridin, licocoumarone, hispaglabridin B, and gancaonin I. These components are all connected to multiple targets and are used on 148, 53, 44, 28, and 28 targets, respectively, and represent important bioactive components in GQD. This finding also proved the synergistic effect of different TCMs and components in the GQD prescription.
Construction of the GQD Protein Interaction Network and Diabetic Protein Interaction Network
Information for 207 target protein interactions was acquired from the String database. This information was imported into Cytoscape to construct a protein interaction network of GQD targets, with a total of 184 nodes and 1009 edges, as shown in Figure S2.
A total of 441 diabetes-related targets were obtained based on OMIM. The abovementioned information for 441 target protein interactions was acquired based on the String database, and a GQD target protein interaction network was constructed. The network comprised a total of 441 nodes and 2802 edges, which is shown in Figure S3.
Analysis of the Hub Targets of GQD in the Treatment of Diabetes Mellitus
To further explore the hub pharmacological mechanism of GQD in treating diabetes mellitus, 2 PPI networks were combined to obtain the intersection target of GQD and diabetes mellitus, that is, the hub target of GQD in the treatment of diabetes mellitus. We eventually obtained 24 hub targets, as shown in Table 1.
Detailed Information on the 24 Hub Targets.

According to the MCODE algorithm, 5 functional modules were obtained from the GQD protein interaction network, which are presented in Figure 3. Using BinGO for the functional enrichment analysis of the module, the results are shown in Figure S4. The main performance includes the G-protein-coupled receptor protein signaling pathway, oxidation reduction, positive regulation of muscle contraction, gamma-aminobutyric acid signaling pathway, and synaptic transmission.

GQD protein–protein interaction network module diagram. The pink node is the intersection target of the GQD protein interaction network and the diabetes protein interaction network, and the green node is the other target of GQD. GQD, Gegen Qinlian decoction.
Based on the MCODE algorithm, 6 functional modules were obtained from the diabetic protein–protein interaction network, as shown in Figure 4. The functional enrichment of the module was analyzed by BinGO, as shown in Figure S5. The main results were as follows: G-protein-coupled receptor protein signaling pathway, response to hormone stimulus, response to stress, regulation of response to stimulus, positive regulation of peptide hormone secretion, and monocarboxylic acid metabolic process. These findings are basically the same as the current pathological mechanism of diabetes mellitus. 36 -40

Diabetic protein–protein interaction network module. The pink node is the intersection target of the Gegen Qinlian decoction. protein–protein interaction network and the diabetes protein–protein interaction network, and the blue node is the other target of diabetes mellitus.
As shown in Figure 3 of the previous study, in the GQD protein–protein interaction network, the hub targets are located in modules 1 and 3, respectively. These targets are involved in G-protein-coupled receptor signaling pathways, the positive regulation of muscle contraction in the biological process. Recent studies have shown that G-protein-coupled receptor agonists can play a role in controlling diet and reducing body weight and have the function of protecting pancreatic islets. At present, many drugs based on the G-protein-coupled receptor signaling pathway have entered the clinical research stage. 36 In addition, medical experts have proved that more than 98% of patients with diabetes have weakened skeletal muscles. Blood sugar can enter muscle tissue cells so smoothly that normal peripheral blood sugar will be stable. With the reduction in muscle tissue, sugar does not go smoothly into the muscle, resulting in insulin resistance. 41 Our module enrichment analysis at the network pharmacology level showed that diabetes and diabetic complications can be treated by GQD through the regulation of the G-protein-coupled receptor signaling pathway and muscle contraction. In addition, in the GQD protein interaction network, modules 2, 4, and 5 enrichment analyses proved that in addition to the treatment of diabetes, GQD can also participate in the redox reaction, the γ-aminobutyric acid signaling pathway, and the synaptic transmission process.
As shown in Figure 4, in the diabetes protein–protein interaction network, the hub targets are located in modules 1, 2, 3, 4, and 6. The vast majority of biological processes involved in diabetes reflect the role of GQD in multiple targets, a wide range of functions but also reflects the overall characteristics of multitarget, multipathway TCM. For example, in module 1, enrichment analysis suggested that it played a role in regulating the G-protein-coupled receptor signaling pathway, which was in agreement with the mechanism of GQD in the treatment of diabetes mellitus by adjusting the G-protein-coupled receptor signaling pathway.
The topological parameters of all nodes in the GQD network and the diabetes network were calculated and Betweenness, Closeness, Degree, and Stress were selected as the indexes to assess the significance of the nodes in the network. 42 The results are presented in Table S2, Table S3, and Figure S6 (all targets are listed in Table S4). The 4 topological parameter values for most hub nodes exceeded the median. This finding suggests that these hub nodes play an important role in both the GQD–protein interaction network and the diabetes protein–protein interaction network. This result may be because the key target of GQD is also the key therapeutic target of diabetes.
We conducted an interaction analysis of the selected 24 hub targets and analyzed their major biological functions, as shown in Figure S7. These results suggest that these interactions include coexpression, prediction, colocalization, physical interactions, shared protein domains, pathway, and genetic interactions. They are mainly involved in the biological function of the cyclic guanosine monophosphate (cGMP) biosynthetic process, cGMP metabolic process, lipid transport, ribose phosphate biosynthetic process, positive regulation of the mitogen-activated protein kinase (MAPK) cascade, and regulation of MAPK activity, which are closely related to diabetes. For instance, CGMP activators have been proven to be promising new drugs for diabetes mellitus treatment and its related complications. 43 In addition, lipid metabolism and ribose phosphate biosynthesis have also been shown to be closely related to the formation and development of diabetes mellitus. 44,45
Hub Gene Verification
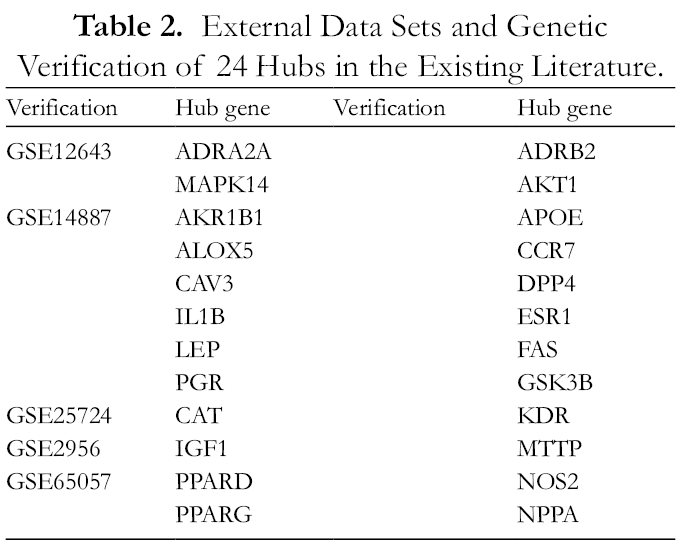
The 24 core genes were verified in the GEO database and existing literature studies. 46 -56 As shown in Table 2, 12 of these genes were differentially expressed in GSE12643, GSE14887, GSE25724, GSE2956, and GSE65057 dataset. In the existing literature, another 12 genes were extensively related to the mechanism of diabetes mellitus.
External Data Sets and Genetic Verification of 24 Hubs in the Existing Literature.
GSEA Analysis
A GSEA analysis was carried out by using the GSE25724 data set to study the metabolic pathways in which hub genes may participate in high or low expression. The results showed that there were 10 genomes enriched in high expression groups of hub genes, including “ubiquitin mediated proteolysis, small cell lung cancer, lysosome, protein export, fructose and mannose metabolism, spliceosome, biosynthesis of unsaturated fatty acids, citrate cycle TCA cycle, glycosaminoglycan biosynthesis chondroitin sulfate, and inositol phosphate metabolism.” There were 10 genomes that were enriched by low expression groups, including “primary immunodeficiency, neuroactive ligand receptor interaction, asthma, olfactory transduction, intestinal immune network for IGA production, systemic lupus erythematosus, linoleic acid metabolism, autoimmune thyroid disease, calcium signaling pathway, and allograft rejection, fructose and mannose metabolism, biosynthesis of unsaturated fatty acids, citrate cycle TCA cycle, glycosaminoglycan biosynthesis chondroitin sulfate inositol phosphate metabolism, and linoleic acid metabolism,” which were closely associated with glucose metabolism in humans, as shown in Figure S8. Therefore, hub genes may be involved in the regulation of metabolic pathways affecting the occurrence and development of diabetes.
Analysis of Molecular Docking Results
According to the calculation results of AutoDock Vina docking, 3D conformations of 24 compounds were successfully generated, as shown in Table 3. A total of 10 target protein crystal structures were downloaded from the PDB database. These crystal structures are compounded with corresponding biologically active ligands and have high resolution, which can be used for molecular docking (Table 4). According to the calculation and analysis of AutoDock Vina, the best composite results of the 3D structure of the protein with 24 chemical components and 10 hub nodes were obtained, and the corresponding binding free energy was obtained. The calculation results of the compounds and targets are shown in Table 5. The interaction of each target with the highest scoring compound is shown in Figure 5.
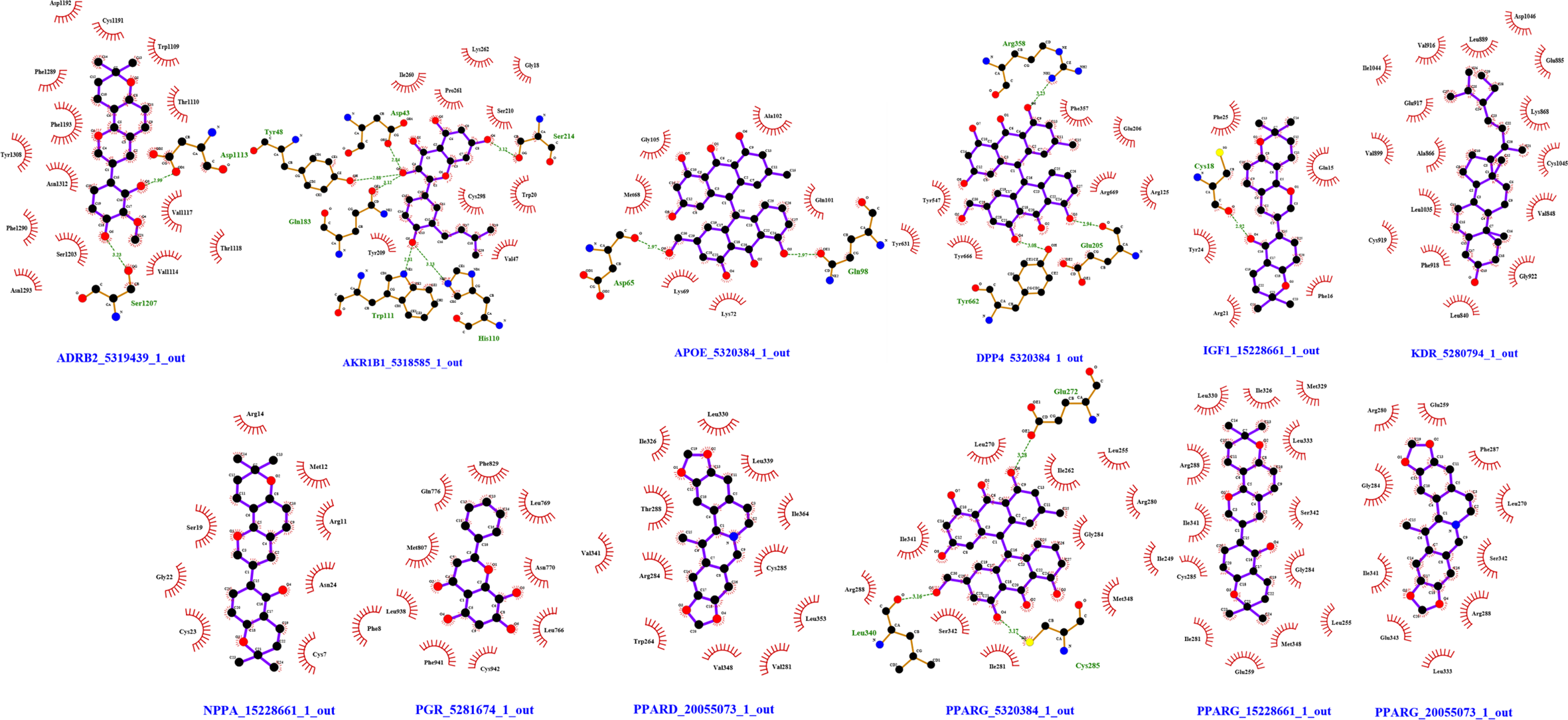
Interaction between each target and the highest-scoring compound. The green dashed lines represent hydrogen bonding and the red semicircles represent hydrophobic interactions.
Twenty-four Compounds That Successfully Generated Conformations in AutoDock Vina.

Protein Crystal Structures of 10 Hub Nodes Downloaded From the PDB Database.

Binding Free Energy of the 3-Dimensional Structure of the 24 Chemical Components and the Protein of 10 Hub Nodes.
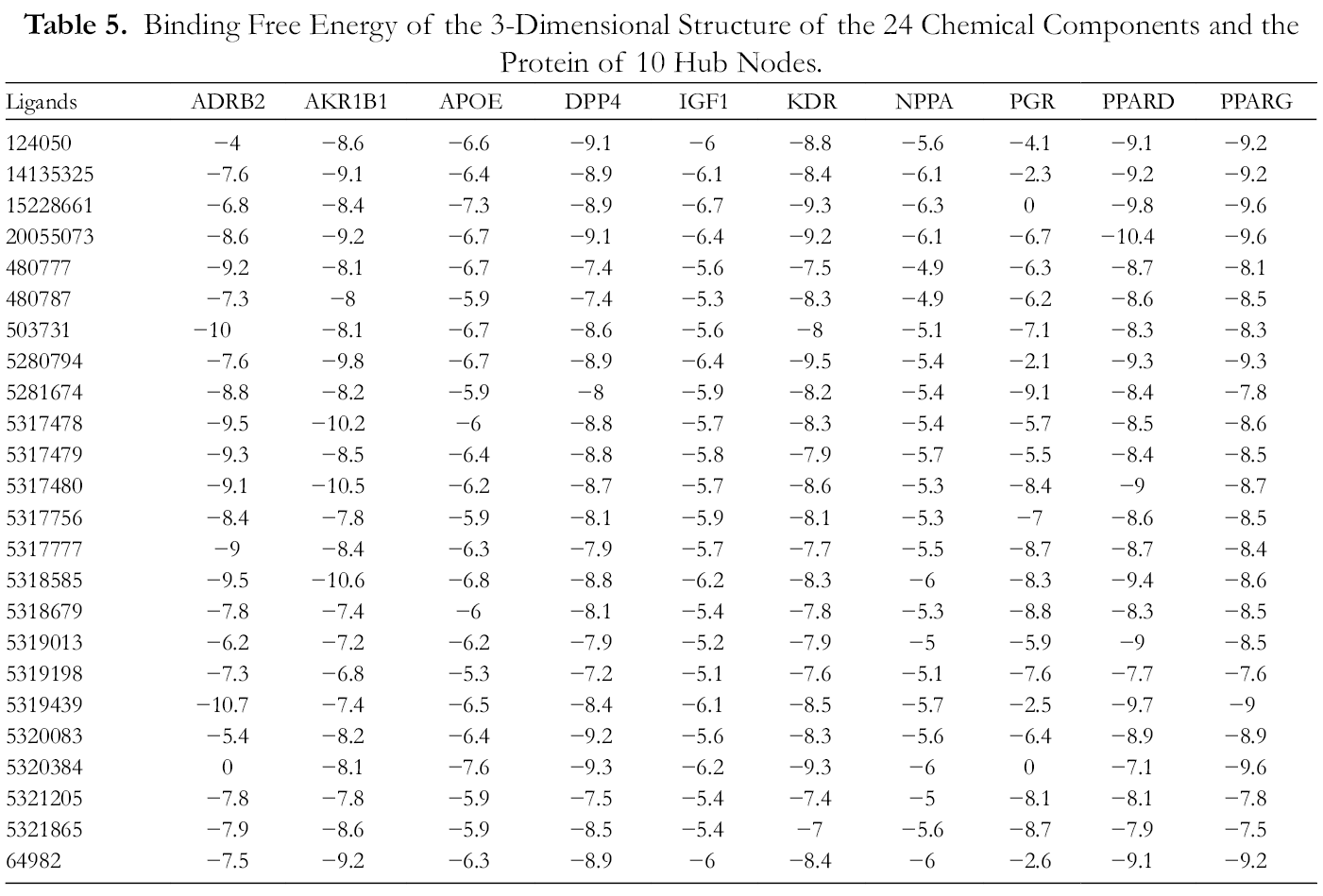
The molecular docking showed that the protein structure of the 10 hub nodes and the highest-scoring compound all formed hydrogen or hydrophobic bonds. For example, worenine and ADRB2 form 2 hydrogen bonds: Ser1207 and Asp1113, worenine and Asp1192, Cys1191, Trp1109, Thr1110, Val1117, Thr1118, Val1114, Asn1293, Ser1203, Phe1290, Asn1312, Tyr1308, Phe1193, Phe1289. Among 24 compounds, 5,7,2', 6'-tetrahydroxyflavone, the best combination with IGF1, NPPA, PPARG, magnograndiolide with PPARD, PPARG the best, gancaonin I with KDR the best, isoglycyrol the best combination of PGR, the best combination of gancaonin A and AKR1B1, the best combination of worenine and ADRB2, the best combination of glyzaglabrin and APOE, DPP4, and PPARG.
Discussion
Diabetes mellitus is a chronic disease that is very common and complex and characterized by high morbidity and mortality rates. To date, drugs have been used primarily to control the associated symptoms of this disease. In contrast, TCM not only controls symptoms but also reduces side effects due to its multitarget and multipathway mode of action. Over a thousand years of use has shown practical efficacy of GQD in diabetes, but the pharmacological mechanisms have not yet been systematically examined and fully discovered. This study uses a network pharmacology method to further investigate the effect of GQD on diabetes mellitus.
Some of the results of the experiment caught our attention: (1) In the GQD component−target network, the top 5 components cover almost all the targets in the network, which shows the universality of the effects of these components. Among the components, stigmasterol covers 24 hub targets and this compound may be the main active component of the prescription. Studies have shown that stigmasterol can inhibit the progression of diabetes. 57 (2) Clustering analysis helps us to better understand the correlation between the functions of nodes in the network. (3) The complex interaction between the 24 hub nodes reflects the characteristics of the multitarget synergy of TCM. (4) Comparing the topology parameters of the hub nodes in the 2 networks revealed that these networks are consistent. AKT1, APOE, CAT, CCR7, ESR1, FAS, GSK3B, IGF1, IL1B, KDR, LEP, MAPK14, NOS2, PGR, and PPARG have all 4 topology parameters above the median in both networks. All 4 topology parameters of NPPA and PPARD are below the median in both networks. The results of the 4 topology parameters selected in this study are consistent and can reflect the importance of the nodes in the network. The importance of the 24 selected hub targets is consistent in the 2 networks. The 4 parameter values of most of the 24 selected hub nodes were high, significantly higher than the median value. This finding shows that the 24 hub targets selected in the present study have a high accuracy. Based on the GEO database, we found that the hub genes were differentially expressed between diabetic samples and normal samples. GSEA analysis demonstrated that these genes played a role in metabolic pathways closely associated with glucose metabolism, which may be the way by which GQD participates in regulating diabetes.
Multitarget drugs for diabetes mellitus, which is a multigene complex disease, can have synergistic effects on each target. In this way, the total effect is greater than the single effect, and thus, the best therapeutic effect can be achieved. This idea coincides with the holistic concept of treatment considered by TCM theory. This paper confirmed the therapeutic effect of GQD on diabetes mellitus by adopting a network pharmacology analysis. Finally, this result was verified by molecular docking. Among them, 5,7,2',6'-tetrahydroxyflavone, magnograndiolide, gancaonin I, isoglycyrol, gancaonin A, worenine, glyzaglabrin, and 10 hub nodes produced the strongest binding effect. The compound−target interaction may be the main mechanism for the curative effect of the compound. In addition to the compounds that have the best binding effect with the 10 hub nodes, there are compounds such as lupiwighteone, isolicoflavonol, licoricone, and baicalin. Although they do not have the strongest binding to a certain target, they have a strong binding effect to each target. A good interaction has been produced, reflecting the synergistic mode of multicomponent and multitarget of TCM. Additionally, some scholars have focused on the relationship between multigene complex diseases and the TCM treatment model based on experimental tests and clinical trials. 58 Therefore, multitarget network analysis may be a new research approach to prove the effectiveness of TCM. However, some limitations still exist. First, the collection of bioactive ingredients and targets is not all-encompassing. With the continuous improvement of the separation and extraction technology of TCM and the continuous updating of the database, our experimental data will be further enriched and improved. Second, our network pharmacology studies lack animal experiments and clinical trials to validate this hypothesis. In the future, we will validate the relationship between multigene complex diseases and TCM treatment patterns in animal experiments and clinical trials.
Conclusions
In this paper, the drug target protein interaction network is combined with the disease target protein interaction network. Through the analysis of these 2 networks, the potential target of GQD in diabetes mellitus treatment was acquired. Our findings screened 24 potential therapeutic targets and found that their interactions were synergistic. These targets have been verified in the GEO database and in the literature, and these genes were involved in biological processes closely related to glucose metabolism. This study proposes an efficient and economical method for drug target recognition, which has important clinical application value.
Supplemental Material
Figure S1 - Supplemental material for Studies on the Mechanism of Gegen Qinlian Decoction in Treating Diabetes Mellitus Based on Network Pharmacology
Supplemental material, Figure S1, for Studies on the Mechanism of Gegen Qinlian Decoction in Treating Diabetes Mellitus Based on Network Pharmacology by Xiaodong Deng, Yuhua Liang, Jianmei Hu and Yuhui Yang in Natural Product Communications
Supplemental Material
Figure S2 - Supplemental material for Studies on the Mechanism of Gegen Qinlian Decoction in Treating Diabetes Mellitus Based on Network Pharmacology
Supplemental material, Figure S2, for Studies on the Mechanism of Gegen Qinlian Decoction in Treating Diabetes Mellitus Based on Network Pharmacology by Xiaodong Deng, Yuhua Liang, Jianmei Hu and Yuhui Yang in Natural Product Communications
Supplemental Material
Figure S3 - Supplemental material for Studies on the Mechanism of Gegen Qinlian Decoction in Treating Diabetes Mellitus Based on Network Pharmacology
Supplemental material, Figure S3, for Studies on the Mechanism of Gegen Qinlian Decoction in Treating Diabetes Mellitus Based on Network Pharmacology by Xiaodong Deng, Yuhua Liang, Jianmei Hu and Yuhui Yang in Natural Product Communications
Supplemental Material
Figure S4 - Supplemental material for Studies on the Mechanism of Gegen Qinlian Decoction in Treating Diabetes Mellitus Based on Network Pharmacology
Supplemental material, Figure S4, for Studies on the Mechanism of Gegen Qinlian Decoction in Treating Diabetes Mellitus Based on Network Pharmacology by Xiaodong Deng, Yuhua Liang, Jianmei Hu and Yuhui Yang in Natural Product Communications
Supplemental Material
Figure S5 - Supplemental material for Studies on the Mechanism of Gegen Qinlian Decoction in Treating Diabetes Mellitus Based on Network Pharmacology
Supplemental material, Figure S5, for Studies on the Mechanism of Gegen Qinlian Decoction in Treating Diabetes Mellitus Based on Network Pharmacology by Xiaodong Deng, Yuhua Liang, Jianmei Hu and Yuhui Yang in Natural Product Communications
Supplemental Material
Figure S6 - Supplemental material for Studies on the Mechanism of Gegen Qinlian Decoction in Treating Diabetes Mellitus Based on Network Pharmacology
Supplemental material, Figure S6, for Studies on the Mechanism of Gegen Qinlian Decoction in Treating Diabetes Mellitus Based on Network Pharmacology by Xiaodong Deng, Yuhua Liang, Jianmei Hu and Yuhui Yang in Natural Product Communications
Supplemental Material
Figure S7 - Supplemental material for Studies on the Mechanism of Gegen Qinlian Decoction in Treating Diabetes Mellitus Based on Network Pharmacology
Supplemental material, Figure S7, for Studies on the Mechanism of Gegen Qinlian Decoction in Treating Diabetes Mellitus Based on Network Pharmacology by Xiaodong Deng, Yuhua Liang, Jianmei Hu and Yuhui Yang in Natural Product Communications
Supplemental Material
Figure S8 - Supplemental material for Studies on the Mechanism of Gegen Qinlian Decoction in Treating Diabetes Mellitus Based on Network Pharmacology
Supplemental material, Figure S8, for Studies on the Mechanism of Gegen Qinlian Decoction in Treating Diabetes Mellitus Based on Network Pharmacology by Xiaodong Deng, Yuhua Liang, Jianmei Hu and Yuhui Yang in Natural Product Communications
Supplemental Material
Table S1 - Supplemental material for Studies on the Mechanism of Gegen Qinlian Decoction in Treating Diabetes Mellitus Based on Network Pharmacology
Supplemental material, Table S1, for Studies on the Mechanism of Gegen Qinlian Decoction in Treating Diabetes Mellitus Based on Network Pharmacology by Xiaodong Deng, Yuhua Liang, Jianmei Hu and Yuhui Yang in Natural Product Communications
Supplemental Material
Table S2 - Supplemental material for Studies on the Mechanism of Gegen Qinlian Decoction in Treating Diabetes Mellitus Based on Network Pharmacology
Supplemental material, Table S2, for Studies on the Mechanism of Gegen Qinlian Decoction in Treating Diabetes Mellitus Based on Network Pharmacology by Xiaodong Deng, Yuhua Liang, Jianmei Hu and Yuhui Yang in Natural Product Communications
Supplemental Material
Table S3 - Supplemental material for Studies on the Mechanism of Gegen Qinlian Decoction in Treating Diabetes Mellitus Based on Network Pharmacology
Supplemental material, Table S3, for Studies on the Mechanism of Gegen Qinlian Decoction in Treating Diabetes Mellitus Based on Network Pharmacology by Xiaodong Deng, Yuhua Liang, Jianmei Hu and Yuhui Yang in Natural Product Communications
Supplemental Material
Table S4 - Supplemental material for Studies on the Mechanism of Gegen Qinlian Decoction in Treating Diabetes Mellitus Based on Network Pharmacology
Supplemental material, Table S4, for Studies on the Mechanism of Gegen Qinlian Decoction in Treating Diabetes Mellitus Based on Network Pharmacology by Xiaodong Deng, Yuhua Liang, Jianmei Hu and Yuhui Yang in Natural Product Communications
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
