Abstract
The coronavirus disease 2019 (COVID‐19) outbreak caused by the 2019 novel coronavirus (2019-nCOV) is becoming increasingly serious. In March 2019, the Food and Drug Administration (FDA) designated remdesivir for compassionate use to treat COVID-19. Thus, the development of novel antiviral agents, antibodies, and vaccines against COVID-19 is an urgent research subject. Many laboratories and research organizations are actively investing in the development of new compounds for COVID-19. Through in silico high-throughput virtual screening, we have recently identified compounds from the compound library of Natural Products Research Laboratories (NPRL) that can bind to COVID-19 3Lpro polyprotein and block COVID-19 3Lpro activity through in silico high-throughput virtual screening. Curcuminoid derivatives (including NPRL334, NPRL339, NPRL342, NPRL346, NPRL407, NPRL415, NPRL420, NPRL472, and NPRL473) display strong binding affinity to COVID-19 3Lpro polyprotein. The binding site of curcuminoid derivatives to COVID-19 3Lpro polyprotein is the same as that of the FDA-approved human immunodeficiency virus protease inhibitor (lopinavir) to COVID-19 3Lpro polyprotein. The binding affinity of curcuminoid derivatives to COVID-19 3Lpro is stronger than that of lopinavir and curcumin. Among curcuminoid derivatives, NPRL-334 revealed the strongest binding affinity to COVID-19 3Lpro polyprotein and is speculated to have an anti-COVID-19 effect. In vitro and in vivo ongoing experiments are currently underway to confirm the present findings. This study sheds light on the drug design for COVID-19 3Lpro polyprotein. Basing on lead compound development, we provide new insights on inhibiting COVID-19 attachment to cells, reducing COVID-19 infection rate and drug side effects, and increasing therapeutic success rate.
In December 2019, several unexplained and very serious cases of pneumonia were reported in Wuhan, China. 1,2 Many governments and researchers worldwide carried out investigations. 3 -6 On February 11, 2020, the World Health Organization named the disease caused by the 2019 novel coronavirus (2019‐nCoV) as coronavirus disease 2019 (COVID‐19). 7,8 As of July 23, 2020, more than 15 1000 000 cases and 620 000 deaths have been reported around the world (https://www.cdc.gov.tw/En). Coronaviruses are a family of viruses that target the respiratory system of humans. 9,10 Previous outbreaks of coronavirus (CoV) infection include severe acute respiratory syndrome (SARS)-CoV and Middle East respiratory syndrome-CoV. 11,12 CoVs are positive-sense single-stranded ribonucleic acid (RNA) viruses containing glycoproteins on the envelope. In human hosts, CoVs cause a wide array of respiratory, body infection, gastrointestinal, neurological, and brain diseases. 13,14 A genomic analysis of 2019‐nCoV in a COVID-19 patient from Wuhan, China, revealed a nucleotide identity of 89% to bat SARS-like-CoVZXC 21% and 82% to human SARS-CoV. 15 The single-stranded RNA genome of 2019‐nCoV contains 29 891 nucleotides that encode for 9860 amino acids. 8,16 Studies showed that 2019‐nCoV invades and infects human lung cells through the receptor angiotensin-converting enzyme II (ACE2). 17,18
The following antiviral agents against 2019-nCoV may represent suitable therapy options: (1) ACE2 receptor antagonist blocks coronavirus-host interactions and attachments 19,20 ; (2) favipiravir and remdesivir inhibit RNA-dependent RNA polymerase; (3) Kaletra (lopinavir/ritonavir) suppresses main protease protein COVID-19 3CLpro polyprotein 21,22 ; (4) lysosomotropic base chloroquine phosphate disrupts intracellular trafficking and viral fusion events 23,24 ; (5) ribavirin interferes with RNA metabolism required for viral replication 25 ; (6) vaccine and immunotherapeutic agents (interferon [IFN]-α, pegylated interferon alfa-2a and -2b). 3,25 -27 On March 23, 2020, the Food and Drug Administration (FDA) designated remdesivir for compassionate use to treat COVID-19. RNA-dependent RNA polymerase inhibitors appear nonspecific, display low potency, and cause common adverse reactions in patients. 28 -30 ACE2 receptor antagonist agents have significant adverse reactions and are not advanced clinically. 31,32 However, the FDA has yet to approve remdesivir for coronavirus treatment. Nevertheless, drug repurposing of remdesivir as a promising antiviral agent against COVID-19 and clinical trials are currently underway to determine the complete efficacy of the compound in patients (clinicaltrials.gov, NCT04280705). 26,33 In addition, COVID-19 patients administered with Kaletra (lopinavir/ritonavir; a protease inhibitor) have improved health outcomes, suggesting the potential of the main protease (COVID-19 3CLpro) as an important target in COVID-19 therapy. 22,26
Curcumin (1,7-bis[4-hydroxy 3-methoxy phenyl]−1,6-heptadiene-3,5-dione) is a polyphenolic compound found in the rhizome of Curcuma longa (usually known as turmeric). Curcumin has been used as a traditional Chinese medicine and as a coloring and flavoring agent in Taiwan, China, and India, and it is increasingly recognized as a potential therapeutic agent. 34 -36 Curcumin is used to treat various disorders, including chronic diseases, inflammatory disorders, and infection, because of its several bioactivities, including antioxidant, anti-inflammatory, and immune-regulatory activities; preventive effects against cancers and diabetes; and protective effects on the liver, brain, and nervous and cardiovascular systems. 37 Curcumin can also fight against herpes simplex virus, 38 Zika virus, 39 Epstein-Barr virus, 40 human immunodeficiency virus (HIV), 41,42 and respiratory syncytial viruses, such as tobacco mosaic virus and cucumber mosaic virus. 43 -46 The antivirus activity of curcumin includes protease inhibition in HIV. 47,48 Recently, curcumin has been demonstrated to block the entry of COVID-19. 49 -51
Molecular docking techniques aim to predict the best matching binding mode of a ligand to a macromolecular partner, which plays a central role in all biological processes. 52 The present study aimed to determine whether COVID-19 3CLpro can be a major target protein with strong binding affinities to lopinavir, curcumin, and compounds from the compound library of Natural Products Research Laboratories (NPRL) by molecular docking techniques. Our results demonstrated that all curcuminoid derivatives from the NPRL compound library could form H-bonds or hydrophobic interactions with COVID-19 3CLpro residues among His3304, Cys3408, Met3428, Pro3431, Gln3452, and Glu3429. NPRL342 has the best docking performance by CDOCKER, suggesting that NPRL342 has a more stable docked structure than lopinavir and curcumin.
Materials and Methods
Collection of Data Set in Compounds of NPRL
The NPRL compound library is a collaboration between Professor Kuo-Hsiung Lee’s NPRL of the University of North Carolina and China Medical University Hospital to share these natural products and their synthesized derivatives. More than 3000 compounds have been produced by Professor Lee and the researchers in NPRL. In this study, curcumin and lopinavir (an antiretroviral protease inhibitor) were selected as positive ligand molecules from the compounds of NPRL from the in-house database. A 2-dimensional (2D)-3D conversion of images was generated using the Discovery Studio 2020 program (DS 2020), followed by energy minimization using the Smart Minimizer algorithm in the CHARMm force field (Dassault Systemes BIOVIA, Discovery Studio Modeling Environment, Release 2020, San Diego, CA, USA: Dassault Systemes, 2020). These minimized molecules were subjected to the following molecular docking experiments. An experimental design was created to discover the potential compound for binding COVID-19 3CLpro polyprotein in in silico studies. The experimental process includes sequence alignment for determining similar regions between the SARS-CoV 3CLpro sequence and the COVID-19 3CLpro polyprotein orf1ab sequence. COVID-19 3CLpro homology models were constructed, and the best homology model among all generated models was selected. Database compounds from the NPRL compound library were prepared. Ligand binding site and molecular docking were defined. The binding interaction and residue in each binding pose were compared. The top 9 ranked compounds in the database were selected for further analysis (Figure 1).

Experimental design to discover potential compounds for binding COVID-19 3CLpro polyprotein in silico study. Details in text. COVID-19, coronavirus disease 2019; NPRL, Natural Products Research Laboratories; SARS-CoV, severe acute respiratory syndrome-coronavirus.
Homology Modeling
The template structure of SARS-CoV 3CLpro and the target protein sequence of COVID-19 3CLpro polyprotein orf1ab were obtained from the Protein Data Bank (PDB code: 2ZU4) and the GenBank (Accession: YP_009724389.1), respectively. Subsequently, sequence alignment in DS 2020 was applied to align the polyprotein sequence of SARS-CoV 3CLpro with the polyprotein orf1ab sequence of COVID-19 3CLpro. The best-aligned region in COVID-19 3CLpro polyprotein orf1ab to SARS-CoV Mpro was then used to generate COVID-19 3CLpro polyprotein homology models in accordance with the Build Homology Models protocol. Finally, the qualities of generated homology models were examined with the discrete optimized potential energy (DOPE) score. The homology model with the lowest DOPE score was selected for the docking calculation.
Molecular Docking
The binding site of COVID-19 3CLpro was identified by homology comparison to this study. After energy minimization, the molecules were docked into COVID-19 3CLpro by using the CDOCKER protocol embedded in DS 2020. For each molecule, 9 docking poses were generated and ranked using the CDOCKER score (-CDOCKER interaction energy). On the basis of the CDOCKER score, from the top 9 Hits of compounds from NPRL in the in-house database, curcumin and lopinavir were selected for further binding interaction analysis.
Results
Construction of COVID-19 3CLpro Homology Models
Before the COVID-19 3CLpro homology models were constructed, COVID-19 3CLpro polyprotein orf1ab residues 3264-3569 were aligned with a template sequence of SARS-CoV 3CLpro in accordance with the sequence alignment protocol in DS 2020. Sequence alignment shows that the sequence identity and sequence similarity between SARS-CoV 3CLpro and COVID-19 3CLpro are 96% and 99%, respectively (Figure 2). On the basis of the sequence alignment results, 10 models were built using the Build Homology Models protocol in DS 2020 and ranked with the DOPE score value. The most suitable COVID-19 3CLpro homology model for docking was selected based on the lowest DOPE score (−34992.3), as shown in Figure 3A. As shown in Figure 3B, 99.3% (302/304) of residues fall in either favored or allowed regions of the Ramachandran plot, suggesting that the backbone conformation predictions from the homology model were within the normal range. With these substantial validation results, our homology model can be used for molecular docking.

Best sequence alignment area of COVID-19 3CLpro polyprotein orf1ab with the crystal structure of SARS-CoV Mpro (protien data bank code: 2ZU4). COVID-19, coronavirus disease 2019; SARS-CoV, severe acute respiratory syndrome-coronavirus.

(A) Most suitable COVID-19 3CLpro homology model obtained in our study. The protein structures are shown as solid ribbon. (B) The Ramachandran phi-psi diagram is used to calculate for the COVID-19 3CLpro homology model. COVID-19, coronavirus disease 2019.
Molecular Docking
All molecules were docked into the binding site of the COVID-19 3CLpro homology model by using the CDOCKER protocol to determine the binding affinities of curcumin and lopinavir to COVID-19 3CLpro. The top 9 Hits from the in-house database were achieved based on their CDOCKER score: NPRL342 (Figure 4), NPRL415 (Figure 5), NPRL339 (Figure 6), NPRL346 (Figure 7), NPRL407 (Figure 8), NPRL420 (Figure 9), NPRL472 (Figure 10), NPRL334 (Figure 11), and NPRL473 (Figure 12). All compounds are curcuminoid derivatives. The most suitable docking site for each compound shown in Figure 13 shows a similar binding position to that of lopinavir. The detailed docking results of the top 9 Hits (Table 1 and Figures 5 -15), lopinavir (Figure 14), and curcumin (Figure 15) with COVID-19 3CLpro are listed in Table 1. As shown in Table 1, all compounds could form H-bonds or hydrophobic interactions with residues among His3304, Cys3408, Met3428, Pro3431, Gln3452, and Glu3429 (Figures 5 -15). In addition, NPRL342 has the best docking performance by CDOCKER (Figure 4), suggesting that NPRL342 has a more stable docked structure than lopinavir (Figure 14) and curcumin (Figure 15). The 2D and 3D interaction diagrams of NPRL342 shown in Figure 4 verify the understanding of the interactions between NPRL342 and COVID-19 3CLpro. For NPRL342, multiple H-bond interactions formed with residues His3304, Phe3403, Glu3429, His3427, Gln3452, Asn3405, and Thr3453. In addition, the alkyl and aromatic ring groups participated in hydrophobic interactions with residues Cys3408, His3304, Met3428, Leu3430, and Pro3431 (Figure 4). Docking results show that NPRL342 plays a key role in H-bond and hydrophobic interactions with crucial residues in the binding site of COVID-19 3CLpro, suggesting that NPRL342 has a similar binding affinity to SARS-CoV Mpro compared with lopinavir.

NPRL342 selectively targets the COVID-19 3CLpro binding site. (A) 2D structure of NPRL342. (B) Best docking pose of NPRL342 with COVID-19 3CLpro. (C) 2D ligand interaction diagram of NPRL342 with COVID-19 3CLpro. COVID-19, coronavirus disease 2019; 2D, 2 dimensional; NPRL, Natural Products Research Laboratories.
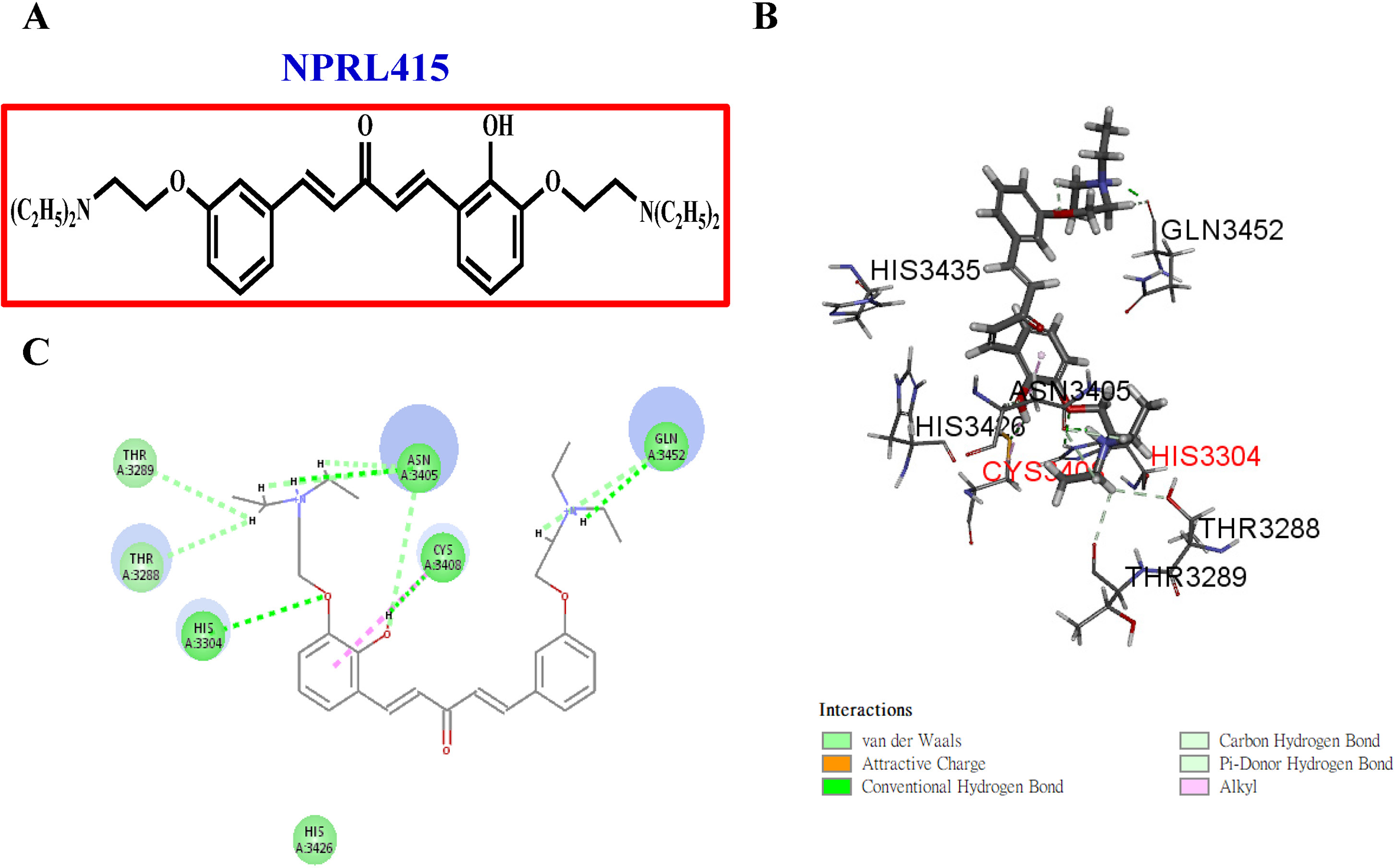
NPRL415 selectively targets the COVID-19 3CLpro binding site. (A) 2D structure of NPRL415. (B) Best docking pose of NPRL415 with COVID-19 3CLpro. (C) 2D ligand interaction diagram of NPRL415 with COVID-19 3CLpro. COVID-19, coronavirus disease 2019; 2D, 2 dimensional; NPRL, Natural Products Research Laboratories.

NPRL339 selectively targets the COVID-19 3CLpro binding site. (A) 2D structure of NPRL339. (B) Best docking pose of NPRL339 with COVID-19 3CLpro. (C) 2D ligand interaction diagram of NPRL339 with COVID-19 3CLpro. COVID-19, coronavirus disease 2019; 2D, 2 dimensional; NPRL, Natural Products Research Laboratories.

NPRL346 selectively targets the COVID-19 3CLpro binding site. (A) 2D structure of NPRL346. (B) Best docking pose of NPRL346 with COVID-19 3CLpro. (C) 2D ligand interaction diagram of NPRL346 with COVID-19 3CLpro. COVID-19, coronavirus disease 2019; 2D, 2 dimensional; NPRL, Natural Products Research Laboratories.

NPRL407 selectively targets the COVID-19 3CLpro binding site. (A) 2D structure of NPRL407. (B) Best docking pose of NPRL407 with COVID-19 3CLpro. (C) 2D ligand interaction diagram of NPRL407 with COVID-19 3CLpro. COVID-19, coronavirus disease 2019; 2D, 2 dimensional; NPRL, Natural Products Research Laboratories.
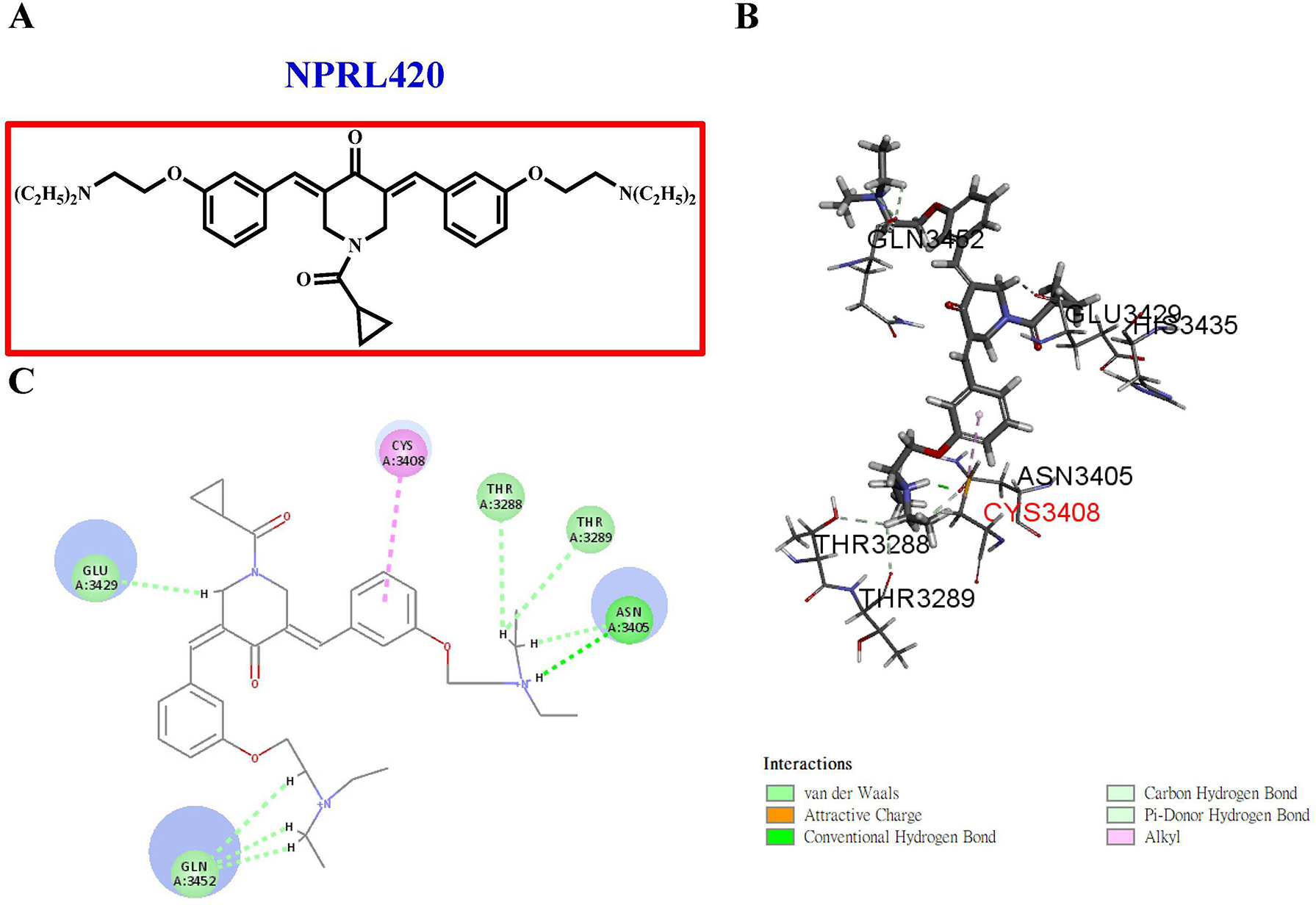
NPRL420 selectively targets the COVID-19 3CLpro binding site. (A) 2D structure of NPRL420. (B) Best docking pose of NPRL420 with COVID-19 3CLpro. (C) 2D ligand interaction diagram of NPRL420 with COVID-19 3CLpro. COVID-19, coronavirus disease 2019; 2D, 2 dimensional; NPRL, Natural Products Research Laboratories.

NPRL472 selectively targets the COVID-19 3CLpro binding site. (A) 2D structure of NPRL472. (B) Best docking pose of NPRL472 with COVID-19 3CLpro. (C) 2D ligand interaction diagram of NPRL472 with COVID-19 3CLpro. COVID-19, coronavirus disease 2019; 2D, 2 dimensional; NPRL, Natural Products Research Laboratories.

NPRL334 selectively targets the COVID-19 3CLpro binding site. (A) 2D structure of NPRL334. (B) Best docking pose of NPRL334 with COVID-19 3CLpro. (C) 2D ligand interaction diagram of NPRL334 with COVID-19 3CLpro. COVID-19, coronavirus disease 2019; 2D, 2 dimensional; NPRL, Natural Products Research Laboratories.

NPRL473 selectively targets the COVID-19 3CLpro binding site. (A) 2D structure of NPRL473. (B) Best docking pose of NPRL473 with COVID-19 3CLpro. (C) 2D ligand interaction diagram of NPRL473 with COVID-19 3CLpro. COVID-19, coronavirus disease 2019; 2D, 2 dimensional; NPRL, Natural Products Research Laboratories.

Overlap of ligand binding surfaces and the protein surface. The best binding pose for each compound is in blue. The best binding pose for lopinavir is in purple. The protein surface is in gray. NPRL, Natural Products Research Laboratories.

Lopinavir selectively targets the COVID-19 3CLpro binding site. (A) 2D structure of lopinavir. (B) Best docking pose of lopinavir with COVID-19 3CLpro. (C) 2D ligand interaction diagram of lopinavir with COVID-19 3CLpro. COVID-19, coronavirus disease 2019; 2D, 2 dimensional; NPRL, Natural Products Research Laboratories.

Curcumin selectively targets the COVID-19 3CLpro binding site. (A) 2D structure of curcumin. (B) Best docking pose of curcumin with COVID-19 3CLpro. (C) 2D ligand interaction diagram of curcumin with COVID-19 3CLpro. COVID-19, coronavirus disease 2019; 2D, 2 dimensional; NPRL, Natural Products Research Laboratories.
The Docking Results of 9 Hits From NPRL Compounds, Curcumin, and Lopinavir With COVID-19 3CLpro.

Abbreviations: COVID-19, coronavirus disease 2019; 2D, 2-dimensional; NPRL, Natural Products Research Laboratories.
The bold letters: It is the same interaction residues with curcumin.
Discussion
Since the outbreak of COVID-19 pneumonia, the number of confirmed cases and deaths has increased in various countries, which has a great impact on medical, economic, and social activities around the world. 1,2,5,8 The prevention and treatment of COVID-19 infection are important in current medical research. 53 -55 Medical research scientists around the world are continuing to work day and night. According to many private estimates from the Chinese government, the number of COVID-19-infected Indians evacuated from China is 0, which may be related to their eating habits. Curry, which is the staple food of Indians, contains a large amount of curcumin and curcuminoids. 56,57 Previous reports have indicated that curcumin and curcuminoids exhibit antiviral, antibacterial, anticancer, and regulating immune response abilities. 36,58,59 Curcumin and related supplement nanoparticles are useful agents to help treat HIV infection. 42,60,61 Antiviral effects of curcumin include (1) inhibiting the attachment of the gp-120 protein to T cells 62,63 ; (2) blocking virus protease activity 64 -67 ; (3) reducing tissue inflammation and inflammatory molecules (interleukins, tumor necrosis factor-α, nuclear factor kappa-light-chain-enhancer of activated b cells, and cyclooxygenase-2 release) 47,68 ; (4) increasing the antiviral effect of splenic IFN-γ and regulatory T cells, 47 and (5) increasing natural killer cell activation. 69 In the present study, DS 2020 software was used for the first time to screen the NPRL compound library. Our results showed that curcumin (Figure 15) and curcuminoid derivatives, including NPRL342 (Figure 4), NPRL415 (Figure 5), NPRL339 (Figure 6), NPRL346 (Figure 7), NPRL407 (Figure 8), NPRL420 (Figure 9), NPRL472 (Figure 10), NPRL334 (Figure 11), and NPRL473 (Figure 12), have strong binding abilities to COVID-19 3CLpro protease. The exact position shares the clinically used HIV protease inhibitor lopinavir (Figure 14). Recently, curcumin has been demonstrated to block the entry of COVID-19 and viral protease, as well as modulating various cellular signaling pathways. 50 Curcumin and lopinavir could form H-bonds or hydrophobic interactions in common with residues among Ala3454, Gln3452, His3304, and Pro3431 of COVID-19 3CLpro. Lopinavir and selected curcuminoids from the NPRL compound library could form H-bonds or hydrophobic interactions with residues among Ala3454, Gln3452, His3304, Leu3430, Met3428, Pro3431, Cys3408, Met3312, and Glu3429 of COVID-19 3CLpro protease (Table 1 and 5 -15). NPRL342 has the best docking performance by CDOCKER, suggesting that NPRL342 may be a more potent agent against COVID-19 3CLpro protease than lopinavir and curcumin. Further experiments will be performed to confirm the anti-COVID-19 viral activity of the potential lead compounds. Residues located on Ala3454, Gln3452, His3304, Gln3455, His3426, Pro3431, His3427, and Thr3453 of COVID-19 3CLpro are crucial positions for binding curcumin and curcuminoid derivatives from the NPRL compound library (Table 1). These residues are labeled in red. Our preliminary result demonstrated that curcumin inhibits COVID-19 RNA-dependent RNA polymerase (RdRp) activity. In addition, related experiments will be performed to confirm the anti-COVID-19 viral activity of the potential lead compounds. We look forward to discovering new and effective agents as soon as possible for providing clinical trials to reduce the infectivity of COVID-19 and treat COVID-19.
Among the many potential targets against SARS-CoV and other CoVs, protease is highly conserved. 70 Protease inhibitors can prevent virus replication and proliferation through interfering with the post-translational processing of viral polypeptides. 70,71 Lopinavir is an HIV-1 protease inhibitor capable of inhibiting SARS-CoV Mpro. 21,72,73 In Figure 2, the COVID-19 3CLpro homology model shows that the sequence identity and sequence similarity between SARS-CoV 3CLpro and COVID-19 3CLpro are 96% and 99%, respectively. Our results suggest that COVID-19 3CLpro can be used as a homologous target for screening compounds that inhibit the replication and proliferation of 2019-nCoV.
In the future, we will conduct in vitro and in vivo experiments to confirm the findings of our in silico studies. We will also use DS 2020 software and Synthia Organic Retrosynthesis Software for the structure optimization to de novo synthesize the most effective compounds for COVID-19 in our artificial-intelligence-based drug discovery platform. Related experiments will be performed to confirm the anti-COVID-19 viral activity of the potential lead compounds. We look forward to discovering new and effective agents as soon as possible for providing clinical trials to reduce the infectivity of COVID-19 and treat COVID-19 pneumonia.
Footnotes
Acknowledgments
We wish to acknowledge the work of Mr Cheng-Li Chou, Yu-Hsun Cheng, and Wain-Ling Huang (VtR Incorporated, Taipei, Taiwan) for the excellent technique and equipment support in this study.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by the project from the compound library of Natural Products Research Laboratories (NPRL) of China Medical University Hospital (CMUH) and University of North Carolina (UNC), Chapel Hill Eshelman School of Pharmacy. This work was also supported by a grant from China Medical University Hospital, Taichung, Taiwan (DMR-109-147).
