Abstract
Introduction:
Prediabetes is a prevalent condition in which early detection and lifestyle interventions can prevent or delay progression to diabetes. Artificial intelligence (AI) and machine learning (ML) offer enhanced tools for diagnosis, risk stratification, and scalable delivery of lifestyle interventions. This review synthesizes current applications of AI/ML in patients with prediabetes.
Methods:
We conducted a scoping review using PubMed, EMBASE, and Web of Science (through May 2025) to identify original studies applying AI/ML to prediabetes prediction or management. Population-level forecasting and models combining prediabetes with other conditions were excluded. Data were extracted via structured REDCap instruments and validated through secondary review. Descriptive statistics summarized findings.
Results:
Of 2072 records screened, 149 studies met criteria: 118 prediction model studies, 20 intervention studies, and 11 miscellaneous. Machine learning models primarily targeted prediction of prediabetes, progression to diabetes, diabetic complications, and glucose metrics. Overall model performance was favorable (mean C-statistic 0.81), with random forests, neural networks, and support vector machines showing better performance. Only 20 studies reported external validation, few compared ML to standard risk tools, and data/code availability was limited. Six AI-based diabetes prevention programs showed positive clinical outcomes, though randomized controlled trial (RCT) evidence was limited. Three personalized nutrition interventions showed mixed efficacy.
Conclusion:
Most AI/ML research in prediabetes focused on predictive modeling, which shows promise but limited translation to real-world settings. Artificial intelligence-based interventions may scale behavioral change support but need further evaluation versus standard care. Future efforts should prioritize external validation, assess added value over standard tools, and address barriers to integration into care.
Keywords
Introduction
Prediabetes is characterized by elevated blood glucose levels below diabetes thresholds and is diagnosed through impaired fasting glucose (100-125 mg/dL), impaired glucose tolerance (140-199 mg/dL at 2 hours post-75g glucose load), or A1C levels of 5.7% to 6.4%. Its global prevalence is projected to reach 1 billion by 2045. 1
Approximately 10% of individuals with prediabetes progress to type 2 diabetes (T2D) each year. 2 Prediabetes also poses health risks, including macrovascular complications such as stroke, myocardial infarction, heart failure, and peripheral vascular,3,4 disease. Despite its prevalence, prediabetes often remains asymptomatic (81% of adults with prediabetes are unaware of their condition), 5 leading to inadequate screening and care, 6 further compounded by underutilization of risk assessment tools and lifestyle interventions. 7 As such, there is an urgent need for improved prevention, early detection, and management strategies for prediabetes.
In recent years, the use of artificial intelligence (AI) and machine learning (ML) has expanded to chronic disease prevention and management, 8 including prediabetes. Leveraging diverse data sources, such as large electronic health records (EHR), continuous glucose monitoring (CGM), and wearable devices, these tools can predict disease onset,9,10 identify key risk factors, 11 and track progression to T2D or diabetes-related complications. By detecting patterns potentially missed by conventional tests like fasting glucose and A1C, AI can enhance early detection through glucose trend analysis, personalized risk stratification, and image-based diagnostics.12-15
Beyond predictive analytics, AI can enhance lifestyle interventions for prediabetes by offering scalable and cost-effective methods to support lifestyle changes, which may address challenges like low participation and limited access to in-person diabetes prevention programs (DPPs).16,17 These tools can emulate the personalization of human coaching through feedback loops, goal setting, behavioral tracking, and reminders.18-20 More recently, an increasing number of direct-to-consumer digital products have emerged leveraging CGM data,21-23 or photo-based meal scanning,23,24 to identify and personalize lifestyle choices.
The rapid growth of AI/ML in prediabetes underscores the need for a comprehensive review detailing its applications. As such, this scoping review aims to describe how AI/ML are being applied in the context of prediabetes.
Methods
Definitions of AI and ML
We broadly defined AI as computational systems that simulate human intelligence processes such as learning, reasoning, and decision-making. ML, a subfield of AI, was defined as the use of data-driven algorithms that identify patterns and generate predictions or classifications. Within the context of prediabetes, AI/ML applications include predictive analytics to support clinical decision-making and patient-specific risk assessment (i.e., ML models), as well as for the personalized delivery of behavioral lifestyle change support (i.e., AI-based interventions).
Study Identification and Selection Process
This review was conducted in accordance with the Preferred Reporting Items for Systematic reviews and Meta-Analyses extension for Scoping Reviews (PRISMA-ScR) guidelines. 25 As this was a scoping review and not a systematic review, our review protocol was not prospectively registered.
We conducted a systematic literature search in PubMed, Web of Science, and Embase from database inception to May 1, 2025. Search queries incorporated terms related to AI/ML and prediabetes. Studies representing original articles employing AI/ML for the clinical care of patients with prediabetes were included. Further details, including the complete search strategy, are provided in the Supplemental Methods.
Data Extraction and Validation
Data extraction was performed using REDCap electronic data capture tools hosted at Johns Hopkins University.28,29 Studies were categorized into predictive modeling, AI-based interventions, or miscellaneous. Further details, including extraction methods, data validation, and REDCap data extraction instruments, are provided in the Supplemental Methods.
Data Analysis
Descriptive statistics were used to summarize extracted data, including counts of predicted outcomes, ML techniques, performance metrics, study designs, and validation methods. Top-performing ML techniques were identified based on internal validation C-statistics or accuracy. For studies reporting prediabetes-specific metrics, model performance was aggregated by technique, with forest plots generated when sufficient data were available. Intervention-related studies were summarized using counts and frequencies of platforms, data sources, and study designs.
Results
Included Studies
Figure 1 shows the PRISMA flow diagram and study selection process. A total of 2072 records were identified through database searching: 845 from PubMed, 626 from Embase, and 601 from Web of Science. After removing duplicates, 1359 records remained for title and abstract screening. Of these, 1190 were excluded for the following reasons: not an original article (n = 270), not meeting study definitions for AI or ML (n = 579), or not relevant to prediabetes (n = 341). The remaining 169 articles underwent full-text review, of which 149 met the eligibility criteria and were included in the final review. These comprised 118 prediction model studies, 20 intervention studies, and 11 miscellaneous articles.
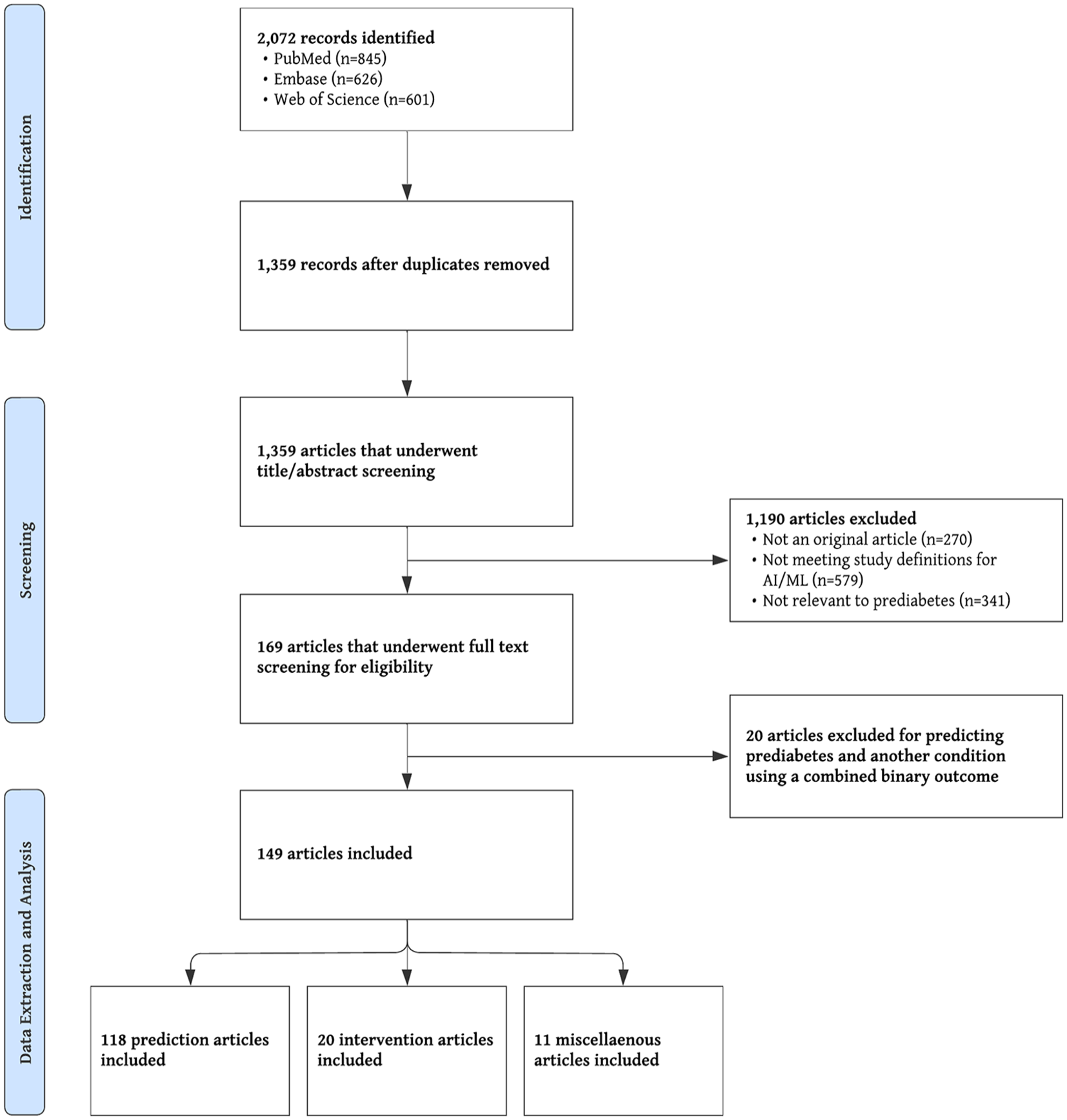
PRISMA flow diagram.
Figure 2 shows the distribution of included studies by year and application type, with most studies published within the past 5 years. Figure 3 highlights the primary areas of AI/ML use in prediabetes: lifestyle interventions and predictive analytics. Artificial intelligence-based interventions include personalized nutrition and automated DPPs, while ML-driven prediction models target transitions between glycemic states (e.g., normal glucose tolerance, prediabetes, diabetes, complications) and key metabolic metrics, including glucose levels, glycemic response to interventions, A1C trajectories, and insulin resistance.
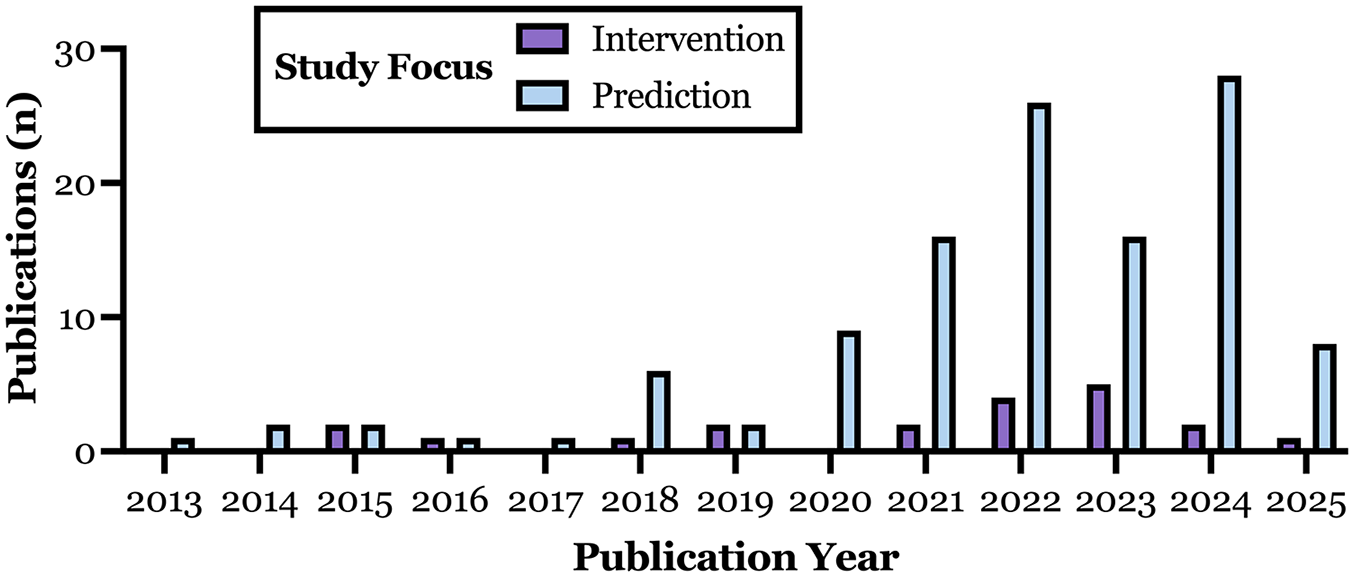
Annual number of publications describing AI-based interventions (purple) and ML-based predictive models (blue) focused on prediabetes from 2013 to 2025. Data reflect publication counts from the final dataset, which included 20 intervention studies and 118 predictive modeling studies.
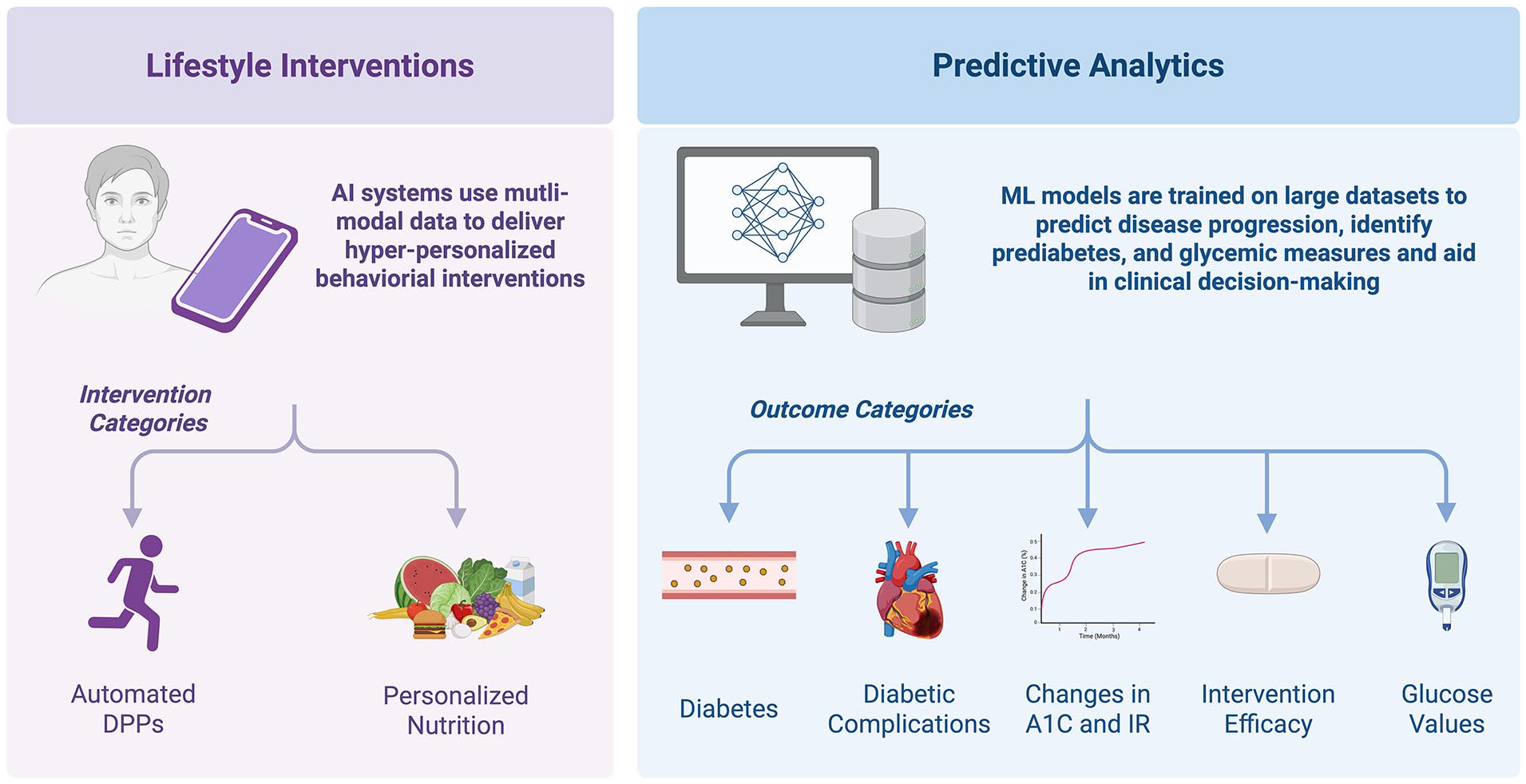
Study focus areas.
Predictive Modeling Using ML
A total of 145 ML models were developed across 118 unique studies. Table 1 categorizes these studies by predicted outcomes, with Figure 4a illustrating their distribution. As displayed in Figure 4b, most models used k-fold cross-validation (n = 71, 49.0%) or split-sample validation (n = 52, 35.9%), but only 20 models (13.8%) underwent external validation. Among the 17 models that reported both internal and external validation C-statistics, the mean internal C-statistic was 0.81 (SD: 0.08), while the mean external C-statistic was 0.77 (SD: 0.08).
Classification of Prediction Models by Outcomes.
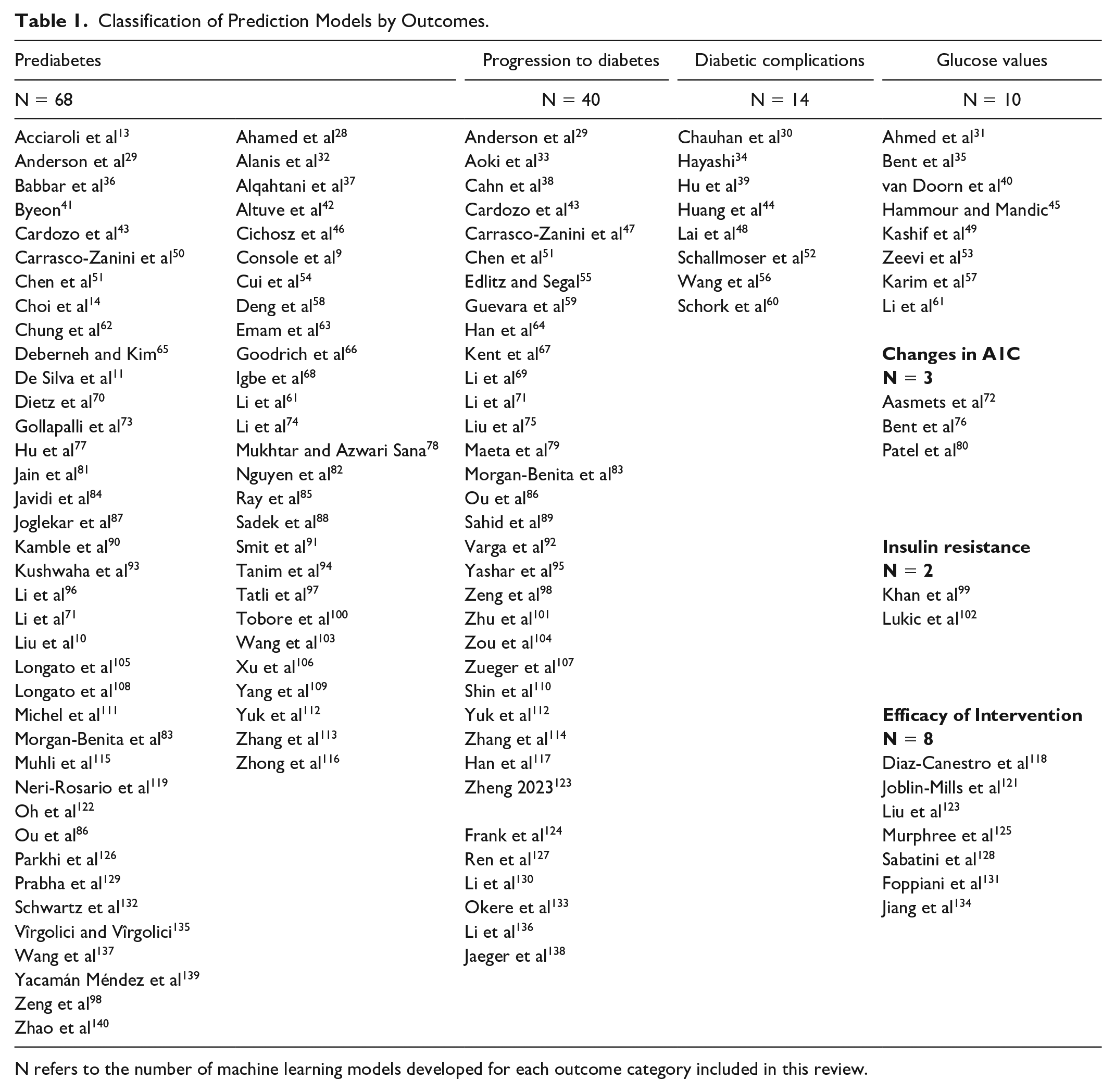
N refers to the number of machine learning models developed for each outcome category included in this review.
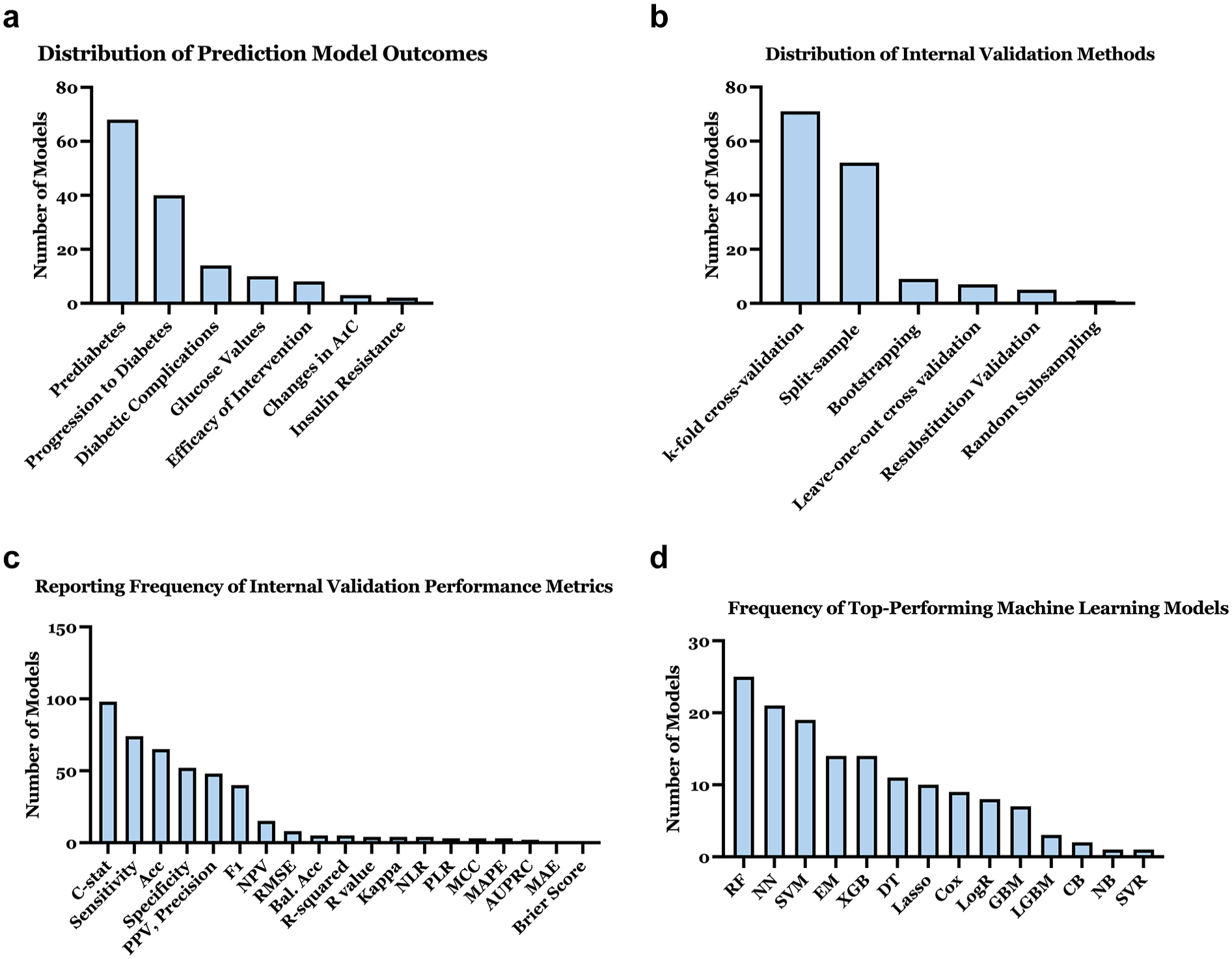
Characterization of ML-based predictive models: (a) Distribution of prediction model outcomes. (b) Distribution of internal validation methods used in models. (c) Reporting frequency of internal validation performance metrics. (d) Frequency of top-performing machine learning models.
Internal validation metrics (Figure 4c) were most frequently reported as C-statistic (n = 98, 67.6%), sensitivity (n = 74, 51.0%), and accuracy (n = 65, 44.8%). Sixty-five percent of studies developed multiple ML models using different techniques to predict the same outcome, typically to compare their relative performance; on average, three models were constructed per study. Figure 4d shows that the most common top-performing techniques were random forest (RF; n = 25, 17.2%), neural networks (NN; n = 21, 14.5%), and support vector machines (SVM; n = 19, 13.1%). Derivation datasets and ML model or code resources were available in 27.1% (n = 32) and 17.8% (n = 21) of studies, respectively.
Diagnosing or predicting prediabetes
Supplementary Table 1 summarizes 68 ML models predicting real-time prediabetes diagnosis (diagnostic models, n = 48, 70.6%), future prediabetes development (prognostic models, n = 19, 27.9%), and EHR-based prediabetes discussions (n = 1). Prediabetes was predicted as a binary outcome in 35 studies (51.5%) and as part of a multi-class outcome (e.g., normal, prediabetes, diabetes) in 32 studies (47.1%). Study designs included retrospective cohort (n = 26, 38.2%), prospective cohort (n = 16, 23.5%), and cross-sectional (n = 26, 38.2%), with sample sizes ranging from 18 to 381,077 (median = 1,461).
The most common top-performing methods were NN (n = 12), RF (n = 9), and SVM (n = 10). Internal validation C-statistics were reported in 43 studies; mean was 0.81 (SD: 0.10; Figure 5a). Diagnostic models (n = 27) demonstrated higher performance than prognostic models (n = 16), with a mean C-statistic of 0.84 (SD: 0.09) compared to 0.75 (SD: 0.10), respectively. Figures 5b and 5c show RF and decision tree (DT) performing best for prognostic models, and extreme gradient boosting (XGB) and ensemble methods (EM) for diagnostic models. External validation was reported in 10 studies (14.7%).
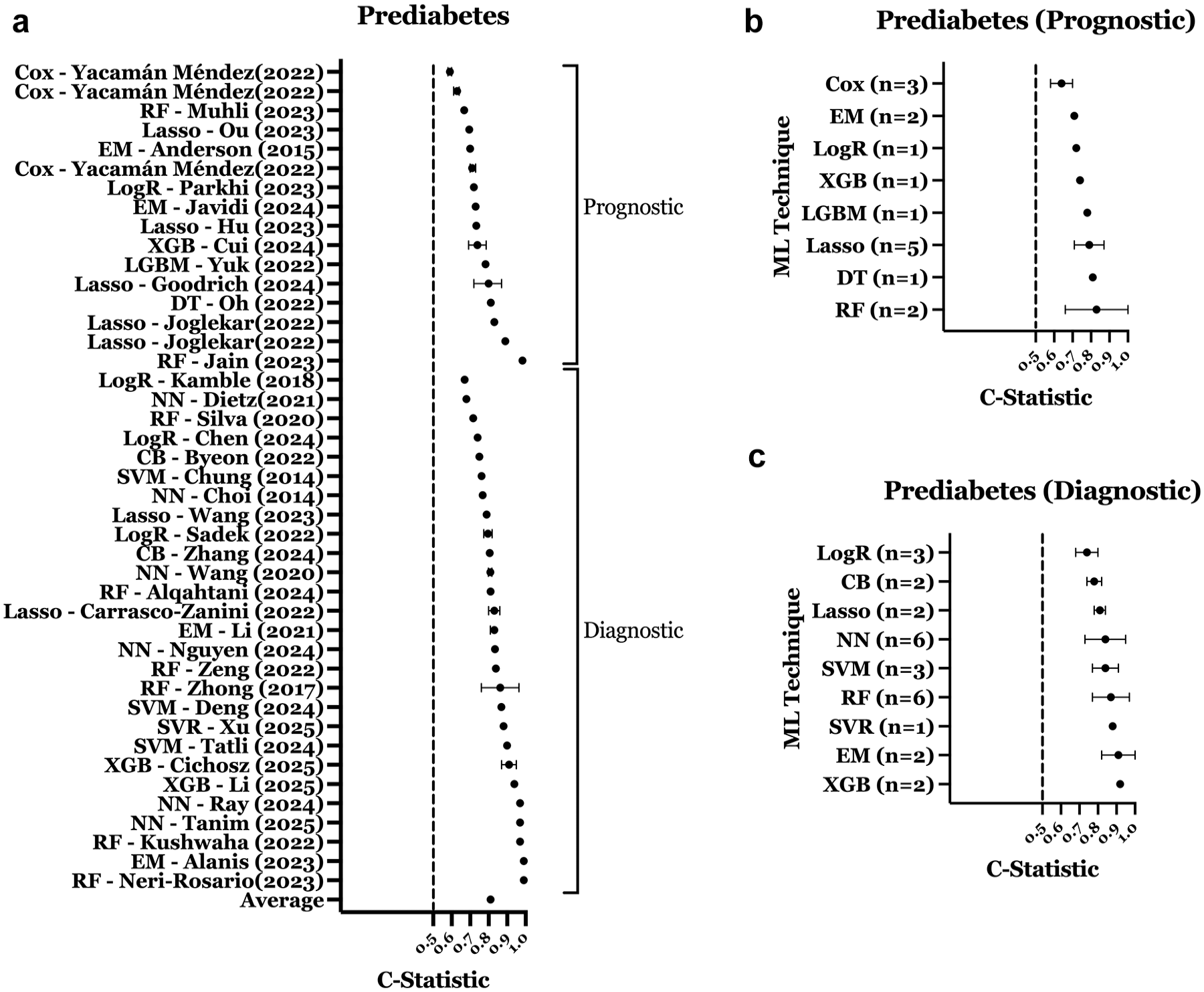
Performance of ML models predicting prediabetes. (a) C-statistic distribution for prediabetes prediction models. (b) Performance of prognostic prediabetes models by ML technique. (c) Performance of diagnostic prediabetes models by ML technique. Error bars represent confidence intervals (if reported) for panel a and standard deviation for panels b and c. All plotted values reflect model performance in prediabetes-specific cohorts or subgroups within mixed cohorts.
Most studies used ADA criteria to define prediabetes, with some applying World Health Organization (WHO) criteria.17,25 Common predictors included age, body mass index (BMI), sex, waist circumference, fasting plasma glucose, and A1C. Unconventional predictors included microRNAs, 87 tongue images and aquaphotomics, 71 biochemical metabolites, 83 and gut microbiome profiling. 119
Diagnosing or predicting progression to diabetes
Supplemental Table 2 summarizes 40 models predicting diabetes progression in patients with prediabetes. Most were prognostic (n = 32), with the rest being diagnostic (n = 8). Study designs included retrospective cohort (n = 25, 62.5%), prospective cohort (n = 7, 17.5%), cross-sectional (n = 5, 12.5%), and randomized clinical trial (RCT; n = 3, 7.5%). Derivation cohort sizes ranged from 80 to 1.89 million (median = 2615). Models were derived from prediabetes-only cohorts (n = 22, 55.0%) or mixed cohorts (n = 18, 45.0%). Overall mean C-statistic in studies reporting performance in prediabetes-specific populations was 0.81 (SD: 0.08), with diagnostic models performing better than prognostic models (Figure 6a). The most common top-performing ML techniques were RF (n = 7), XGB (n = 7), SVM (n = 6), and gradient boosting machine (GBM; n = 5). Figure 6b shows RF and LogR models achieving the highest C-statistics for this category. External validation was reported for 7 models (17.5%).
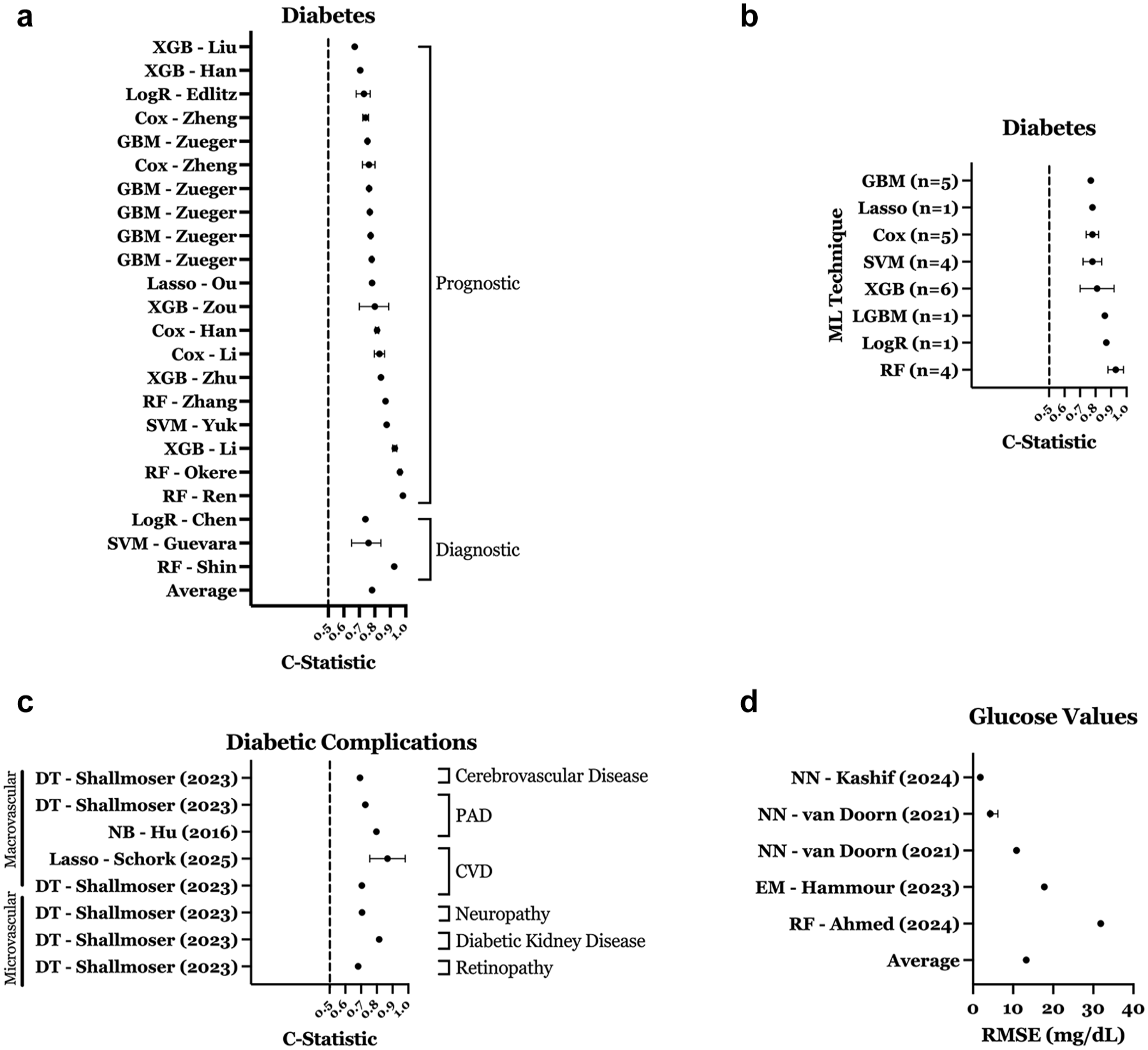
Performance of ML models predicting diabetes, diabetic complications, and glucose values. (a) C-statistic distribution for diabetes prediction models. (b) Performance of diabetes models by ML technique. (c) C-statistic distribution for diabetic complication models predicting microvascular and macrovascular outcomes. (d) RMSE distribution for glucose value prediction models. Error bars represent confidence intervals (if reported) for panels a, c, and d, and standard deviation for panels b. All plotted values reflect model performance in prediabetes-specific cohorts or subgroups within mixed cohorts.
Frequent predictors included triglycerides, age, high-density lipoprotein (HDL) cholesterol, and A1C. Some studies used uncommon features in their ML models, such as pectoral muscle mass, 95 microbiota analysis, 98 Raman spectroscopy on the skin, 59 plasma proteins, 101 high-performance liquid chromatography, 101 and Ultra-Performance Liquid Chromatography. 101
Predicting diabetic complications
Supplementary Table 3 summarizes 14 ML models predicting diabetic complications in patients with prediabetes. Common top-performing ML techniques included DT (n = 7) and NN (n = 2). Complications predicted were microvascular (neuropathy [n = 4], diabetic kidney disease [n = 3], retinopathy [n = 1]) and macrovascular (peripheral artery disease [n = 3], cerebrovascular disease, cardiovascular disease, heart failure [each n = 1]). As displayed in Figure 6c, the mean C-statistic for seven models was 0.79 (SD: 0.1), with it higher in microvascular (0.81, SD: 0.1) versus macrovascular models (0.77, SD: 0.1).
Predicting glucose values, glycemia, and insulin resistance
Supplementary Table 4 summarizes 23 ML models glucose values (n = 10), intervention efficacy (n = 8), A1C changes (n = 3), and insulin resistance (n = 2). Most used prospective cohort designs (n = 15, 65.2%) or RCTs (n = 4, 17.4%). Cohort sizes ranged from 1 to 12,147 (median = 38).
The most common top-performing techniques for predicting glucose values were NN (n = 4) and RF (n = 2). Among models reporting prediabetes-specific root mean square error (RMSE), the mean was 13.3 mg/dL (SD: 10; Figure 6d). Prediction intervals varied: 2-hour postprandial glucose (RMSE: 31.87 mg/dL; 48 R = 0.68), 53 5-minute, 35 15-minute, and 60-minute, 40 and 3-hour interstitial glucose (R² = 0.99) using physiological signals. 49 One study evaluated an in-ear photoplethysmography-based monitor for glucose prediction. 45
Eight ML models predicted the efficacy of interventions in patients with prediabetes, including metformin response, 125 exercise-induced changes in insulin sensitivity, 123 changes in insulin resistance and fasting insulin following high-intensity interval training, 118 and metabolic responses to whey protein intake. 121 Three studies predicted A1C changes using noninvasive wearables for activity patterns, 80 physiological metrics (skin temperature, heart rate), 76 and gut microbiome data. 72 Two models predicted insulin sensitivity/resistance: RF for insulin resistance in obese Hispanic adolescents 99 and NN for hyperinsulinemia in school-age adolescents. 102
AI/ML-Based Interventions
Interventions directed toward patients with diabetes fell into two broader categories: automated DPPs that aim to replicate human coaching for behavior change (e.g., weight loss, increased physical activity) using AI; and personalized nutrition platforms that use AI to generate individualized dietary guidance. Table 2 presents nine AI-based interventions: six automated DPPs and three personalized nutrition programs. These interventions utilize various AI methodologies, including reinforcement learning to adapt interventions in real time, natural language processing for interactive, conversational support, and deep learning (e.g., recurrent neural network and long-short-term memory network) to predict future blood glucose levels and tailor nutritional recommendations.
Identified AI-Based Interventions.
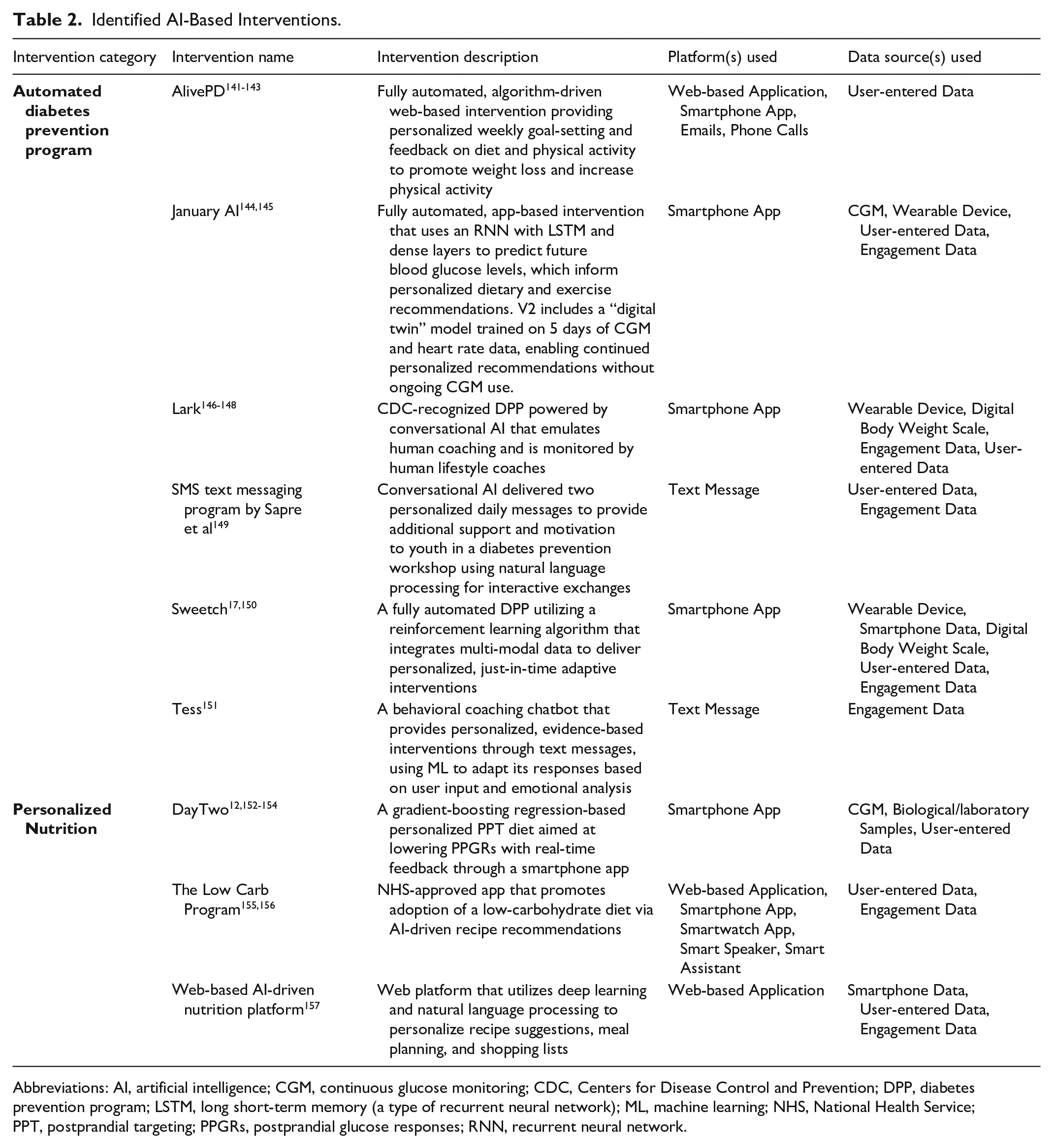
Abbreviations: AI, artificial intelligence; CGM, continuous glucose monitoring; CDC, Centers for Disease Control and Prevention; DPP, diabetes prevention program; LSTM, long short-term memory (a type of recurrent neural network); ML, machine learning; NHS, National Health Service; PPT, postprandial targeting; PPGRs, postprandial glucose responses; RNN, recurrent neural network.
Platforms for diabetes prevention (Figure 7a) primarily included smartphone apps, followed by web applications and text messaging. Less common platforms were emails, phone calls, and smart devices. Most interventions used multimodal data (Figure 7b), with data sources mainly being user-entered data (e.g., food or activity logs or health goals manually inputted by users), engagement data (e.g., app usage patterns used to optimize behavioral nudges through reinforcement learning), wearables, CGM, and body weight scales. Occasionally, smartphone data and biological samples were incorporated.
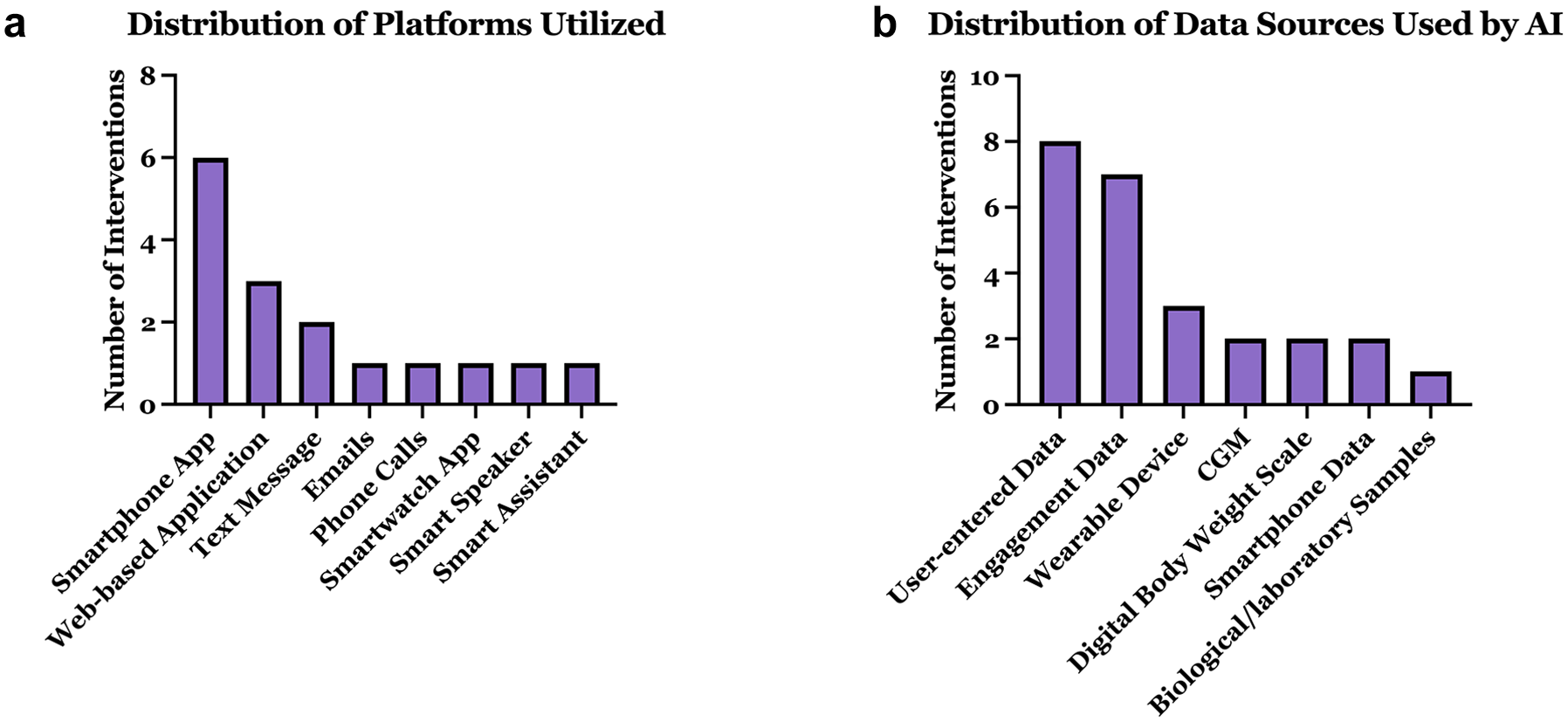
Characterization of AI-based interventions: (a) distribution of platforms utilized and (b) distribution of data sources used by AI.
Supplementary Tables 5 and 6 summarize studies evaluating the performance of nine interventions in cohorts that included, at least partially, patients with prediabetes. Of the 19 studies identified, three were published RCT protocols. Because two of these protocols had completed RCTs with published results, the protocol papers were not included in the data extraction tables; instead, only the publications presenting the corresponding results were included.141,153
Automated DPPs
Supplementary Table 5 summarizes 11 publications (10 clinical studies) evaluating six automated DPPs. Most used prospective (n = 4) or retrospective cohorts (n = 4), with only two RCTs—one for Sweetch and one for Alive-PD. The comparator arms differed: Sweetch was tested against a human coach-based DPP, 17 while Alive-PD was compared to usual care. 142 Study phases were evenly split across pilot, postimplementation, and full-scale evaluations. Average study duration was 7 months (most lasting 12 months), with cohorts ranging from 13 to 16 327 participants, primarily prediabetes-only, except January AI, which included mixed glucose-control populations.144,145 Two chatbot interventions targeted youth.149,151
Automated DPPs consistently improved clinical outcomes, including glycemic control, weight management, and physical activity. January AI significantly improved glucose levels, glycemic variability, body weight (−2.5 lbs), dietary intake (calories, carbohydrates, sugars), and physical activity (+73 min/day), with fewer hyperglycemic episodes at 28 days. 144 A follow-up retrospective study of January V2—an updated version of the intervention that used a 5-day CGM and heart rate-based training period to generate a digital twin simulation model—reported greater time-in-range and weight loss of 4 lbs over ~1 month among users with prediabetes, with more favorable results in more engaged users. 145
Lark DPP demonstrated sustained 12-month weight loss (−4.96%), with effectiveness consistent across rural, urban, and health care shortage areas.146-148 Alive-PD showed greater reductions than usual care in A1C (−0.26%), fasting glucose, refined carbohydrate intake, and increased aerobic activity (+1.21 days/week) at 6 months in an RCT.141-143 Sweetch pilot results (12 weeks) indicated high usability, retention, weight loss (−1.6 kg), improved physical activity (+2.8 MET-hours/week), and reduced A1C (−0.1%),146,147 a larger Sweetch RCT has concluded with published results not yet available. 17
Personalized nutrition
Supplementary Table 6 summarizes seven publications evaluating three personalized nutrition interventions across five clinical studies (three full-scale, two pilot/feasibility). Designs included prospective cohorts (n = 3) and RCTs (n = 2, both of which tested the DayTwo intervention). Study durations were typically 12 months, except one 2-month trial. 157 Cohorts ranged from 45 to 269 participants (average: 153), featuring mostly mixed populations except one prediabetes-only RCT. 12
The DayTwo intervention, applying an ML model developed by Zeevi et al 53 predicting postprandial glucose response (PPGR), significantly reduced PPGR and improved microbiome alpha-diversity compared to a Mediterranean diet in one RCT. 12 However, another RCT found no difference in weight loss, glycemic variability, or A1C change compared to standard calorie restriction. 154 In addition, the web-based AI-driven nutrition platform evaluated by Bul et al 157 achieved significant short-term weight loss (−4.5 kg at 8 weeks), though participant engagement and usability were limited. Finally, the Low Carb Program, assessed in two prospective cohorts, significantly improved weight (−2.82 kg) and A1C (−2.35 mmol/mol) at 12 months, with outcomes closely linked to program completion.155,156
Two interventions reported A1C outcomes: the Low Carb Program consistently reduced A1C,155,156 whereas DayTwo showed mixed results (−0.16% vs. no significant change) across two separate RCTs.12,158 Weight-loss results varied similarly: the Low Carb Program demonstrated meaningful weight loss,155,156 DayTwo did not differ significantly from controls, 154 and the web-based AI platform showed substantial short-term weight loss. 157
Miscellaneous Studies
Supplemental Table 7 summarizes the 11 articles that were classified as Miscellaneous. Studies employed diverse methodologies, such as latent Markov models and cluster analysis, to classify individuals into risk categories based on obesity indices, psychosocial factors, metabolic phenotypes, and risk groups, while AI-based techniques were used to identify metabolic biomarkers, clinical predictors, and retinal vascular features, to enhance understanding of prediabetes and its implications for diabetes and cardiovascular risk.
Discussion
Most AI/ML applications identified in this review focused on predictive modeling. In contrast, relatively few studies evaluated AI-based lifestyle interventions, despite growing consumer and commercial interest in digital approaches for diabetes prevention. 159 However, clinical integration remains limited in both areas. Predictive models often lacked external validation and data/code accessibility, limiting their generalizability and clinical utility. Meanwhile, AI-driven lifestyle interventions were hindered by a lack of rigorous evaluations, with only a few studies directly comparing them to standard care. Together, these gaps suggest that while AI is creating promising tools for prediabetes prediction and management, their integration into routine clinical care is still in its early stages.
Across the 145 predictive models reviewed, outcome selection was skewed toward glycemic-state transitions—normoglycemia to prediabetes (n = 68) or prediabetes to diabetes (n = 40)—while fewer studies predicted continuous metrics such as glucose values, A1C, or insulin resistance. This emphasis on disease progression and diagnosis is likely more clinically actionable, as it aligns with established diagnostic thresholds used in treatment decisions and risk-based screening. In contrast, predicting surrogate markers such as HOMA-IR may have limited clinical utility, as these metrics are not direct endpoints. For example, Khan et al 99 used a breath-based random forest model to predict HOMA-IR in adolescents with prediabetes, but its relevance remains unclear without evidence of impact on diagnosis, monitoring, or intervention decisions.
One promising application of ML to improve clinical care in prediabetes is through more personalized and efficient risk screening. This review identified 40 models that predict progression of diabetes in patients with prediabetes. When embedded into EHRs, such models could operate behind the scenes to flag at-risk individuals based on routinely collected data—helping to close screening gaps (only 50%-60% of U.S. adults receive recommended A1C screening) 160 without adding burden to clinician workflows. By facilitating earlier diagnosis and timely intervention, these tools have the potential to improve patient outcomes and reduce progression of diabetes. In addition, ML algorithms predicting patient-specific health metrics are also being applied to tailor interventions; for instance, Zeevi et al 53 developed a model predicting postprandial glucose levels based on meal content, activity, CGM data, microbiome, and lifestyle habits, demonstrating benefits over the Mediterranean diet in one RCT.12,154
Overall, diagnostic models in both the prediabetes and diabetes categories demonstrated better C-statistic performance than their prognostic counterparts, likely due to stronger contemporaneous associations between predictors and disease status. Commonly top-performing ML techniques—RF, NN, SVM, Lasso, DTs, LightGBM, and XGB—align with findings from diabetes and chronic disease prediction literature.161-164
In our review, we observed that less than 15% of studies performed external validation on their ML models. Several factors may explain this shortfall: (1) the field is still nascent, and many research groups remain in an algorithm-development phase; (2) truly independent, well-phenotyped prediabetes cohorts are scarce, partly because prediabetes is often undocumented in routine care; (3) privacy regulations and fragmented EHR infrastructures complicate cross-institutional data sharing; and (4) many journals do not require external validation for publication. Without testing in diverse populations, even technically strong models risk overfitting to local practice patterns and will likely fail to gain regulatory approval given the Food and Drug Administration (FDA)’s requirement for external validation within its regulatory framework for AI/ML-based medical devices. 165
In prediction-focused studies, we observed considerable variability in the reporting of performance metrics, with over 20 distinct metrics used across studies. This heterogeneity hampers cross-study comparisons and limits benchmarking of model performance in clinically meaningful ways. The TRIPOD-AI statement provides guidance on reporting standard metrics for ML models predicting binary outcomes, including AUC, sensitivity, specificity, positive predictive value (PPV), and negative predictive value (NPV). 166 In the context of prediabetes diagnosis, where disease prevalence is high, concerns about overstated performance due to class imbalance are reduced. Although considered optional, positive and negative likelihood ratios can offer valuable diagnostic insight. Calibration metrics (e.g., Brier scores), which assess the agreement between predicted probabilities and observed outcomes, are often underreported but strongly recommended.
The selection of performance metrics should align with the model’s intended use. Diagnostic models may prioritize sensitivity to reduce false negatives, particularly in screening contexts, while specificity may be emphasized when false positives carry risks of overtreatment or unnecessary testing. Prognostic models, aimed at forecasting disease progression, benefit from metrics assessing both discrimination (e.g., C-statistic) and calibration for effective risk stratification. Models predicting outcomes along the prediabetes continuum should also be benchmarked against existing clinical tools to clarify their added value—an approach largely absent from the literature. In rare cases where this has been done, ML models demonstrated improved reclassification over A1C and outperformed tools like the Finnish Diabetes Risk Score, highlighting the importance of direct comparisons for clinical relevance.55,130 The Net Reclassification Index (NRI) can further quantify these improvements over reference models. 167
Beyond the lack of external validation and benchmarking, many ML models face integration barriers due to reliance on impractical predictors—such as tongue images, gut microbiome data, or mass spectrometry—that are not routinely collected in clinical care. 168 This limits their real-world applicability. For ML tools to be useful in clinical workflow, they must be practical and operate in harmony with existing approaches. Models should therefore prioritize accessible inputs, rather than introducing new burdens.
Furthermore, the black-box nature of many ML models has been cited as a barrier to clinical adoption, particularly when providers cannot understand how predictions are made.169,170 This challenge is compounded by the frequent lack of publicly available code, data, or trained models, which limits independent validation and prevents other researchers from replicating or building upon the work. For tools intended to support frontline decision-making, models must be understandable and interpretable. Yet this review highlights a lack of model transparency, potentially limiting clinician trust and real-world adoption.
This review also highlights several findings regarding AI-based interventions for prediabetes. We found that AI interventions are understudied as most are evaluated in observational studies without comparison to standard care (i.e., the human-coach-based DPP). 171 The lack of RCT data in this space may be driven by underlying regulatory procedures. Most AI-driven lifestyle interventions fall outside FDA oversight, as the agency generally exempts general wellness tools from regulation. 172 Instead, these programs pursue validation through Centers for Disease Control and Prevention (CDC) recognition, which relies on observational data and a per-protocol evaluation in program completers—an approach exemplified by Lark’s recognition based on retrospective data alone. 159 The broader literature has highlighted the need for more RCTs to assess AI tools in clinical care, 173 and comparisons to human-delivered interventions are especially important given that many AI systems, including the interventions in this review, aim to replicate functions traditionally performed by health professionals. However, a large-scale RCT evaluating Sweetch, which concluded in late 2024 (results not yet available), is the first to compare an AI-based intervention directly to the standard-of-care DPP. Demonstrating comparable performance between AI and human coaching could unlock scalable delivery of lifestyle interventions and help address the widespread underuse of preventive care in prediabetes. 174
While these interventions are generally well-received and show potential for promoting lifestyle changes, their effectiveness depends on user engagement, which is inconsistently measured. 175 For instance, the Lark DPP study defined engagement via lesson completion, coaching exchanges, and weigh-in frequency—each independently linked to ≥5% weight loss—whereas the January V2 study used food logging frequency to identify “power users,” who achieved greater improvements in time in range and modestly greater weight loss.145,146 This engagement-outcome association is analogous to that observed in human-coach DPPs, 176 suggesting that future work should focus on identifying strategies that reliably sustain user engagement in AI-based interventions.
In addition, our findings reveal that AI-based interventions for diabetes prevention are integrating diverse data inputs like CGM, photo-based meal analysis, and data from wearables to provide hyper-personalized recommendations. This trend fits well with the growing availability of over-the-counter CGMs, which are making real-time glucose monitoring more accessible to individuals with prediabetes as well as healthy adults seeking wellness insights. 177 As these devices become more widely used in this population, they open new opportunities for AI tools to integrate dynamic glucose data into personalized feedback loops. Interventions like January AI have already begun leveraging CGM data to guide individualized dietary and activity recommendations.
The most common delivery modalities for AI-based interventions were mobile applications, web-based platforms, and text messaging. Each offers distinct advantages and limitations. Mobile apps may be the most promising due to their portability, bidirectional communication capabilities, and potential for real-time, context-aware feedback. Web platforms support structured content delivery and are conducive to in-depth education but may be less accessible for users seeking on-the-go support. Text messaging is broadly accessible and low-cost yet limited in interactivity and capacity for personalized engagement.
Integration of AI-based tools for diabetes prevention faces several implementation barriers. First, the lack of rigorous effectiveness data compared to standard-of-care lifestyle programs raises questions about whether these tools should be adopted at scale or if resources would be better spent expanding the CDC’s established National DPP. Second, although asynchronous delivery can lower geographic barriers, real-world uptake still depends on reliable internet, smartphone access, and digital literacy—factors that can vary widely by age, income, and rurality. Consequently, future work must conduct head-to-head trials with appropriate endpoints linked to diabetes incidence (e.g., weight loss, A1C reduction) and pragmatic studies of access, engagement, and adherence to determine where and for whom AI interventions add the greatest value.
Strengths of our review include a comprehensive multidatabase literature search, clear inclusion and exclusion criteria that promoted the relevance of included studies, and a rigorous multireviewer process that reduced bias and errors in data extraction. Structured data extraction further supported consistency and reliability in reporting. However, our study also had limitations. Relying on published studies may have introduced publication bias, as negative results are often underreported, and some performance data may remain in company white papers rather than peer-reviewed literature. Gray literature and conference proceedings were not reviewed, and no non-English articles were identified or included. In addition, by focusing only on top-performing ML models within studies, we may have overlooked potentially useful algorithms.
Future research should focus on strengthening the clinical relevance and implementation potential of AI/ML tools for prediabetes. For predictive models, priorities include conducting external validation in diverse populations, improving model transparency through open code and interpretable features, and benchmarking performance against existing risk scores to clarify added value. A systematic synthesis of top-performing predictors across outcome categories could enhance interpretability and guide the selection of clinically practical variables that consistently contribute to model performance. For AI-based interventions, randomized trials comparing digital programs to human-coach DPPs are needed, alongside implementation studies that address disparities in access, digital literacy, and sustained engagement.
Conclusions
This scoping review provides an overview of the applications of AI/ML for prediabetes. The majority of research focuses on predictive analytics, with fewer studies evaluating AI-based interventions. While ML-based prediction models demonstrated good discrimination across various outcomes, their clinical utility and practicality remain largely unvalidated. For interventions, AI-driven lifestyle programs show promising results; however, their real-world effectiveness remains uncertain.
Supplemental Material
sj-docx-1-dst-10.1177_19322968251351995 – Supplemental material for Applications of Artificial Intelligence and Machine Learning in Prediabetes: A Scoping Review
Supplemental material, sj-docx-1-dst-10.1177_19322968251351995 for Applications of Artificial Intelligence and Machine Learning in Prediabetes: A Scoping Review by Benjamin Lalani, Rohan Herur, Daniel Zade, Grace Collins, Devin M. Dishong, Setu Mehta, Jalene Shim, Yllka Valdez and Nestoras Mathioudakis in Journal of Diabetes Science and Technology
Supplemental Material
sj-xlsx-2-dst-10.1177_19322968251351995 – Supplemental material for Applications of Artificial Intelligence and Machine Learning in Prediabetes: A Scoping Review
Supplemental material, sj-xlsx-2-dst-10.1177_19322968251351995 for Applications of Artificial Intelligence and Machine Learning in Prediabetes: A Scoping Review by Benjamin Lalani, Rohan Herur, Daniel Zade, Grace Collins, Devin M. Dishong, Setu Mehta, Jalene Shim, Yllka Valdez and Nestoras Mathioudakis in Journal of Diabetes Science and Technology
Supplemental Material
sj-xlsx-3-dst-10.1177_19322968251351995 – Supplemental material for Applications of Artificial Intelligence and Machine Learning in Prediabetes: A Scoping Review
Supplemental material, sj-xlsx-3-dst-10.1177_19322968251351995 for Applications of Artificial Intelligence and Machine Learning in Prediabetes: A Scoping Review by Benjamin Lalani, Rohan Herur, Daniel Zade, Grace Collins, Devin M. Dishong, Setu Mehta, Jalene Shim, Yllka Valdez and Nestoras Mathioudakis in Journal of Diabetes Science and Technology
Supplemental Material
sj-xlsx-4-dst-10.1177_19322968251351995 – Supplemental material for Applications of Artificial Intelligence and Machine Learning in Prediabetes: A Scoping Review
Supplemental material, sj-xlsx-4-dst-10.1177_19322968251351995 for Applications of Artificial Intelligence and Machine Learning in Prediabetes: A Scoping Review by Benjamin Lalani, Rohan Herur, Daniel Zade, Grace Collins, Devin M. Dishong, Setu Mehta, Jalene Shim, Yllka Valdez and Nestoras Mathioudakis in Journal of Diabetes Science and Technology
Supplemental Material
sj-xlsx-5-dst-10.1177_19322968251351995 – Supplemental material for Applications of Artificial Intelligence and Machine Learning in Prediabetes: A Scoping Review
Supplemental material, sj-xlsx-5-dst-10.1177_19322968251351995 for Applications of Artificial Intelligence and Machine Learning in Prediabetes: A Scoping Review by Benjamin Lalani, Rohan Herur, Daniel Zade, Grace Collins, Devin M. Dishong, Setu Mehta, Jalene Shim, Yllka Valdez and Nestoras Mathioudakis in Journal of Diabetes Science and Technology
Supplemental Material
sj-xlsx-6-dst-10.1177_19322968251351995 – Supplemental material for Applications of Artificial Intelligence and Machine Learning in Prediabetes: A Scoping Review
Supplemental material, sj-xlsx-6-dst-10.1177_19322968251351995 for Applications of Artificial Intelligence and Machine Learning in Prediabetes: A Scoping Review by Benjamin Lalani, Rohan Herur, Daniel Zade, Grace Collins, Devin M. Dishong, Setu Mehta, Jalene Shim, Yllka Valdez and Nestoras Mathioudakis in Journal of Diabetes Science and Technology
Supplemental Material
sj-xlsx-7-dst-10.1177_19322968251351995 – Supplemental material for Applications of Artificial Intelligence and Machine Learning in Prediabetes: A Scoping Review
Supplemental material, sj-xlsx-7-dst-10.1177_19322968251351995 for Applications of Artificial Intelligence and Machine Learning in Prediabetes: A Scoping Review by Benjamin Lalani, Rohan Herur, Daniel Zade, Grace Collins, Devin M. Dishong, Setu Mehta, Jalene Shim, Yllka Valdez and Nestoras Mathioudakis in Journal of Diabetes Science and Technology
Footnotes
Abbreviations
A1C, hemoglobin A1C; ADA, American Diabetes Association; AI, artificial intelligence; BMI, body mass index; CatBoost, categorical boosting; CDC, Centers for Disease Control and Prevention; CGM, continuous glucose monitoring; Cox, Cox proportional hazards model; DPP, diabetes prevention program; DT, decision tree; EHR, electronic health record; EM, ensemble methods; FDA, Food and Drug Administration; GBM, gradient boosting machine; GMI, glucose management indicator; ITT, intention-to-treat; Lasso, least absolute shrinkage and selection operator; LGBM, light gradient boosting machine; LogR, logistic regression; MET, metabolic equivalent of task; ML, machine learning; NB, naïve bayes; NN, neural network; PPGR, postprandial glucose response; PRISMA, Preferred Reporting Items for Systematic Reviews and Meta-Analyses; RCT, randomized controlled trial; RF, random forest; RMSE, root mean squared error; SVM, support vector machine; T2D, type 2 diabetes; WHO, World Health Organization; XGB, eXtreme gradient boosting.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The study was funded by a grant (grant no. R01DK125780) from the National Institute of Diabetes and Digestive and Kidney Diseases (NIDDK), National Institutes of Health (NIH;
ORCID iDs
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
