Abstract
Introduction:
Current methods to detect hypoglycemia in type 1 diabetes (T1D) require invasive sensors (ie, continuous glucose monitors, CGMs) that generally have low accuracy in the hypoglycemic range. A forward-looking alternative is to monitor physiological changes induced by hypoglycemia that can be measured non-invasively using, eg, electrocardiography (ECG). However, current methods require extraction of fiduciary points in the ECG signal (eg, to estimate QT interval), which is challenging in ambulatory settings.
Methods:
To address this issue, we present a machine-learning model that uses (1) convolutional neural networks (CNNs) to extract morphological information from raw ECG signals without the need to identify fiduciary points and (2) ensemble learning to aggregate predictions from multiple ECG beats. We evaluate the model on an experimental data set that contains ECG and CGM recordings over a period of 14 days from ten participants with T1D. We consider two testing scenarios, one that divides ECG data according to CGM readings (CGM-split) and another that divides ECG data on a day-to-day basis (day-split)
Results:
We find that models trained using CGM-splits tend to produce overly optimistic estimates of hypoglycemia prediction, whereas day-splits provide more realistic estimates, which are consistent with the intrinsic accuracy of CGM devices. More importantly, we find that aggregating predictions from multiple ECG beats using ensemble learning significantly improves predictions at the beat level, though these improvements have large inter-individual differences.
Conclusion:
Deep learning models and ensemble learning can extract and aggregate morphological information in ECG signals that is predictive of hypoglycemia. Using two validation procedures, we estimate an upper bound on the accuracy of ECG hypoglycemia prediction of 81% equal error rate and a lower bound of 60%. Further improvements may be achieved using big-data approaches that require longitudinal data from a large cohort of participants.
Introduction
A critical aspect in managing type 1 diabetes (T1D) is preventing hypoglycemic events. While minimally invasive devices, such as continuous glucose monitors (CGMs) exist, they generally have lower accuracy in the hypoglycemic range (4 mmol/L; 72.07 mg/dL). Continuous glucose monitor use for insulin dosing decisions is generally considered to be feasible for mean absolute relative differences (MARDs) less than 10%. 1 However, Food and Drug Administration (FDA) approval letters for the four leading selling CGMs in the United States show MARD in the hypoglycemic range (interstitial fluid glucose concentration below 54 mg/dL) for adults are 10% to 16% for the Abbott Freestyle Libre3, 2 1% to 14% for the Dexcom G7, 3 13% to 23% for the Medtronic Simplera System, 4 and 13% to 20% for the Senseonics Eversense E3. 5
Several physiological variables have been investigated as potential indirect indicators of hypoglycemia. 6 Early work shows that skin temperature and skin conductivity decrease at the onset of hypoglycemia. 7 Several commercial instruments were developed in the 1980s,8,9 but they suffered from several issues, such as false alarms due to perspiration that was unrelated to hypoglycemia—a reported 3:1 ratio of false alarms to true alarms, 10 so that, they never received FDA approval. Most of the work on hypoglycemia detection using noninvasive physiological sensors has focused on electrocardiography (ECG). Several changes in cardiac signals have been robustly associated with hypoglycemia, most notably a lengthened QT interval. 11 However, this requires extracting fiduciary points from ECG recordings (eg, T wave), which lacks robustness due to motion artifacts. To avoid these issues, Porumb et al 12 have shown that convolutional neural networks (CNNs) can be used to extract beat-level morphological changes in raw ECG signals that are associated with hypoglycemia, thus avoiding the error-prone problem of identifying fiduciary points.
Borrowing from Porumb et al, 12 this study examines whether aggregating beat-level predictions at the timescale of CGM reading (ie, 5 minutes, or about 300-600 heart beats, depending on the heart rate) can improve the accuracy of hypoglycemia prediction. For this purpose, we propose a CNN model that predicts the probability of hypoglycemia for each ECG heartbeat and combines them into a single prediction at 5-minute intervals. Our approach is based on ensemble learning, 13 a machine-learning approach that combines multiple models to improve prediction performance. Our model computes a percentile plot of the beat-level probabilities associated with each CGM reading, and uses them to train a second, smaller neural network. We also propose a strategy to estimate the upper and lower bounds of accuracy that may be expected when predicting hypoglycemia from ECG.
Methods
Physiological Recordings
Data for this article were collected at Baylor College of Medicine under institutional review board (IRB) protocol H-49867. Participants were eligible to participate in the study if they had a clinical diagnosis of T1D with a duration greater than 1 year and were 13 years or older. In an effort to obtain sufficient CGM recordings to train the prediction models, all participants were verified to have at least 80% CGM use with a history of glycemic excursions (<70 and >180 mg/dL) in the month before enrollment. Ten subjects were enrolled between the ages of 29 and 41 years old and their body mass index (BMI) was in the range of 21.8 and 34.1 kg/m2. Participant demographics are included in Table 1. All participants provided written consent prior to initiating the study.
Overview of the Data Set Used for This Study:

Abbreviations: BMI = body mass index; CGM = continuous glucose monitor.
As part of their regular medical treatment, all subjects were using a hybrid closed-loop insulin pump with a Dexcom G6 CGM. In addition, participants wore three commercial wearable devices: (1) an Empatica E4 wristwatch that measures photoplethysmography (PPG) and electrodermal activity (EDA), 14 (2) an Oura Ring that measures heart rate via PPG, 15 and (3) a Zephyr Bioharness that measures ECG and respiration. 16 Upon enrollment, subjects had at least 14 days of data collection on all devices. Figure 1 shows CGM recordings for the ten subjects over the study period. Two subjects (c1s02 and c1s04) experienced few hypoglycemic events during the 14 days (see Table 1), so that, they were not included in the analysis given the lack of sufficient hypoglycemic recordings to train the prediction models.
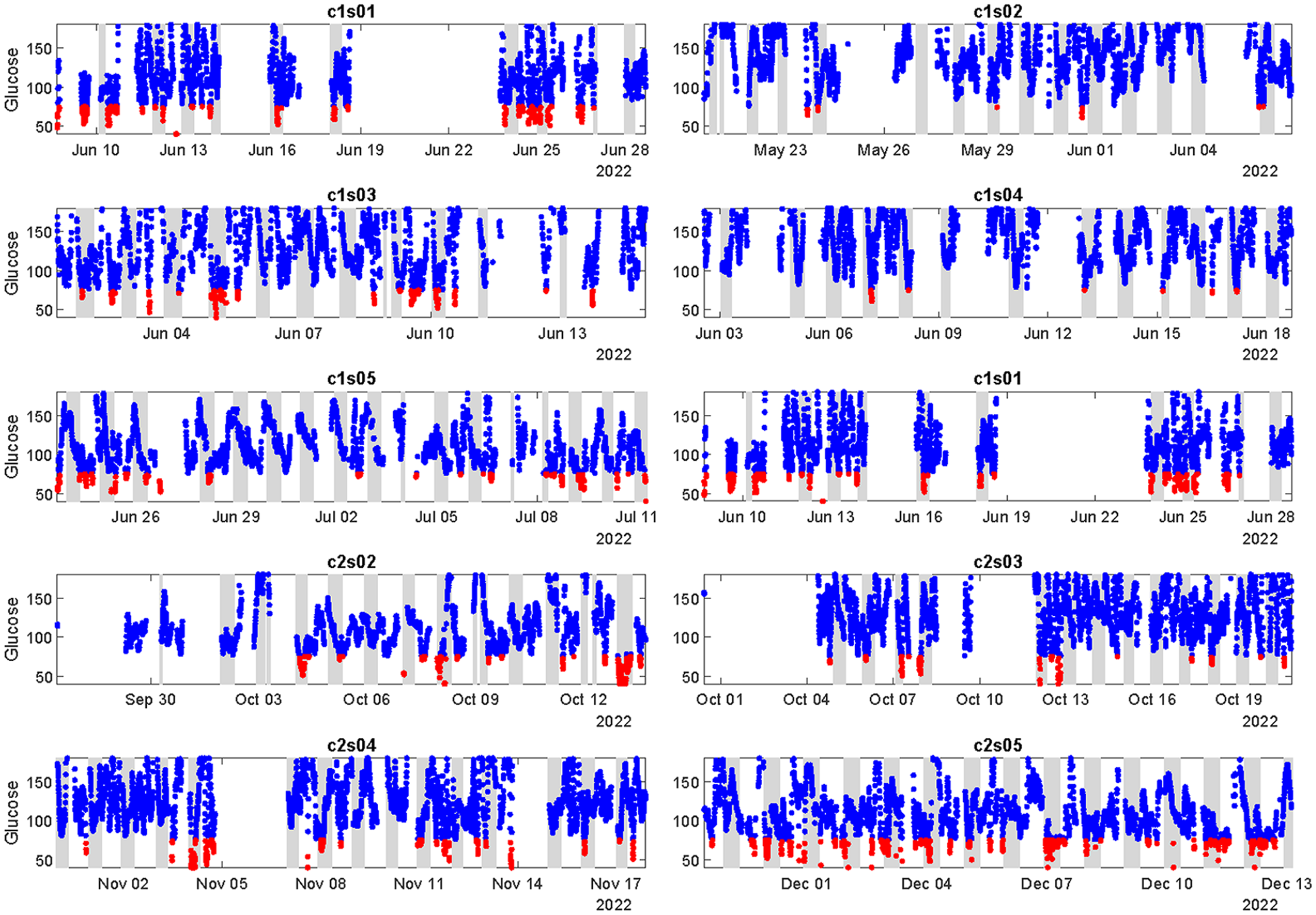
CGM recordings for the ten participants in the study.
Furthermore, we only considered data from the Zephyr Bioharness, which records ECG at 250 Hz, and from the Dexcom G6 CGM, which reports interstitial glucose every 5 minutes. Following our prior work, 17 we used metadata from the Bioharness to identify “good quality” ECG segments, defined as having heart-rate confidence (HRC) greater than 199 and ECG sensor noise (ECG-N) less than 0.001. Then, we used Neurokit2 18 to detect R peaks and extracted a variable-length window as a percentage of the RR interval (33% back, 66% forward). Using a variable window allows us to account for changes in beat morphology due to heart rate (eg, QT interval prolongation with increased RR interval). Finally, we zero-padded ECG beats into a fixed length for the CNN models. We labeled each ECG beat according to the next (closest) CGM reading.
Soft Labeling
Given that the Dexcom G6 has an MARD of 12% to 14% in the hypoglycemic range, 19 we do not use 70 mg/dL as a hard threshold for hypoglycemia, as this would make it difficult for the CNN model to learn. Instead, we convert CGM readings into an estimated probability of hypoglycemia that accounts for the intrinsic error of the device. In this fashion, we convert the problem of hypoglycemia prediction (ie, binary classification) into one of predicting a continuous variable (ie, regression). Similar techniques have been used in the machine-learning literature to avoid overfitting and are known as label smoothing. 20 Illustrated in Figure 2a, label smoothing maps CGM readings into an estimated probability of hypoglycemia using a piece-wise linear function, with a probability of 1.0 at 40 mg/dL, 0.9 at 63 mg/dL, 0.5 at 70 mg/dL, 0.1 at 76 mg/dL, and 0.0 at 100 mg/dL. We remove CGM readings above 180 mg/dL from the training set, since hyperglycemia does not pose an immediate threat to the patient. In doing so, we reduce class imbalance and allow the model to better detect hypoglycemia, the more significant clinical problem in the short term.
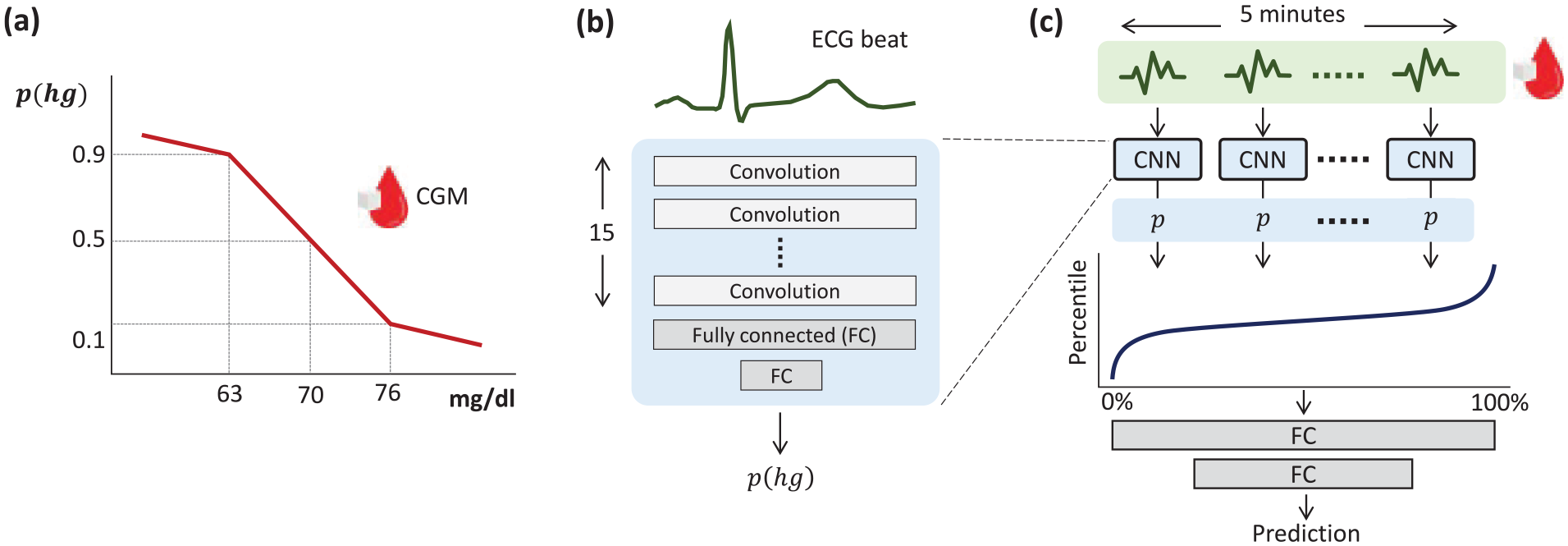
(a) Generating soft labels from CGM readings. (b) Basic architecture of the Convolutional Neural Network. (c) Ensemble predictions from the percentiles of CNN output probabilities.
Beat-Level Prediction of Hypoglycemia
Our beat-level prediction model is a CNN that consumes fixed-length zero-padded ECG waveforms and produces a glucose estimate in the form of a soft label, as described above. The CNN consists of 15 convolutional layers, each with a kernel size of 3, a stride of 1, and 50 filters (see Figure 2b). We apply an ReLU (rectified linear unit) activation function and one-dimensional batch normalization within each convolution layer. The CNN layers are followed by two fully connected (FC) layers with 250 and 30 neurons, respectively, which predict the probability of hypoglycemia from the embeddings of the CNN. We apply a dropout layer with a dropout rate of 20%, along with an ReLU activation function between the two layers. We train the model for ten epochs using the ADAM optimizer with a learning rate of 0.00005. We implement early stopping with a patience of 7 to prevent overfitting. Given the large class imbalance (hypoglycemic readings represent 1%-5% of all CGM readings), we use weighted cross-entropy as the loss function:
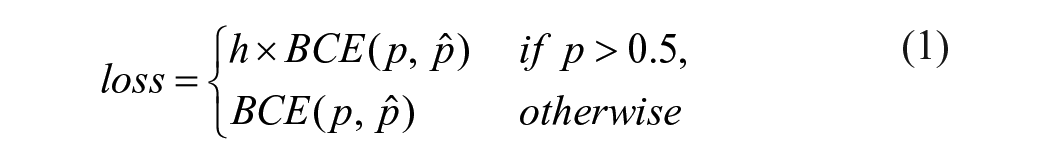
where
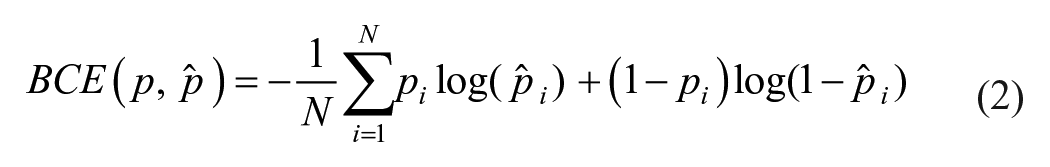
and
Ensemble Prediction of Hypoglycemia
Given that glucose dynamics are significantly slower that cardiac dynamics, we use an ensemble learning technique to aggregate beat-level predictions from all ECG beats associated with a CGM reading (ie, a 5-minute window, or about 300-600 heart beats, depending on the heart rate) into a single prediction. Under the assumption that prediction errors at the ECG beat level are independent identically distributed (i.i.d.), ensemble learning has been shown to improve the performance of a “lazy learner” algorithm (a learner that performs just above the chance level). 21
Our ensemble method is based on stacked generalization, a machine-learning approach that trains a second-level model to combine predictions from two or more first-level models.
22
In our case, the first-level models are the CNNs that predict hypoglycemia at the beat level. Given the CNN output probabilities
Validation Approaches
We evaluate two approaches to split the data into training, validation, and test sets. The first approach (CGM-splits) randomly splits a participant’s CGM recordings (and the prior 5 minutes of ECG recordings) into a training set (70%) and a test set (30%) in a stratified fashion to ensure that both sets have the same proportion of hypoglycemic beats. The training set is then further divided into a training subset (70%) to train the CNN models, and a validation subset (30%). For the CNN beat-level prediction model, we use the validation set to find the optimum threshold for the posterior probability at the output of the CNN that minimizes the equal error rate (EER), defined as the point where the true-positive rate (correct hypoglycemia alarms) equals the false negative rate (false hypoglycemia alarms). For the stacked-generalization model, we combine the validation and training subsets to train the FC network that consumes the percentile curve (see Figure 2c), using weighted cross-entropy loss to balance the proportion of training and validation data. Given the potential confounding effect of time (ie, ECG beats from the same period can have similar beat morphology), CGM-splits can lead to overly optimistic results. Thus, we use these results as an optimistic estimate of the EER.
The second approach (day-splits) partitions each participant’s data set into training, validation and test sets on a day-by-day basis, rather than by CGM recordings. Namely, given
Statistical Analysis
We compare CGM-splits vs day-splits using a two-sample t test on EERs, with EER for CGM-splits based on the average across five separate runs (each run with a random 70/30 split), and EER for day-splits based on the average of
We compare beat-level vs ensemble-level predictions using a two-sample t test on day-wise splits. To examine individual differences across subjects, we then conducted two-way analysis of variance (ANOVA) without replication using subject and model type (beat vs ensemble level) as independent factors.
Results
Predictions From CGM-Splits vs Day-Splits
In a first step, we evaluate the performance of the CNN models when partitioning according to CGM readings vs experimental days. As noted earlier, we only consider beat-level predictions for this comparison, since the relative performance of both splitting approaches is likely to generalize to the ensemble model.
Results across participants are illustrated in Figure 3. As expected, predictive accuracy at EER is higher with CGM-splits (81%) than with day-splits (60%), a difference that is statistically significant (P < .01). Considering that the sensitivity of CGM devices for hypoglycemia detection is around 85%, 23 it is highly likely the estimated accuracy of 81% when using CGM-splits is, in part, due to the confounding effect of time. For example, when a hypoglycemic event is longer than 5 minutes (which is generally the case), splitting the data at the CGM level leads to that hypoglycemic event to be used both for training and testing, which results in unrealistically high predictive accuracy. For this reason, we consider the estimated 81% EER from CGM-splits and the estimated 60% EER from day-splits as the upper and lower bounds, respectively, of hypoglycemia prediction from ECG beat morphology.
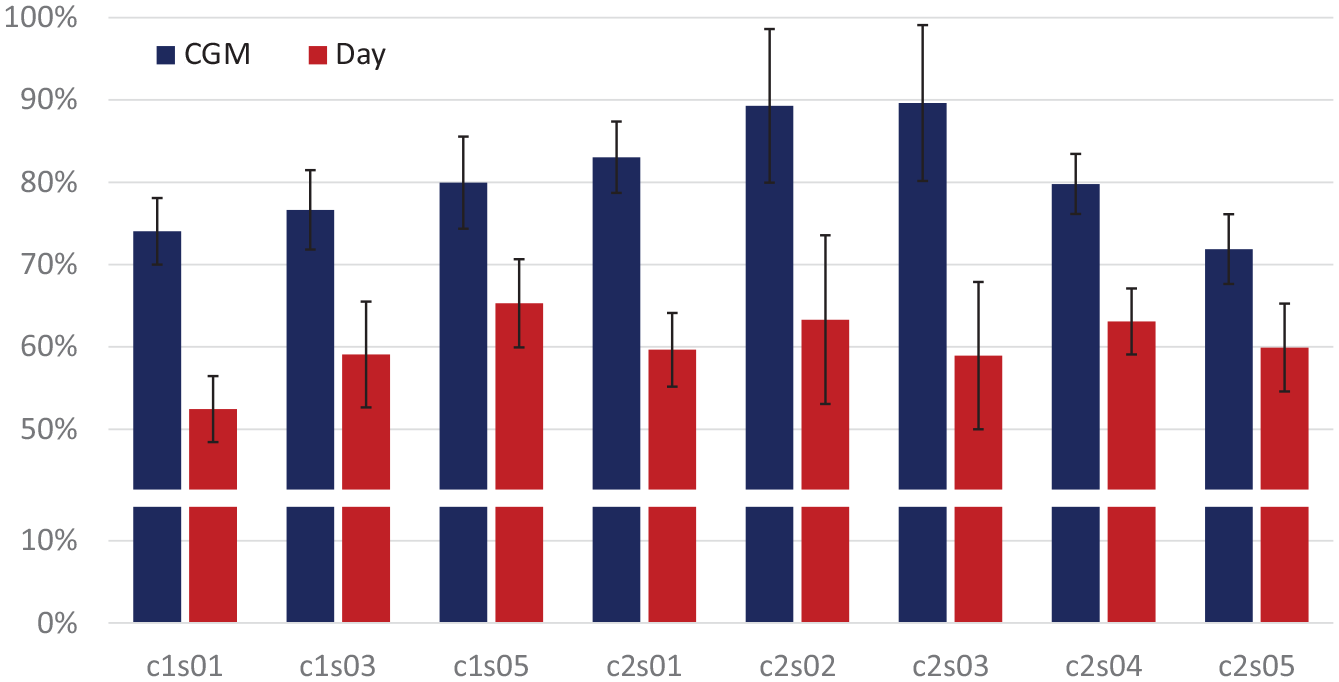
Accuracy at EER for the beat-level CNN model, with data partitioned based on CGM readings (blue) and days of collection (red). Error bars represent the standard error.
Beat-Level vs Ensemble Prediction
Given that CGM-split predictions lead to overly optimistic results, for the second experiment, we compared the beat-level and ensemble-level predictions using only day-splits. Results across subjects are summarized in Figure 4. The stacked-generalization ensemble achieves higher accuracy at EER (65%) than the vote-level model (60%), though in this case differences are only statistically significant at the P = .07 level. To examine individual differences across subjects, we then conducted two-way ANOVA without replication using subject and model type (beat vs ensemble level) as independent factors. We find a main effect for subject (P = .021) and for model type (P = .026), indicating that, when considering individual differences, the ensemble method provides significantly higher predictive accuracy than the beat-level model.
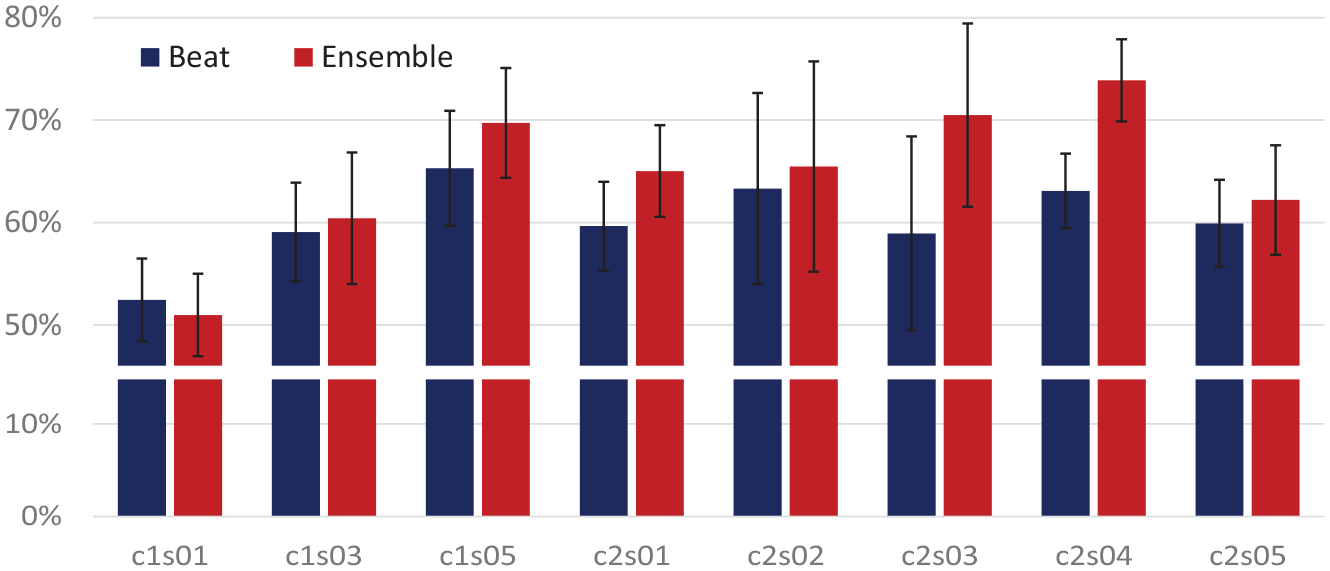
Accuracy at EER for the beat-level CNN (blue) and stacked generalization (red). Error bars represent standard error.
To illustrate the information consumed by the ensemble model, Figure 5a shows the percentile plots of CNN probability estimates (over 5-minute windows) for hypoglycemic and euglycemic CGM readings on one of the test days for subject c2s04, whereas Figure 5b shows the average percentile plot over all test days. As shown, for percentiles above 60%, probabilities are significantly higher for hypoglycemic readings and have significantly lower variability than those for euglycemic readings. The larger variability to the latter may be due to the large class imbalance.
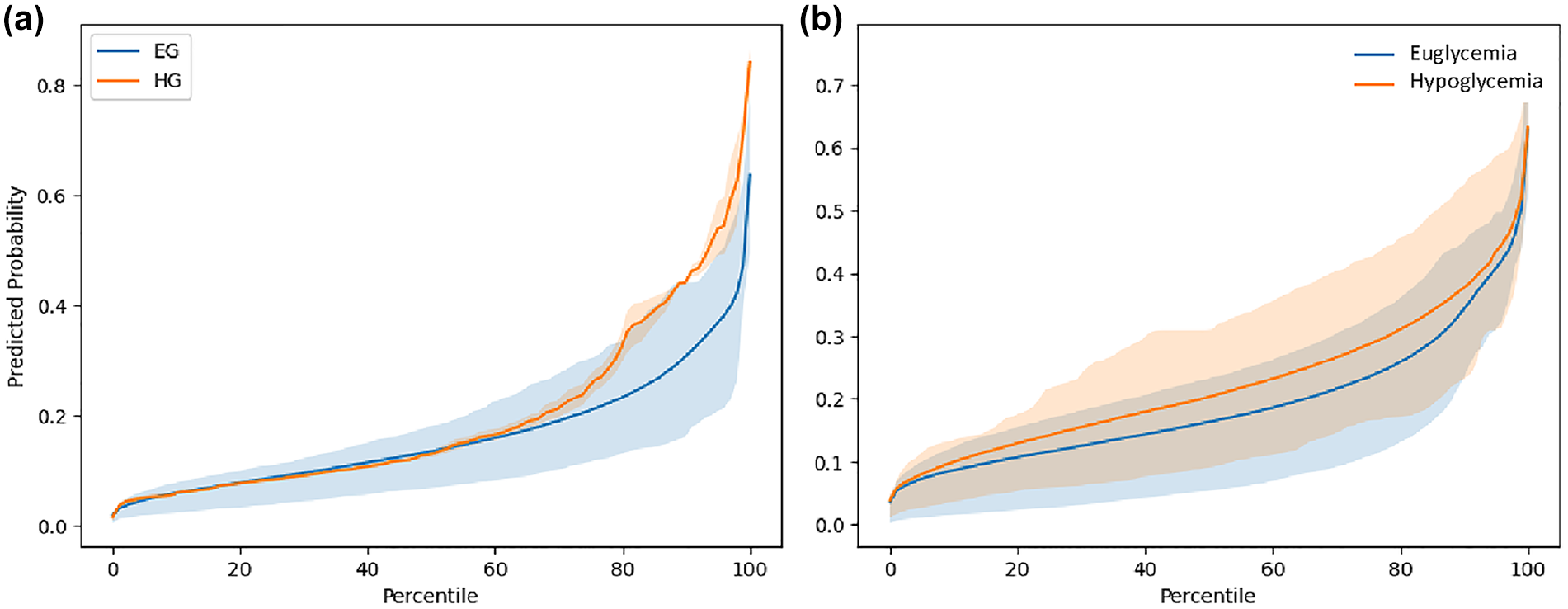
Percentile plot for hypoglycemic and euglycemic beats for subject c2s04.
Discussion
Depending on the subject, the ensemble methods improve the predictive accuracy from 3% to 10%, when compared with beat-level predictions, though these improvements are only statistically significant for three of the eight subjects. A potential explanation for the relatively low accuracy of the models is the presence of pressure artifacts in CGM readings, 24 particularly during the night. As patients inadvertently roll over the CGM, interstitial fluid is pushed away from the CGM electrode, resulting in false hypoglycemic readings when the actual ECG beat morphology is that of euglycemic glucose levels. To address this issue, data-driven techniques 24 may be used to detect such pressure artifacts, and remove those measurements form the training set.
Predictive accuracy may also be increased by replacing the CNN with a model better suited to analyze time series, such as the InceptionTime architecture, 25 which incorporates residual blocks as well as convolutional layers with varying kernel sizes, allowing the model to capture morphological features at different scale factors. Additional gains in performance may also be obtained by combining beat morphology with additional information in the ECG signal, such as measures of heart rate variability and timing information in the beat-to-beat time series, as well as other contextual information, such as time of day, physical activity and posture, all of which can be easily obtained from commercial wearable sensor devices. Our prior work 17 has shown that a combination of these various sources of information achieves higher accuracy that predictions from each of them in isolation.
Simpler models (in terms of number of parameters) may also be used. While conducting this study, we also evaluated boosted regression trees. However, this type of model is very sensitive to the alignment of the ECG R peak. In contrast, the proposed CNN is shift invariant. With poor R-peak alignment, our CNN generally outperforms boosted regression trees. Thus, while the CNN has a higher risk of overfitting, in our experience, this has not been the case.
We focused on a 5-minute ECG window to match the sampling period of the Dexcom G6, but our study can be easily extended to longer windows, simply by associating each CGM reading with the prior 10 or 15 minutes of ECG recordings. Our expectation is that a longer analysis window will improve model accuracy, up to the point where the ECG window is longer than the time constant of glucose dynamics. We are currently developing a model that examines a multi-beat window (3c-30 seconds) to capture not only beat morphology but also beat-to-beat information, such as heart rate variability. Our preliminary results indicate that this multi-beat window size does improve accuracy.
Our proposed can be used for real-time detection as is, since it predicts hypoglycemia at the same rate as the fastest CGM devices on the market. This would require carefully optimizing the model (ie, by reducing the number of model parameters) to reduce the lag of the predictions. We believe this would not be an issue since our CNN model is relatively smaller than those use in speech research. Another possibility we have not yet examined is whether our model can be used to forecast future glucose readings, eg, predicting glucose levels 5 to 30 minutes into the future. Our expectation is that accuracy would decrease as the forecasting horizon is increased.
Limitations of the Study
One limitation of this study is the relatively small number of patients in the study (10 patients). To our knowledge, however, the only publicly available data set containing both ECG and CGM recordings is the D1NAMO data set, and only contains 4 days of recordings from nine patients with T1D (36 days). In comparison, our data set contains 10 to 15 days of recording from ten participants (roughly three times as large). Furthermore, while our current number of participants is objectively small and needs to be validated on data from a larger set of participants, this limitation does not affect the results of our study since our models are participant dependent. As we extend our data set with data from new participants being recruited at the time of this writing, it then becomes possible to train and evaluate subject-independent models in a leave-one-subject-out cross-validation procedure. Our experience with other diabetes-related data sets we have collected in the past 26 is that subject-independent models only improve the performance of subject-dependent models when the number of participants is larger than 50. This is largely due to the large individual differences in physiology and glucose regulation.
A second limitation of this study is that hypoglycemic readings were not validated with capillary or venous blood samples, since this was not specified in the protocol. At the time of this writing, we are developing a new protocol that will include verification of CGM hypoglycemic readings with capillary blood samples, as well as validated self-report measures of hypoglycemia, such as the Edinburgh Hypoglycemia Scale 27 and Clarke and Gold scores. 28
Finally, our hypoglycemia prediction models were trained using participant data from up to 14 days, so that, we are unable to examine the long-term stability of the models. Given that participants were required to wear two additional devices (which included charging them every night and uploading their data to the cloud for the research team), increasing the study period beyond 2 weeks would have put significant burden on participants and likely reduced adherence to the protocol. As part of the grant that funded the study, we have developed a compact sensing device (to be worn in the upper arm) that measures ECG, PPG, EDA, temperature, and acceleration simultaneously. Once this device is thoroughly validated against the commercial devices used in the current study, conducting longitudinal studies becomes feasible. Longitudinal data will allow us to examine the stability of our models and potentially improve model accuracy with the additional training data that become available.
Conclusions
Convolutional neural networks can be used to extract morphological information from ECG recordings that is predictive of glucose readings in the hypoglycemic range without the need to detect fiduciary points in ECG recordings, which is challenging in ambulatory settings due to motion artifacts. Aggregating hypoglycemia predictions for individual beats within a CGM recording improves predictive accuracy at EER over beat-level predictions, though there are large inter-individual differences in the magnitude of these improvements.
Footnotes
Abbreviations
ECG, electrocardiography; FDA, Food and Drug Administration; CNN, convolutional neural networks; CGM, continuous glucose monitor; T1D, type 1 diabetes; EDA, electrodermal activity; IRB, institutional review board; QT, interval between the Q and T waves on an ECG recording; MARDs, mean absolute relative differences; PPG, photoplethysmography;
Declaration of Conflicting Interests
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: Daniel J. DeSalvo has served as an independent consultant for Dexcom and Insulet separate from this work. The remaining authors declare no competing interests.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by NSF award #2037383.
