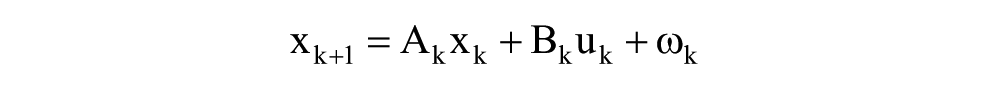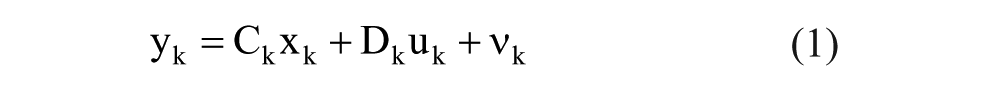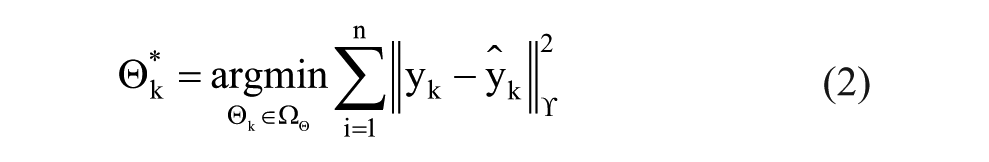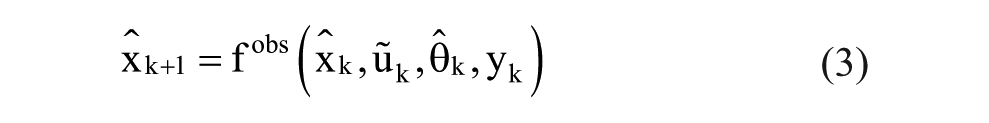Abstract
Background:
Despite the recent advancements in the modeling of glycemic dynamics for type 1 diabetes mellitus, automatically considering unannounced meals and exercise without manual user inputs remains challenging.
Method:
An adaptive model identification technique that incorporates exercise information and estimates of the effects of unannounced meals obtained automatically without user input is proposed in this work. The effects of the unknown consumed carbohydrates are estimated using an individualized unscented Kalman filtering algorithm employing an augmented glucose-insulin dynamic model, and exercise information is acquired from noninvasive physiological measurements. The additional information on meals and exercise is incorporated with personalized estimates of plasma insulin concentration and glucose measurement data in an adaptive model identification algorithm.
Results:
The efficacy of the proposed personalized and adaptive modeling algorithm is demonstrated using clinical data involving closed-loop experiments of the artificial pancreas system, and the results demonstrate accurate glycemic modeling with the average root-mean-square error (mean absolute error) of 25.50 mg/dL (18.18 mg/dL) for six-step (30 minutes ahead) predictions.
Conclusions:
The approach presented is able to identify reliable time-varying individualized glucose-insulin models.
Keywords
Improved glycemic control in individuals with type 1 diabetes mellitus (T1DM) via the use of artificial pancreas (AP) systems, which integrate a continuous glucose monitoring (CGM) sensor, continuous subcutaneous insulin infusion (CSII) pump, and insulin dosing control algorithm, reduces the risks of immediate life-threatening conditions, such as severe hypoglycemia and ketoacidosis, and long-term health complications, such as cardiovascular disease, nephropathy, neuropathy, and retinopathy.1-13 A fully automated AP system eliminating the need for patients with T1DM to enter user inputs for meal and exercise announcements represents a substantial step toward achieving better insulin delivery systems. Although AP systems are shown to be effective compared to conventional multiple daily insulin injections or sensor augmented pump therapy, improvements in the insulin dosing algorithms are needed if the AP systems are to compensate for meals and exercise without requiring patients to manually interact with the systems. Model-based predictive control has emerged as an effective insulin dosing algorithm for AP systems as it inherently considers future projections of the glycemic dynamics for predictive hypo- and hyperglycemia alarms and manipulating insulin delivery when glucose concentrations are forecast to deviate from the desired outcomes. The ability of the predictive controllers is dependent on accurate models of the metabolic system that can effectively encode a comprehensive understanding of the physiology of T1DM, handle the long dynamical effects of the insulin action, and characterize the effects of various diurnal disturbances on glycemic dynamics. Furthermore, many mathematical models of the glucose and insulin dynamics employ real-time feedback solely from the CGM signal, which limits their ability to automatically accommodate meals and exercise without manual interventions by people with T1DM.
One of the challenges to achieve a fully automated and reliable AP is the lack of an accurate model to represent the dynamic changes in physiology under various conditions. The future glucose measurements are difficult to predict accurately as glycemic dynamics vary substantially due to the effects of numerous factors such as current and historical glucose trends, carbohydrate consumption, previously administered insulin, exercise or physical activity levels, and concentrations of certain hormones. Moreover, metabolic processes vary substantially among subjects and temporally within people due to the diversified lifestyles and erratic routines of individuals. The diversity of physiology and behaviors in people causes the glucose-insulin dynamics and insulin sensitivities of individuals to vary over time, while large perturbations such as meals and physical activity cause significant excursions in glucose concentrations that may not be described accurately by generalized models. Hence, the necessity in AP systems of personalized glucose-insulin models rather than models with generalized parameters that do not reflect the dynamic characteristics of subjects in different situations.14-19
The glycemic models proposed in literature can be categorized as either physiological or data-driven models, often with carbohydrate intake and infused insulin as inputs. Physiological models, usually based on compartmental models, consist of simultaneous differential equations describing the insulin and glucose metabolism with a number of physiological parameters to be identified. Detailed physiological models are very useful in simulations, though model complexity and computational load impede their real-time implementation in AP systems. In contrast to physiological models, data-driven models with relatively simpler structures that effectively characterize the intricate relationships among the measured variables generally require less computational effort. 15 Among the various empirical modeling approaches proposed to predict future glycemic measurements, subspace-based system identification methods are capable of efficiently identifying linear state-space models from multi-input, multi-output sampled data of a dynamic system. Nevertheless, a fixed empirical model may not predict glycemic measurements well in all scenarios and across all subjects due to the significant variability in glycemic dynamics. Therefore, the empirical models need to be appropriately adapted on-line to characterize the current dynamics of the individuals and make accurate short-term predictions of glucose concentration measurements. For this purpose, adaptive system identification approaches are proposed to determine linear, time-varying models and effectively characterize the evolving glycemic dynamics, thus allowing the adaptive models to be valid over a diverse range of daily conditions.
Despite the predictive ability of the adaptive system identification approach, the discrete basal changes and acute bolus impulses present a challenging configuration for empirical modeling techniques. Further complicating the model identification procedure is the fact that administered insulin (basal or bolus) gradually accumulates in the bloodstream and is eventually utilized by the body. One of the factors that prolongs the utilization of administered insulin is the significant time delays involved in the diffusion and absorption of the subcutaneously injected insulin analogues. For instance, the plasma insulin response to subcutaneously infused fast-acting insulin has a time lag of approximately 15 minutes, peak effect at about 45-90 minutes, and an overall effective duration of 4-6 hours. The protracted effects of the administered insulin may cause glucose concentration measurements to continue rising in response to carbohydrate consumption even though sufficient insulin is already administered. Under such postprandial circumstances where insulin effects are not observed in CGM measurements for lengthy periods of time after insulin infusion, methods relying entirely on data-driven models may lead to inappropriate generalizations with regards to the effects of insulin. This well-recognized artifact of modeling glycemic measurements can be addressed by filtering the administered insulin dose into a newly constructed variable that readily accommodates the effects of previously administered insulin. It is well recognized that filtering the input variables has a significant effect on the predictive performance of the identified model. Although such schemes are shown to improve glycemic prediction accuracy, the filtering algorithms utilized, whether based on numerical signal processing techniques or derived from compartment models describing the underlying physiological phenomena, are often time-invariant. The application of fixed time-invariant filters thus may be suboptimal and may diminish the potential for improvement, especially when employed simultaneously with adaptive system identification techniques. Utilizing PIC estimates from an adaptive and personalized PIC estimator as the newly established input for the identification algorithm provides a filtered variable that improves prediction ability. In contrast to the conventional insulin on board approach that estimates the amount of active insulin in the body through static appropriations of action profiles and decay curves, the estimated PIC is a physically meaningful and quantifiable measure of insulin the bloodstream that may benefit control design as well. The estimated plasma insulin concentration (PIC) can be considered as a more appropriate input variable in the identification of a dynamic model for predicting future glucose concentrations because it directly quantifies the insulin in the bloodstream that affects the glycemic evolution.20-24
Besides the effects of administered insulin, another limitation of current AP systems is the requirement of manual meal and exercise announcements. Several studies have incorporated unannounced meals through the estimation of time-varying parameters or analysis of glucose trends.25-30 Concerning the automation of AP systems, additional physiological variables related to physical activity are also considered to automatically accommodate exercise.31-34 Despite this progress, automatically handling of unannounced meals and exercise in adaptive and personalized glycemic models for AP systems is not sufficiently studied.
Motivated by the above considerations, in this work, an adaptive and personalized compartment model that translates the abrupt bolus and discrete basal changes into estimates of PIC is integrated with an adaptive system identification approach to characterize the transient dynamics of glycemic measurements. The adaptations by the recursive system identification assist in handling stochastic disturbances like the effects of meal consumption and physical activities. Furthermore, the system identification algorithm is extended to improve reliability for use in fully automated artificial pancreas systems. The proposed approach involves the modification of the recursive predictor-based subspace identification (PBSID) algorithm to incorporate constraints on the fidelity and accuracy of the identified models, correctness of the sign of the input-to-output gains, and the integration of heuristics to ensure the stability of the recursively identified models. To achieve this, the proposed adaptive models also include estimated meal effects and the reported exercise related biometric variables by BodyMedia SenseWear armband as additional inputs to automatically accommodate unannounced meals and exercise. The proposed adaptive and personalized modeling approach considering the effects of unannounced meals and exercise on the transient glycemic dynamics is applied to 15 clinical data sets involving closed-loop experiments of the AP systems.
Methods
The predictor-based subspace identification (PBSID) approach is used to identify a recursively updated state-space model characterizing the glucose-insulin dynamics. The PBSID algorithm estimates a vector autoregressive with exogenous inputs model and uses it to construct matrices related to the state-space model parameters. A low-rank approximation then identifies a state sequence, and the state-space model parameters are recursively updated. The PBSID technique is further extended to provide stable, time-varying, and individualized models for glycemic predictions by using estimates of PIC and meal effects, and measured physiological signals. The estimates of PIC and meal effects are obtained by-using an adaptive and personalized PIC estimator designed based on the Hovorka’s glucose-insulin dynamic model. In this section, a review of the PBSID algorithm for the identification of linear, time-varying state-space models is provided, followed by a brief overview of the adaptive and personalized PIC estimator. Figure 1 shows a flowchart of the proposed personalized and adaptive modeling algorithm.
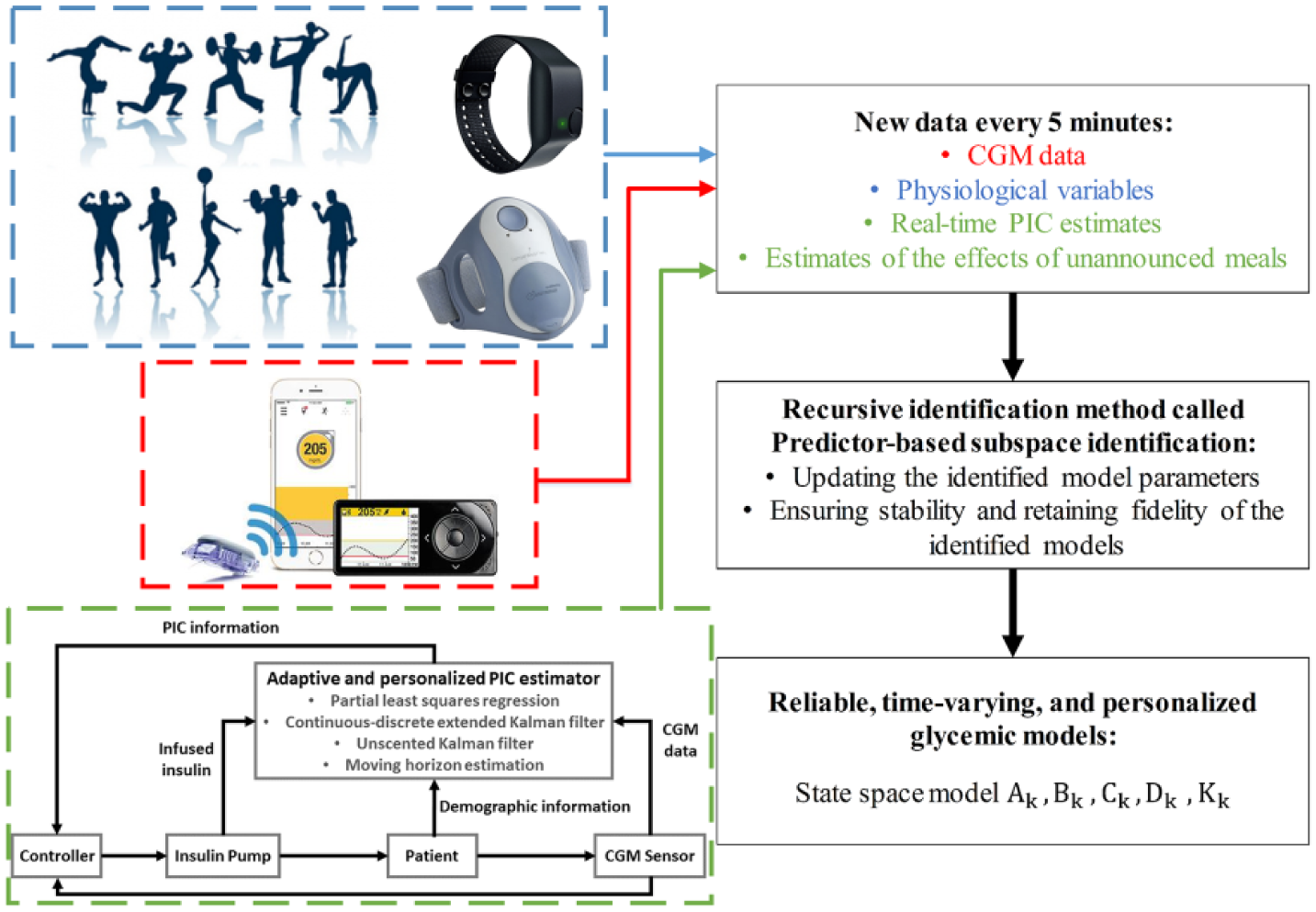
Flowchart of the proposed personalized and adaptive modeling algorithm.
Adaptive Modeling Approach
In this work, a recursive subspace-based system identification approach is used to build linear, time-varying state-space models of the form:
where
where a weighted quadratic norm of the deviation of the predicted CGM outputs is penalized from the actual CGM measurements while ensuring that the estimated model coefficients satisfy the imposed constraints
One modification compared to traditional glycemic models is the use of PIC as an input to the system identification procedure. The insulin present in the bloodstream is derived from the insulin subsystem of Hovorka’s model, which translates the abrupt basal and discrete bolus inputs into PIC estimates with time-varying model parameters simultaneously estimated using a nonlinear observer.36,37 The observer is of the form
where
Subjects and Clinical Study Experiments
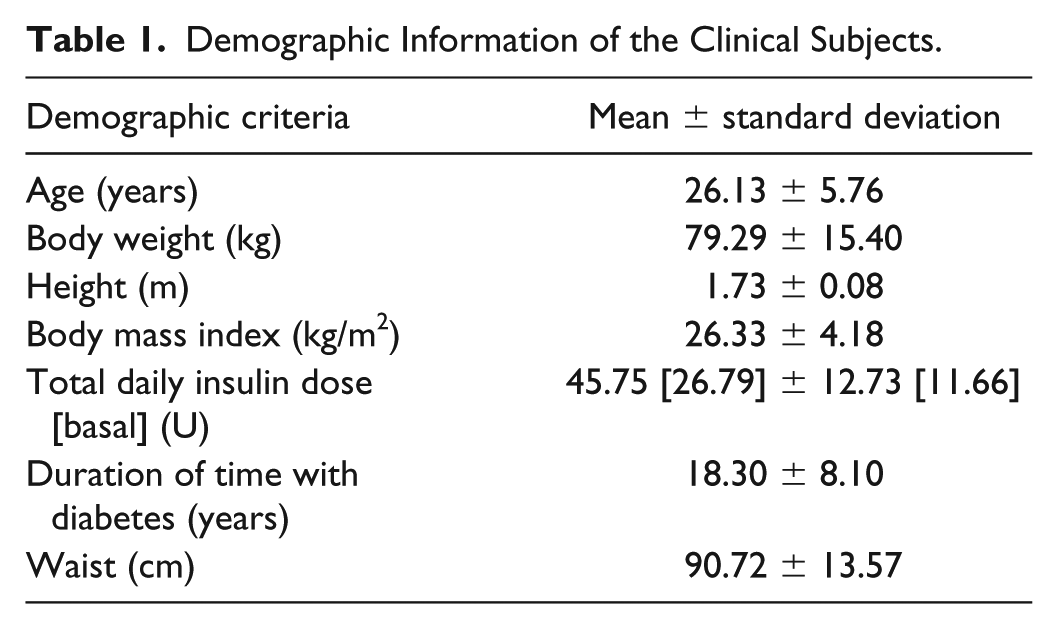
The subjects involved in this study were recruited by the Kovler Diabetes Center, University of Chicago Medical Center, Chicago, Illinois, and were scheduled for a visit at the University of Chicago General Clinical Research Center. The subjects included adults 19-39 years with T1DM. All subjects used CSII pump therapy by adjusting the insulin infusion flow rate based on suggestions from a generalized predictive controller.6,30,31,38 Subjects wore their personal insulin pump, a continuous glucose monitor (CGM; Medtronic Guardian-Real Time Continuous Glucose Monitors [Medtronic, Northridge, CA] or Dexcom G4 Platinum [Dexcom, San Diego, CA]) and a BodyMedia SenseWear Pro3 (BodyMedia, Pittsburgh, PA) armband reporting physiological signals. Each patient’s visit was approximately 60 hours long during the closed-loop experiment using a multimodule multivariable adaptive artificial pancreas system.24,29,39-42 The subjects’ own insulin type and pumps were used during the experiments. Subjects participated in two exercise bouts of 20- to 30-minute sessions before and after lunch. Overall 10 subjects participated in the 15 clinical closed-loop experiments. Table 1 shows the characteristics of the participants of the closed-loop studies.
Demographic Information of the Clinical Subjects.
Results
The efficacy of the proposed adaptive system identification algorithm for identifying high fidelity glycemic models is demonstrated using 15 clinical datasets. The output of the model is the predicted CGM measurements and the inputs are the estimates of the PIC, meal effect (
The MAE and RMSE are calculated based on the following equations:
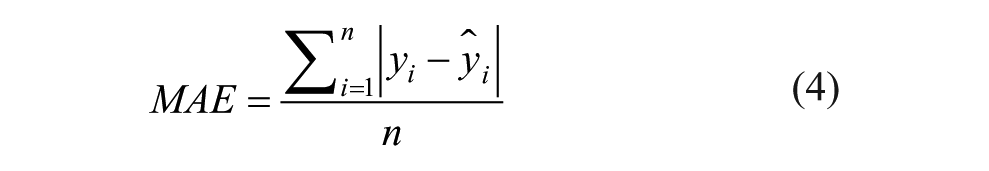
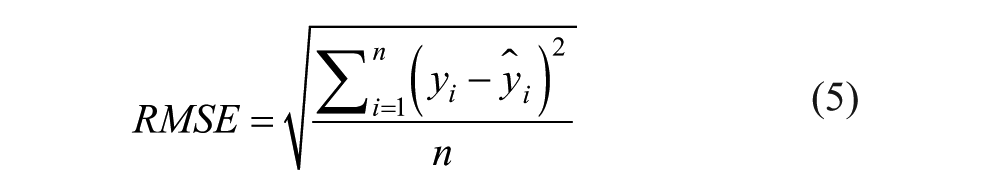
The MAE and RMSE are two of the most common metrics used to measure the prediction accuracy. MAE measures the average magnitude of the errors in a set of predictions, without considering their direction. It’s the average over all samples of the absolute differences between prediction and actual observation where all individual differences have equal weight. RMSE is a quadratic scoring rule that also measures the average magnitude of the error. It is the square root of the average of squared differences between prediction and actual observation. Both the MAE and RMSE express average model prediction error in units of the variable of interest. Both metrics can range from 0 to ∞ and are indifferent to the direction of errors. They are negatively oriented scores, which means lower values are better. Taking the square root of the average squared errors has some interesting implications for the RMSE. Since the errors are squared before they are averaged, the RMSE gives a relatively high weight to large errors. This means the RMSE should be more useful when large errors are particularly undesirable.
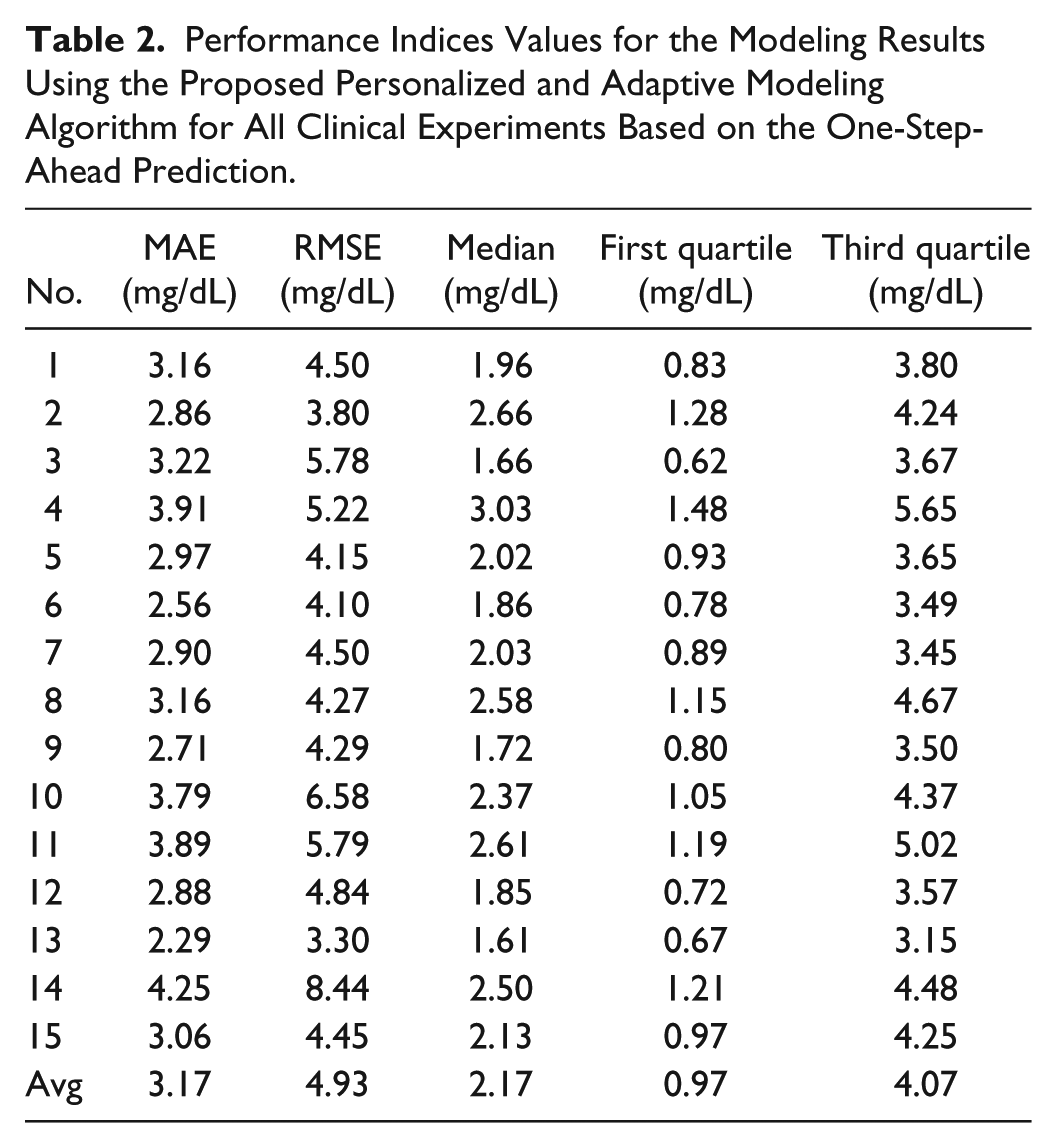
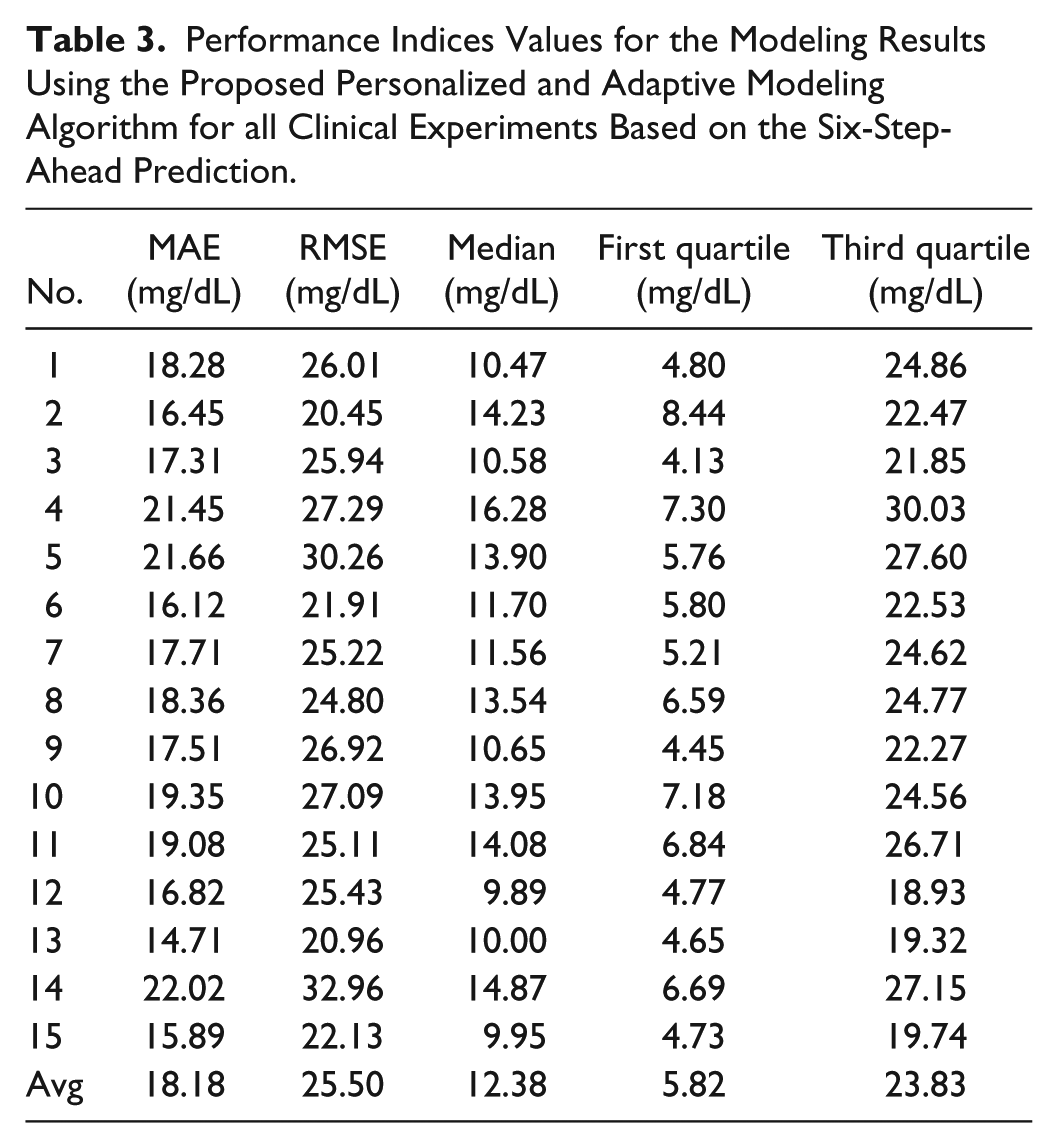
The quantitative results for all 15 clinical experiments are presented in Tables 2 and 3. These performance indices are computed based on one- and six-step-ahead predictions. The one- and six-step-ahead prediction are used to show the performance of the proposed technique in predicting the future outputs. The average RMSE (MAE) for one- and six-step-ahead CGM prediction are 4.93 (3.17) and 25.50 mg/dL (18.18 mg/dL), respectively. The RMSE (MAE) values for the one-step-ahead predicted CGM are less than 8.44 mg/dL (4.25 mg/dl), and for the six-step-ahead predicted CGM is less than 32.96 mg/dL (22.02 mg/dL), which demonstrates the effective performance of the adaptive system identification technique to model the glycemic dynamics.
Performance Indices Values for the Modeling Results Using the Proposed Personalized and Adaptive Modeling Algorithm for All Clinical Experiments Based on the One-Step-Ahead Prediction.
Performance Indices Values for the Modeling Results Using the Proposed Personalized and Adaptive Modeling Algorithm for all Clinical Experiments Based on the Six-Step-Ahead Prediction.
The proposed adaptive system identification method is able to provide stable and adaptive state-space models at each sampling time. Figure 2 shows the predicted and actual CGM values along with the inputs of the model (the estimates of the PIC and meal effect parameter, and the energy expenditure values) and maximum norm of model eigenvalues for a selected experiment. The results for the clinical experiments (Figure 2) show that the maximum norm of eigenvalues of the models is maintained within the unit circle and evolve over time to best describe the available input and output data, thus ensuring stability of the identified models. Furthermore, the estimated and predicted CGM values closely track the actual CGM measurements. The state-space model is identified such that the available input-output data is well characterized by the system realization while the stability and fidelity of the identified models is guaranteed. Compared to only using the PIC as a model input, additional inputs capturing the effects of meals and physical activity, such as the estimated meal effect parameter and energy expenditure, significantly improve the prediction ability of the modeling approach (Figure 3). Although the model using PIC and meal effect as inputs does not show statistical significance compared to the proposed model using PIC, meal effect and energy expenditures as input variables, the standard deviation is biased by the outliers, which influences the statistical hypothesis test. To elucidate this bias caused by the outliers, a one-sided test for statistical significance is used to compare the model (b) with the proposed model (d), which shows statistical significance (P value = 0.04 for MAE and 0.029 for RMSE). The results demonstrate that the use of the PIC as a filtered insulin input, the incorporation of estimates of the meal effect obtained through the nonlinear observer algorithm, and the inclusion of energy expenditure as an auxiliary input from the BodyMedia SenseWear armband improve the identified glucose-insulin dynamic models.
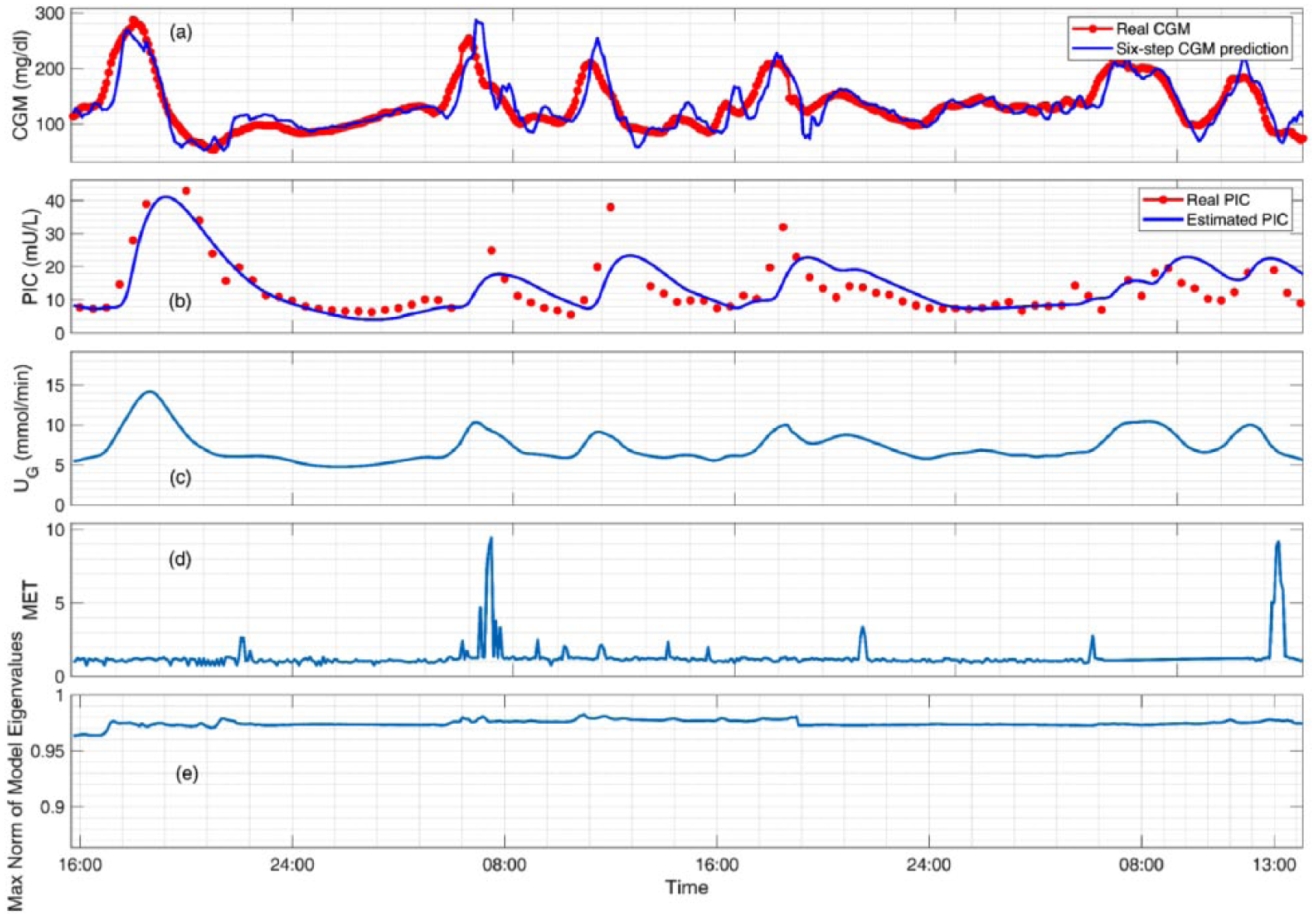
Modeling results for a select experiment (subject 7) showing (a) the six-step (30 minutes ahead) predicted and actual CGM values, (b) the estimated plasma insulin concentration and actual PIC values (not used in model development), (c) the estimated meal effect parameter for capturing unannounced meals, (d) the energy expenditure in units of metabolic equivalent (MET) for incorporating unannounced exercise, and (e) the maximum norm of eigenvalues of the stable time-varying identified models.
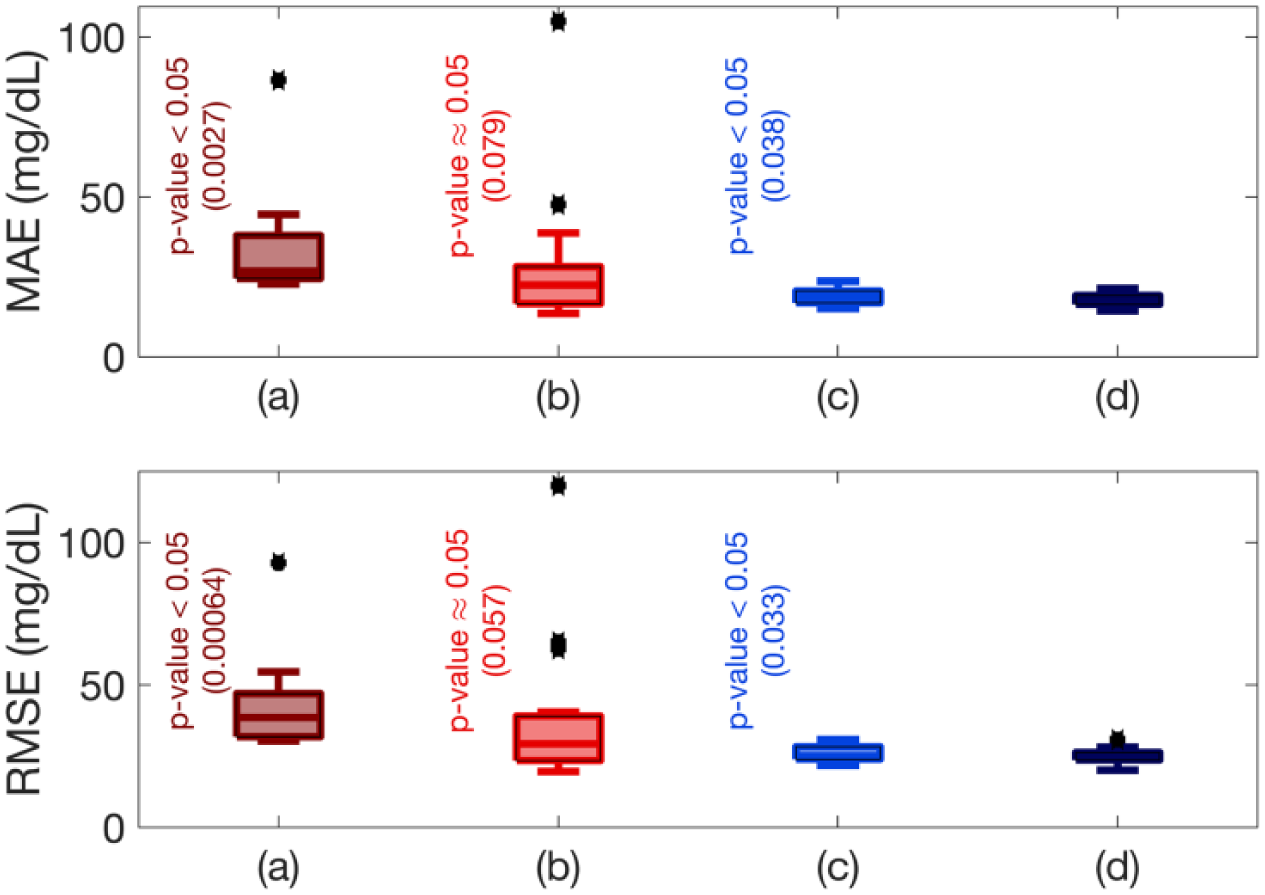
Comparison of six-step (30 minutes ahead) prediction accuracies across all experiments using the mean absolute error (MAE) and root mean square error (RMSE) for prediction with different model inputs: (a) PIC; (b) PIC and energy expenditure; (c) PIC and meal effect; and (d) PIC, energy expenditure, and meal effect. All hypothesis tests computed using two-sided paired t-tests at 5% significance level and are relative to the proposed model (d), which considers PIC, energy expenditure, and meal effect as input variables.
The Clarke error grid approach is used to assess the clinical significance of differences between glucose predictions generated by the proposed technique and measured values.43-46 The method uses a Cartesian diagram, in which the six-step-ahead CGM predictions obtained by the presented technique are displayed on the y-axis, whereas the real (measured) CGM values are displayed on the x-axis. The diagonal represents the perfect agreement between the two, whereas the points below and above the line indicate, respectively, overestimation and underestimation of the actual values. Zone A (acceptable) represents the glucose values that deviate from the real values by ±20% or are in the hypoglycemic range (<70 mg/dl), when the reference is also within the hypoglycemic range. The values within this range are clinically exact and are thus characterized by correct clinical treatment. Zone B (benign errors) is located above and below zone A; this zone represents those values that deviate from the reference values, which are incremented by 20%. The values that fall within zones A and B are clinically acceptable, whereas the values included in areas C-E are potentially dangerous, and there is a possibility of making clinically significant mistakes. Based on the results presented in the Table 4, we can see that 98.87% of values fall within zones A and B. There is no cases fall within zone E that is the most dangerous case and just 1.13% of values fall within zones C and D. Figure 4 shows of the Clarke error grid analysis for a select experiment (subject 7).
Percentages in Each Zone Based on Clarke Error Grid Analysis for the Modeling Results Using the Proposed Personalized and Adaptive Modeling Algorithm for All Clinical Experiments Based on the Six-Step-Ahead Prediction.
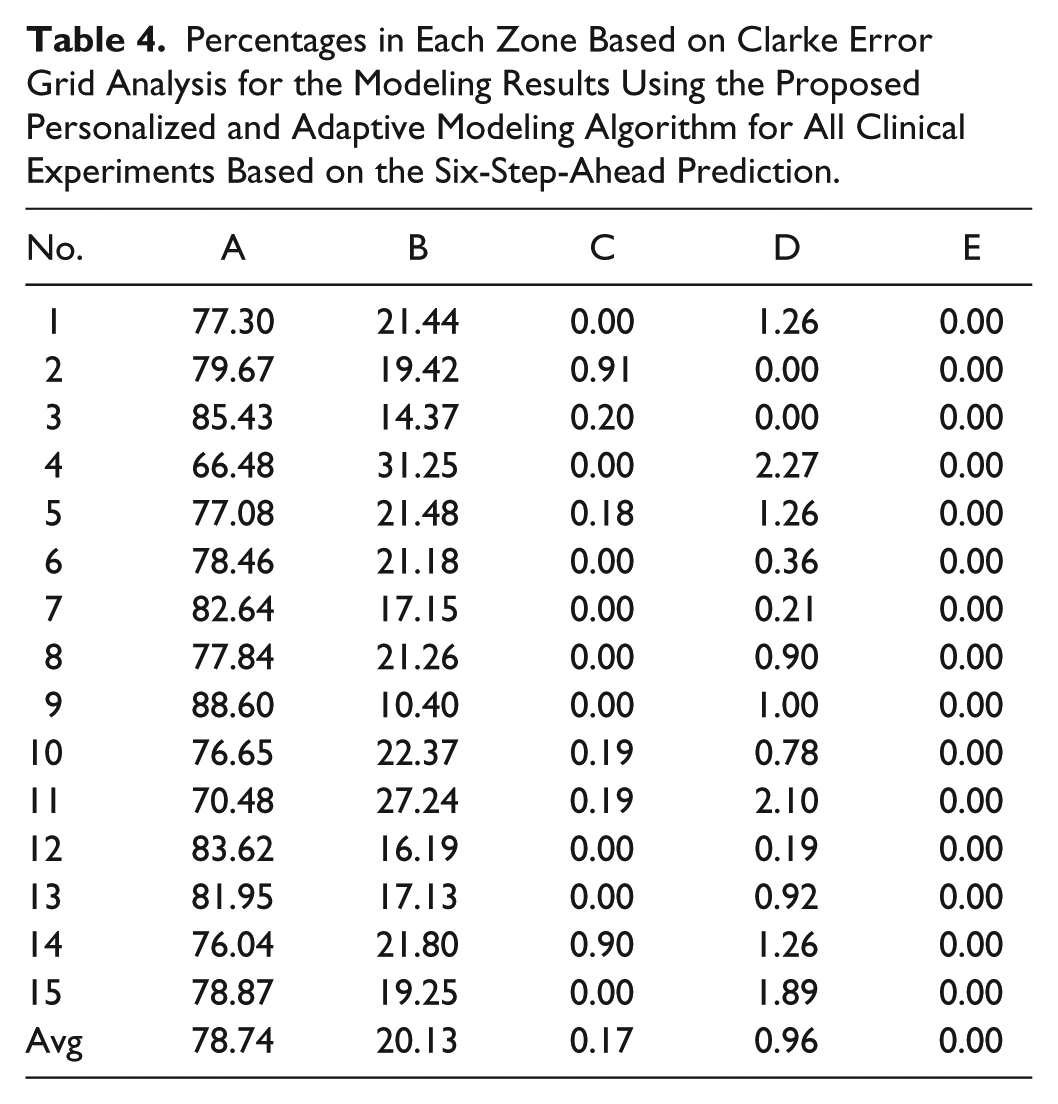
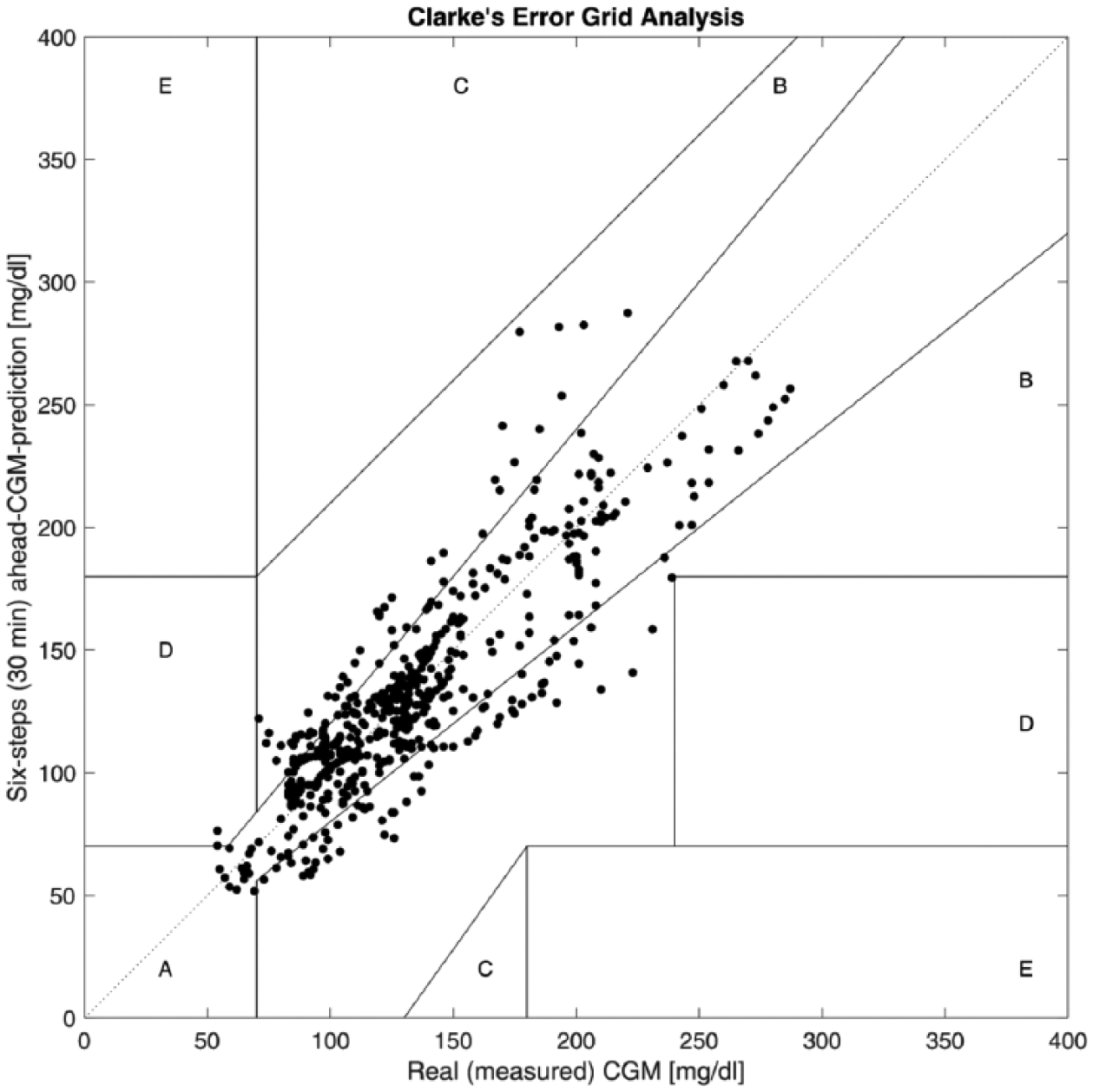
Figure of the Clarke error grid analysis for a select experiment.
In addition to the MAE and RMSE performance indices and Clarke error grid analysis, the low risk of missing hypoglycemia (below 70 mg/dL) or hyperglycemia (above 200 mg/dL) is a key feature of an efficient and reliable identified glucose-insulin dynamic model. We have also calculated the results for the percentage of the cases which the hypoglycemia or hyperglycemia are missed. Nine out of 15 experiments have CGM values below 70 mg/dl, and the average and standard deviation of missed hypoglycemia for these 9 experiments are 24.09% ± 16.50%. Thirteen out of 15 experiments have CGM values above 200 mg/dl, and the average and standard deviation of missed hyperglycemia for these 13 experiments are 07.83% ± 05.29%.
Discussion
A physiological model derived from the insulin compartment subsystem of Hovorka’s model is incorporated in this work with adaptive data-driven models developed using a recursive subspace identification technique. A computationally efficient nonlinear observer is used to estimate the time-varying parameters and the meal effect, thus facilitating the explicit incorporation of unannounced meals. The results for the proposed integrated compartment model with adaptive subspace identification techniques are promising. The proposed adaptive models may be used in the design of a model-based predictive control algorithm.
There are several benefits to the integration of compartment models with adaptive models. First, the PIC estimates are adaptive and individualized to particular patients, thus providing accurate real-time estimates of the amount of active insulin present in the bloodstream. The plasma insulin information can be readily used to impose constraints on the insulin dosing computation algorithm. Second, in contrast to the discrete basal and acute bolus insulin variations, the estimated PIC from the compartment model readily provides a more appropriate and filtered input variable for model identification. Third, the adaptive subspace identification technique renders the recursively identified glycemic models valid over a diverse range of daily activities without requiring onerous and obscure information such as the amount of carbohydrate consumption or quantification of physical activity levels.
Although meal and exercise announcements may improve the prediction ability of the proposed approach, unannounced meals and exercise are considered in this work to refrain from the inconvenience of user inputs and avoid the potential hazards of erroneous entries. The proposed adaptive modeling approach with an integrated compartment model utilizes additional auxiliary input variables such as the energy expenditure bio-signals to consider the effects of physical activity on glycemic dynamics.
Any insulin model capable to translate the abrupt bolus and discrete basal changes into estimates of PIC can be used. In this work, insulin subsystem of Hovorka’s model has been used since the PIC estimator is designed based on the Hovorka’s glucose-insulin dynamic model and we have developed personalization techniques based on the PLS models to individualize the parameters of this insulin subsystem for each specific patient.
PIC measurements are available from the clinical experiments, however, this information is not used in the development of the PIC estimation model and the on-line and real-time prediction of future CGM values. We use the clinical measurements of PIC to validate the estimates of the PIC obtained through the nonlinear simultaneous state and parameter estimation technique. As the PIC estimates are accurate, we employ the PIC estimates in the identification of adaptive state-space models. This ensures that only CGM measurements, infused insulin information, and physiological measurements from an nonintrusive wearable activity tracker are used in the on-line modeling and predictions of future glucose values.
For any data-driven modeling approaches where the model is obtained based only on input and output data measured form the underlying system, the quality and accuracy of the data play an important role on the performance of identified model. The time and frequency of calibrations have a significant effect on accuracy of the CGM measurements and consequently on the predictions accuracy. Subjects calibrated the CGM at least twice each day according to manufacturer instructions. The participants used their personal glucometers, and the CGM calibration may have occurred in nonideal glucose trends. CGM accuracy is greatest when the CGM is calibrated during a stable glucose period (flat arrows on CGM), but this was not always the case when the subjects chose to calibrate.
The steady-state value of the gut appearance rate (UG) parameter should be zero during fasting time, provided all model parameters are personalized for each subject and the basal insulin rate stabilizes the BGC around a steady-state value. The purpose of considering the UG in the PIC estimator is to handle any unknown disturbances present in the system, including the effects of unannounced meals (such as main meals or fast-acting carbohydrates for treatment of hypoglycemia) or any other uncertainties in the model parameters (such as EGP0,
The most challenging periods for the CGM predictions are the times that exercise and meal affect the glucose-insulin dynamics. Since, any information about these unknown disturbances such as their time, intensity, amount and duration are not known in advance, the accuracy of multistep-ahead predictions that is just based on current and past data decreases during these periods until the model parameters are updated based on measurements in those specific periods. One of the reasons that a recursively updated model is necessary for a fully automated artificial pancreas system (unannounced meal and exercise) is to update the model parameters at each sampling time using new measurements to adapt the model for that specific period. As shown in Figure 3, including the MET values and meal effects estimates improves the performance of the identified model for predicting the CGM values. In addition to analyzing the global error metrics, investigating the local error variabilities around the unannounced meals and exercises can be useful.
To design a fully automated artificial pancreas system that can operate without any user input, in addition to CGM data, information that could help the artificial pancreas control system to detect exercise and act properly for that period is needed. This information can be physical activity signals measured by wearable devices. To achieve this, different bio-signals have been used in artificial pancreas systems. The metabolic equivalent task (MET), or simply metabolic equivalent, is a physiological measure expressing the energy cost of physical activity (PA) and is defined as the ratio of metabolic rate (and therefore the rate of energy consumption) during a specific PA to a reference metabolic rate. MET is used as a means of expressing the intensity and energy expenditure of activities in a way that is comparable among people. We have developed a real-time MET estimation algorithm by using noninvasive measurements of physiological variables by our research group at Illinois Institute of Technology. In this approach, these MET values are derived using heart rate, galvanic skin response, skin temperature, blood volume pulse, and accelerometer.
The performance of proposed technique has been verified using different indices based on one-step-ahead predictions and six-step-ahead predictions. Different indices are considered to show the ability of the identified model in tracking the real measurements as well as predicting the future trend in CGM measurements. As we extend the prediction horizon, the accuracy of the predicted values decreases (as it can be seen from 1 step to 6 steps ahead of prediction) due to several reasons including unknown phenomena like exercise, stress or meal that may take place. So choosing the prediction horizon is a trade-off between knowing long-term information about the future measurements and their accuracy.
For few clinical experiments (e.g. No. 14 in Table 3) the relative high value of performance indices (MAE and RMSE values) is observed in comparison to the average values. The proposed technique is a data-driven technique and the quality of the measured data (eg, CGM and physiological variables that may be corrupted with noise and outliers) have a significant effect of the performance, as well as the fact that any information about the meal and exercises are not used. Predicting such abnormal measurements is very challenging and updating the glucose-insulin dynamic models using poor quality of data may result in bad performance in predicting the future trend of measurements.
One of the limitations of our clinical experiments is the exercise times are close to the meal times. Although people with type 1 diabetes usually consume carbohydrates before, during, and after exercise to avoid hypoglycemia and the proposed approach demonstrates that the effects of meals and exercise have well been captured by the modeling technique, in our future clinical work, the clinical experiments will be conducted to test the algorithm under different conditions such as. no meals very close to the exercise time.
Conclusion
An adaptive system identification algorithm is proposed to determine reliable dynamic glycemic models for use in AP systems. The proposed modifications to the recursive system identification algorithm assist in handling uncertain disturbances such as unannounced meals and exercise, stochastic measurement noise, and other adverse phenomena. The efficacy of the proposed approach is demonstrated using case studies involving the modeling of time-varying glucose-insulin dynamics in clinical experiments, and the results demonstrate the approach is able to identify reliable, time-varying, and personalized glycemic models.
Footnotes
Appendix A
Appendix B
Appendix C
Appendix D
Abbreviations
AP, artificial pancreas; CGM, continuous glucose monitoring; CSII, continuous subcutaneous insulin infusion; MAE, mean absolute error; MET, metabolic equivalent task; PA, physical activity; PAST, projection approximation subspace tracking; PBSID, predictor-based subspace identification; PIC, plasma insulin concentration; PLS, partial least squares; RLS, recursive least squares; RMSE, root mean square error; SVD, singular value decomposition; T1DM, type 1 diabetes mellitus; UKF, unscented Kalman filter; UT, unscented transformation; VARX, vector autoregressive with exogenous inputs.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Financial support by NIH with grants NIDDK DP3 DK101075-01 and DP3 DK101077-01 is gratefully acknowledged.