Abstract
Previous studies have shown differences in the microbiota of patients with Parkinson's disease (PD) compared to healthy controls (HC). To deduce a possible causality, it is highly relevant to examine changes in the prodromal phase. This study investigated the microbiome in stool samples of individuals with isolated REM sleep behavior disorder (iRBD, n = 32) compared to clinical PD (n = 23) and HC (n = 34) and showed significant changes of beta-diversity in PD and iRBD patients compared to HC (p = 0.025; p = 0.003), with an increase in proinflammatory species in iRBD and PD and decrease in SCFA-producing bacteria in PD.
Plain language summary
Previous studies have shown that the bacteria in the gut of patients with Parkinson's disease (PD) differ from those of healthy controls (HC). However, it is still unclear whether these changes are a cause or consequence of the disease. Therefore, we aimed to examine also the gut bacteria in individuals with a high risk to develop PD in the future, i.e., individuals with isolated REM sleep behavior disorder (iRBD). We were indeed able to confirm that differences in the gut microbiome are already present in people with iRBD, as important evidence that these changes are already present before the onset of clinical symptoms and that they may be important for the development of the disease.
Introduction
Current research suggests that the gut-brain-axis plays an important role in the etiology and progression of Parkinson's disease (PD). 1 Many studies have shown significant differences in the microbiota of PD patients compared to healthy controls (HC), leading to pro-inflammatory dysbiosis.2,3 However, it remains unclear whether dysbiosis is the cause of the disease or a mere consequence of PD-related factors, such as constipation or intake of dopaminergic medication. Since most studies to date have been conducted in clinical PD and thus in the presence of these multiple influencing factors, it remains difficult to deduce causality. Therefore, it is of high importance to include potentially influencing factors in case-control analysis of gut microbiota and to investigate changes already in the prodromal phase of the disease. In this respect, initial studies have shown that differences in microbiota composition already occur in individuals with isolated REM sleep behavior disorder (iRBD). This study aimed to further investigate group differences of the microbiota composition as well as the effects of potentially influencing factors in individuals with iRBD, clinical PD and HC.
Methods
Study population
IRBD and PD patients were recruited from the Department of Neurology, Kiel, Germany. Spouses and volunteers served as HC. Inclusion criteria were: (1) diagnosis of PD according to the Movement Disorder Society (MDS) clinical diagnostic criteria for PD 4 or diagnosis of polysomnographic proven iRBD, (2) ability to give informed consent. Exclusion criteria included: (1) Presence of chronic inflammatory bowel disease, (2) intake of antibiotics in the last six weeks, (3) condition after organ transplantation, (4) presence of active cancer, (5) diagnosis of a neurological disorder in HC. This study was approved by the local ethics committee of the University of Kiel. All participants gave informed consent (ethical vote D440/18).
Clinical assessments
Every participant provided information about sex, age, symptom onset and time of diagnosis of iRBD or PD, height and weight (used to calculate the body mass index, BMI). Participants answered a questionnaire on comorbidities, medication and intake of antibiotics and indicated to what extent they suffered from constipation (“not at all”, “sometimes”, “often”, “very often”). Motor symptoms in the iRBD group were assessed using the MDS Unified Parkinson's Disease Rating Scale part III (MDS-UPDRS-III). 5 IRBD patients also filled out the RBD screening questionnaire (RBDSQ). 6 Severity of depression was assessed using the Beck depression inventory (BDI). 7
Stool sampling and processing
All stool samples were taken between 2018 and 2021. The native samples were collected by the patients at home and sent back via postal service to the University Hospital Kiel. After arrival they were stored in a freezer (−80°C) until further processing. DNA extraction and 16S rDNA sequencing were performed as described before. 8
Sequence data processing
Data processing was performed using the DADA2 9 workflow resulting in abundance tables of amplicon sequence variants (ASVs) according to a workflow adjusted for the V1–V2 region. 10 Resulting ASVs underwent taxonomic annotation using the Bayesian classifier provided in DADA2 and using the Ribosomal Database Project (RDP) version 16 release. 11 One sample with fewer than 10,000 sequences was not considered for further analysis.
Statistical analysis
The R package “vegan” (v2.5-1) 12 was used to investigate alpha-diversity. Alpha-diversity of the samples was measured by Chao1 and Shannon index. The association between microbial diversity and compared groups was tested via Wilcoxon rank-sum tests. For analysis on beta-diversity, non-metric multidimensional scaling (NMDS) was performed with Bray–Curtis dissimilarity as implemented in the R package vegan v2.5-1. A permutational multivariate analysis of variance (PERMANOVA) 13 was then performed on the distance matrix to assess the group effects on the variance between microbial communities. The PERMANOVA was performed based on the Adonis function of the R package vegan v2.5-1, with 999 permutations.
Differential abundance of taxa between PD, iRBD and HC was determined at the ASV level using the “DESeq2” package in R, 14 for each ASV which was covered by at least 50 read pairs and showed a minimum prevalence of 5% in all samples. Missing data were included as partial data in “DESeq2” models. Age and sex were included as covariates as the groups differed significantly in these factors (see Table 1). Additionally, BMI was included as covariate. Results were expressed as log2 fold change in PD and iRBD relative to HC. False discovery rate (FDR) was used for multiple testing correction. To calculate the variation explained by each of our collected host factors, an Adonis test was performed, implementing the R package vegan v2.5-1 with 999 permutations. In addition, we carried out a correlation analysis for the iRBD-specific significant variables to determine which ASVs were significantly influenced by them.
Demographic and clinical data.
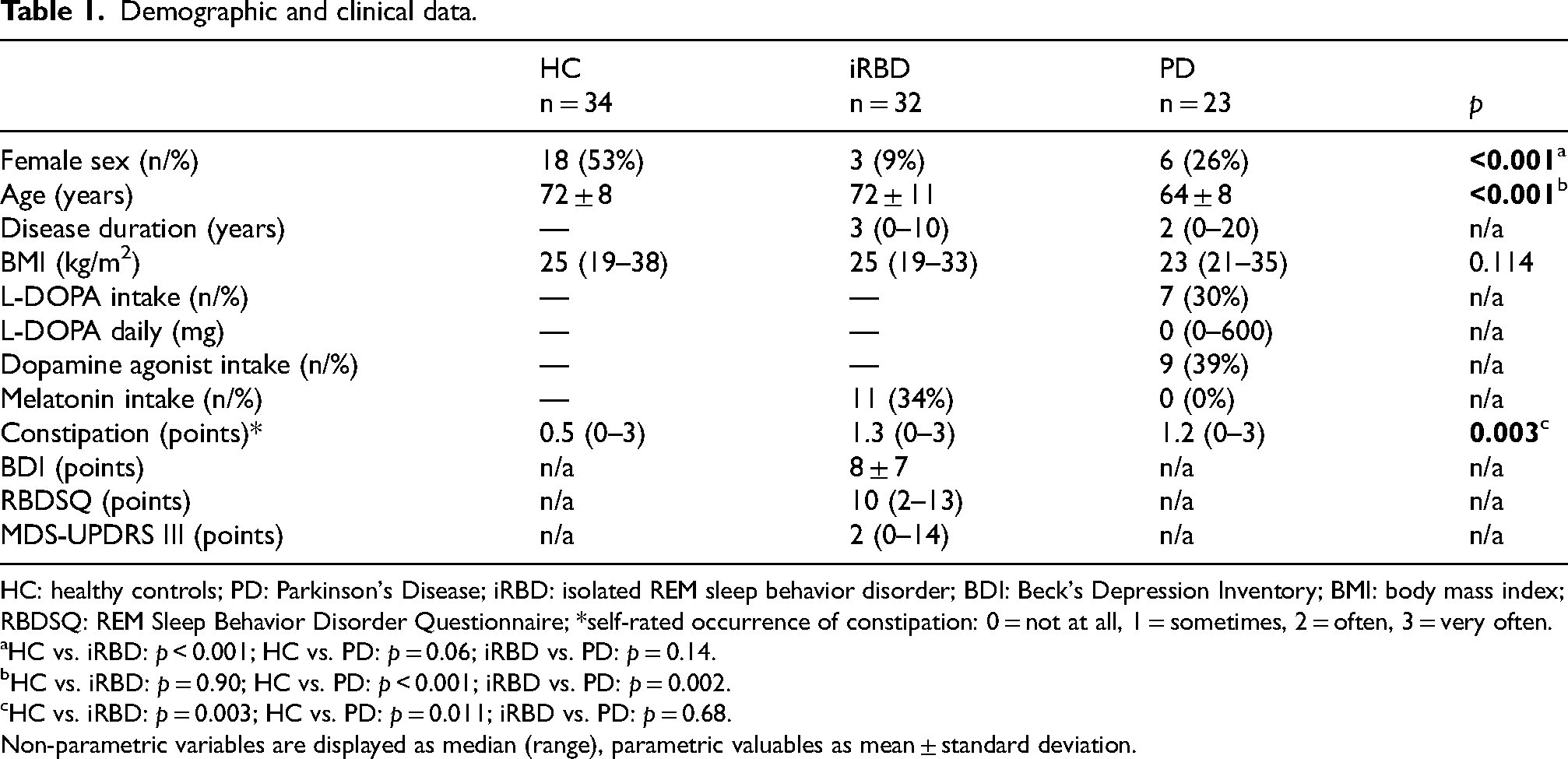
HC: healthy controls; PD: Parkinson's Disease; iRBD: isolated REM sleep behavior disorder; BDI: Beck's Depression Inventory; BMI: body mass index; RBDSQ: REM Sleep Behavior Disorder Questionnaire; *self-rated occurrence of constipation: 0 = not at all, 1 = sometimes, 2 = often, 3 = very often.
HC vs. iRBD: p < 0.001; HC vs. PD: p = 0.06; iRBD vs. PD: p = 0.14.
HC vs. iRBD: p = 0.90; HC vs. PD: p < 0.001; iRBD vs. PD: p = 0.002.
HC vs. iRBD: p = 0.003; HC vs. PD: p = 0.011; iRBD vs. PD: p = 0.68.
Non-parametric variables are displayed as median (range), parametric valuables as mean ± standard deviation.
Results
Study cohort
Stool samples from 32 iRBD patients, 23 PD patients and 34 HC were analyzed. Demographic and clinical data are shown in Table 1.
Alpha-diversity
No consistent significant differences in alpha-diversity were seen between the groups. Chao1-Index differed significantly between HC and PD (p = 0.027), however, analysis of Shannon diversity, taking also the abundance of microbes into account, revealed no significant differences between the groups.
Beta-diversity
Significant differences of beta-diversity were seen in PD patients compared to HC (p = 0.025) and in iRBD patients compared to HC (p = 0.003) (Figure 1). There were no significant differences in the comparison of PD and iRBD patients (p = 0.24).
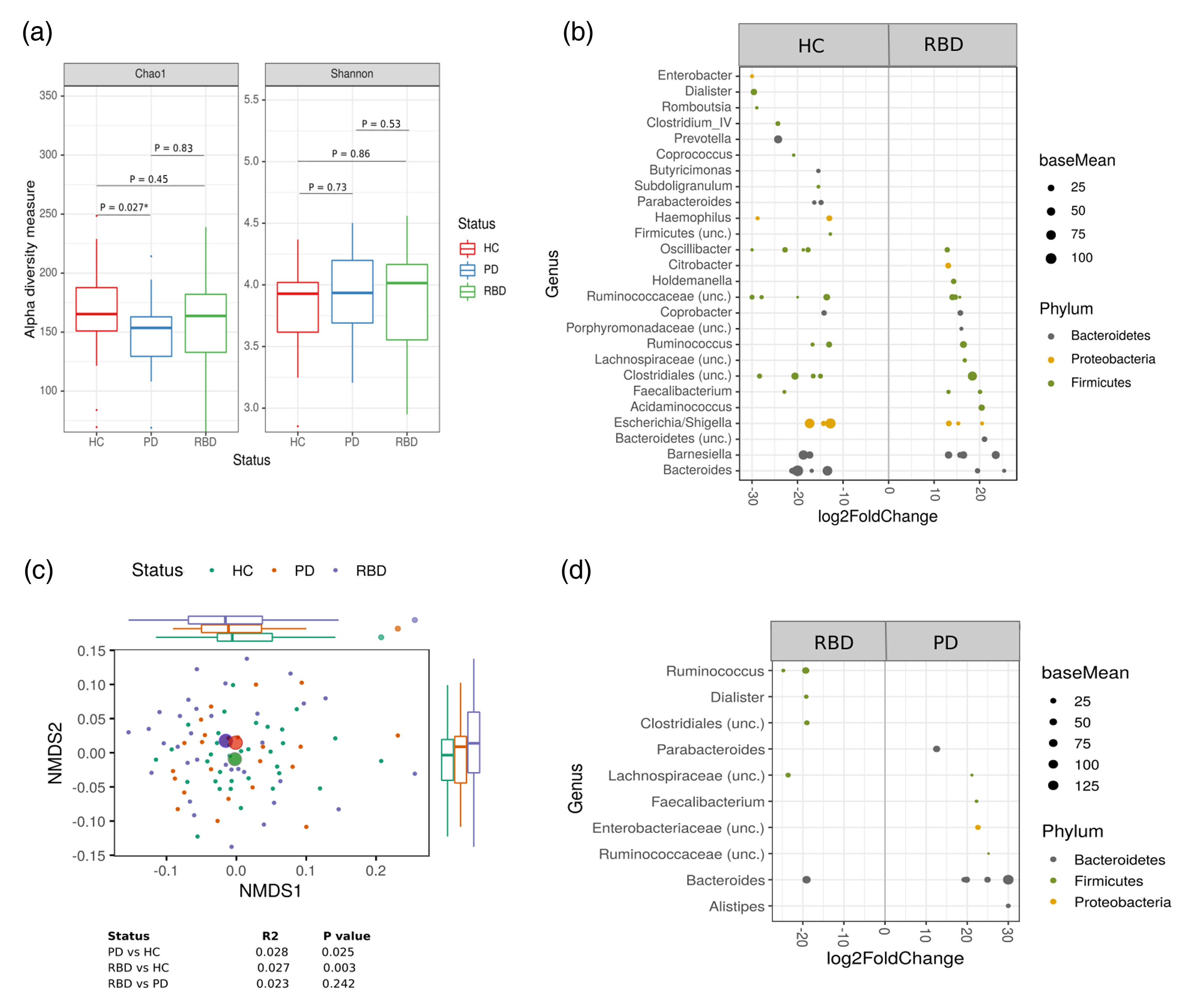
Alteration of fecal microbiota among groups. (a) Alpha-diversity measurements (Chao1 and Shannon index) for patients with PD and iRBD and versus HC. (b) NMDS plot of Bray–Curtis dissimilarities display sample-wise differences in community composition between groups (indicated by color, group centroids indicated by large symbols). Marginal boxplots display the grouped distribution of individuals/samples along the respective dimension of the NMDS plot analysis of relative sample ASV composition. (c) Differential abundance analysis identified 25 ASVs that were increased and 45 ASVs that were decreased in iRBD relative to HC (p(FDR) < 0.05). ASVs to the right of the zero line are more abundant and ASVs to the left of the zero line are less abundant in iRBD compared with control groups. (d) 10 ASVs increased and 6 ASVs decreased in patients with PD relative to iRBD participants. ASVs to the right of the zero line are more abundant and ASVs to the left of the zero line are less abundant in PD compared to iRBD. Each point represents an ASV. ASV: amplicon sequence variants; NMDS2: non-metric multidimensional scaling; PD: Parkinson's disease; HC: healthy controls; iRBD: isolated REM sleep behavior disorder.
Differential abundance
Analysis of differential abundance showed reduced levels of Clostridium Cluster IV and Prevotella in PD and iRBD compared to HC, while some bacterial members of Bacteroides as well as Citrobacter and Acidaminococcus were more abundant in iRBD and PD compared to HC. Moreover, short-chain fatty acids (SCFA)-producing Lachnospiraceae and Faecalibacterium were more common in iRBD compared to HC, but not in PD patients. On the contrary, Faecalibacterium was significantly reduced in PD patients compared to HC. Additionally, Escherichia/Shigella were more abundant in PD compared to HC.
Comparing the microbiota of iRBD and PD, slight differences were observed. In iRBD, SCFA-producing bacteria like Ruminococcus, Dialister, Clostridiales and some bacterial members of Lachnospiraceae were more abundant compared to PD. Enterobacteriacea, Bacteroides and Alistipes were more abundant in PD compared to iRBD. However, there were also some SCFA-producing bacteria like Faecalibacterium and Ruminococcaceae more abundant in PD.
Factors influencing microbiota composition
Of all investigated variables, sex, age, RBDSQ score as well as dopamine agonist intake significantly influenced beta-diversity, while Levodopa medication, disease duration, MDS-UPDRS 3 sum score, BDI sum score, melatonin intake, BMI and self-reported constipation had no significant influence.
Correlation analysis
Correlation analysis of the iRBD-specific significant variables showed that the RBDSQ score was positively correlated with approximately 30 ASVs. No significant correlations were seen for the intake of dopamine agonists in the PD cohort.
Discussion
This study confirms the previously reported finding that microbiota composition already shows significant differences in iRBD patients compared to HC and that these differences partly resemble the microbiota composition in clinical PD patients.2,3 Factors influencing the microbiota composition were age, sex, intake of dopamine agonists and the self-perceived severity of iRBD (RBDSQ score), while, surprisingly, levodopa use and presence of self-reported constipation had no influence.
Comparison of microbiota composition revealed no consistent changes in alpha-diversity between groups; however, analysis of beta-diversity significantly differed between PD and iRBD patients compared to HC. This finding is consistent with previous small studies in iRBD patients, that detected similar differences in beta-diversity.15,16 Interestingly, differences in beta-diversity were even more pronounced in the iRBD group. These results are particularly relevant when considering the currently much discussed hypothesis of different prodromal subtypes, the “brain-first” vs. “gut-first” subtype. 17 According to this hypothesis, α-synuclein pathology in iRBD patients follows a gut to brain progression. Thus, a more peripheral (gut/microbiome) pathology would be expected when analyzing an isolated iRBD cohort, than in a “mixed” clinical PD group that includes both “brain-first”- and “gut-first” individuals (as assumed here, since no stratification for iRBD was performed).
Similar to previous studies15,18 iRBD and PD patients showed increased levels of proinflammatory genera compared to HC (bacterial members of Bacteroides, Citrobacter, Escherichia/Shigella). In addition, the SCFA-producing genera Prevotella and Clostridium Cluster IV were reduced in both iRBD and PD patients. In this regard, previous studies also observed a change of microbiota associated with SCFA-production, but not the same genus (among others Haemophilus and Roseburia).15,19 Interestingly, when comparing iRBD and PD patients, some SCFA-producing genera (e.g., Ruminococcus, Dialister, Clostridiales, Lachnospiraceae) were more abundant in iRBD than in PD, which is consistent with one of the conclusions of a recent meta-analysis stating that decreases in SCFA-producing bacteria may indicate the conversion from prodromal to clinical PD. 20
As a further objective, this study investigated potential confounders. While age, sex, intake of dopamine agonists and RBDSQ sum score were confirmed as confounding factors, levodopa intake, BMI and self-reported constipation were not significant influencing factors. Like previous contradictory study results, this study emphasizes once again that neither levodopa nor constipation are proven factors influencing the microbiome in PD.19,21–24 It was shown before that the intake of dopamine agonists could influence microbiota composition. 25 However, correlation analysis did not reveal significant correlations with any ASV. In contrast, the RBDSQ score was positively correlated with the abundance of some bacteria. Among others, an increase in the RBDSQ score (reflecting more pronounced RBD symptoms) was associated with increased concentrations of proinflammatory Bacteroides and Escherichia/Shigella. In this context, it must be mentioned that this is a very preliminary result, as the RBDSQ score is actually intended as a screening tool and not as a measurement tool for the severity of iRBD. However, in light of previous studies on the bidirectional relationship between the gut microbiome and sleep,26,27 it may be of great interest to further investigate whether changes in the microbiome could be a consequence of iRBD or, conversely, proinflammatory changes in the gut microbiome could promote iRBD and an associated more malignant course of PD and might thus be a future target for interventional therapies.
There are several limitations to this study. First, this analysis only included a small sample size and studies with larger sample sizes are urgently needed to validate our findings. Second, iRBD patients convert to PD over many years. Hence, longitudinal studies are crucial to evaluate the influence of microbiome changes on disease progression. Additionally, metagenomic or metatranscriptomic data sets instead of 16S rDNA sequencing data are needed to evaluate metabolic pathways of the gut microbiota. Furthermore, as the focus of this study was on the iRBD cohort, we have limited clinical data from PD patients. Particularly in view of the here found potential influence of the severity of iRBD on the microbiome it would be highly interesting for future studies to investigate the effects of specific sleep disorders i.e., iRBD on the microbiome in clinical manifest PD patients.
In summary, this study emphasizes previous observations that an increase in pro-inflammatory and a decrease in SCFA-producing bacteria can already be observed in the prodromal phase of PD. However, considering especially the different species observed in previous studies, it remains essential to validate our findings in larger and longitudinal cohorts.
Supplemental Material
sj-docx-1-pkn-10.1177_1877718X251354931 - Supplemental material for Differences in intestinal microbiota in Parkinson's disease and isolated REM sleep behavior disorder
Supplemental material, sj-docx-1-pkn-10.1177_1877718X251354931 for Differences in intestinal microbiota in Parkinson's disease and isolated REM sleep behavior disorder by Alba Troci, Julienne Haas, Anna Weiß, Sebastian Heinzel, Andre Franke, Daniela Berg, Corinna Bang and Eva Schaeffer in Journal of Parkinson's Disease
Supplemental Material
sj-docx-2-pkn-10.1177_1877718X251354931 - Supplemental material for Differences in intestinal microbiota in Parkinson's disease and isolated REM sleep behavior disorder
Supplemental material, sj-docx-2-pkn-10.1177_1877718X251354931 for Differences in intestinal microbiota in Parkinson's disease and isolated REM sleep behavior disorder by Alba Troci, Julienne Haas, Anna Weiß, Sebastian Heinzel, Andre Franke, Daniela Berg, Corinna Bang and Eva Schaeffer in Journal of Parkinson's Disease
Footnotes
Acknowledgements
We would like to thank all participants involved in sampling of biomaterial for this study—without them such a study would not be possible. Further, we would like to thank the staff of the IKMB microbiome laboratory for processing samples as well as IKMB sequencing facilities for excellent technical support. We additionally acknowledge financial support by Land Schleswig-Holstein within the funding programme Open Access Publikationsfonds.
ORCID iDs
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: J.H. has been supported by the Faculty of Medicine, University of Kiel, Germany. Microbiota sequencing and data analysis received infrastructure support from the DFG Research Unit 5042 “miTarget” and the DFG Excellence Cluster 2167 “Precision Medicine in Chronic Inflammation” (PMI).
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability statement
Data supporting the findings of this study are available on request from the corresponding author. The data are not publicly available due to privacy and ethical restrictions.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
