Abstract
Purpose
This study investigated causal relationships and underlying mechanisms between serum DNA repair proteins and liver cancer to identify biomarkers for clinical management.
Methods
A two-sample bidirectional Mendelian randomization (MR) design was employed. We performed inverse variance weighted (IVW) analysis, followed by protein quantitative trait loci (pQTL), colocalization, and pathway enrichment analyses to explore biological mechanisms.
Results
MR analysis revealed significant associations: NBR1 (IVW: OR = 2.28, 95%CI: 1.17–4.45, P = 0.015) and RAD51 (IVW: OR = 2.16, 95%CI: 1.12–4.15, P = 0.021) were risk factors. PARP11 was protective for hepatocellular carcinoma (OR = 0.48, 95%CI: 0.25–0.93, P = 0.030) but a risk for intrahepatic cholangiocarcinoma (OR = 1.86, 95%CI: 1.04–3.33, P = 0.038). Mechanistically, pQTL and colocalization identified rs2793568 as a key regulator of PARP1, which was enriched in base excision repair pathways. Sensitivity analyses confirmed the robustness of these findings.
Conclusion
Serum NBR1, RAD51, and PARP11 are potentially causal in liver cancer pathogenesis. Specifically, the genetic regulation of PARP1 highlights a critical DNA repair mechanism, supporting their utility as predictive biomarkers.
Keywords
Introduction
Liver cancer remains a major global health burden despite advances in cancer detection, molecular classification, and targeted therapy. Worldwide, cancer continues to be a leading cause of morbidity and mortality, with research increasingly focusing on genetic susceptibility, tumor microenvironment, and molecular targets that underlie cancer initiation and progression. Recent progress in precision oncology and high-throughput genomic profiling has revealed complex genetic and epigenetic landscapes across cancers, driving the development of molecularly guided therapies. Nonetheless, hepatocellular carcinoma (HCC) and biliary tract cancers continue to exhibit poor prognoses and limited therapeutic options, underscoring an urgent need for improved understanding of their underlying molecular determinants.1–3
Liver cancer is the sixth most common cancer, with an estimated 865,350 new cases in 2022 worldwide (4.3% of all cancers), but the third leading cause of cancer-related mortality, with an estimated 757,948 deaths (7.8% of all cancer-related deaths). 1 HCC is a highly lethal invasive carcinoma that accounts for the majority of liver cancers.2,4 The most important risk factors for HCC include preexisting liver cirrhosis and hepatitis B infection (due to both direct oncogenic effect and its contribution to cirrhosis). Other major risk factors contributing to cirrhosis (and therefore HCC) are hepatitis C infection, alcohol use, and nonalcoholic steatohepatitis.2,4–6 The 5-year survival rate for HCC remains approximately 18%. 7 Among non-HCC liver cancers, intrahepatic bile duct carcinoma constitutes a rare, aggressive malignancy (20%-25% of all biliary tract cancers, representing < 1% of all cancers) associated with older age, cholelithiasis, primary sclerosing cholangitis, obesity, and diabetes.3,8–10
DNA repair is central to eukaryotic life, as mammalian DNA in a single cell can experience approximately 10,000 lesions daily from endogenous and exogenous sources. 11 DNA repair pathways preserve genomic integrity by correcting these lesions, enabling accurate gene transmission and normal cellular function. 12 DNA repair factors (i.e., proteins directly involved in these pathways) are critical for genome maintenance. 13 Improper repair can inactivate tumor suppressor genes or activate oncogenes, promoting malignant transformation. 14 In liver cancer, recurrent genetic alterations involve TERT, TP53, CTNNB1, ARID1A, AXIN1, ARID2, CCND1, and VEGFA.2,6 In contrast, biliary tract cancers display frequent mutations in KRAS, TP53, and C-ERBB2. 15 However, the potential influence of DNA repair factor expression on liver cancer risk remains insufficiently understood, largely due to the scarcity of large-scale longitudinal data.
To address this research gap, the present study utilized genome-wide association study (GWAS) data to investigate potential causal relationships between genetically predicted serum levels of DNA repair factors and the risk of liver cancer.16,17 Mendelian randomization (MR) was applied as an analytical framework to infer causality while minimizing confounding and reverse causation.18–20 This method relies on three assumptions: (1) genetic variants are strongly associated with the exposure, (2) they are independent of confounders, and (3) they affect the outcome only through the exposure. 21
Therefore, the purpose of this study was to evaluate the causal association between genetically predicted serum DNA repair factors and liver cancer risk using large-scale GWAS data. By elucidating the mechanistic role of DNA repair processes in liver carcinogenesis, this research aimed to provide new molecular insights that may inform early detection strategies and potential therapeutic targets for liver cancer.
Methods
Study design
This study used publicly available data from GWASs to investigate the causal associations between DNA repair factors and liver cancer (Figure 1). Ethical approval was not necessary since the data were from studies that were already in compliance with the Declaration of Helsinki. The assumptions for MR studies are 1) the relevance assumption (i.e., the genetic variant(s) being used as an instrument for the exposure is associated with the exposure), 2) the independence assumption (i.e., there are no common causes of the genetic variant(s) and the outcome of interest), and 3) the no horizontal pleiotropy assumption (i.e., there is no independent pathway between the genetic variant(s) and the outcome other than through the exposure).
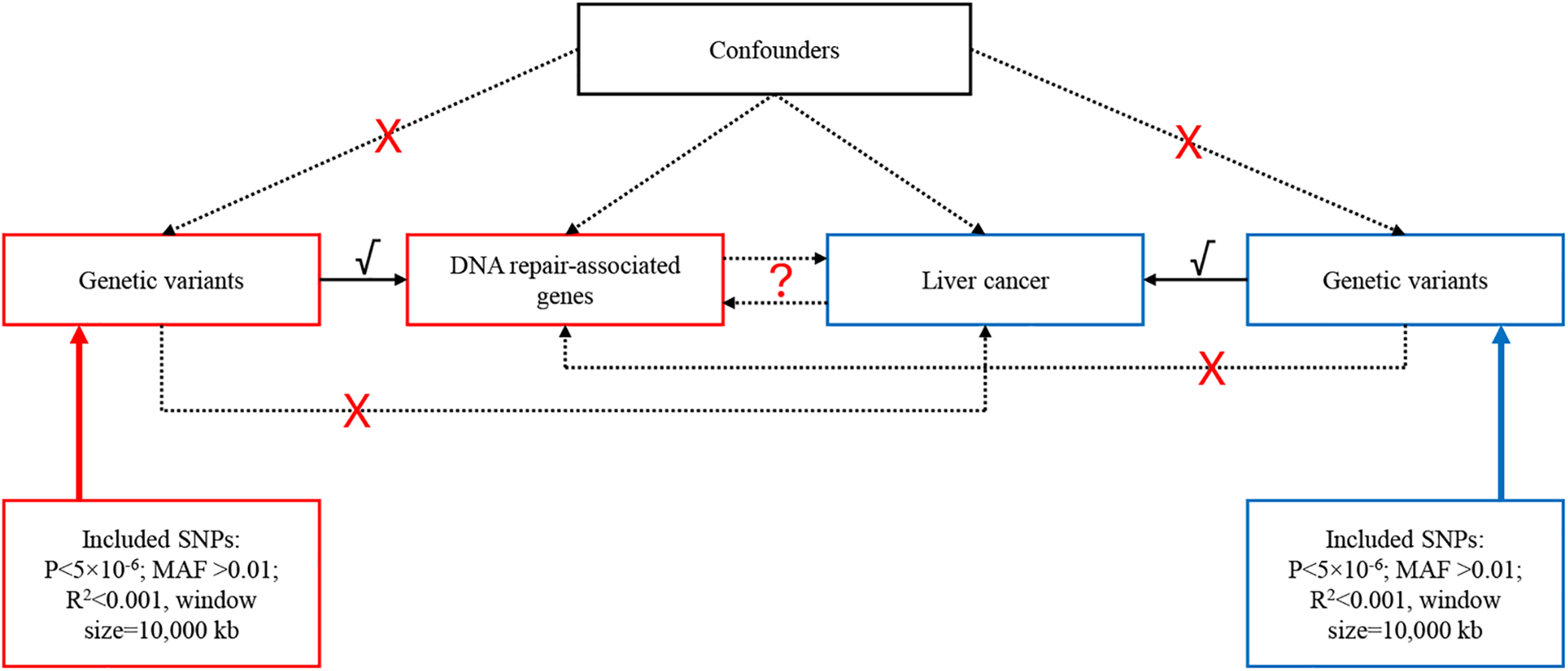
Schematic representation of the study.
Data source
The dataset for HCC, intrahepatic cholangiocarcinoma (IHCC), and malignant neoplasm of the liver was derived from the genomic data of the UK Biobank (UKB), which was used for a previous study. The malignant neoplasm of the liver, primary (PheCode 155.1) (ID GCST90041812) dataset contains 128 patients and 456,220 controls. The intrahepatic bile duct carcinoma (ICD-10 C22.1) (ID GCST90043859) contains 104 cases and 456,244 controls. The HCC (ICD-10 C22.0) (ID GCST90043858) dataset contains 123 cases and 456,225 controls. All individuals were of European ancestry. The exposure data were also from the GWAS catalog: serum levels of protein RAD51 (ID GCST90088117, 5354 individuals), Next to BRCA1 gene 1 (NBR1) protein levels (ID GCST90242103; 3301 individuals), DNA polymerase levels (ID GCST90240928; 3301 individuals), DNA repair protein XRCC4 levels (ID GCST90240935; 3301 individuals), A/G-specific adenine DNA glycosylase levels (ID GCST90240167; 3301 individuals), DNA repair endonuclease XPF measurement (ID GCST90240931; 3301 individuals), Poly (ADP-ribose) polymerase 1 (PARP1) levels (ID GCST90242303; 3301 individuals), and Poly (ADP-ribose) polymerase 11 (parp11) levels (ID GCST90242304; 3301 individuals). Those DNA repair factors were selected because they are available in GWAS datasets.
Instrumental variable selection
The instrumental variables (IVs) included in this study had to meet the following three criteria. 1) The entire genome was screened for SNPs significantly associated with the exposure, i.e., selecting SNPs associated with each exposure at P < 5 × 10−8, but that threshold yielded too few SNPs to allow an MR analysis, and the threshold had to be relaxed to P < 5 × 10−6.22,23 Second, SNPs with a minor allele frequency (MAF) > 0.01 were selected. Third, linkage disequilibrium (LD) between SNPs was removed based on R2 < 0.001 and a window size of 10,000 kb. 24 Hence, SNPs located within 10,000 kb of each other in the genome sequence and showing a strong association (R2 > 0.001) were excluded. If a SNP associated with an exposure was not found in the outcome summary data, SNPs with high LD (R2 > 0.8) with the original SNP were selected as a proxy. Finally, to ensure the absence of weak instrumental bias, the F-value was calculated for each SNP using F = R2 × (N-2)/(1-R2). F-values > 10 indicated the absence of weak instrumental bias and that the SNP could be kept for analysis.
Mendelian randomization analysis
This primary analysis to assess the causal association between exposure and outcome risk was the inverse variance weighted (IVW) method, which provides results as odds ratios (ORs) and 95% confidence intervals (CIs). Other MR methods were used to test the robustness of the IVW results, including the MR-Egger, weighted median, and weighted mode methods. Statistically significant IVW results, accompanied by statistically significant results from the other tests, were considered robust; otherwise, they were considered with caution. A causal association shown by IVW and not by the other methods, or by one of the other methods and not by IVW, had to be taken cautiously. All analyses were carried out using the “TwoSampleMR” package in R 4.3.1 (The R Project for Statistical Computing, www.r-project.org). The results were depicted using scatter and forest plots.
Sensitivity analysis
Potential heterogeneity among IVs was detected using Cochran's Q test, which was used to assess heterogeneity. A P-value > 0.05 indicated low heterogeneity, meaning that the estimates among IVs were randomly distributed and had little impact on the IVW results. Due to the risk of genetic variation heterogeneity influencing the estimation of association effects, the MR-Egger regression method was employed to detect the presence of horizontal pleiotropy. The absence of heterogeneity was considered when the intercept term of the MR-Egger regression tended toward zero or lacked statistical significance. In addition, an MR pleiotropy residual sum and outlier (MR-PRESSO) analysis was used to detect potential outliers (i.e., SNPs with P < 0.05). Such SNPs were removed to recalibrate the causal associations and correct for horizontal pleiotropy. A leave-one-out analysis was used to assess the robustness and consistency of the results.
Protein quantitative trait loci
A protein quantitative trait loci (pQTL) analysis was performed to determine whether SNPs could be associated with variations in protein levels or expression. Using IVW MR, DNA repair proteins significantly associated with liver cancer (P < 5 × 10−8) were identified. After extracting the corresponding SNPs and cross-referencing with the plasma pQTL database, 25 only the PARP1-related loci with a consistent direction and P < 0.05 were retained. Bayesian colocalization analysis (COLOC, PPH4 ≥ 0.8) confirmed that the PARP1 locus shared causal variants with liver cancer risk. Finally, PARP1 was imported into the KEGG pathway enrichment analysis to determine whether it drove liver cancer development through the base excision repair pathway (hsa03410).
Bioinformatics
The proteins identified as causally associated with liver cancer were investigated using Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) annotation analyses.
Results
Instrument variable selection
Our study performed an extensive series of MR analyses, focusing on diverse exposures implicated in DNA damage and repair pathways, using liver cancer as the primary outcome of interest. First, we identified IV sets with varied potency corresponding to each exposure. All F-values were > 10, affirming the exclusion of weak instrumental biases across all selected IVs, thereby reinforcing the reliability of our investigation into the intricate dynamics connecting DNA integrity, its restoration, and the onset of liver cancer (Supplementary Table 1).
Mendelian randomization analysis results
Then, the IVs identified for each exposure were tested for association with the outcomes. The genetic prediction results suggested a causal association between Next to BRCA1 gene 1 protein levels and primary malignant neoplasm of the liver (IVW: OR = 2.28, 95%CI: 1.17–4.45, P = 0.015; weighted median: P = 0.015), between serum levels of protein RAD51 and primary malignant neoplasm of the liver (IVW: OR = 2.16, 95%CI: 1.12–4.15, P = 0.021), between Poly (ADP-ribose) polymerase (PARP) 11 levels and HCC (IVW: OR = 0.48, 95%CU: 0.25–0.93, P = 0.030), and between PARP11 levels and intrahepatic bile duct carcinoma (IVW: OR = 1.86, 95%CI: 1.04–3.33, P = 0.038).
Then, cancers were used as the exposure, and DNA repair factor levels were used as the outcomes. The reverse analysis suggested causal associations between HCC and PARP1 levels (IVW: OR = 0.96, 95%CI: 0.93–0.99, P = 0.007) and between intrahepatic bile duct carcinoma and DNA repair protein XRCC4 levels (IVW: OR = 1.05, 95%CI: 1.02–1.09, P = 0.003) and Poly (ADP-ribose) polymerase 1 levels (IVW: OR = 1.05, 95%CI: 1.01–1.08, P = 0.005 (Table 1 and Figures 2 and 3). The remaining results showed no significant correlations. (All P > 0.05).
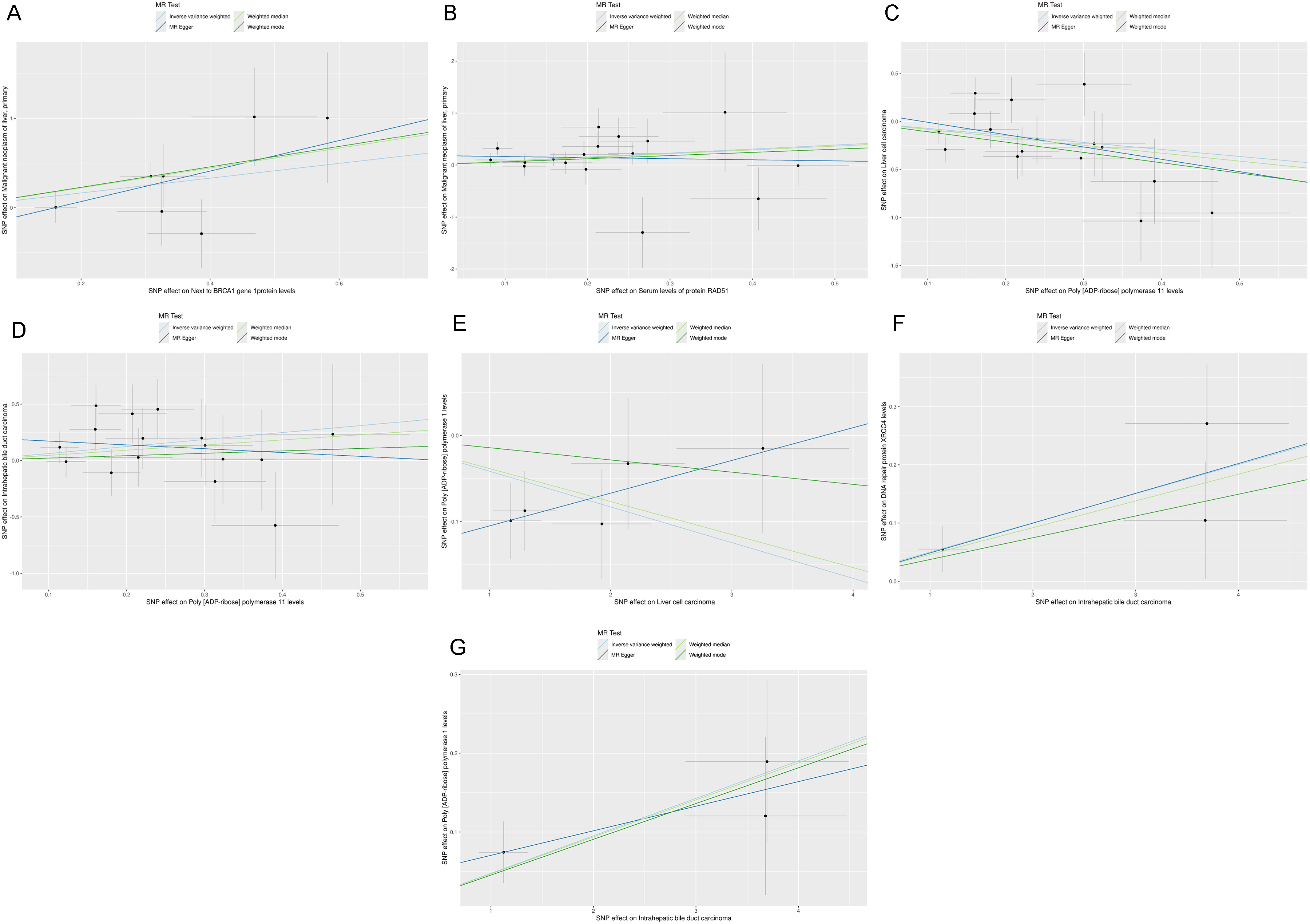
Scatter plots for the Mendelian randomization analysis of (A) next to BRCA1 gene 1 protein (NBR1) levels as exposure and primary malignant neoplasm of the liver as the outcome, (B) serum RAD51 recombinase (RAD51) protein levels as exposure and primary malignant neoplasm of the liver as the outcome, (C) poly(ADP-ribose) polymerase 11 (PARP11) levels as exposure and hepatocellular carcinoma as the outcome, (D) PARP11 levels as exposure and intrahepatic bile duct carcinoma as the outcome, (E) hepatocellular carcinoma as exposure and poly(ADP-ribose) polymerase 1 (PARP1) levels as the outcome, (F) intrahepatic bile duct carcinoma as exposure and DNA repair protein X-ray repair cross complementing 4 (XRCC4) levels as the outcome, and (G) intrahepatic bile duct carcinoma as exposure and PARP1 levels as the outcome.
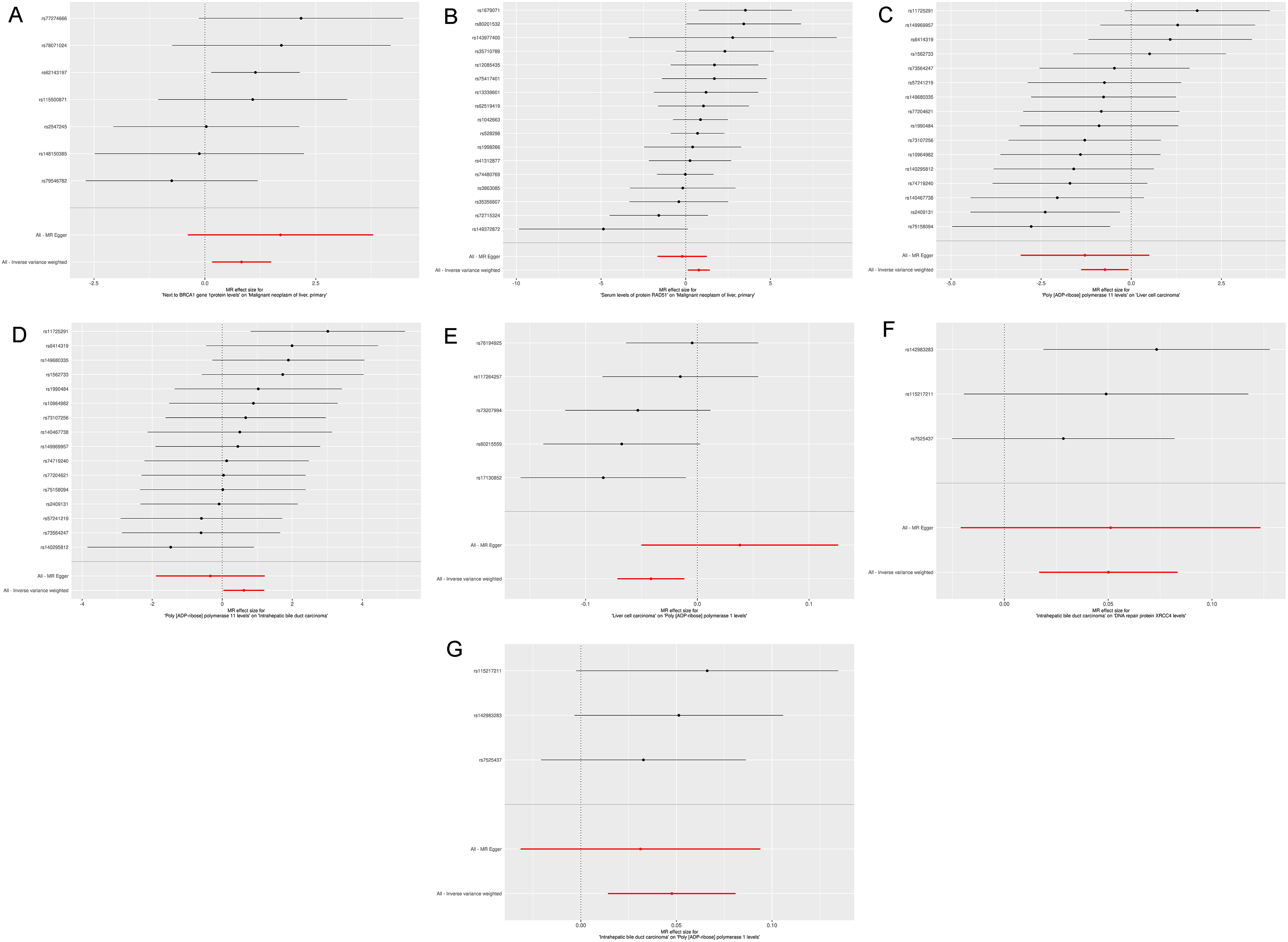
Forest plots for the Mendelian randomization analysis of (A) next to BRCA1 gene 1 protein (NBR1) levels as exposure and primary malignant neoplasm of the liver as the outcome, (B) serum RAD51 recombinase (RAD51) protein levels as exposure and primary malignant neoplasm of the liver as the outcome, (C) poly(ADP-ribose) polymerase 11 (PARP11) levels as exposure and hepatocellular carcinoma as the outcome, (D) PARP11 levels as exposure and intrahepatic bile duct carcinoma as the outcome, (E) hepatocellular carcinoma as exposure and poly(ADP-ribose) polymerase 1 (PARP1) levels as the outcome, (F) intrahepatic bile duct carcinoma as exposure and DNA repair protein X-ray repair cross complementing 4 (XRCC4) levels as the outcome, and (G) intrahepatic bile duct carcinoma as exposure and PARP1 levels as the outcome.
Mendelian randomization (mr) results of the causal associations between liver cancer and DNA repair-related genes.

Sensitivity analyses
Heterogeneity and horizontal pleiotropy can affect MR results, and their presence was investigated. No heterogeneity or pleiotropy was observed (all P > 0.05) (Table 2 and Figure 4). Outliers and single SNPs driving the results can also affect the ME results. The MR-PRESSO analysis revealed no outliers (Table 3). The leave-one-out analyses suggested that no individual SNP drove the results (Figure 5).

Funnel plots for the Mendelian randomization analysis of (A) next to BRCA1 gene 1 protein (NBR1) levels as exposure and primary malignant neoplasm of the liver as the outcome, (B) serum RAD51 recombinase (RAD51) protein levels as exposure and primary malignant neoplasm of the liver as the outcome, (C) poly(ADP-ribose) polymerase 11 (PARP11) levels as exposure and hepatocellular carcinoma as the outcome, (D) PARP11 levels as exposure and intrahepatic bile duct carcinoma as the outcome, (E) hepatocellular carcinoma as exposure and poly(ADP-ribose) polymerase 1 (PARP1) levels as the outcome, (F) intrahepatic bile duct carcinoma as exposure and DNA repair protein X-ray repair cross complementing 4 (XRCC4) levels as the outcome, and (G) intrahepatic bile duct carcinoma as exposure and PARP1 levels as the outcome.

Leave-one-out analysis plots for the Mendelian randomization analysis of (A) next to BRCA1 gene 1 protein (NBR1) levels as exposure and primary malignant neoplasm of the liver as the outcome, (B) serum RAD51 recombinase (RAD51) protein levels as exposure and primary malignant neoplasm of the liver as the outcome, (C) poly(ADP-ribose) polymerase 11 (PARP11) levels as exposure and hepatocellular carcinoma as the outcome, (D) PARP11 levels as exposure and intrahepatic bile duct carcinoma as the outcome, (E) hepatocellular carcinoma as exposure and poly(ADP-ribose) polymerase 1 (PARP1) levels as the outcome, (F) intrahepatic bile duct carcinoma as exposure and DNA repair protein X-ray repair cross complementing 4 (XRCC4) levels as the outcome, and (G) intrahepatic bile duct carcinoma as exposure and PARP1 levels as the outcome.
Sensitivity and horizontal pleiotropy analyses of the causal associations between liver cancer and DNA repair-related genes.

Results for pleiotropy in the causal relationships between liver cancer and DNA repair-associated genes.

Protein quantitative trait loci analysis
The pQTL analysis showed that the rs2793568 SNP was associated with the expression levels of the PARP1 protein (Figure 6A). PARP1 was associated with several GO terms related to DNA replication and DNA repair (Figure 6B). PARP1 was also associated with the KEGG terms of diabetic cardiomyopathy, necroptosis, apoptosis, NF-κB signaling, and base excision repair (Figure 6C).
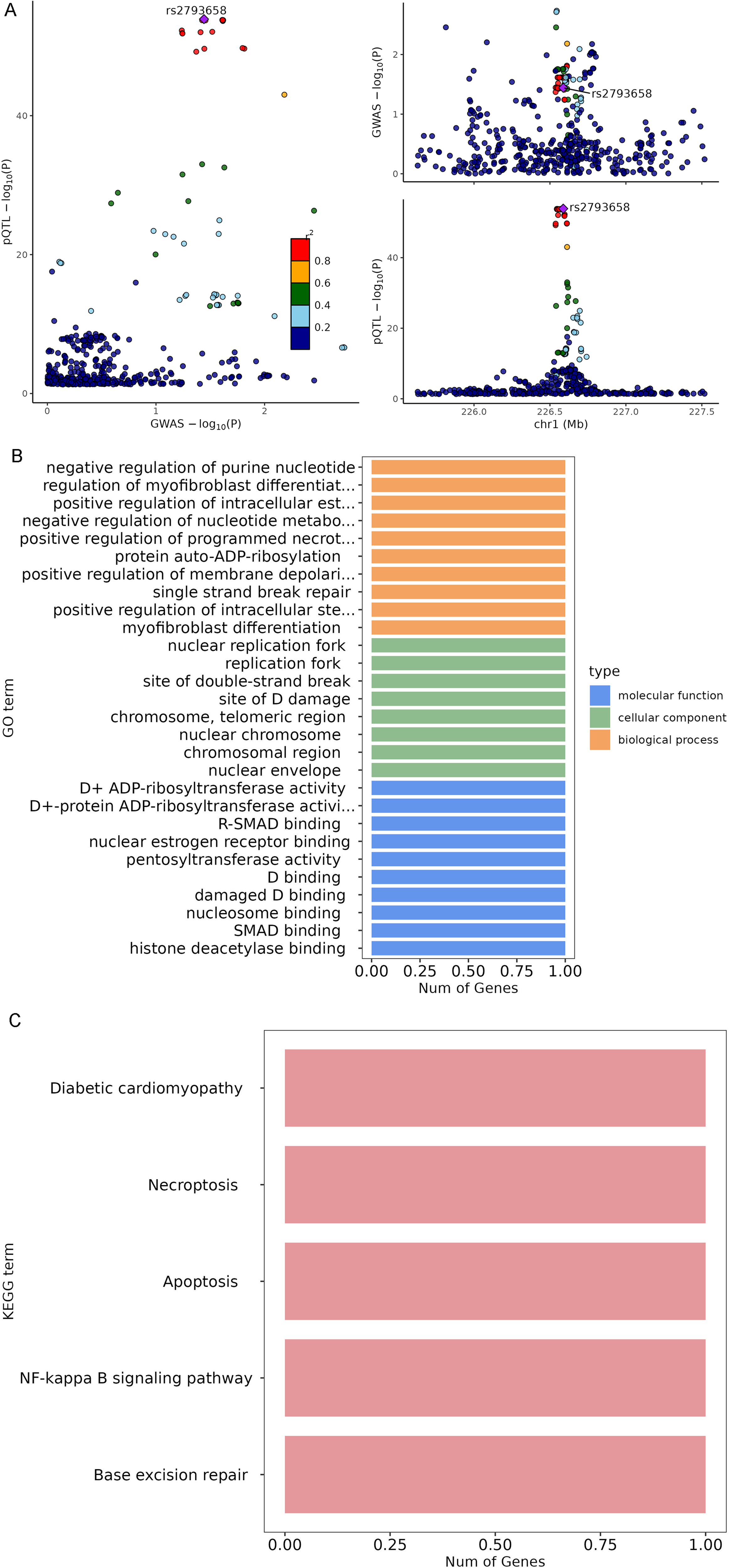
Protein quantitative trait loci analysis. (A) protein quantitative trait loci (pQTL) analysis showing that the rs2793568 single-nucleotide polymorphism (SNP) was associated with Poly(ADP-ribose) polymerase 1 (PARP1) protein levels. (B) Gene Ontology (GO) analysis of PARP1 and DNA replication and repair. (C) Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis of PARP1.
Discussion
This study aimed to explore the causal association between genetically predicted DNA repair factors and liver cancer. The results suggest that Next to BRCA1 gene 1 protein levels, serum levels of protein RAD51, PARP1 levels, PARP11 levels, and DNA repair protein XRCC4 levels may be involved in the pathogenesis of liver cancers. Attention should be given to those factors in future studies.
Next to BRCA1 gene 1 protein is encoded by the NBR1 gene and is an ubiquitin-binding autophagy adapter that participates in various processes, including host defense and intracellular homeostasis.26,27 It acts as a shuttle, bringing ubiquitinated proteins to autophagosomes and participating in the formation of protein aggregates.28,29 Next to BRCA1 gene 1 protein also regulates the innate immune response by modulating type I interferon production and targeting ubiquitinated IRF3 for autophagic degradation. 30 In response to oxidative stress, Next to BRCA1 gene 1 protein promotes an increase in SQSTM1, phosphorylation, and body formation by preventing its autophagic degradation. In turn, it activates the KEAP1-NRF2/NFE2L2 antioxidant pathway. Next to BRCA1 gene 1 protein is involved in the metastatic progression of ovarian cancer 31 and breast cancer. 32 In liver cancer, Next to BRCA1 gene 1 protein is involved in the specific degradation of MHC-1, thereby reducing CD8+ T cell-mediated antitumor immunity.33,34 In the present study, Next to BRCA1 gene 1 protein levels showed a causal association with primary malignant neoplasm of the liver, as supported by the above studies.
RAD51 plays a key role in DNA repair, especially in homologous strand exchange through homologous recombination.35–37 RAD51 binds to single-stranded DNA in an ATP-dependent manner to form nucleoprotein filaments essential for the homology search and strand exchange processes during DNA repair to form a joint molecule between a processed DNA break and the repair template.35,36,38 RAD51 is also involved in the resolution of stalled replication forks during replication stress, is part of a PALB2-scaffolded HR complex containing BRCA2 and RAD51C, plays a role in DNA repair,39,40 is involved in regulating mitochondrial DNA copy number under conditions of oxidative stress in the presence of RAD51C and XRCC3, 41 and plays a role in interstrand crosslink repair. 42 Several cancer types, including liver cancer, overexpress RAD51. 43 Inhibiting RAD51 in HCC displays antitumor effects. 44 The present MR study suggests a causal association between serum levels of the protein RAD51 and primary malignant neoplasms of the liver.
PARP is a family of proteins involved in many cellular processes, such as DNA repair, genomic stability, and programmed cell death. 45 The primary role of Poly (ADP-ribose) polymerases is to detect and initiate an immediate cellular response to single-strand DNA breaks by signaling the enzymatic machinery involved in repair. 46 Although the involvement of Poly (ADP-ribose) polymerase 1 in the pathogenesis of cancer is relatively well-known, 47 the role of Poly (ADP-ribose) polymerase 11 remains elusive. 48 The present MR analysis suggested a negative causal association between Poly(ADP-ribose) polymerase 11 and HCC, but a positive association with intrahepatic bile duct carcinoma. Hence, Poly (ADP-ribose) polymerase 11 exhibited strikingly divergent associations with HCC and IHCC, reflecting potential tissue-specific mechanisms in hepatocytes versus bile duct epithelial cells. HCC and IHCC, although both originating from the liver, arise from distinct cellular lineages (hepatocytes and biliary epithelial cells (BECs)) that differ transcriptionally, metabolically, and immunologically. This cellular divergence provides a biologically plausible framework for understanding the opposing statistical associations of Poly (ADP-ribose) polymerase 11 (protective in HCC, risk-enhancing in IHCC).49,50 In hepatocytes, Poly(ADP-ribose) polymerase 11 may play a homeostatic or protective role by maintaining genomic integrity and controlling lipid metabolism. Poly (ADP-ribose) polymerases are central to single-strand DNA break repair and replication fork stabilization. In hepatic tissue, particularly under oxidative stress, moderate PARP activity supports survival by repairing damage and avoiding necroinflammatory cascades. Excessive activation of poly (ADP-ribose) polymerase 1/11, however, may shift metabolism toward NAD + depletion; conversely, controlled Poly(ADP-ribose) polymerase 11 activity can protect hepatocytes from oncogenic transformation. Conversely, in bile duct epithelial cells, Poly(ADP-ribose) polymerase 11 expression may potentiate oncogenic pathways, possibly through an interplay with biliary-specific transcriptional regulators (HNF1β, SOX9) and distinct immune microenvironments that favor chronic inflammation and fibrosis, which are well-known drivers of cholangiocarcinogenesis.49–52 Several molecular mechanisms may explain the opposite effects, including the context of DNA repair pathways,53,54 immune modulation, 55 and the epigenetic and metabolic contexts. 56 In addition, both the IVW and weighted median methods support the causal association between Poly(ADP-ribose) polymerase 11 and HCC, whereas the weighted mode estimator did not reach statistical significance. This pattern is common in MR analyses and arises from the differing statistical assumptions and power characteristics of these estimators. The potential reasons for the discrepancy include weaker instrument strength, reduced number of valid clusters, sampling variability and heterogeneity, and low total IV strength.
XRCC4 plays important roles in DNA repair: DNA non-homologous end joining core factor required for double-strand break repair and V(D)J recombination,57,58 recruitment of other proteins to DNA double-strand breaks, formation of alternating helical filaments that bridge DNA like a bandage,59,60 and direct interaction with DNA ligase IV. 61
Many proteins are involved in DNA repair and cancer, and the relationships often vary among cancer types.62–64 Although some results were negative in the present study, it does not indicate that those proteins are not involved in liver cancer. MR analyses are based on the genetic prediction of protein expression, and other factors can influence the expression and activity of proteins. Importantly, the present study was conducted using serum levels of DNA repair factors, and whether these serum indicators accurately reflect what occurs in cells remains unknown. In addition, the investigated proteins are nuclear proteins, and their half-life, once released into circulation, may not accurately represent their actual fate within the cells. Nuclear proteins are typically found in the blood circulation due to cell death and the release of their contents.30,65 Those factors could explain why some results were positive and others were negative. MR studies do not reveal the specific mechanisms responsible for the observed associations. Nevertheless, this study suggests that the serum levels of some DNA repair factors are associated with liver cancer, but whether the proteins themselves are involved in liver cancer or their release through cell death indicates that the development of liver cancer remains to be investigated.
The integration of DNA repair-related serum proteins, such as BRCA1, RAD51, XRCC4, Poly(ADP-ribose) polymerase 1, and Poly(ADP-ribose) polymerase 11, into established HCC and IHCC screening workflows could markedly improve early detection and risk stratification. These biomarkers reflect homologous recombination repair (HRR) dysfunction and DNA damage response dysregulation, both of which are increasingly recognized in hepatobiliary tumorigenesis.66–68 Alpha-fetoprotein (AFP) remains a cornerstone of liver cancer screening, though it has limited sensitivity and specificity for early-stage HCC or IHCC. Recent studies show that combining AFP with alternative molecules (such as AFP-L3, des-γ-carboxy prothrombin (DCP), or CA19-9) improves diagnostic accuracy. Serum proteins like RAD51 and Poly (ADP-ribose) polymerase 1 could be evaluated alongside AFP to identify molecular subsets of HCC that exhibit HRR defects, improving sensitivity where AFP is negative or borderline.69–71 Routine imaging (ultrasound, computed tomography, and magnetic resonance imaging) can miss small or atypical lesions, particularly in cirrhotic livers. Serum panels incorporating Poly(ADP-ribose) polymerase 1, Poly(ADP-ribose) polymerase 11, and RAD51 may serve as pre-selection or triage tools, flagging high-risk patients for intensive imaging. Elevated DNA repair-related protein levels could also help interpret indeterminate imaging findings.54,72 Serum testing is minimally invasive and easy to implement in surveillance programs, particularly for high-risk groups such as chronic hepatitis B/C or cirrhosis patients. It aligns well with established paradigms of semiannual blood-based and imaging surveillance.68,72 Serial monitoring of RAD51 and Poly (ADP-ribose) polymerase 1 levels may reveal evolving genomic instability before lesion detectability by imaging, thus supporting longitudinal tracking of disease dynamics.66,69 Biomarkers such as Poly(ADP-ribose) polymerase 1 and RAD51 not only hold diagnostic potential but also predict therapeutic responsiveness to Poly(ADP-ribose) polymerase inhibitors and DNA damage-targeted agents, linking detection with precision treatment opportunities.53,67
In the context of reverse MR, the observed associations of HCC with Poly (ADP-ribose) polymerase 1 and IHCC with XRCC4 likely represent downstream biological effects of the cancer process rather than primary causal mechanisms. Nevertheless, these reverse associations provide insight into tumor-driven feedback pathways, cellular stress signaling, and systemic alterations that accompany disease progression, rather than indicating direct etiologic roles. Reverse MR linking hepatocellular carcinoma to increased Poly (ADP-ribose) polymerase 1 levels likely reflects feedback upregulation of the DNA damage response. Poly (ADP-ribose) polymerase 1 is a nuclear enzyme essential for recognizing and repairing DNA strand breaks. During hepatocarcinogenesis, genomic instability, oxidative stress, and chronic inflammation trigger the excessive activation of Poly(ADP-ribose) polymerase 1, which maintains cell survival under genotoxic conditions.73–75 Elevated serum Poly(ADP-ribose) polymerase 1 levels are often observed in advanced or aggressive tumors due to increased cell turnover and necrosis, leading to the passive release of nuclear proteins into the circulation. Therefore, this reverse association plausibly represents both tumor-induced transcriptional activation of Poly(ADP-ribose) polymerase 1 within malignant hepatocytes and protein leakage into the serum during hepatic injury.74,76 The association between IHCC and XRCC4 levels, identified by reverse MR, may also denote secondary molecular responses to DNA damage or stromal remodeling. XRCC4 is a key scaffolding factor in non-homologous end joining (NHEJ) repair, often upregulated in biliary tumors exposed to oxidative DNA insults or chemoresistance pressure.77,78 In cholangiocarcinoma, chronic inflammation and bile acid-induced genotoxicity activate XRCC4-mediated repair pathways, which support the survival of malignant cells but may concurrently lead to detectable elevations of XRCC4 in serum due to tumor leakage or exosome-mediated secretion.78,79 Taken together, the elevated circulating PARP1 and XRCC4 levels in reverse MR analyses should be interpreted as markers of cancer-associated tissue stress or damage response activation, rather than as drivers of tumor initiation. They likely reflect both intracellular feedback regulation sustaining malignant viability and extracellular release due to progression-related cell death.
This study had several strengths. It used a two-sample MR design and GWAS data from thousands of patients. It also evaluated the causal association between DNA repair proteins and liver cancers. On the other hand, the study also had limitations. The GWAS data in the present study primarily originated from a European population, and their generalizability to other populations remains unknown. Liver cancer etiologies exhibit pronounced geographic diversity, primarily driven by differences in viral hepatitis prevalence, alcohol consumption, and metabolic disease patterns worldwide. HCC accounts for about 90% of primary liver cancers, and its underlying causes vary significantly between regions and populations. In Asia and Africa, hepatitis virus B or C infection is the most common etiology, while in Europe and North America, alcohol and metabolic liver diseases are the most common culprits.80–82 Second, some participants could have been included in both the exposure and the outcome, but it is impossible to determine the exact number. Still, using stringent statistics should minimize the impact of an eventual overlap. Third, several SNPs could be selected as IVs, affecting the causal associations. Finally, the DNA repair proteins we studied were selected based on their established roles in DNA repair and oncogenesis, as documented in the literature. MR studies are entirely statistical studies based on available datasets, and no experiments were performed in the present study. The mechanistic roles of the DNA repair factors identified in this study will need to be confirmed in future studies.
In conclusion, next to BRCA1 gene 1 protein levels, serum levels of proteins RAD51, XRCC4, Poly(ADP-ribose) polymerase 1, and Poly(ADP-ribose) polymerase 11 may be involved in the pathogenesis of liver cancers. Attention should be given to those factors in future studies.
Supplemental Material
sj-docx-1-cbm-10.1177_18758592261429328 - Supplemental material for Bidirectional Mendelian randomization analysis reveals significant associations between Serum DNA repair proteins and liver cancer
Supplemental material, sj-docx-1-cbm-10.1177_18758592261429328 for Bidirectional Mendelian randomization analysis reveals significant associations between Serum DNA repair proteins and liver cancer by Sihao Lin, Kunpeng Fang and Li Geng in Cancer Biomarkers
Supplemental Material
sj-docx-2-cbm-10.1177_18758592261429328 - Supplemental material for Bidirectional Mendelian randomization analysis reveals significant associations between Serum DNA repair proteins and liver cancer
Supplemental material, sj-docx-2-cbm-10.1177_18758592261429328 for Bidirectional Mendelian randomization analysis reveals significant associations between Serum DNA repair proteins and liver cancer by Sihao Lin, Kunpeng Fang and Li Geng in Cancer Biomarkers
Supplemental Material
sj-docx-3-cbm-10.1177_18758592261429328 - Supplemental material for Bidirectional Mendelian randomization analysis reveals significant associations between Serum DNA repair proteins and liver cancer
Supplemental material, sj-docx-3-cbm-10.1177_18758592261429328 for Bidirectional Mendelian randomization analysis reveals significant associations between Serum DNA repair proteins and liver cancer by Sihao Lin, Kunpeng Fang and Li Geng in Cancer Biomarkers
Footnotes
Acknowledgements
Not applicable.
Ethical statement
Not applicable.
Consent for publication
Not applicable.
Author contributions
Conception and design: Li Geng
Administrative support: Li Geng Provision of study materials or patients: Kunpeng Fang Collection and assembly of data: Sihao Lin Data analysis and interpretation: Sihao Lin Manuscript writing: All authors Final approval of manuscript: All authors.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Availability of data and materials
All data generated or analyzed during this study are included in this published article.
Supplemental material
Supplemental material for this article is available online.
