Abstract
Background
Considering the significance of circRNA-miRNA network underlying cervical cancer (CC) development, this investigation was devised to explore whether and how 6-methyladinosinek (m6A)-adjusted hsa_circ_0101308/miR-224 axis participated in altering chemo-resistance in CC.
Methods
Forty-nine pairs of CC tissues and para-cancerous normal tissues were gathered, and CC cell lines, comprising HeLa, HeLa/DDP, HeLa/ADM and HeLa/TAX cell lines, were pre-prepared. Expressions of circRNAs, miRNAs and mRNAs were determined using quantitative reverse transcription-polymerase chain reaction (qRT-PCR), and m6A-modification of hsa_circ_0101308 was verified based on methylated RNA immunoprecipitation sequencing (MeRIP-Seq) assay. Among CC cell lines, their chemo-resistance was evaluated through CCK8 assay, and their viability was assessed via MTT assay.
Results
Hsa_circ_0101308 expression markedly dwindled, accompanied by notably elevated expression of miR-224, within CC tissues, when compared with para-cancerous normal tissues (P < 0.05). Hsa_circ_0101308 sponed miR-224 and suppressed its expression in HeLa cell line (P < 0.05), and either under-expressed m6A-adjusted hsa_circ_0101308 or over-expressed miR-224 strengthened viability of HeLa, HeLa/DDP, HeLa/ADM and HeLa/TAX cell lines (P < 0.05). Additionally, miR-224 targeted CADM1 and down-regulated its mRNA level (P < 0.05), which influenced p-PI3K/PI3K or p-Akt/Akt ratio (P < 0.05).
Conclusion
The network combined by m6A-adjusted hsa_circ_0101308 and miR-224 interfered with chemo-resistance in CC via acting upon CADM1 and PI3K/AKT pathway, which was conducive to optimizing CC treatment.
Introduction
Cervical cancer (CC), induced by human papillomavirus (HPV) and some cofactors, was a frequently-found malignancy that occurred in females’ reproductive system.1,2 Though HPV vaccine and screening test have been widely employed to substantially prevent against CC onset, there were, as documented by GLOBOCAN2020, still up to 0.604 million new cases and approximately 0.342 million deaths per year around the globe. 3 Responding to CC progression, International Federation of Gynecology and Obstetrics (FIGO) designated surgery to tackle early-stage CC, and radiotherapy coupled with chemotherapy to cope with late-stage CC,4–6 nevertheless, a variety of treatment puzzles arose. It has been emphasized that multi-drug resistance (MDR) largely restricted favorable treatment outcomes of chemotherapies, which have been proved to drastically prolong survival of CC patients.7,8 To overcome this obstacle, it was vital to figure out biomarker-focused mechanisms underlying MDR in CC, thereby ameliorating CC treatment.
Features by covalently-closed ring structures that displayed resistance to RNase R, 9 circRNAs surpassed linear RNAs in terms of precisely reflecting happening, exacerbation or chemo-resistance of neoplasms, which incorporated gastric cancer, colorectal cancer, breast cancer, hepatocellular carcinoma and CC.10–13 For instance, a series of circRNAs (e.g., hsa_circ_0101308), retrieved from Gene Expression Omnibus Series (GSE)102686, were distinctly expressed between CC tissues and para-cancerous normal tissues by a wide margin, 14 whereas certain circRNAs, such ashsa_circ_0004488 15 or hsa_circ_0001589, 16 were inclined to spur resistance of CC cells against paclitaxel or DDP. It was noteworthy that N6-methyladenosine (m6A) modification, a RNA-methylation process, 17 has been recognized to affect biogenesis, stability or cytoplasmic export of circRNAs underlying cancer etiology.18–20 Taking CC for instance, hsa_circ_0000069, whose stability was maintained on account of m6A modification, was found to sponge miR-4426, so as to accelerate proliferation and migration of CC cells. 21 Besides, since being modified by m6A, circular E7 RNA (circE7), produced by HPV, was apt to localize in cytoplasm, and enhance production of E7 oncoproteins after combination with polysome. 22 Even if the tight linkage between m6A modification and CC pathogenesis has been stressed,23–28 there were scarce studies that connected m6A-adjusted circRNAs, encompassing hsa_circ_0101308 14 verified to be adjusted by m6A in this research, with MDR inherent in CC etiology.
MiRNAs, 19∼25 nt-long linear non-coding RNAs, could be sponged by particular circRNAs that encompassed miRNA response elements (MREs), 29 thereby adjusting progression or drug-resistance in neoplasms.30,31 For example, hsa_circ_0004872 sponged miR-224, which was adept at modifying docetaxel 32 or cisplatin resistance, 33 to impede tumor aggravation, 34 and growth of tumor cells was greatly demotivated after hsa_circ_0043280 sponged miR-203a. 35 Additionally, it has been accepted that miRNAs bound to MREs of peculiar mRNAs, such as miR-486 targeting CADM1 36 and miR-512-3p targeting Livin, 37 thus influencing cancer progression or MDR. Due to the competing endogenous RNA (ceRNA) network mentioned above, it was pivotal to screen out downstream miRNA/mRNA axes of hsa_circ_0101308, such as miR-224 32 /CADM1 36 axis confirmed in this investigation, so as to broadening molecular mechanisms in fighting against CC progression or chemo-resistance.
Hence, this investigation focused on exploring the involvement of m6A-adjustedhsa_circ_0101308-miR-224-CADM1 network in altering CC chemo-resistance, which was conducive to implementing efficacious measures against MDR in CC and ameliorating treatment of CC.
Materials and methods
Subjects
Altogether forty-nine CC patients who received radical operation from July 2014 to October 2016 were recruited from People's Hospital affiliated to Hubei University of Medicine. Pairs of CC tissues and para-cancerous normal tissues, ≥ 2 cm distant from the boundary of CC tissues, were excised from each patient. The CC cases 1) were diagnosed by means of post-operative histopathology, and they 2) hardly underwent radiotherapy or chemotherapy before the surgery. Nonetheless, they were excluded in case 1) they possessed histories of other tumors, 2) they suffered from severe diseases in heart, liver, kidney and other organs, 3) they were subjected to hematological diseases or immune system disorders; 4) they were affected by systemic infection, and 5) they were pregnant or lactating. All patients have signed informed consents, and this retrospective program received approval from the ethics committee of People's Hospital affiliated to Hubei University of Medicine (approval no.: 20240715072).
Drug treatment and follow-up of CC patients
TP regimen, composed by paclitaxel and cisplatin (DDP), was arranged for treating post-surgery CC patients, who were treated by 1) intravenous drip of 0.137–0.175 g/m2 paclitaxel injection (Hospira Australia Pty Ltd, Australia) for 3 h, and then 2) intravenous drip of 75 g/m2 DDP and sodium chloride injection (Guizhou Hanfang Pharmaceutical Co. Ltd, China) for once. The treatment course was implemented every fourth week, and each CC patient received 6 times of the treatment. Following the chemo-therapy, the subjects were followed up for 2 years, and 2-year overall survival (OS) of the CC patients was tracked via correspondence, telephone or visit.
Cell culture and cell screening
Human CC cell lines, comprising HeLa cell line, SiHa cell line, C-33A cell line and CaSki cell line, and normal cervical epithelial cell line (i.e., End1/E6E7 cell line) were supplied from cell bank of Chinese Academy of Sciences (China). The cell lines were suspended within 10% fetal bovine serum (FBS)-inclusive Roswell Park Memorial Institute (RPMI) 1640 culture solution (Gibco, USA), ensured at 37 °C under the circumstance of 5% CO2.For the sake of harvesting HeLa/DDP cell line, HeLa/ADM cell line and HeLa/TAX cell line, gradually rising concentrations of cisplatin (DDP) (Shanghai Xinyu Biological Technology Co. Ltd, China), adriamycin (ADM) (Shanghai Fortunei Biotech Co. Ltd, China) and Taxol (Shanghai Acmec Biochemical Co. Ltd, China) were prepared to cultivate HeLa cell lines at the logarithmic phase for 1–2 d, until the cell lines grew stably. The experimental procedures were repeated for 5 times.
Quantitative reverse transcription-polymerase chain reaction (qrt-PCR)
In line with the guidance of One Step TB Green® PrimeScriptTM RT-PCR kit (Takara Bio, Japan), qRT-PCR was performed to detect and quantify RNAs, separated from tissue samples or cell lines using Trizol reagent (Sangon Biotech, China), through relying on real-time PCR instrument (Applied Biosystems, USA). Primers of circRNAs, miRNAs and mRNAs were listed, respectively, in Supplementary Tables 1–3. The reaction program of PCR consisted of 1) 42 °C for 5 min and 95 °C for 10 s, and 2) 40 cycles of 95 °C for 5 s and 60 °C for 32 s. Relative expressions of target genes were calculated by virtue of 2−ΔΔCt method. 38 GAPDH was set as the internal reference for circRNAs and mRNAs, whereas U6 was designated as the internal reference for miRNAs. The experimental procedures were repeated for 5 times.
Cell transfection
After being digested by appropriate amounts of trypsin (Deebio, China), HeLa cell line, HeLa/ADM cell line, HeLa/DDP cell line and HeLa/TAX cell line at the logarithmic growth phase, resuspended within complete medium, were separately inoculated witin 6-well plates at the concentration of 1 × 105 per well for overnight. Then serum-free Opti-MEM medium (Gibco, USA) was arranged to cultivate above-mentioned cell lines for 2 h. Subsequently, si-hsa_circ_0101308 (Sangon Biotech, China), pcDNA3.1-hsa_circ_0101308 (Sangon Biotech, China), miR-224 mimic (GenePharma, China), miR-224 inhibitor (GenePharma, China), si-CADM1, 39 si-negative control (NC) (GenePharma, China) and NC (GenePharma, China), which have been been mixed with LipofectamineTM 3000 transfection reagent (Invitrogen, USA) at room temperature for 25 min, were transfected into HeLa cell line, HeLa/ADM cell line, HeLa/DDP cell line or HeLa/TAX cell line. Six hours later, the cell lines were cultivated within fresh complete medium, placed within a 5% CO2–contained incubator, at 37 °C for 48 h. The experimental procedures were repeated for 5 times.
Dual-luciferase reporter gene assay
In accordance with CircInteractome software (https://circinteractome.nia.nih.gov/), there existed specific binding sites between hsa_circ_0101308 and miR-224. Aided by LipofectamineTM 3000 reagent (Invitrogen, USA), the wide-type (WT) or mutant-type (MUT) plasmid of hsa_circ_0101308 (GenePharma, China) was co-transfected with miR-224 mimic or miR-NC (GenePharma, China) into HeLa cell line for 12 h. On the other hand, miR-224 mimic or miR-NC (GenePharma, China) and the wide-type (WT) or mutant-type (MUT) plasmid of CADM1 were jointly transfected into HeLa cell line, which lasted for 12 h. Then luciferase activity of HeLa cell line was detected, assisted by dual-luciferase reporter assay kit (Promega, USA). The experimental procedures were repeated for 5 times.
Methylated RNA immunoprecipitation (MeRIP)
After transfection of miR-224 mimic or NC, total RNAs separated from HeLa cell line using TRIzol reagent (Invitrogen, USA) were purified and then fragmented through employing Poly(A)Tract mRNA purification kit (Sigma, USA). After diluting RNA fragments by addition of immunoprecipitation (IP) solution (Thermo Fisher, USA), magnetic beads that were combined with both RNAs and m6A antibody (Abcam, USA) or IgG antibody (Abcam, USA) were incubated at 4 °C for overnight. In the wake of digestion by protease K (Sigma, USA), RNAs that united with m6A were extracted and sedimented via addition of phenol-chloroform-isoamyl alcohol mixture (Shanghai Bioshun Cor., China), after which qRT-PCR was conducted to determine expressions of m6A-adjusted circRNAs. The experimental procedures were repeated for 5 times.
Cell counting kit 8 (CCK8) assay for assessing drug resistance
HeLa cell line, HeLa/DDP cell line, HeLa/ADM cell line and HeLa/TAX cell line at the logarithmic growth phase were cultivated in 96-well plates at the density of 2 × 104 per well for 24 h. Afterwards, DDP, ADM or TAX was supplemented to cultivate cell lines for 48 h, followed by 4-h incubation of cell lines utilizing 10 μl CCK8 solution (Keygentec, China). Then optical density (OD) values at the wavelength of 450 nm were measured on a microplate reader (Bio-Rad, USA). The inhibition rate (%) of DDP, ADM or TAX on survival of HeLa cell line, HeLa/DDP cell line, HeLa/ADM cell line or HeLa/TAX cell line was computed, from which half maximal inhibitory concentration (IC50) values were figured out. The experimental procedures were repeated for 5 times.
3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) assay
HeLa cell line, HeLa/DDP cell line, HeLa/ADM cell line or HeLa/TAX cell line, mixed with 10 μmol/L LY294002 (MedChemExpress, USA) or not, were cultured within 96-well plates at the concentration of 1 × 104/well, guaranteed within a 5%-CO2 incubator for 24 h. After utilizing MTT solution (Sigma, USA) to cultivate cells at 37 °C for 4 h, dimethyl sulphoxide (DMSO) was added to dissolve insoluble products. OD values of the samples were determined on a microplate reader (Bio-Rad, USA) at the wavelength of 490 nm. The experimental procedures were repeated for 5 times.
Statistical analyses
The whole data were analyzed by feat of GraphPad Prism 9.5.0 (GraphPad Inc., La Jolla, CA, USA). Differential analyses relevant to two sets of measurement data [mean ± mstandard deviation (SD)] were implemented based on student's t test, while multi-group comparisons were performed via one-way analysis of variance (ANOVA) with Turkey's post hoc test. Chi-square test was adopted to compare enumeration data (n, %). Kaplan-Meier method with Log-rank test was applied to estimate the correlation between hsa_circ_0101308/miR-224 expression and 2-year survival of CC patients. Founded upon spearman's correlation analysis, the relationship between hsa_circ_0101308 expression and miR-224 expression was analyzed. The difference was considered as statistically significant in case of P < 0.05.
Results
Association of hsa_circ_0101308/miR-224 expression with clinical and pathological characteristics of CC subjects
As per GSE102686 dataset stored within Gene Expression Omnibus (GEO; https://www.ncbi.nlm.nih.gov/geo/), 40 28 circRNAs were significantly under-expressed within CC tissues in comparison to normal tissues. 14 In this retrospective study, we firstly compared expressions of the 28 circRNAs among 3 pairs of CC tissues and para-cancerous normal tissues (Supplementary Figure 1), and then further confirmed that hsa_circ_0101308 was prominently under-expressed in CC tissues compared with para-cancerous normal tissues, considering 46 CC patients (P < 0.05) (Figure 1(a)). Furthermore, aside from age, tumor size and pathological type (all P > 0.05), over-expressed (>mean expression) hsa_circ_0101308 was markedly associated with fine/moderate differention (OR = 6.48, 95%CI: 1.68–21.17, P < 0.05), International Federation of Gynecology and Obstetrics (FIGO) stage IA&IB (OR = 3.56, 95%CI: 1.00–13.03, P < 0.05), lymph node non-metastsis (OR = 7.27, 95%CI: 1.80–26.96, P < 0.05) and vascular non-invasion (OR = 5.60, 95%CI: 1.00–28.11, P < 0.05) of CC patients, in contrast to under-expressed (≤mean expression) hsa_circ_0101308 (Supplementary Table 4). More than that, CC patients in the over-expressed hsa_circ_0101308 group survived longer than ones in the under-expressed hsa_circ_0101308 group (P < 0.05) (Supplementary Figure 2).
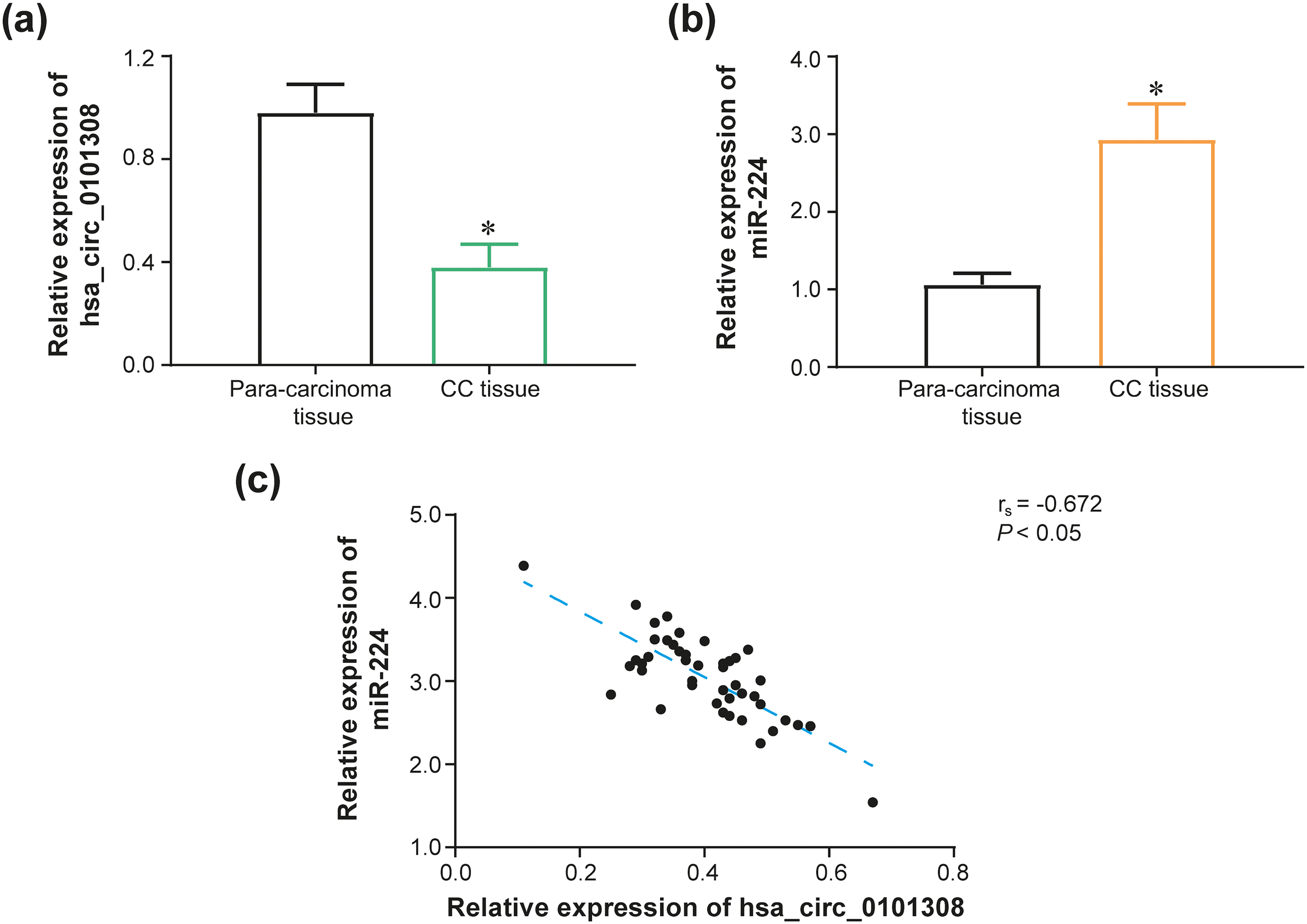
Clinical values of hsa_circ_0101308 or miR-224 expression in cervical cancer (CC). (a–b) Hsa_circ_0101308 expression (a) or miR-224 expression (b) was compared between CC tissues and para-cancerous normal tissues. *: P < 0.05 in comparison to para-cancerous normal tissues.(c) Hsa_circ_0101308 expression was negatively correlated with miR-224 expression among 46 CC tissues.
In addition, initially expressions of hsa_circ_0101308-sponged miRNAs, summarized from Circinteractome database (https://circinteractome.irp.nia.nih.gov/), were determined within CC tissues and para-cancerous normal tissues among 3 CC patients (Supplementary Figure 3). In the next step, among 46 CC patients, it was validated that miR-224 expression in CC tissues remarkably exceeded that in para-cancerous normal tissues (P < 0.05) (Figure 1(b)). What's more, over-expressed (>mean expression) miR-224 was dramatically correlated with poor differentiation of CC patients (OR = 4.29, 95%CI: 1.19–13.72, P < 0.05), as relative to under-expressed (≤mean expression) miR-224, except for age, tumor size, pathological type, FIGO stage, lymph node metastasis and vascular invasion (all P > 0.05) (Supplementary Table 5). Concerning overall survival (OS), over-expressed miR-224 was significantly interconnected with undesirable 2-year survival in comparison to under-expressed miR-224 (P < 0.05) (Supplementary Figure 4). Furthermore, among 46 CC tissues, hsa_circ_0101308 expression was negatively linked with miR-224 expression (rs = -0.672, P < 0.05) (Figure 1(c)).
M6A-adjusted hsa_circ_0101308 sponged mir-224 to degrade its expression in CC cells
Compared with normal cervical epithelial cell line (i.e., End1/E6E7), hsa_circ_0101308 expression signally declined (Figure 2(a)), along with distinctly elevated miR-224 expression (Figure 2(b)), within CC cell lines (i.e., HeLa, SiHa, C-33A and CaSki) (P < 0.05). Within HeLa cell line, hsa_circ_0101308 expression was down-regulated significantly in the si-hsa_circ_0101308 group (P < 0.05), while it ascended greatly in the pcDNA3.1-hsa_circ_0101308 group (P < 0.05), compared with respective NC group (Figure 2(c)). Regarding miR-224 expression in HeLa cell line, it was notably heightened in the miR-224 mimic group (P < 0.05), yet it descended significantly in the miR-224 inhibitor group, in comparison to NC groups (P < 0.05) (Figure 2(d)).
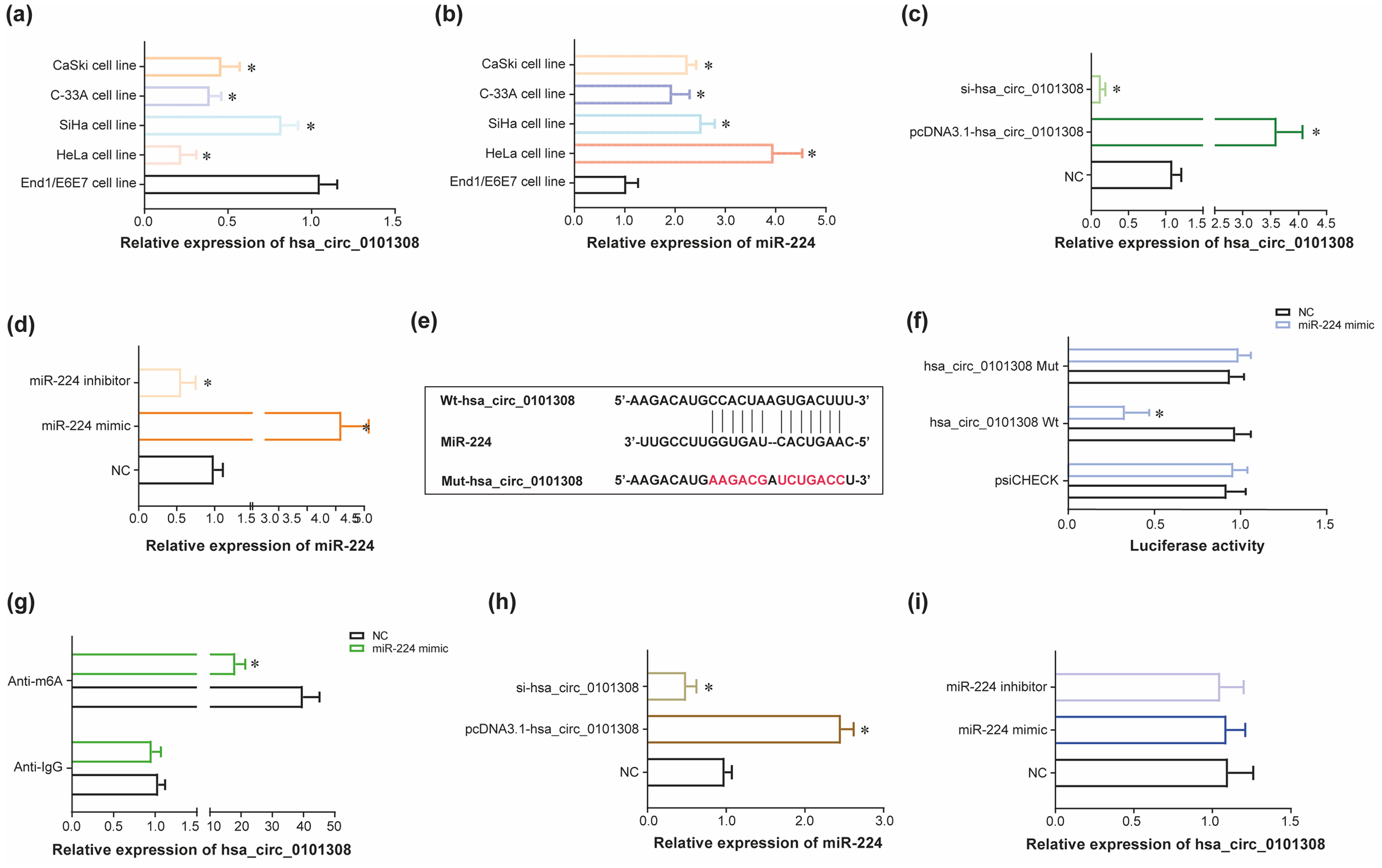
Influence of N6-methyladenosine (m6A)-adjustedhsa_circ_0101308 on miR-224 expression in cervical cancer (CC) cells. (a) Hsa_circ_0101308 expression was determined within CaSki cell line, C-33A cell line, SiHa cell line, HeLa cell line and End1/E6E7 cell line. *: P < 0.05 in comparison to End1/E6E7 cell line. (b) MiR-224 expression was detected within CaSki cell line, C-33A cell line, SiHa cell line, HeLa cell line and End1/E6E7 cell line. *: P < 0.05 in comparison to End1/E6E7 cell line. (c) Hsa_circ_0101308 expression was drawn after transfection of pcDNA3.1-hsa_circ_0101308 or si-hsa_circ_0101308 into HeLa cell line. *: P < 0.05 in comparison to negative control (NC) group. (d) MiR-224 expression was evaluated after transfection of miR-224 mimic or miR-224 inhibitor into HeLa cell line. *: P < 0.05 in comparison to NC group. (e-f) Hsa_circ_0101308 bound to miR-224 in specific sites (e), and luciferase activity of HeLa cell line was compared among hsa_circ_0101308 Mut + miR-224 mimic group, hsa_circ_0101308 Wt + miR-224 mimic group and hsa_circ_0101308 Wt + NC group (f). *: P < 0.05 in comparison to hsa_circ_0101308 Mut + miR-224 mimic group and hsa_circ_0101308 Wt + NC group. (g) N6-methyladenosine (m6A)-modified hsa_circ_0101308 in HeLa cell line was compared between miR-224 mimic group and NC group. *: P < 0.05 in comparison to NC group. (h) MiR-224 expression in HeLa cell line was assessed after transfection of si-hsa_circ_0101308 or pcDNA3.1-hsa_circ_0101308. *: P < 0.05 in comparison to NC group. (i) Hsa_circ_0101308 expression in HeLa cell line was detected after transfection of miR-224 inhibitor or miR-224 mimic.
In addition, hsa_circ_0101308 sponged miR-224 in certain sites (Figure 2(e)), and relative luciferase activity of HeLa cell line in the hsa_circ_0101308 WT + miR-224 mimic group became down-regulated as relative to NC group (P < 0.05) (Figure 2(f)). Nonetheless, relative luciferase activity of HeLa cell line in the hsa_circ_0101308 MUT + NC group remained nearly equivalent to that in thehsa_circ_0101308 MUT + miR-224 mimic group (P > 0.05). What's more, the expression level of anti-m6A, which symbolized expression of m6A-adjusted hsa_circ_0101308, in the miR-224 mimic group was pronouncedly reduced in comparison to NC group (P < 0.05) (Figure 2(g)). The m6A-adjusted sites of hsa_circ_0101308 were concretely presented in Supplementary Figure 5a–c, based on its sequence (Supplementary Table 6) obtained from CircRase database (http://circbase.org/), and predictions implemented by sequence-based RNA adenosine methylation site predictor (SRAMP) database (http://www.cuilab.cn/sramp). Furthermore, transfection of pcDNA3.1-(m6A-adjusted) hsa_circ_0101308 elevated miR-224 expression in HeLa cell line (P < 0.05), but on the contrary, silencing of m6A-adjusted hsa_circ_0101308 triggered significant decline of miR-224 expression within HeLa cells, as compared with NC group (P < 0.05) (Figure 2(h)). What's more, expression of m6A-adjusted hsa_circ_0101308 stayed nearly invariant in HeLa cell line, no matter miR-224 mimic (P > 0.05) or miR-224 inhibitor (P > 0.05) was transfected, as relative to NC group (Figure 2(i)).
M6A-adjusted hsa_circ_0101308/miR-224 axis affected viability of drug-resistant CC cells
In response to DDP, the IC50 value of HeLa cell line reached 1.93 μg/ml, yet the IC50 value of HeLa/DDP cell line achieved as high as 43.03 μg/ml (Figure 3(a)). In terms of combating against ADM, the IC50 value of HeLa/ADM cell line was equivalent to 3.14 mg/L, which transcended that of HeLa cell line (i.e., 0.73 mg/L) (Figure 3(b)). With respect to fighting against TAX the IC50 value of HeLa/TAX cell line, namely 98.09 μmol/L, seemed, to a great extent, over that of HeLa cell line (i.e., 30.42 nmol/L) (Figure 3(c)).
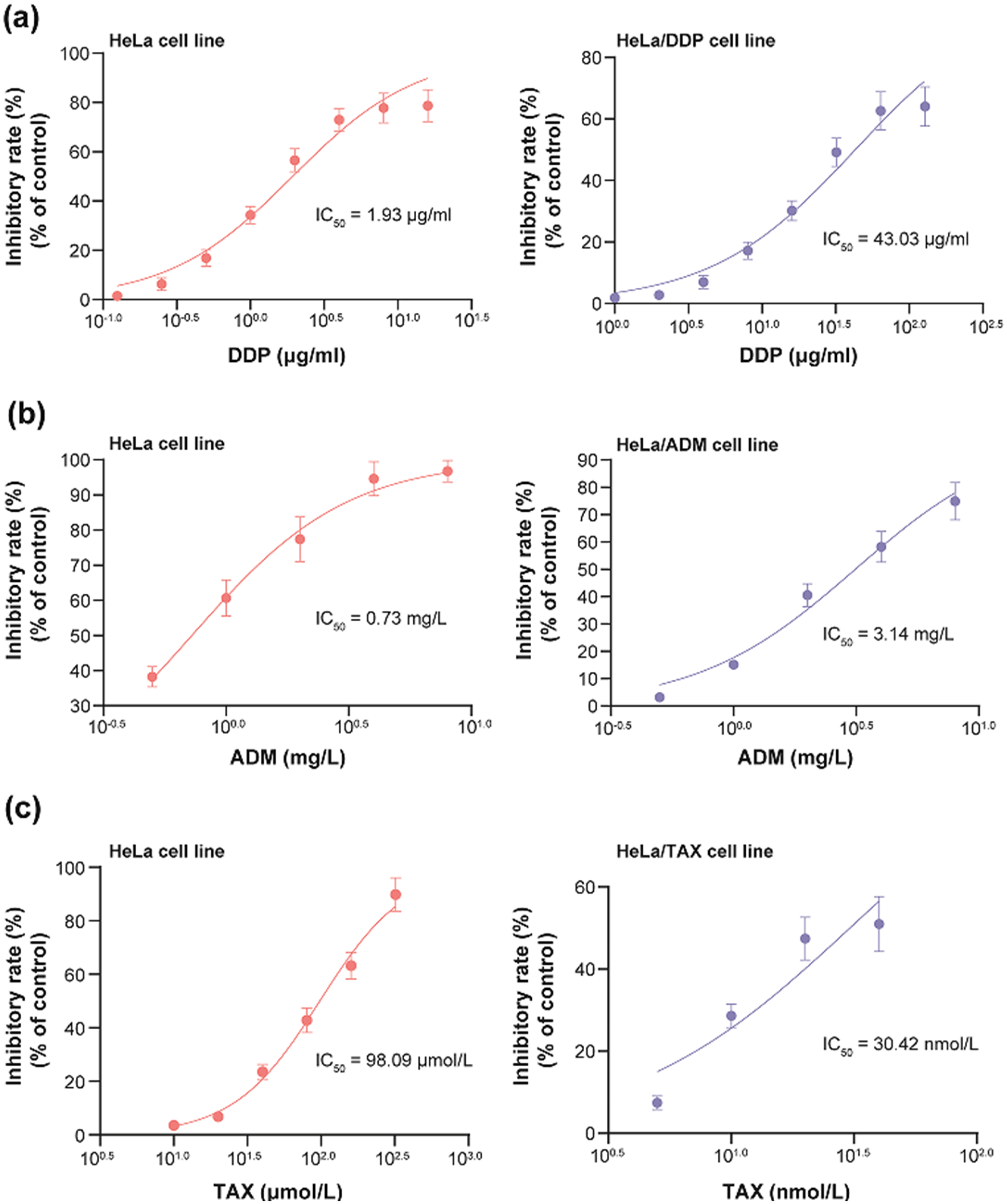
Resistance of cervical cancer (CC) cell lines against cisplatin (DDP) (a), Adriamycin (ADM) (b) and taxol (TAX) (c).
In comparison to respective NC groups, over-expressed m6A-adjusted hsa_circ_0101308 (Figure 4(a)) or under-expressed miR-244 (Figure 4(d)) attenuated viability of both HeLa cell line and HeLa/DDP cell line, which, on the contrary, was strengthened when m6A-adjusted hsa_circ_0101308 was silenced (Figure 4(a)) or miR-224 mimic was transfected (Figure 4(d)) (P < 0.05). Besides, viability of HeLa cell line and HeLa/ADM cell line was weakened as pcDNA3.1-(m6A-adjusted hsa_circ_0101308) (Figure 4(b)) or miR-224 inhibitor (Figure 4(e)) was transfected (P < 0.05), yet was motivated in case si-(m6A-adjusted hsa_circ_0101308) (Figure 4(b)) or miR-224 mimic (Figure 4(e)) was transfected (P < 0.05), as relative to corresponding NC groups. Eventually, HeLa cell line and HeLa/TAX cell line were associated with weakened viability supposing pcDNA3.1-(m6A-adjusted hsa_circ_0101308) (Figure 4(c)) or miR-224 inhibitor (Figure 4(f)) was transfected (P < 0.05), however, their viability was stimulated, to a large degree, after transfection of si-(m6A-adjusted hsa_circ_0101308) (Figure 4(c)) or miR-224 mimic (Figure 4(f)), compared with separate NC groups (P < 0.05).
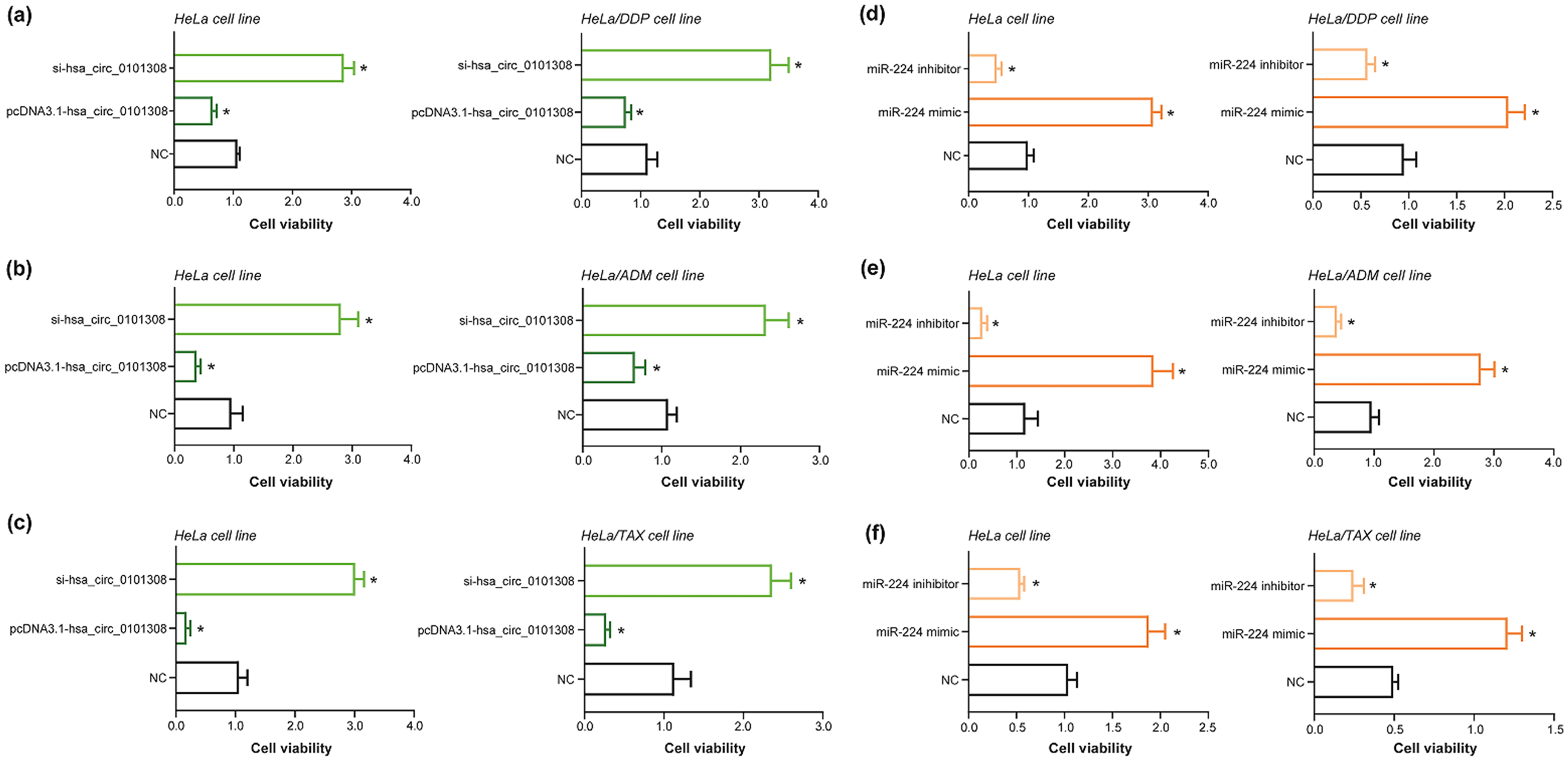
Impact of hsa_circ_0101308 or miR-224 on viability of chemo-resistant cervical cancer (CC) cell lines. (a–c) Viability of HeLa cell line, HeLa/DDP cell line, HeLa/ADM cell line or HeLa/TAX cell line was compared among si-hsa_circ_0101308 group, pcDNA3.1-hsa_circ_0101308 group and negative control (NC) group. *: P < 0.05 in comparison to NC group. (d–f) Among miR-224 mimic group, miR-224 inhibitor group and NC group, the viability of HeLa cell line, HeLa/DDP cell line, HeLa/ADM cell line or HeLa/TAX cell line was contrasted. *: P < 0.05 in comparison to NC group.
PI3K/AKT pathway mediated the effect of M6A-adjusted hsa_circ_0101308/miR-224 axis on viability of drug-resistant CC cells
It was indicated that LY294002 (i.e., PI3K/Akt inhibitor) dramatically suppressed the viability of HeLa cell line and HeLa/DDP cell line that have been transfected by si-(m6A-adjusted hsa_circ_0101308) (Figure 5(a)) or miR-244 mimic (Figure 5(d)) (P < 0.05). Moreover, the enhancive viability of HeLa cell line and HeLa/ADM cell line, due to transfection of si-(m6A-adjusted hsa_circ_0101308) (Figure 5(b)) or miR-244 mimic (Figure 5(e)), became debilitated if LY294002 was replenished to culture cells (P < 0.05). Analogously, LY294002 also significantly brought down the viability of si-(m6A-adjusted hsa_circ_0101308)-transfected (Figure 5(c)) or miR-244 mimic-transfected (Figure 5(f)) HeLa cell line and HeLa/TAX cell line (P < 0.05).
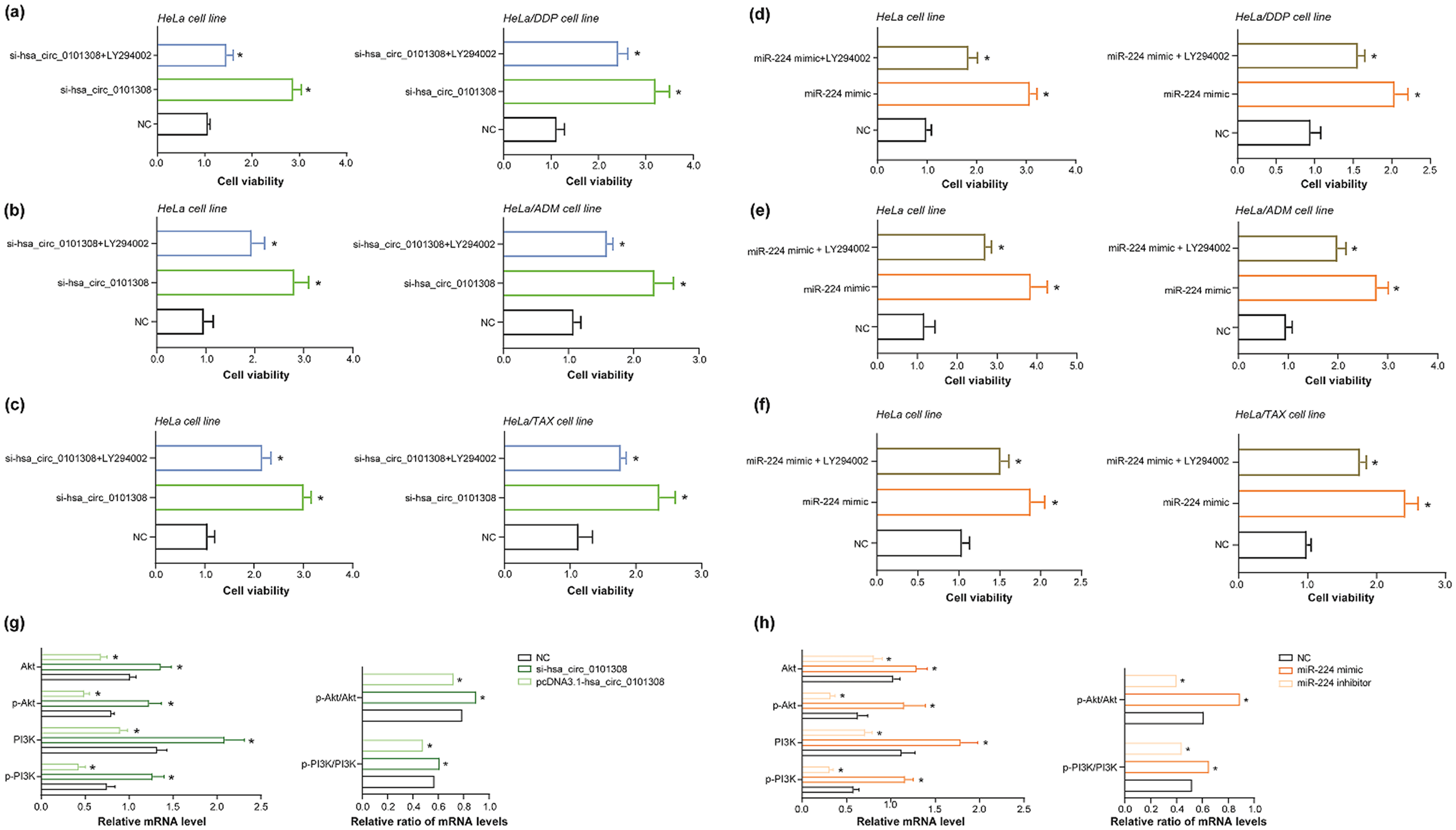
Phosphatidylinositol 3-kinase/phosphorylated protein kinase B (PI3K/Akt) pathway repressed effects of hsa_circ_0101308 or miR-224 on viability of drug-resistant cervical cancer (CC) cell lines. (a–c) Viability of HeLa cell line, HeLa/DDP cell line, HeLa/ADM cell line or HeLa/TAX cell line was contrasted among si-hsa_circ_0101308 + LY294002 group, si-hsa_circ_0101308 group and negative control (NC) group. *: P < 0.05 in comparison to NC group. (d–f) Among miR-224 mimic + LY294002 group, miR-224 mimic group and NC group, viability of HeLa cell line, HeLa/DDP cell line, HeLa/ADM cell line or HeLa/TAX cell line was compared. *: P < 0.05 in comparison to NC group. (g) The mRNA levels of PI3K, p-PI3K, Akt and p-Akt, along with p-PI3K/PI3K ratio and p-Akt/Akt ratio, within HeLa cell line were compared among pcDNA3.1-hsa_circ_0101308 group, si-hsa_circ_0101308 group and NC group. *: P < 0.05 in comparison to NC group. (h) Among HeLa cell lines transfected by miR-224 mimic, miR-224 inhibitor or NC, mRNA levels of PI3K, p-PI3K, Akt and p-Akt, as well as p-PI3K/PI3K ratio and p-Akt/Akt ratio, were compared. *: P < 0.05 in comparison to NC group.
Either silencing of m6A-adjusted hsa_circ_0101308) (Figure 5(g)) or transfection of miR-224 mimic (Figure 5(h)) not merely raised mRNA levels of p-PI3K, PI3K, p-Akt and Akt (P < 0.05), but also boosted p-PI3K/PI3K ratio and p-Akt/Akt ratio in HeLa cell line (P < 0.05), compared with respective NC groups. In contrast, as relative to NC group, transfection of pcDNA3.1-(m6A-adjusted hsa_circ_0101308) was apt to lower both mRNA levels of p-PI3K, PI3K, p-Akt or Akt and ratios of p-PI3K/PI3K or p-Akt/Akt (P < 0.05) (Figure 5(g)), so was transfection of miR-224 inhibitor (P < 0.05) (Figure 5(h)).
MiR-224 intervened in PI3K/AKT pathway via targeting CADM1
TargetScan database (https://www.targetscan.org/vert_80/) predicted that miR-224 targeted CADM1 in specific sites (Figure 6(a)), and the CADM1 has been documented to obstruct cancer development by impeding PI3K/AKT pathway. 41 Luciferase activity of HeLa cell line in the CADM1 WT + miR-224 mimic group overstepped that in the CADM1 MUT + miR-224 mimic group and that in the CADM1 WT + NC group (P < 0.05) (Figure 6(b)). It was further manifested that transfection of miR-224 mimic diminished mRNA levels of CADM1 in HeLa cell line when compared with NC group (P < 0.05), nevertheless, mRNA levels of CADM1 followed a rising trend after transfection of miR-224 inhibitor (P < 0.05) (Figure 6(c)).
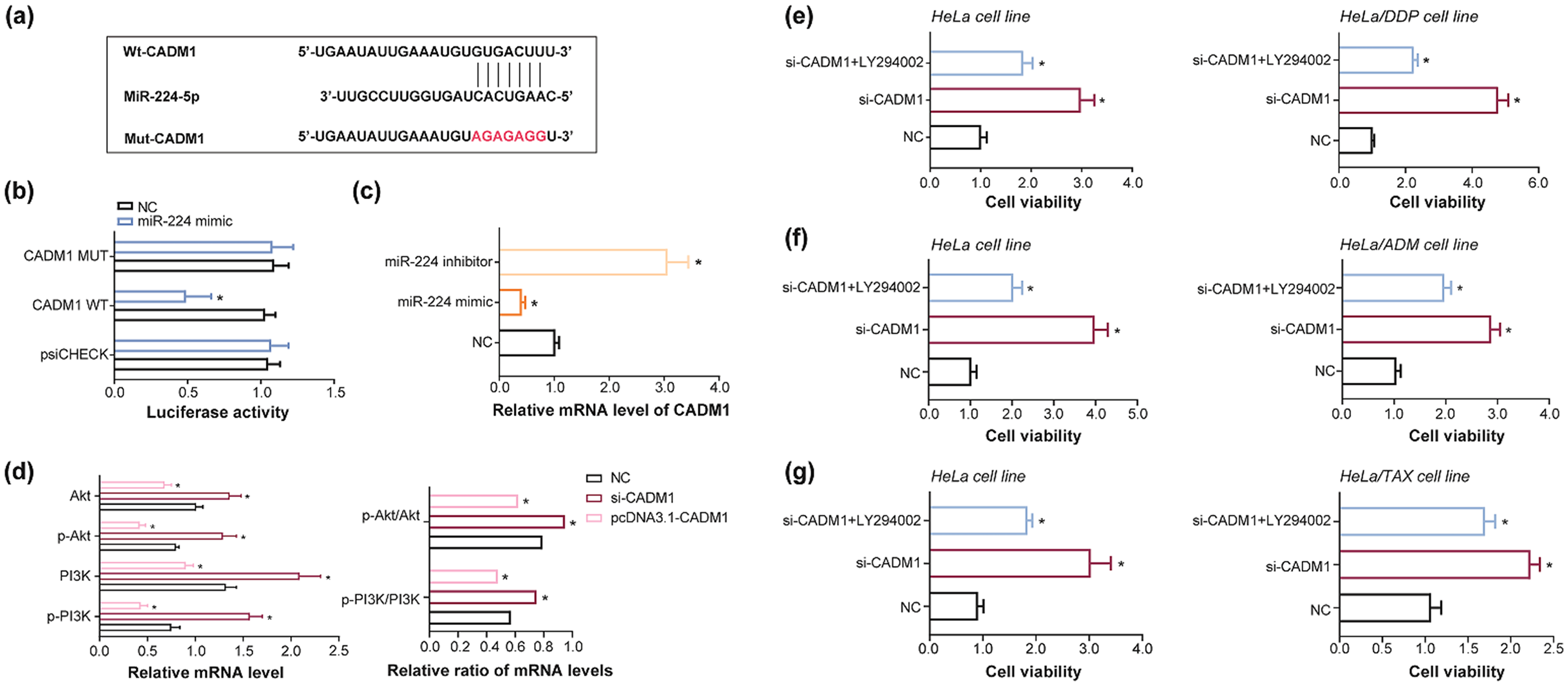
Effect imposed by CADM1-PI3K/Akt pathway network on viability of drug-resistant cervical cancer (CC) cell lines. (a–b) MiR-224 bound to CADM1 in specific sites (a), and luciferase activity of HeLa cells, transfected by miR-224 mimic or negative control (NC), was contrasted among CADM1 Mut + miR-224 mimic group, CADM1 Wt + miR-224 mimic group and CADM1 Wt + NC group group. *: P < 0.05 in comparison to CADM1 Wt + miR-NC group and CADM1 Mut + miR-224 mimic group. (c) The mRNA levels of CADM1 were compared among miR-224 inhibitor group, miR-224 mimic group and NC group. *: P < 0.05 in comparison to NC group. (d) The mRNA levels of p-PI3K, PI3K, p-Akt and Akt, as well as p-PI3K/PI3K ratio and p-Akt/Akt ratio, were contrasted among HeLa cells transfected by pcDNA3.1-CADM1, si-CADM1 and NC. *: P < 0.05 in comparison to NC group. (e–g) Viability of HeLa, HeLa/DDP, HeLa/ADM or HeLa/TAX cell line was compared among si-CADM1 + LY294002 group, si-CADM1 group and NC group. *: P < 0.05 in comparison to NC group.
Furthermore, silencing of CADM1 uplifted mRNA levels of p-PI3K, PI3K, p-Akt and Akt (P < 0.05), accompanied by up-regulation of p-PI3K/PI3K ratio and p-Akt/Akt ratio (P < 0.05), in comparison to NC group, whereas mRNA levels of p-PI3K, PI3K, p-Akt and Akt, as well as p-PI3K/PI3K and p-Akt/Akt ratios, dwindled markedly in the event of pcDNA3.1-CADM1 transfection (P < 0.05) (Figure 6(d)). Concerning LY294002, it imposed contraints over viability of si-CADM1-transfected HeLa cell line and HeLa/DDP cell line (P < 0.05) (Figure 6(e)), and viability of si-CADM1-transfected HeLa cell line and HeLa/ADM cell line fell off greatly under the action of LY294002 (P < 0.05) (Figure 6(f)). Under the influence of LY294002 treatment, viability of si-CADM1-transfected HeLa cell line and HeLa/TAX cell line also dropped significantly (P < 0.05) (Figure 6(g)).
Discussion
On account of persistent HPV infection, as well as premature sexuallife, fecundity, abortion, smoking, mal-nutrition, long-term usage of oral contraceptive and so on, massive females were confronted with severe CC onset. 42 Despite surgery or chemoradiotherapy has been employed for CC treatment, prognosis of CC patients remained far from desirable, partially due to MDR. 43 Hence, illuminating biomolecular mechanisms behind MDR was consequential for CC treatment.44,45
CircRNAs, embodied as stable loops without 5′ cap and 3′-poly(A) tail, 46 could subtly mirror the progression stage of cancers, attributed to their steady expression and their involvement in cancer pathogenesis. 14 For instance, circ9119 expression in CC samples far outweighed that in healthy tissues, suggesting the availability of circ9119 in diagnosing CC onset. 47 Moreover, over-expressed circUBAP2 in CC tissues might be predictive of poor prognosis among CC patients. 48 In this investigation, we observed that under-expressed hsa_circ_0101308, a novel circRNA, in CC tissues was markedly associated with unfavorable clinicopathological features of CC patients, which consisted of poor differentiation, FIGO stage IIA, lymph node metastasis and vascular invasion (Figure 1(a), Supplementary Table 4). For another, highly-expressed hsa_circ_0101308 in CC tissues could imply a favorable prognosis among CC patients, compared with lowly-expressed hsa_circ_0101308 (Supplementary Figure 2).
M6A, a frequently-observed RNA-modification mode within eukaryon, was adept at modifying transcription, splicing, degradation and translation of RNAs. 49 It has been verified that some m6A writers [e.g., methyltransferase-like 3 (METTL3) and RNA-binding motif protein (RBM15)] and certain m6A readers [e.g., YTH N6-methyladenoosine RNA binding protein 2 (YTHDF2)]) became over-expressed, while several m6A erasers [e.g., fat mass and obesity-associated protein (FTO)] turned to be under-expressed within cervical squamous cell carcinoma (CESC) tissues as relative to normal tissues, 50 suggesting that elucidating m6A modification-centered mechanisms was beneficial for estimating cancer progression. Interestingly, m6A broadly appeared within peculiar circRNAs underlying cancer etiology. 51 For instance, m6A modification could accommodate molecular mechanisms inherent in liver metastases of colorectal cancer by virtue of impacting sub-cellular localization of particular circRNAs. 52 Regarding CC, a portion of m6A-adjusted circRNAs (e.g., hsa_circ_0000069) have been identified to express dissimilarly between CC cells or tissues and normal cells or tissues. 53 Nevertheless, investigations relevant to the correlation between m6A-adjusted circRNAs and drug resistance in cancers, especially CC, were still limited. To remedy this part, this study revealed that m6A-adjusted hsa_circ_0101308 sponged miR-224, which insinuated CC exacerbation, 54 and down-regulated its expression (Figure 2(h)), restraining viability of drug-resistant CC cell lines (Figure 4(a)–(c)), on the strength of which CC treatment might be ameliorated.
MiR-224, belonging to post-transcriptional modifiers that enabled degradation or blocked translation of mRNAs, 55 was reported to boost DDP-resistance in ovarian carcinoma partially by means of targeting PRKCD 56 or in lung adenocarcinoma through targeting p21WAF1/CIP1. 57 It was also documented that miR-224 activated 5-fluorouracil (5FU)-resistance in colorecter cancer, owing to effects of its targeted genes on proliferation, invasion or epithelial-mesenchymal transition (EMT) of cancer cells. 58 Over-expression of miR-224, additionally, was found to reinforce docetaxel (DOC)-resistance of breast cancer cells through targeting apoptosis inhibitor 5 (API-5). 32 According to previous investigations mentioned above, miRNA/mRNA axes played pivotal roles in altering resistance of cancer cells. Within this study, we discovered that miR-224 targeted CADM1 to reduce its expression (Figure 6(a)–(c)), and silencing of CADM1 heightened viability of drug-resistant cells (Figure 6(e)–(g)). Furthermore, assisted by miRPath v4.0 database (http://62.217.122.229:3838/app/miRPathv4), Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways of miR-224, made up of adenosine monophosphate activated protein kinase (AMPK) signaling pathway (hsa04152), 59 phosphatidylinositol 3-kinase-serine/threonine-specific protein kinase (PI3K-Akt) signaling pathway (hsa04151), 60 p53 signaling pathway (hsa04115), 61 Ras-proximate-1 (Rap1) signaling pathway (hsa04015), 62 Hippo signaling pathway (hsa04390), 63 Insulin signaling pathway (hsa04910) 64 and FoxO signaling pathway (hsa04068) 65 (Supplementary Figure 6), were summed up. Among them, PI3K-Akt signaling pathway has been highlighted regarding its intimate correlation with CC chemo-resistance. 66 In this study, we observed that m6A-adjusted hsa_circ_0101308, miR-224 and CADM1, linked with PI3K/Akt pathway during cancer exacerbation, 41 attached huge influences on p-PI3K/PI3K or p-Akt/Akt ratio in drug-resistant CC cells (Figures 5 and 6), which expounded the participation of m6A-adjusted hsa_circ_0101308-miR-224-CADM1-PI3K/Akt pathway network in MDR of CC.
Conclusively, m6A-adjusted hsa_circ_0101308 sponged miR-224, which targeted CADM1 and regulated PI3K/AKT pathway, to modify MDR in CC, which was conducive to eradicating drug resistance in CC treatment. Nonetheless, there existed a series of shortcomings. Firstly, pathways that urged m6A modification of hsa_circ_0101308 remained vague, so that mechanisms relevant to CC MDR were not exquisite. Secondly, in-depth studies correlated with animal models were demanded, in order to certify consequences of this study. Thirdly, a wide range of radically diverse CC tissues were demanded to explore the linkage of m6A-adjusted hsa_circ_0101308-miR-224-CADM1-PI3K/AKT pathway network with drug resistance in CC.
Supplemental Material
sj-docx-1-cbm-10.1177_18758592251341157 - Supplemental material for Hsa_circ_0101308 adjusted by N6-methyladenosine (m6A) impacts chemo-resistance in cervical cancer via sponging miR-224
Supplemental material, sj-docx-1-cbm-10.1177_18758592251341157 for Hsa_circ_0101308 adjusted by N6-methyladenosine (m6A) impacts chemo-resistance in cervical cancer via sponging miR-224 by Li Shang, Ruchun Yan, Heng Wang and Zhuyan Li in Cancer Biomarkers
Footnotes
Acknowledgements
None.
Author contributions
ZL, LS and RY: conception. ZL, LS, RY and HW: interpretation or analysis of data. ZL, LS and RY: preparation of the manuscript. ZL, LS and RY: revision for important intellectual content. ZL, LS, RY and HW: supervision. All authors read and approved the final manuscript.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
