Abstract
Background
At present, there are no universal markers of tumor stem cells known, including for B-lymphomas. Previously, we have shown that Epstein–Barr virus-induced B-cell lymphoma culture contains cells capable of internalizing TAMRA-labeled DNA. These cells form sphere-forming centers and are essential for the development of xenografts genetically identical to the initial culture.
Objective
To analyze the stem characteristics of cells that internalize DNA.
Methods
Sorting and RNA sequencing of two subpopulations (TAMRA + and TAMRA–) of Epstein–Barr virus-induced B-cell lymphoma culture and a series of quantitative real-time reverse transcription PCR were performed.
Results
TAMRA + cells were shown to have increased synthesis of mRNA of genes associated with the maintenance of a poorly differentiated state (SOX2, NANOG, POU5F1, CYP26A1), self-renewal (FZD5, FZD7, TCF3, LEF1) and epithelial–mesenchymal transition (MMP2, ITGB7). Transcriptomic analysis revealed that in TAMRA + cells, the synthesis of mitochondrial genes, as well as caspases and some apoptosis inhibitors, is reduced. TAMRA + cells possess clonogenic properties, increased level of synthesis of mRNA for key genes associated with self-renewal and poorly differentiated state maintenance.
Conclusions
Internalization of the TAMRA-DNA probe is the marker of B-lymphoma cancer stem cells and can be used to detect tumor stem cells and develop new approaches to targeted treatment of B-lymphoma.
Keywords
Introduction
B-cell lymphomas. Disease heterogeneity and classification
Non-Hodgkin B-cell lymphomas are a group of heterogeneous diseases that are believed to be caused by recombination events in immune system cells responsible for the molecular diversity of immune responses. The most common lymphoma is diffuse large B-cell lymphoma; it is characterized by distinctive biological features, clinical manifestations, and response to standard treatment. The disease heterogeneity does not allow for adequate use of new targeted therapeutic approaches in clinical practice. For this reason, constant attempts to find the causes of disease onset and determine the molecular mechanisms underlying it that will allow for systematization and adequate application of treatment strategies are being made.1–3
The generally accepted classification of B-cell lymphomas is based on three parameters. They are the type of cell from which the oncological transformation originated,4,5 genotyping results, 6 and molecular properties of the tumor. 7 The new classification of B-cell lymphomas based on molecular profile characterization 2 divides heterogeneous diseases into several distinct subtypes. This makes it possible to use both the available and new therapeutic approaches in a biologically reasonable and more effective fashion.
Malignant transformation of B cells. Existence of cancer stem cells of B-lymphoma
The efforts to classify heterogeneous diseases are made mainly because the original cancer cell and its characteristic surface markers that allow its genotyping among other cancer cells have not been defined for B-cell lymphoma. And the identification of the most malignant clone is the basis of every targeted therapy.
B-cell lymphomas are believed to develop from both committed cells and cells with stem cells properties due to translocations (which are the natural mechanism of B-cell maturation), involving oncogenes. Molecular analysis of breakpoints indicates that some lymphomas develop from bone marrow myeloid precursors or pro/pre-B-cells. 8 Other cases show involvement of somatic hypermutation, a mechanism characteristic of B-cells proliferating in the germinal center after interaction with T-helper cells. 9 The recombinogenic situation in maturing B-lymphocytes with RAG recombinase errors leads to the activation of oncogenes located at the recombination sites. For instance, it has been noted for BCL2 subtypes. 10 However, oncogenes activated due to translocations are known to regulate the signaling pathways responsible for lymphocyte development, proliferation, differentiation, and apoptosis in the majority of cases; however, they do not have a crucial role in acquisition of stem cell self-renewal properties by cells. 11 This means that the premalignant B-cell lymphoma-inducing cell should restore the required stemness, which is the basis for cancer onset and progression, and therefore become a lymphoma-initiating cell. This reprogramming should be the result of secondary and tertiary mutations in genes regulating cell stemness and pluripotency such as the MYC oncogene, which dysregulates the self-renewal program and is recruited in the mechanisms inducing the development of Burkitt lymphoma. 11 There are exceptions for cells that already have stem cell properties such as centroblasts and centrocytes, memory lymphocytes that do not require the second mutation and can induce the tumor in its current state.12,13
Two other ways of transformation of normal B-cells into malignant B-cell lymphoma cells have been reported. They are large-scale remodeling of epigenetic states14–16 and virus-induced B-cell transformation. 17
In general, it appears that B-cell lymphomas do not have pronounced stem cells and that a significant number of clustered malignant B-cells with the above oncogenic changes are involved in cancer development.16,18
Brief history of the research into the Epstein–Barr virus-induced B-cell lymphoma HH47 culture generated by us
For several years, our laboratory has been analyzing the properties of cells obtained from the bone marrow aspirate of a patient with multiple myeloma. Main results have been presented in two papers.19,20 We established that the multiple myeloma clone was replaced by Epstein-Barr virus (EBV)-induced B-cell lymphoma cells. In the initial cell culture, 70% of the cells expressed B-cell markers CD20+/CD45+/CD19dim on their surface, and 30% of the cells expressed myeloid cell markers CD33+/ CD34+/ CD13+/ CD7+. After culturing, 100% of the cells became positive for B-cell markers CD20+/CD45+/CD19dim. Furthermore, the culture acquired a new property: the ability to form sphere aggregates, and dispersed cells formed free-floating spheres during culturing. Genetic analysis, secretome analysis, and EBV infection testing evidenced the presence of EBV-induced B-cell lymphoma. The mechanism of this transformation is described in detail in17,19 and is due to the presence of EBV-transformed B-cells in the primary multiple myeloma culture, which have previously been under immune pressure in the patient's body and have replaced the multiple myeloma clone of the primary culture.
Our numerous studies have demonstrated that stem cells of different genesis can internalize extracellular dsDNA via a natural mechanism. Those can be linear fragments of different length and origin as well as supercoiled plasmids. For historical reasons, we employ the TAMRA + PCR-labeled AluI probe in our studies. Any dsDNA fragment carrying any fluorochrome can be used as a marker probe.
A cell capable of capturing the TAMRA-labelled DNA probe (TAMRA + PCR AluI double-stranded DNA) through the natural mechanism was found to always present in the sphere aggregate (3.3% or an average of 3 cells per sphere). 19 This cell has stem cell characteristics: (1) it is a cell that induces the formation of a sphere-forming center; (2) xenograft is not formed in its absence; (3) the xenograft resulting from transplantation of TAMRA + cell-containing spheres but not TAMRA + depleted spheres is histologically and genetically identical to the transplanted culture. In other words, this is a clonotypic and clonogenic B-lymphoma cell. It secretes specific chemokines IP-10 and MCP-1 that aggregate committed cells into a sphere. An analysis showed that TAMRA + cells are related to neither CD34 + hematopoietic stem cells nor CD90 mesenchymal stem cells. A total of ∼90% of TAMRA + cells express the CD133 marker, and 100% of CD133 + cells internalize the TAMRA DNA probe, which is typical for cancer stem cells (CSCs) of a vast number of cancers. 21 Taken together, the results suggest that it is a stem cell of EBV-induced B-cell lymphoma culture.19,20
Detection of such CSC marker as an ability to internalize extracellular dsDNA fragments for lymphomas allows one to use this new trait as a therapeutic target in treatment of lymphoid hemoblastosis using the Karanahan technology (see detailed description in the last section of Discussion).
Experimental design. General notes
In order to finally identify the hierarchical position of TAMRA + EBV-induced B-cell lymphoma cells, the following experimental approach has been developed. The first stage included cloning of TAMRA+/– cells, which once again confirmed that TAMRA + cells are clonogenic cells of EBV-induced B-cell lymphoma. Genotyping of clone cells showed that they are B-cells. Thus, all unbiased biological tests confirmed that TAMRA + EBV-induced B-cell lymphoma cells are CSCs. The second stage of the study included analysis of mRNA levels of that genes active in TAMRA+/– cells characterizing signaling pathways characteristic of CSCs.22,23
Cells capable of capturing dsDNA fragments carrying a fluorescent label are always detected in EBV-induced B-cell lymphoma cell culture and free-floating sphere aggregates of these cells. 20 A dsDNA probe labeled with the TAMRA fluorochrome was synthesized to detect and isolate this type of cells. EBV-induced B-cell lymphoma cells were incubated with the DNA probe. Next, a series of FACS sorts was performed to collect TAMRA + and TAMRA– cells. After RNAseq of two subpopulations and read trimming, a list of genes expressed in these cells was formed. Genes were divided into two main clusters. The first group included genes with expression level in TAMRA + genes more than tenfold higher than that in TAMRA– cells. The analysis of these genes has previously been conducted in ref., 24 which showed that they are the genes of surface glycoproteins having a glycocalyx structure. According to the recent data, developed glycocalyx is one of the main characteristics of CSCs.25–28 The second cluster included genes characteristic of CSCs. The genes were divided into functional groups responsible for different characteristics of CSCs. These groups included genes encoding signaling cascade proteins, proteins responsible for self-renewal, establishment and maintenance of poorly differentiated state, epithelial–mesenchymal transition (EMT), anti-apoptotic properties, multidrug resistance (MDR), the Warburg effect, and immunosuppression. Having analyzed the RNAseq results, we selected genes from each designated group for TAMRA+/– cells. Involvement of these genes in the signaling cascades responsible for the stemness of cancer cells was independently and reliably confirmed by bioinformatics database analysis STRING: functional protein association networks (string-db.org).
The genes responsible for self-renewal, establishment, and maintenance of a poorly differentiated state are the most important gene groups that characterize TAMRA + EBV-induced B-cell lymphoma cells as CSCs. From these groups and the group of genes responsible for EMT, we selected genes with the greatest difference in expression between TAMRA + and TAMRA– cells, as well as the most functionally significant genes. RT-qPCR was performed for these genes.
The following approach was used to assess the reliability of the results. We selected genes involved in the signaling cascades of interest based on the RNAseq data. The expression levels of these genes were further confirmed by RT-qPCR followed by a comparative analysis of the results obtained by both approaches. RT-qPCR was performed for main candidate genes of the three groups of factors characterizing the properties of symmetric/asymmetric division, establishment and maintenance of poorly differentiated cell status, and EMT. An analysis of the results of RNAseq obtained in previous studies in our laboratory29,30 suggests that changes in the level of gene mRNA up to twofold are not reliable and require confirmation by RT-qPCR. In addition, according to the selected approach to assess the reliability of results, RT-qPCR was performed in the case of an obvious difference in mRNA level between TAMRA + and TAMRA– cells. Each analysis was repeated at least three times for statistical significance. If no difference in mRNA level was detected, PCR was not repeated.
For groups of genes characterizing anti-apoptotic properties, MDR, general cellular metabolism, and immunosuppression, relative expression was analyzed using RNAseq data. The association of proteins encoded by these genes with the biological processes that determine these properties was also assessed.
Materials and methods
Cell culture
EBV-induced human B-cell lymphoma cell culture (designated name HH47) was obtained from the Collection of Pluripotent Human and Mammalian Cell Cultures for Biological and Biomedical Research of the Federal Research Center Institute of Cytology and Genetics of the Siberian Branch of the Russian Academy of Sciences (registration No. HSCC00081).
EBV-induced B-cell lymphoma cells supplanted multiple myeloma cells during culturing. 19 Cells were cultured in DMEM supplemented with 40 μg/ml gentamicin and 10% fetal bovine serum (Sigma-Aldrich, USA) at 37°C and in an atmosphere of 5% CO2.
Synthesis of TAMRA-labeled double-stranded DNA probe
A 500 bp DNA fragment, which is a human AluI repeat sequence, was used as a double-stranded DNA (dsDNA) probe. The probe was generated by PCR using standard M13 primers and the AluI repeat template cloned into the pBS (SK+) vector. A detailed description of the DNA probe production is presented in. 31
Cell sorting
Part of the cells were collected in a falcon tube and centrifuged for 5 min at 4°C and 400 g. Cells were resuspended in 3 ml of phosphate buffered saline and counted in a Goryaev chamber. A total of 400,000 cells were selected as a control and placed in 500 μl of DMEM containing 10% fetal bovine serum. The volume of the sample containing the remaining cells was adjusted to 5 ml and 0.1 μg of TAMRA-labeled dsDNA was added to 2 × 106 cells. Cells were incubated for 40 min in the dark at room temperature and sorted into two populations (with internalized (TAMRA+) and without internalized TAMRA-DNA probes (TAMRA–)) using a BD FACSAria III cell sorter at the Collective Use Center for Microscopy of Biological Objects of the Siberian Branch of the Russian Academy of Sciences.
RNAseq
We performed a cell sorting procedure five times and collected a total of 5 × 106 cells. To isolate total RNA, TRIzol (Thermo Fisher Scientific, USA) was used according to the manufacturer's instructions. An Illumina TruSeq kit was used to prepare RNA libraries. MiSeq (Illumina) was used to perform the RNAseq. The resulting reads were trimmed of adapter sequences using the TrimGalore package (Babraham Institute) and short reads (<20 nucleotides). Reads were mapped to target sequences using the ReadsMap package. The reads were aligned under strict conditions (no more than one insertion/deletion per alignment). Histone genes were used for normalization. Normalization factor was 1.075. Genes were sorted in descending order of the ratio of the number of unique reads per transcript in TAMRA + and TAMRA– cells. According to the literature, functional gene groups characterizing certain properties of CSCs were identified. Genes with altered expression in TAMRA+/– cells were isolated from these groups based on RNAseq data. Validating qPCR was performed for selected genes. The full list of genes obtained after read mapping is presented in Appendix A.
RT-qPCR
Primers for reverse transcription quantitative real-time PCR (RT-qPCR) were selected using NCBI PrimerBLAST. Primers were selected for a region that included borders between the exons, except for cases when the last exon was more than 1 kb long (primer sequences are presented in Appendix B). Specificity of the primers was confirmed using PCR of cDNA obtained using total RNA isolated from EBV-induced B-cell lymphoma cells. Independent cell sorting was performed for each qPCR round. Total RNA was isolated using the RiZol reagent (diaGene, Russia); cDNA was synthesized using the Evrogen reagent kit (Evrogen, Russia) according to the manufacturer's instructions.
The reaction was performed using the SYBR Green PCR Master Mix kit (Biolabmix, Russia). The housekeeping gene Rplp0 was used as a reference gene. A total of 1–3 ng of cDNA was used per one qPCR reaction sample. Each sample was analyzed in triplicate. A total of 3 to 5 independent qPCRs were performed for each gene. The analysis was performed using QuantStudio 5 system (Thermo Fisher Scientific, USA) and QuantStudio Design & Analysis software. The relative expression of the selected genes was evaluated using the ΔΔCt method. 32
Statistical data analysis
Curves were obtained using GraphPad Prism v.8 (GraphPad Software, USA). The reliability of the results was assessed using the nonparametric Mann–Whitney U test and Statistica 10 software.
Results
Cloning of TAMRA + cells and genotyping of resulting cells
TAMRA + cells were cloned. As a result, three clones were obtained (from 192 seed cells; 1.56%), one of which yielded the transplanted culture. The analysis performed demonstrated that a similar cell line with a typical biological property was obtained, namely, the ability to form aggregates in spheres. Cell genotyping showed that they are immortalized B-cells (Figure 1).
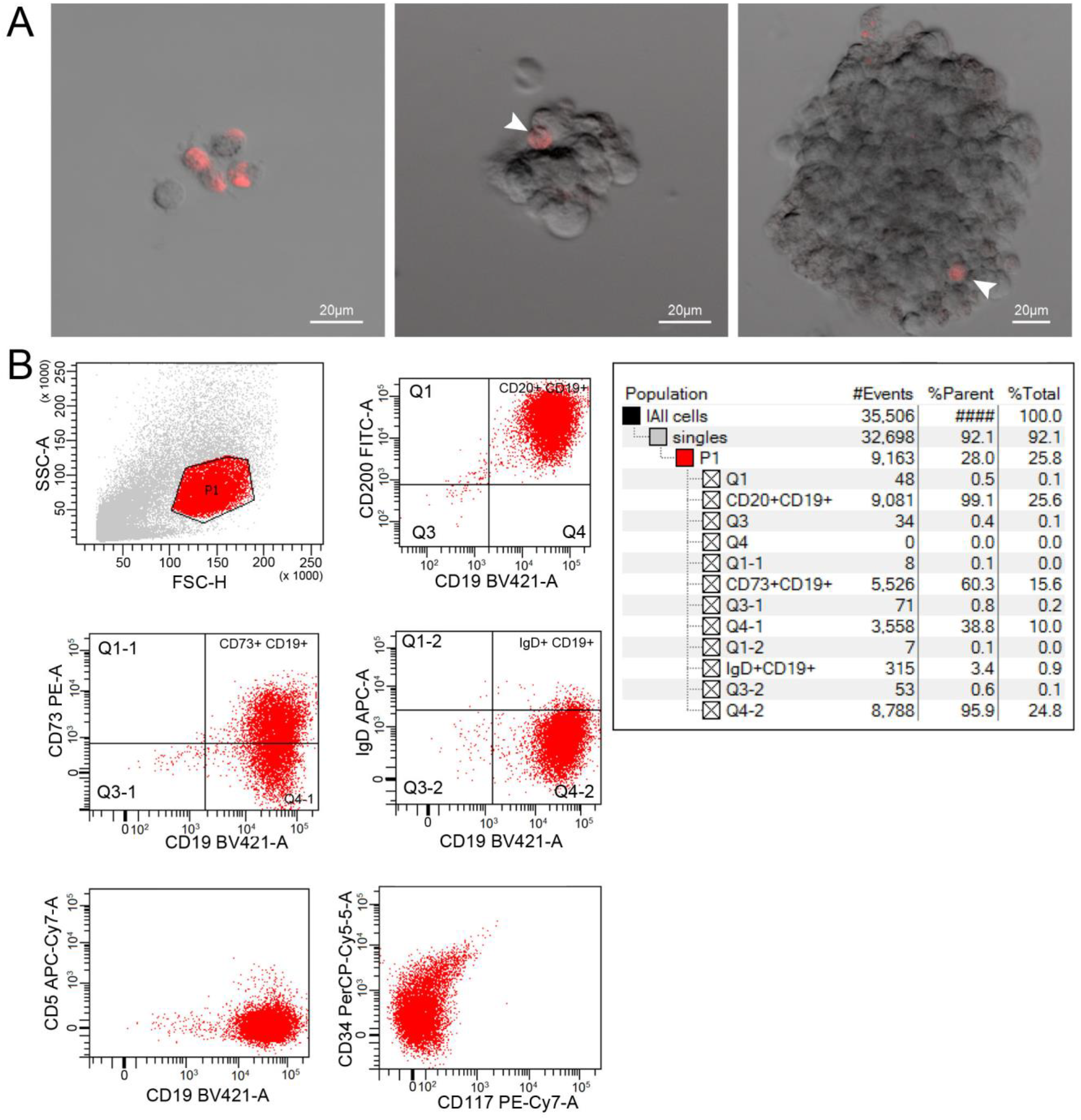
Internalization of TAMRA-DNA probe in clone cells obtained from a TAMRA + EBV-induced B-cell lymphoma cell. A) Different images with individual cells and spheres. B) Clone genotyping for B-cell surface markers.
Characterization of the expression of the genes responsible for the CSC properties of TAMRA+/– cells
Symmetrical/asymmetrical division
RNAseq results showed that expression of the following Wnt signaling pathway genes is increased in TAMRA + cells: Wnt ligands, transcription factors activating transcription of target genes, and some target genes. The expression of the receptors FZD and LRP and genes coding for inhibitors of the canonical Wnt signaling pathway is decreased. Bioinformatics analysis demonstrated that factors of the analyzed group form an interaction network characteristic of the Wnt signaling pathway. According to qPCR data, expression of the Wnt ligand WNT5A, receptors FZD5 and FZD7, and transcription factors TCF3 and LEF1 activating the transcription of target genes for the Wnt signaling pathway is increased (p < 0.05) (Figure 2A).
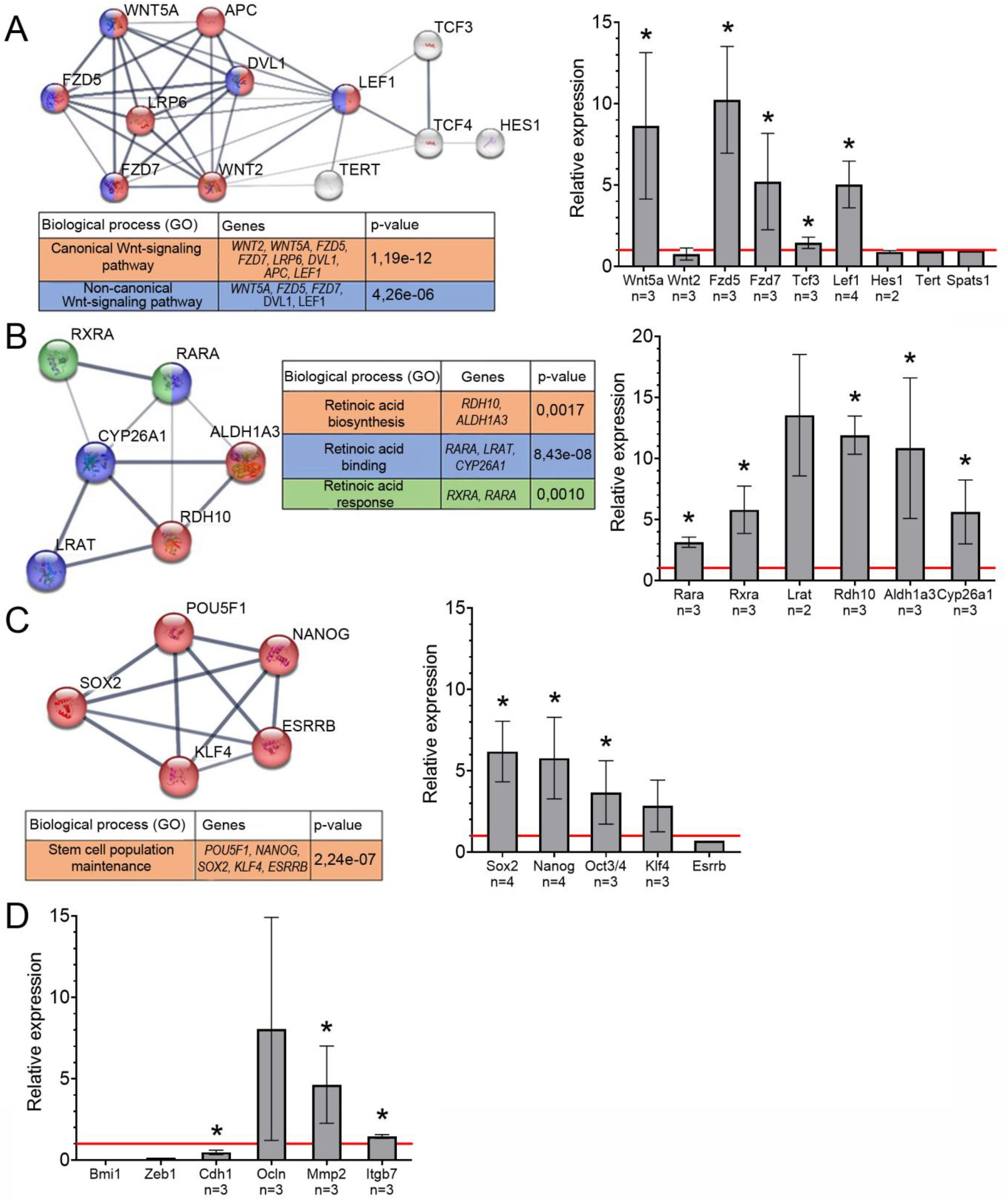
Schematic representation of protein interactions, biological processes associated with them and relative expression of the genes encoding these proteins based on qPCR data. STRING functional protein association networks are presented. The diagrams indicate expression of genes in TAMRA + cells relative to TAMRA– cells (taken as 1, the red horizontal line). The data are means ± SEM, the results from several different qPCR from different sortings (n indicate the number of experiments). The statistically significant differences compared to TAMRA– cells are denoted as *, p-value < 0.05, Mann-Whitney U test. (A) Wnt signaling pathway. (B) Retinoic acid cascade. (C) Transcription factors associated with maintenance of poorly differentiated state. (D) Epithelial-mesenchymal transition.
The results of the RNAseq showed decreased expression of the retinoic acid receptors RARA and the RXRA and enzymes responsible for the conversion of retinol to retinoic acid LRAT, RDH10, and ALDH1A3 in TAMRA + cells compared to TAMRA– cells. At the same time, the expression of the CYP26A1 enzyme responsible for retinoic acid degradation is reduced. Bioinformatics database analysis shows that the analyzed factors form an interaction network characteristic of the retinoic acid signaling pathway. Quantitative PCR showed increased expression of receptors and enzymes involved in retinoic acid biosynthesis and degradation in TAMRA + cells compared to TAMRA– cells (p < 0.05) (Figure 2B).
In addition, qPCR of the mRNA levels of the transcription factor genes responsible for the poorly differentiated state of the CSC such as SOX2, NANOG, OCT3/4, and KLF4 was analyzed.33,34 On the basis of RNAseq data, these genes were included in the list of genes active in the analyzed cells. Quantitative PCR showed elevated expression of SOX2, NANOG, and OCT3/4 and a trend to increase for KLF4 mRNA in TAMRA + cells (p < 0.05) (Figure 2C). The analysis of the bioinformatics database shows that the analyzed group of transcription factors forms a functional network responsible for maintaining the poorly differentiated state of the stem cells.
Epithelial–mesenchymal transition
Based on RNAseq data, the expression of transcription factors BMI1, LEF1, ZEB1, and TWIST, as well as genes encoding proteins that provide apical-basal cell polarity, E-cadherin (CDH1), occluding, and Rho-GTPases, changed insignificantly in TAMRA + cells. Expression of transcription factors BMI1 and ZEB1 and E-cadherin is decreased in TAMRA + cells. The levels of occluding, MMP2 metalloproteinase, and integrin β7 were increased (Figure 2D).
Anti-apoptotic properties
According to RNAseq, the expression of genes associated with induction of apoptosis differs between TAMRA + and TAMRA– cells. In both cells, the expression of some apoptosis inhibitors is increased, while the expression of others is decreased. The expression of the main signaling (CASP8, CASP9) (Figure 3A) and effector caspases 3 and 7 is also reduced in TAMRA + cells
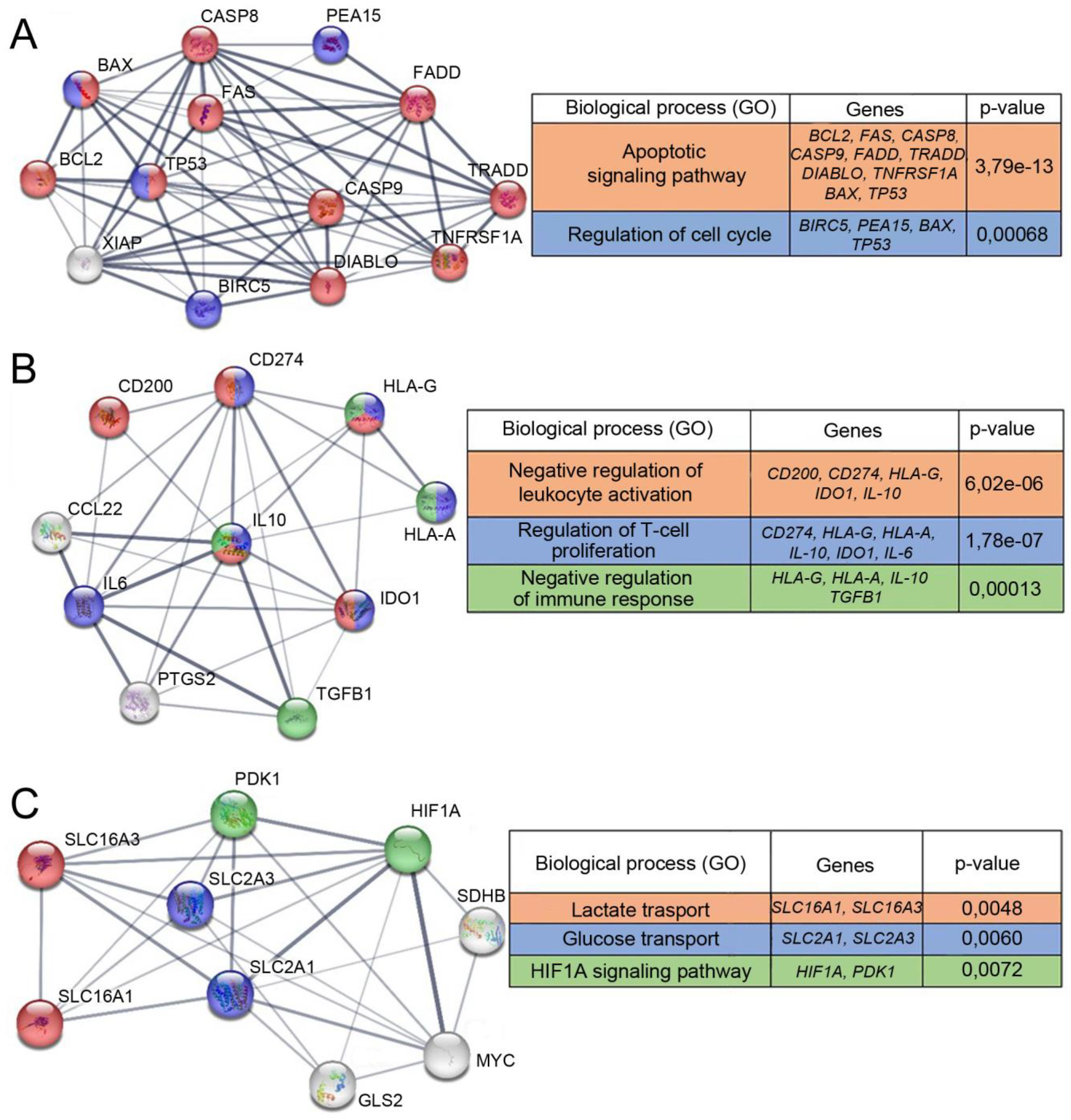
Schematic representation of protein interactions and biological processes associated with them. The STRING functional protein association networks are presented. (A) Proteins responsible for anti-apoptotic properties. (B) Proteins associated with immunosuppression. (C) Proteins characterizing cell metabolism.
Immunosuppression
According to RNAseq data, the expression of effector T cell inhibitors such as IL-10, IL-6, TGFB1, CCL22, CD274, and IDO1 is slightly reduced (within error) in TAMRA + cells. The expression of the T cell inhibitor CD200 and the major histocompatibility complex class I, G, which has an immunosuppressive function, is increased (Figure 3B). Bioinformatics database analysis demonstrates that factors of the analyzed group form a network of interactions characteristic of the body's immune response modulation.
General metabolic activity
According to RNAseq data, the expression of mitochondrial genes is reduced in TAMRA + cells compared to TAMRA– cells. The relative expression of the genes responsible for glycolysis (HIF1A, MYC, SLC2A1, and SLC2A3) is also decreased in TAMRA + cells. This indicates a lower level of metabolic activity in TAMRA + cells. The transcriptional activity of genes responsible for either the Warburg effect or the reverse Warburg effect is slightly altered (Figure 3C). The analysis of the bioinformatics database indicates that the analyzed group of factors forms a network of interactions that unite several metabolic pathways responsible for energy production.
Multidrug resistance
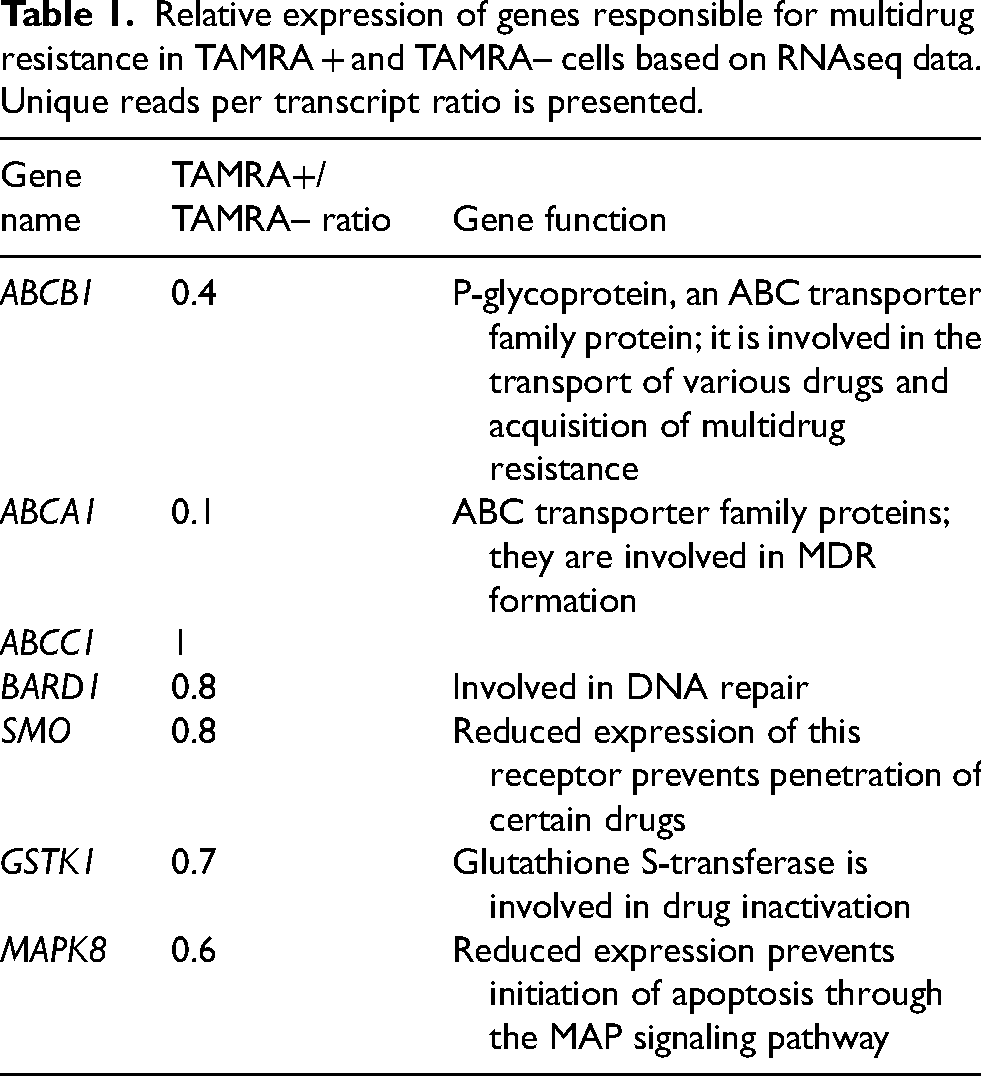
According to RNAseq data, the expression of ABC transporters, the BARD1 and GSTK1 genes, the SMO receptor, and a protein involved in the MAP signaling pathway (MAPK8) is reduced in TAMRA + cells (Table 1). Both MDR and EMT present a set of specific properties that allow the CSC to be insensitive to various xenobiotics and effects of radiation. Each of these properties is regulated by the activity of a specific signaling pathway (Table 1).
Relative expression of genes responsible for multidrug resistance in TAMRA + and TAMRA– cells based on RNAseq data. Unique reads per transcript ratio is presented.

Discussion
Cloning TAMRA + HH47 EBV-induced B-cell lymphoma cells. Characterization of stabilized cell lines generated from a single TAMRA + cell
Our xenograft experiments have clearly demonstrated that a viable graft cannot be obtained from TAMRA + cells per se (like TAMRA– cells). However, when a TAMRA + cell resided within spheres during grafting (portions of spheres not containing TAMRA + cells were used as control), we managed to produce a xenograft whose molecular characteristics were typical of the parental sphere-forming culture.19,20 It was not surprising, since many hematological cancers under experimental conditions are not grafted without preliminary formation of stromal substrate. 20 In this connection, generation of even a small number of clones from a single TAMRA + cell was a hallmark event. Moreover, these cells gave rise to clones capable of forming spheres can carrying markers identical to those in the parental culture (Figure 1). Furthermore, the spheres formed by cells generated from a single TAMRA + cell always contained a TAMRA + cell; the rest of them were TAMRA– lymphocytes (according to cell typing). It meant that the parental TAMRA + cell divided mostly asymmetrically, and in some cases, symmetrically, giving rise to a small population of new TAMRA + cells capable of organizing into a culture and controlling its existence. The ability to divide both symmetrically and asymmetrically is an integral feature of CSCs.
The broad outline of expression of certain groups of genes characterizing TAMRA + cells of EBV-induced B-cell lymphoma HH47 as CSCs
Considering the results presented in studies19,20 altogether, the following conclusion can be drawn: EBV-induced B-cell lymphoma НН47 culture contains a population of cells with all characteristics of the CSC. The dominating characteristic of this cell type is developed glycocalyx that provides a cell with a positive charge due to the proteins located on the outside of the cytoplasmic membrane. The main groups of proteins that form the positively charged cell surface architecture characteristic of only CSCs are presented by the glycoprotein/proteoglycan (GP/PG), glycosylphosphatidylinositol-anchored protein (GPI-AP), and scavenger receptor (SR) groups of factors. Representatives of these groups vary between different CSCs; however, individual proteins belonging to these groups are always present in a sufficient amount on the cytoplasmic membrane of these types of cells. Their presence allows preservation of the main function of these cells: maintenance of the overall positive charge, which is formed due to heparin-binding domains and groups of positively charged amino acids that are found in the structure of the above factors.24,35
The following proteins carrying specific domains that provide internalization of dsDNA fragments into the CSC intracellular space were identified for EBV-induced B-cell lymphoma HH47. The GP/PG group includes CDH17, COL4A5, GPC2, ITGA5, etc.; the GPI-AP group includes CD14 and GDPD5; SR include MEGF6, SCART1, and SCARF2. 24 As noted in our previous studies, this is the most important characteristic of CSCs. The overall positive charge allows the CSC to immediately form a stem niche in any body part due to electrostatic interaction with negatively charged soma cells, as demonstrated in, 36 allows the cell to avoid anoikis 37 and immune surveillance by activated neutrophils, which also carry an overall positive charge. 38 The positive charge of CSCs is a general biological cause of entry of extracellular dsDNA fragments into the cell. It is possible that the characteristic discovered is the basis for interaction of this type of cells with body energy channels. 39
The transcriptome analysis shows an increase in the mRNA level of the genes of certain functional groups in the TAMRA + CSCs of EBV-induced B-cell lymphoma HH47. These genes include surface glycoproteins and factors determining cell interactions (CDH17, CDH11, TMEM178B, TENM4, TREM1, GYPE, etc.), some receptors regulating the activity of molecular pathways (IGF1R, PBX4, and GRIN), transcription factor genes regulating the activity of other genes (TLE1 and DPF3) and tumor suppressor (PLCD1). The expression of IFI27 increases, which is typical of EBV-positive lymphomas originating from germinal B-cells under conditions of chronic inflammation. 40 Furthermore, an analysis followed by real-time PCR validation of mRNA synthesis indicates that CSC-specific gene cascades are activated in TAMRA + cells. They are the molecular pathways that determine stemness, pluripotency, and epithelial-mesenchymal status.
In the group characterized by symmetric and asymmetric division, increased expression of the following genes was shown: WNT5A ligand, main receptors FZD5 and FZD7, and transcription factors LEF1 and TCF3 (Figure 2A), which indicates the activity of the Wnt signaling pathway in TAMRA + EBV-induced B-cell lymphoma cells. This signaling pathway is active in stem cells, including CSCs, and ensures their ability to self-renew. 41 Its activity in TAMRA + cells characterizes them as CSCs.
In the group of genes associated with the poorly differentiated cell state, retinoic acid cascade proteins and transcription factors active in stem cells (SOX2, NANOG, OCT3/4, and KLF4) were analyzed. According to qPCR data, the expression of genes of the retinoic acid metabolic pathway (receptors and enzymes involved in the biosynthesis and degradation), which trigger terminal differentiation of stem cells, turned out to be significantly higher in TAMRA + cells compared to TAMRA– cells (Figure 2B). The activity of the retinoic acid pathway suggests that TAMRA + cells can continuously differentiate into mature B-cells, forming free-floating sphere aggregates. Moreover, qPCR analysis reliably showed increased mRNA level of the genes of transcription factors SOX2, NANOG, OCT3/4, and KLF4 (Figure 2C), which are markers of poorly differentiated cell status. In total, the results of the analysis on the expression of genes responsible for the poorly differentiated status suggest that TAMRA + cells belong to poorly differentiated cells of EBV-induced B-cell lymphoma. 42
The results obtained for the EMT genes indicate a hybrid cell phenotype with coexpression of epithelial and mesenchymal markers, preserved E-cadherin mRNA level, and decreased expression of transcription factors compared to full-scale EMT. Usually, tumor cells do not undergo complete EMT and remain in an intermediate state, while retaining the properties of both epithelial and mesenchymal cells. Tumor cells with a hybrid phenotype often migrate in the form of multicellular clusters while maintaining intercellular contacts. 43
A simple comparative analysis of functional group genes responsible for anti-apoptotic properties, MDR, Warburg effect, immunosuppression, and general metabolic activity demonstrates a trend towards the activity of the genes characteristic of CSCs.
In TAMRA + cells, the metabolic activity of mitochondria appears to be reduced, which is a characteristic of cells under hypoxic conditions and, mainly, clonogenic B-lymphoma cells located in the center of free-floating sphere aggregates. At the same time, no pronounced expression of genes responsible for alternative metabolic pathways of energy production was detected. For example, the expression of genes involved in glycolysis (Warburg effect) is lower in TAMRA + cells compared to TAMRA– cells. It is possible that an alternative mechanism (the reverse Warburg effect) is active in this case, which allows TAMRA + cells to use lactate synthesized and secreted by the other cells in the sphere aggregates.
The association between number of mitochondria and their functional activity has been studied well for hematopoietic stem cells. Hypoxia is known to be one of the key factors determining the hematopoietic stem cell state, and the majority of quiescent and primitive hemopoietic stem cells are found in hypoxic areas of the bone marrow with low blood perfusion. It is also known that hypoxic conditions are maintained in the hemopoietic stem cell directly by regulating mitochondrial number and activity. Mitochondria are the main organelles producing reactive oxygen species. Mitochondrial oxidative phosphorylation is the main source of cellular reactive oxygen species. Primitive quiescent hemopoietic stem cells with a high potential for stemness were shown to have a low level of reactive oxygen species.44,45 Therefore, it is fair to assume that the low metabolic mitochondrial activity in TAMRA + clonogenic B-lymphoma cells located in the center of free-floating sphere aggregates also characterizes the analyzed cell type as primitive CSCs of EBV-induced B-cell lymphoma HH47.
According to RNAseq results, among the MDR group, the expression of major ABC transporters, an enzyme involved in DNA repair, and glutathione S-transferase, is reduced in TAMRA + cells. Expression of the SMO receptor protein and MAPK8 is also reduced. SMO is responsible for the entry of certain drugs into cells, and its reduced expression can result in cell resistance to these drugs. 46
In the group of genes associated with immunosuppression, RNAseq showed increased levels of the membrane glycoprotein CD200, which is found in CSCs of some cancers and is responsible for inhibition of effector T cells and macrophages. 47 Expression of the major histocompatibility complex involved in immunosuppression is also increased (HLA-G). 48
The results obtained are the final piece of evidence in a series of experiments to support the fact that TAMRA + cells of EBV-induced B-cell lymphoma HH47 are CSCs, as confirmed by gene expression levels.
Characterization of the EBV-induced B-cell lymphoma HH47 line as belonging to the ST2 molecular subtype of B-lymphoma
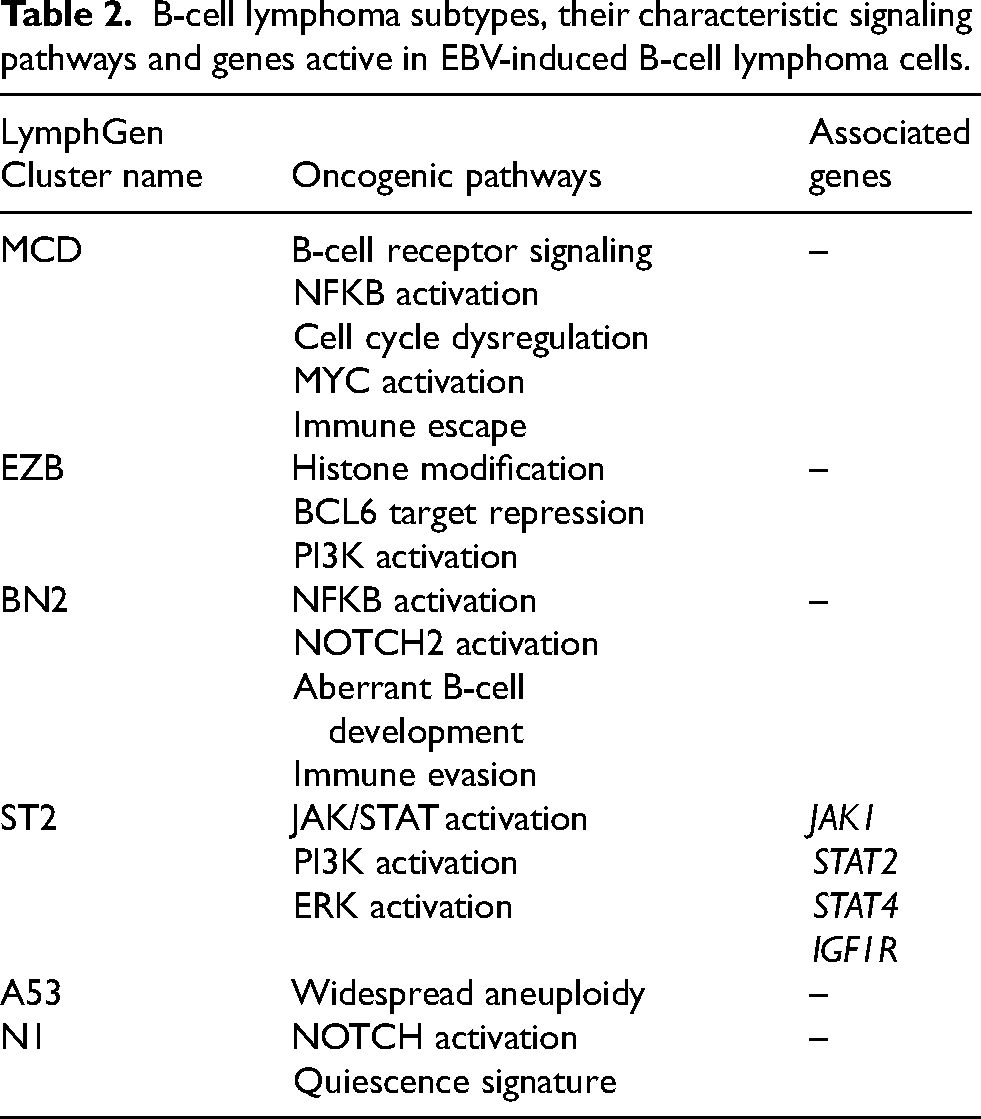
As mentioned above, to date, a large-scale attempt has been made to classify B-lymphomas based on genetic markers. 3 Using RNAseq, we determined the molecular subtype of the EBV-induced B-cell lymphoma HH47 characterized in our studies (Table 2). 2 An increased expression of the following proteins was found in TAMRA + cells: some JAK kinases and STAT kinases (3.4-fold, 1.5-fold, and 1.8-fold increase for JAK1, STAT2, and STAT4, respectively), and IGF1R (25-fold increase), which is involved in positive regulation of the MAPK pathway, has anti-apoptotic activity that promotes the survival of the transformed cell, and is characterized by elevated expression in many tumors. The level of CD14, which participates in the positive regulation of NF-kB, also increased (based on qPCR data). Thus, a pronounced activity of the JAK/STAT kinase genes and the MAPK signaling pathway (based on analysis of mRNA level analysis) is observed in TAMRA + cells. The data obtained suggest that EBV-induced B-cell lymphoma HH47 belongs to the ST2 molecular subtype of B-cell lymphomas.
B-cell lymphoma subtypes, their characteristic signaling pathways and genes active in EBV-induced B-cell lymphoma cells.

Clinical prospects of the new general biological feature of CSCs (ability to internalize extracellular dsDNA via a natural mechanism) for lymphoma treatment. The Karanahan technology
The novel Karanahan technology for treating malignancies of different etiology is based on the ability of CSCs to internalize extracellular dsDNA fragments. The history of developing this technology has been thoroughly described in ref. 49 The original idea was that extracellular fragments (TAMRA + dsDNA probe) delivered into the internal compartments of CSCs may interfere with the chromatin repair process whose damage is caused by cytostatic agents used during chemotherapy. Under certain circumstances, such involvement in repair will have a deleterious effect on CSCs, thus inducing their death. Parameters causing CSC death both under ex vivo conditions and directly in the tumor have been identified in multi-year studies. It was elucidated that if the repair cycle of tumor cells is “locked” by triple treatment with moderate-dose cyclophosphamide, accompanied by treatment of the tumor tissue or culture with a therapeutic dsDNA preparation within selected time periods, most tumor cells, including CSCs, will undergo apoptotic degradation. It turned out that a tiny amount of CSCs survive this treatment and continue the cell cycle, which inevitably causes recurrence. We found that after the aforementioned triple treatment, the survived cells synchronously enter the therapy- susceptible cell cycle G1 phase. According to the Karanahan technology schedule, they undergo the final eradication treatment within this time period, and the tumor nidus loses its tumorigenicity. The remnant tumor is lysed by the activated immune system, and animals are either cured from fulminant malignancies or achieve long-term remission. Cell cycle synchronization is related to sequential repair of cyclophosphamide-induced interstrand cross-links in euchromatin/heterochromatin/centromeric chromatin by final amount of repair complexes. It is believed that the spatial structure of centromeric heterochromatin is repaired in the last turn because repair sites are inaccessible for repair complexes. In this situation, the pooled repair complexes released after repair of simple sites simultaneously start repairing interstrand cross-links of centromeric heterochromatin. Meanwhile, the number of complexes is greater than the number of repair sites. For this reason, both single and multiple damaged sites stochastically present in different cells are repaired simultaneously. Remnant cells synchronously complete the repair and enter the cell cycle, where they undergo final eradication treatment.
The Karanahan technology has been tested in different experimental murine tumor models: Krebs-2 ascites and solid carcinoma,50,51 hepatocellular carcinoma (G-29) with ascites, 52 RLS lymphosarcoma, 53 Lewis carcinoma, 49 as well as human tumor models: U87 glioblastoma 54 and primary glioblastoma cultures. 55 Furthermore, Karanahan technology has been tested in 12 pilot clinical trials focusing on breast cancer treatment. 56
It is noteworthy that the Karanahan technology was used for treating experimentally induced B-cell lymphoma A-20. 57 The aforementioned approach showed a high therapeutic effect, which was significantly more pronounced than that in in situ vaccination with anti-OX40 antibodies within the same experiment.50,57,58
The Karanahan technology was used in a case study involving a patient with large B-cell lymphoma with metastatic spread to the mediastinal, cervical, and axillary lymph nodes. Because patient's condition had critically aggravated during standard chemotherapy, she could not receive further treatment. As tumor was rapidly progressing, the patient underwent six courses of the Karanahan technology with a 3-week interval between the courses. The size of all the lymph nodes decreased during treatment; no intoxication was observed; patient's general condition was stable. Once therapy had been completed, polychemotherapy with rituximab (six courses) became possible. Two and a half years have passed since therapy using the Karanahan technology had been started. The patient is currently in stable remission.
Hence, TAMRA + CSC of B-cell lymphoma is a legitimate therapeutic target, and its eradication from tumor mass can be the basis of a new therapeutic approach to the treatment of B-cell lymphomas.
Conclusions
We conducted a direct experiment that demonstrated the clonogenicity of TAMRA + EBV-induced B-cell lymphoma cells.
A partial analysis of gene expression was performed in two subpopulations of EBV-induced B-cell lymphoma cells. The presence of pronounced differences in the level of mRNA of genes associated with self-renewal, poorly differentiated state, cell metabolism, differentiation, response to apoptosis, etc. shows that TAMRA + and TAMRA– cells belong to two different subpopulations of EBV-induced B-cell lymphoma cells. TAMRA + cells are characterized by activity of signaling cascades characteristic of CSCs. The activity of the Wnt signaling pathway indicates the ability of TAMRA + EBV-induced B-cell lymphoma cells to self-renew. This property is characteristic only for stem cells of various genesis. 41
The active status of signaling cascades responsible for the poorly differentiated state of TAMRA + cells is also a characteristic of stem cells and, in particular, CSCs. 42 TAMRA + cells have an intermediate epithelial-mesenchymal phenotype, which is a characteristic of cancer cells. 43
The obtained findings combined with the results of our previous studies demonstrate that TAMRA + EBV-induced B-cell lymphoma cells belong to CSCs of this cell line.
The results obtained in this study also suggest the feasibility of using the TAMRA + dsDNA probe as a CSC marker. In the future, the ability of CSCs to internalize extracellular dsDNA may become the basis for the development of new therapeutic methods aimed at eradicating CSCs, increasing tumor sensitivity to standard treatment methods, and reducing the risk of disease relapse.
Supplemental Material
sj-xlsx-1-cbm-10.1177_18758592251322040 - Supplemental material for Internalization of extracellular double-stranded DNA as a potential marker of cancer stem cells in Epstein–Barr virus-induced B-cell lymphoma
Supplemental material, sj-xlsx-1-cbm-10.1177_18758592251322040 for Internalization of extracellular double-stranded DNA as a potential marker of cancer stem cells in Epstein–Barr virus-induced B-cell lymphoma by Evgeniya V Dolgova, Sofya G Oshikhmina, Yaroslav R Efremov, Vera S Ruzanova, Anastasia S Proskurina, Svetlana S Kirikovich, Evgeniy V Levites, Genrikh S Ritter, Oleg S Taranov, Olga Y Leplina, Alexandr A Ostanin, Elena R Chernykh, Dmitry N Strunkin, Nikolay A Kolchanov and Sergey S Bogachev in Cancer Biomarkers
Supplemental Material
sj-docx-2-cbm-10.1177_18758592251322040 - Supplemental material for Internalization of extracellular double-stranded DNA as a potential marker of cancer stem cells in Epstein–Barr virus-induced B-cell lymphoma
Supplemental material, sj-docx-2-cbm-10.1177_18758592251322040 for Internalization of extracellular double-stranded DNA as a potential marker of cancer stem cells in Epstein–Barr virus-induced B-cell lymphoma by Evgeniya V Dolgova, Sofya G Oshikhmina, Yaroslav R Efremov, Vera S Ruzanova, Anastasia S Proskurina, Svetlana S Kirikovich, Evgeniy V Levites, Genrikh S Ritter, Oleg S Taranov, Olga Y Leplina, Alexandr A Ostanin, Elena R Chernykh, Dmitry N Strunkin, Nikolay A Kolchanov and Sergey S Bogachev in Cancer Biomarkers
Footnotes
Acknowledgments
The Authors express their gratitude to the Common Use Center for Microscopy of Biologic Objects SB RAS and to Sergey I. Bayborodin for confocal analysis assistance, as well as Collective Use Facility for Flow Cytometry IC&G SB RAS.
Author contributions
Conception: EVD, ASP, SSB
Interpretation or analysis of data: EVD, SGO, YRE, VSR, ASP, SSK, EVL, GSR, OST, OYL, AAO, DNS
Preparation of the manuscript: EVD, SGO, YRE, ASP, SSB
Revision for important intellectual content: YRE, AAO, ERC, NAK, SSB
Supervision: ERC, NAK, SSB
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was funded by the Ministry of Science and Higher Education of the Russian Federation via the Institute of Cytology and Genetics [State Budget Project No. FWNR-2022-0016], as well as Inga N. Zaitseva.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
