Abstract
Objective
Study aims to develop diagnostic and prognostic models for lung adenocarcinoma (LUAD) using Machine learning(ML)algorithms, aiming to enhance clinical decision-making accuracy.
Methods
Data from The Cancer Genome Atlas (TCGA) for LUAD patients were split into training (n = 196) and test sets (n = 133). Feature selection (Least Absolute Shrinkage and Selection Operator (LASSO), Random Forest (RF), and Support Vector Machine (SVM)) identified miRNAs distinguishing stage I LUAD. Six ML algorithms predicted pulmonary node classification. Model performance was evaluated using Receiver Operating Characteristic (ROC) curve, Precision-Recall (PR) curves, and Error Rates (CE). A prognostic model was constructed using Lasso Cox regression. Risk score plots were generated, and model performance was assessed using Kaplan-Meier (K-M) and time-dependent ROC curves. Functional enrichment analyses investigated miRNA function and mechanism.
Results
The feature selection results identified five miRNA molecules as distinguishing characteristics between early-stage LUAD and adjacent non-cancerous tissues. A prognostic model using 13 miRNAs predicted poorer outcomes for patients with higher risk scores, supported by time-dependent ROC curves and a nomogram. Functional enrichment analysis identified cancer-related signaling pathways for the biomarkers.
Conclusion
ML identified a diagnostic five-miRNA signature and a prognostic 13-miRNA model for LUAD, both robust and reliable.
Introduction
Lung cancer is still the main cause of cancer-associated death worldwide. 1 Lung adenocarcinoma (LUAD) is the most common type, accounting for about 40% of all lung cancers. 2 With the wide application of low-dose spiral computed tomography (LDCT), the detection rate of pulmonary nodules is increasing year by year. However, low-dose CT screening for lung cancer often fails to distinguish between benign and malignant early lung nodules. 3 Timely determination of which lung nodules are early lung adenocarcinoma is crucial for early detection and treatment of lung adenocarcinoma. In addition, several factors are routinely used as prognostic indicators for predictive prognosis of LUAD, including tumor size, nodal status, distant metastasis and tumor mutational burden. 4 However, malignant tumors are highly heterogeneous at multiple levels and even patients with the same tumor stage still have a great difference in prognosis and response to therapy.5,6 Sometimes, they do not accurately predict the prognosis of patients. Therefore, it is urgent to find new biomarkers to improve the diagnosis and prognosis of LUAD patients.
The role of miRNAs in the discovery and treatment of tumors is important.7,8 With the advent of high-throughput sequencing technology, which have reduced the per-base sequencing cost, the amount of cancer sequence information submitted to public databases has grown quickly.9,10 Therefore, more and more miRNAs are being explored for cancer diagnosis and prognostic prediction, and their specificity and sensitivity have also been improved. For example, detection of breast cancer serum biomarkers in prediagnostic samples can achieve early detection and prognosis of breast cancer. 11 In addition, autoantigen biomarker signatures can be used for early diagnosis of lung cancer with a sensitivity of 46% and specificity of 83%. 12
In recent years, machine learning (ML), which is a branch of artificial intelligence (AI), has been perceived as a promising tool to analyze high-dimensional omics data sets.13,14 There are two main advantages of machine learning. First, ML can deal with more complex and high-dimensional variables. Second, traditional modeling has poor generalization. 15 Ren Z et al. identified tuberculous pleural effusion using machine learning algorithms. The respective sensitivity and specificity of each of the machine learning TPE diagnostic models were high (logistic regression: 80.5 and 84.8%; KNN: 78.6 and 86.6%; SVM: 83.2 and 85.9%; and RF: 89.1 and 93.6%). Random Forest (RF)model is the most effective diagnostic method. 16 Another study study used machine learning algorithms to classify microarray data. In the first data, Support Vector Machine (SVM)had the highest accuracy rate of 99.23%. 17 Nevertheless, the number of articles that use six machine-learning methods for classification models with high accuracy of early pulmonary nodules is limited. highly sensitive and specific diagnostic models of lung adenocarcinoma developed by machine learning algorithms are still rare.
In this study, the miRNA expression and clinical data was obtained from The Cancer Genome Atlas (TCGA) database. Firstly, the data were randomly divided into a training set and a test set. In the training set, feature selection was conducted by least absolute shrinkage and selection operator (LASSO), random forest and SVM. Subsequently, six machine learning methods including k-nearest neighbors, naive bayes, random forest, decision tree, SVM and extreme gradient boosting (XGBoost) were applied to model and compare the results in our study. We then tested how well these models predict outcomes in test set. Univariate Cox regression analyses was performed to determine predictive miRNAs associated with overall survival. Lasso-Cox regression was used to constructed prognostic model. Subsequently, a prognostic model was constructed through the utilization of Lasso Cox regression. The flowchart for this study was shown in Figure 1.
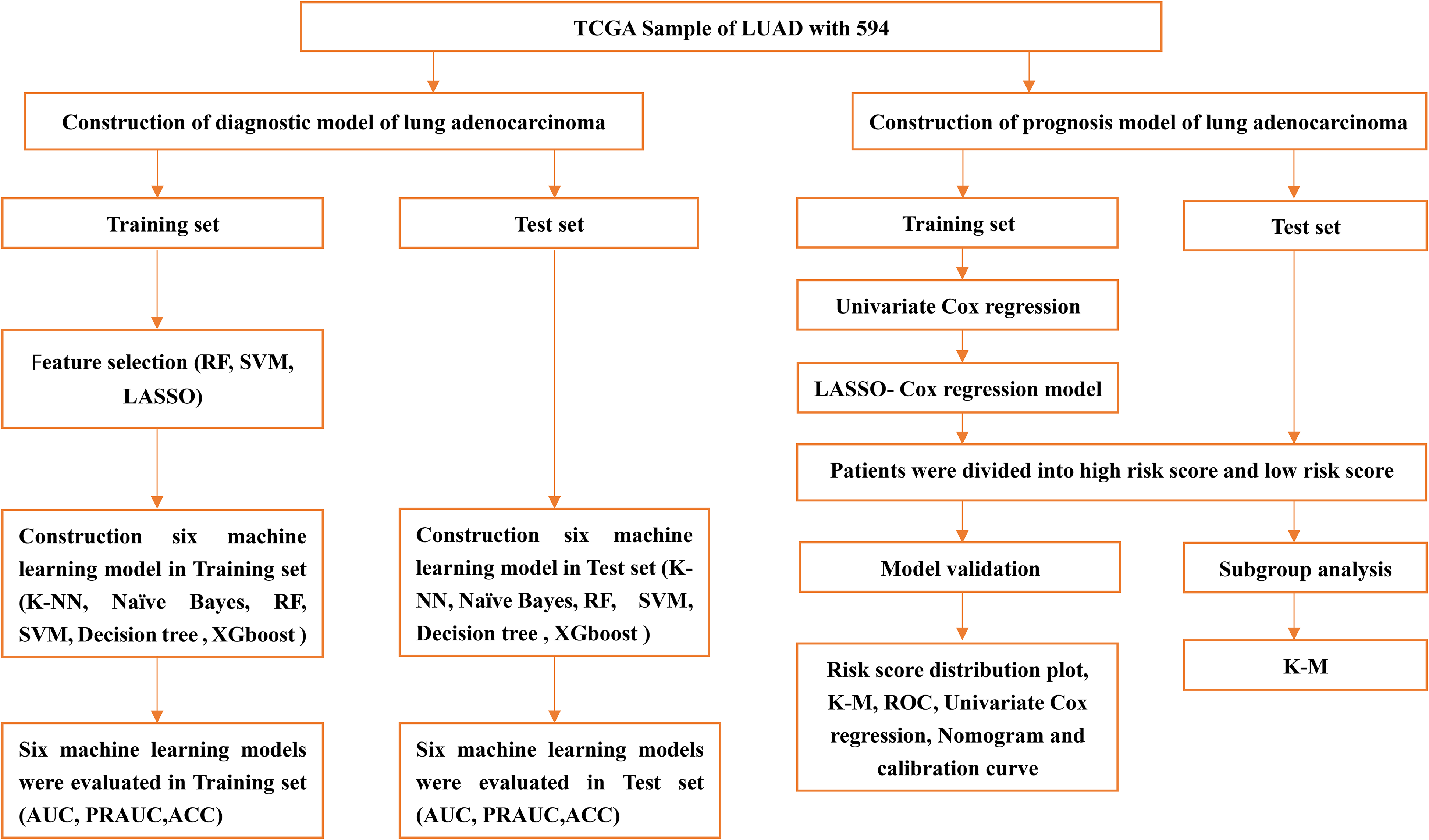
Flowchart of constructing and evaluating diagnostic and prognostic models for lung adenocarcinoma using machine learning algorithms.
Methods
Data sources
The data was downloaded from the TCGA data portal and the miRNA expression and clinical information were determined using the inner_join function from the tidyverse package. Subsequently, the data were randomly divided into a training set and a test set in the ratio of 6:4.
Feature selection
In this study, R language (version 4.2.1) was adopted as the primary tool for data analysis, with the utilization of two R packages, ggplot2 (version 3.3.6) and e1071 (version 1.7.13), to perform SVM analysis and visualize the results. When employing SVM for feature selection, the Radial Basis Function (RBF) kernel was chosen as the kernel type due to its excellent performance in handling nonlinear relationships. Additionally, the model's penalty coefficient (C) and gamma parameters were optimized through grid search and 5-fold cross-validation to identify the optimal model configuration. A random seed number of 2024 was set to ensure the reproducibility of the experiments. By plotting confusion matrices, ROC curves, and precision-recall curves, relevant performance metrics, including classification error rate, precision, and recall, were calculated. Lasso regression method was employed in this study, in conjunction with 10-fold cross-validation and the glmnet package, for feature selection in high-dimensional datasets. By adjusting the regularization parameter lambda (λ) in Lasso regression, variables that significantly contributed to the prediction results were selected. The randomForest package was utilized for feature selection. During the initial training phase, ntree = 800 was set, and a visualization of the error rate versus the number of decision trees was plotted to observe the trend in model performance as the number of decision trees increased. In the formal training phase, ntree = 200 was selected as the final number of decision trees. Simultaneously, the parameters important = TRUE and proximity = TRUE were set to calculate the importance of feature variables and generate a proximity matrix among samples.Construction and evaluation of six machine learning diagnostic models.
Construction prognosis model of lung adenocarcinoma
To identify key variables associated with survival outcomes from a large number of candidate features, the LASSO Cox regression model was adopted. LASSO, an extension of linear regression, achieves feature selection and variable sparsity by introducing an L1 regularization term to compress coefficients. This method facilitates the identification of variables that significantly influence survival time while reducing model complexity. The glmnet package in R was utilized to perform the LASSO Cox regression. The lambda for model fitting was established as the “lambda.1se” value, which was determined through tenfold cross-validation. Subsequently, a risk score was calculated for each sample, determined as a linear combination of expression levels of genes within a signature set. The weighting was provided by their respective LASSO regression coefficients. This calculation followed a previously reported formula, which was: “risk score” = Σ (regression coefficient)×(expression value of each prognostic miRNA). Based on a median risk score, patients were divided into high- and low-risk groups.
Evaluation prognosis model of lung adenocarcinoma
These groups were then subjected to Kaplan-Meier survival analysis, which was conducted using the R survminer and survival packages. The Kaplan-Meier survival analysis method is employed to visually demonstrate the changes in survival probabilities over time for various groups by constructing survival curves, and the log-rank test is used to determine whether the survival differences between different groups are statistically significant. Significance was defined as p < 0.05. The sensitivity and specificity of the prediction model were evaluated through the area under the ROC curve (AUC), utilizing the “timeROC” and “ggplot2” packages. The differential expression of gene signatures between the high-risk and low-risk groups was analyzed using the Mann–Whitney U-test, with statistical significance defined as p < 0.05. The Mann-Whitney U test is applied as a non-parametric test method suitable for handling two groups of data that do not meet the assumptions of normal distribution or homogeneity of variances. By calculating the U statistic and comparing it to critical values, a judgment can be made on whether there are significant differences between the two groups of data. An R software-generated nomogram, incorporating the TNM staging system and risk score, was created utilizing the “rms” and “survival” packages. Graphical assessments of calibration curves involved plotting the observed survival rates alongside the nomogram-predicted survival rates. The discrimination performance of nomograms was quantitatively measured using concordance indices (C-index). For the test set, KM survival curves, nomogram, calibration curves, as well as the distribution of risk scores, survival status, and gene expression heat maps, were all plotted.
Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analyses
To gain a deeper understanding of the potential roles of the identified key genes in biological pathways, KEGG pathway analysis was conducted. KEGG is a database that integrates genomic, chemical, and systems functional information, providing extensive information on biological pathways. By comparing the identified genes with the pathways in the KEGG database, the biological pathways in which these genes participate can be identified, and their potential mechanisms in the initiation and progression of diseases can be explored. The KEGG pathway analysis and GO analysis were conducted using the clusterProfiler R package, along with the R packages org.Hs.eg.db and Goplot. For both GO and KEGG analyses, the significance threshold was set at p.adj < 0.05 and qvalue < 0.2.
Statistical analysis
All analyses were performed using R, version 4.2.1. The diagnostic model was constructed using the mlr3 package of R. Categorical variables were represented through numerical values and percentages. Differences between groups were compared using the non-parametric Wilcoxon signed-rank test. Data visualization was done using the ggplot2 package of R. The ROC plot demonstrated the TPR against FPR. ROC curves and the area under the ROC (AUROC) were calculated using the pROC package for R. Survival analyses were conducted using the statistical packages ‘survival’ and ‘survminer’.
Results
Clinical characteristics
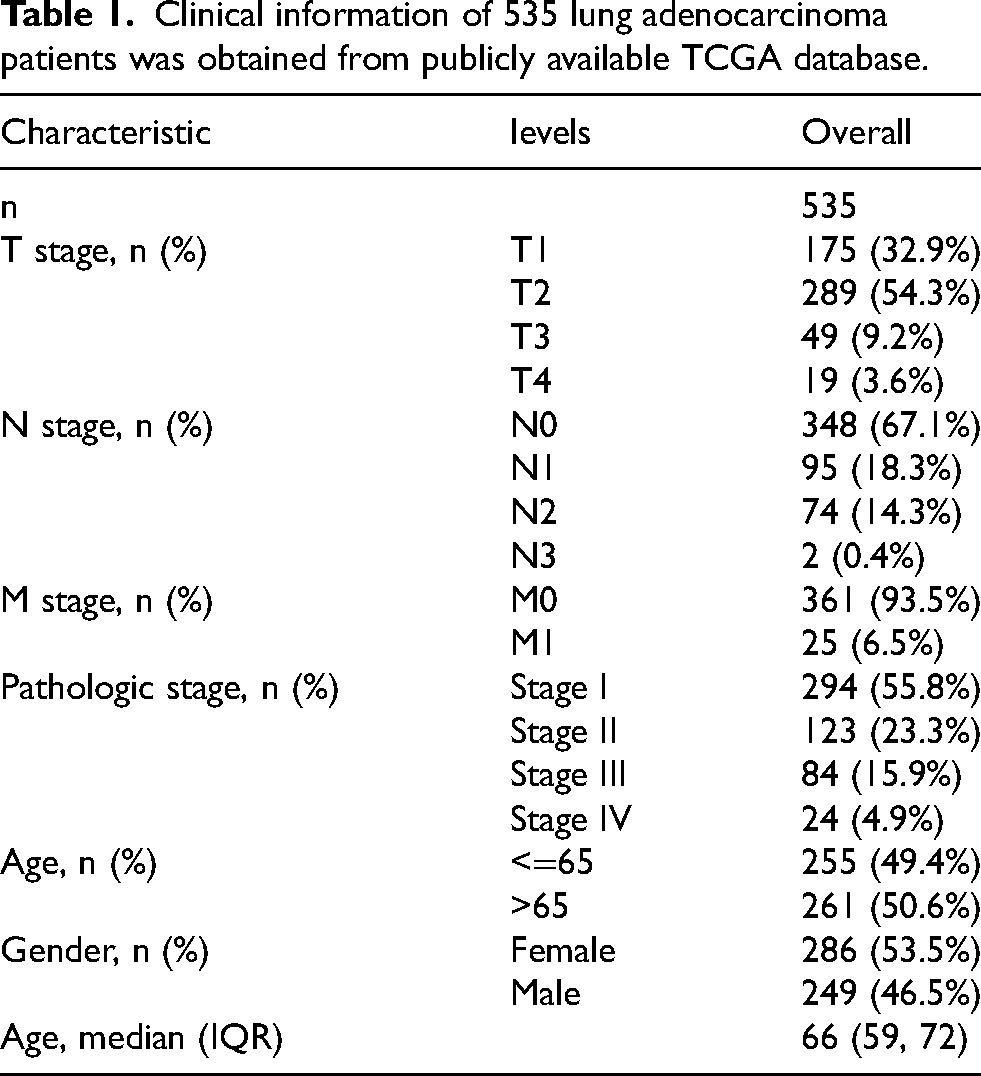
The clinical information of 535 patients was obtained and compiled from TCGA resources. This comprehensive dataset encompassed various parameters such as TNM-stage, clinical stage, gender, race, age, and histologic grade of each patient. These details were carefully recorded and are now presented in Table 1.
Clinical information of 535 lung adenocarcinoma patients was obtained from publicly available TCGA database.
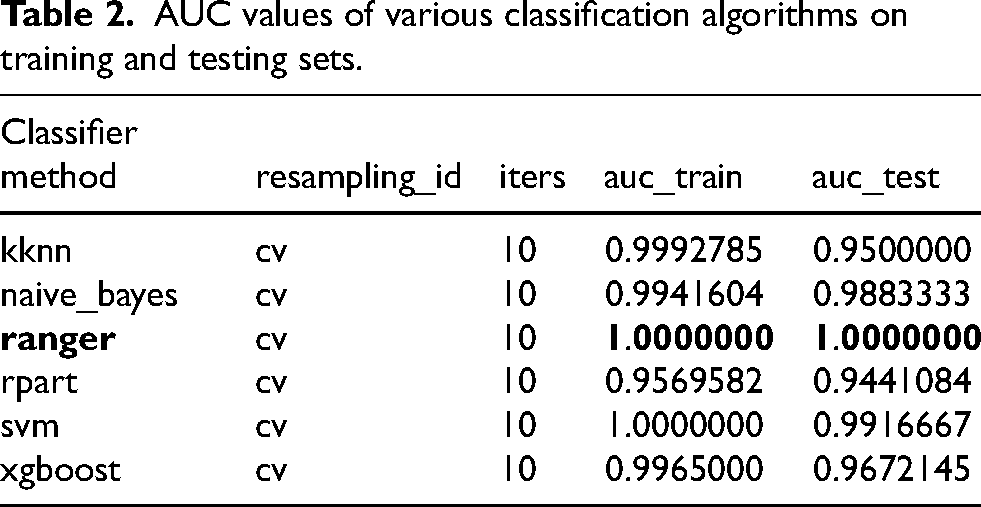
AUC values of various classification algorithms on training and testing sets.
Feature selection
Firstly, a Random Forest model containing 800 decision trees was constructed. It was observed that the error rate stabilized after the number of decision trees reached a certain threshold, indicating model convergence, as shown in Figure 2(A). Subsequently, 200 decision trees were selected as the final model parameter, with the number of variables tried at each split set to 8, and the importance of feature variables was calculated. The result revealed that the estimated error rate of the model on Out-Of-Bag (OOB) data was 1.02%, suggesting high predictive accuracy. The confusion matrix of the model showed that, among the Normal category, 26 samples were correctly classified, and 2 were misclassified as Stage I; while all 168 samples in the Stage I category were correctly classified. Therefore, the model performed well on both categories, especially with very high classification accuracy on the Stage I category. Feature variable selection was conducted using the varSelRF package, selecting the top 20 most important feature variables, with the results presented in Figure 2(B). A SVM model, combined with a Gaussian Radial Basis Function (RBF) kernel, was employed for thorough feature selection and performance evaluation on the given dataset. Through systematic cross-validation and parameter optimization, the optimal model configuration was determined, and the impact of feature variables on model performance was analyzed accordingly. Initially, the kernel function type was fixed as the Gaussian RBF, and the best penalty coefficient (cost) of 2 and the optimal gamma value of 0.00390625 were identified through grid search. These parameter selections aimed to balance the complexity and generalization ability of the model, ensuring optimal performance on unseen data. To assess model performance, a 5-fold cross-validation strategy was adopted, with a random seed number set to 2024 to ensure reproducibility. During cross-validation, the model error rates under different numbers of features/variables were recorded to analyze their impact on model performance. The results indicated that the model achieved the lowest error rate of 0.02013 when the number of features/variables was 50. This not only validated the importance of feature selection but also suggested that 50 feature variables were sufficient to provide adequate information for accurate model predictions on the given dataset. Additionally, it was noted that the minimum number of feature variables corresponding to both the lowest error rate and the highest accuracy was 50, further emphasizing the crucial role of this subset of features in optimizing model performance. For a more intuitive demonstration of model performance, the confusion matrix was also calculated, yielding an error rate of 0.01 and an accuracy of 0.99 under the current kernel configuration. These results demonstrated that the optimized SVM model performed excellently on the given dataset, with high predictive accuracy and robustness. Figure 2(C) displays the SVM feature selection results, with the horizontal axis representing the number of feature variables and the vertical axis representing the feature variable error rate. The annotated points and labels indicate the error rate corresponding to the minimum number of feature variables, representing the minimum number of feature variables associated with the lowest error rate obtained from cross-validation tests. Furthermore, the Lasso regression model combined with 10-fold cross-validation was utilized for variable selection. By setting the seed number to 2024 to ensure reproducibility, cross-validation was performed to obtain the model's prediction accuracy, standard error (SE), and the number of non-zero coefficients under different lambda values. When the lambda value was 0.0019859 (denoted as lambda.min), the model achieved the highest prediction accuracy, with 19 non-zero coefficients. However, considering the model's complexity and generalization ability, a simpler model was preferred. Lambda.1se, with a lambda value of 0.020326, was chosen. At this lambda value, although the model's prediction accuracy declined slightly (with a statistic of 27), the number of non-zero coefficients decreased to 9, indicating a more concise model with potentially better generalization ability. The variable coefficient selection plot is shown in Figure 2(D), and the variable trajectory plot is presented in Figure 2(E). The final result of feature selection, obtained by taking the intersection of the three feature selection methods, is illustrated in the Venn diagram in Figure 2(F). Ultimately, five miRNAs were selected as the result of feature selection from a total of 1881 miRNAs. These selected miRNAs are MIMAT0004597, MIMAT0004584, MIMAT0000243, MIMAT0001620, and MIMAT0000076.
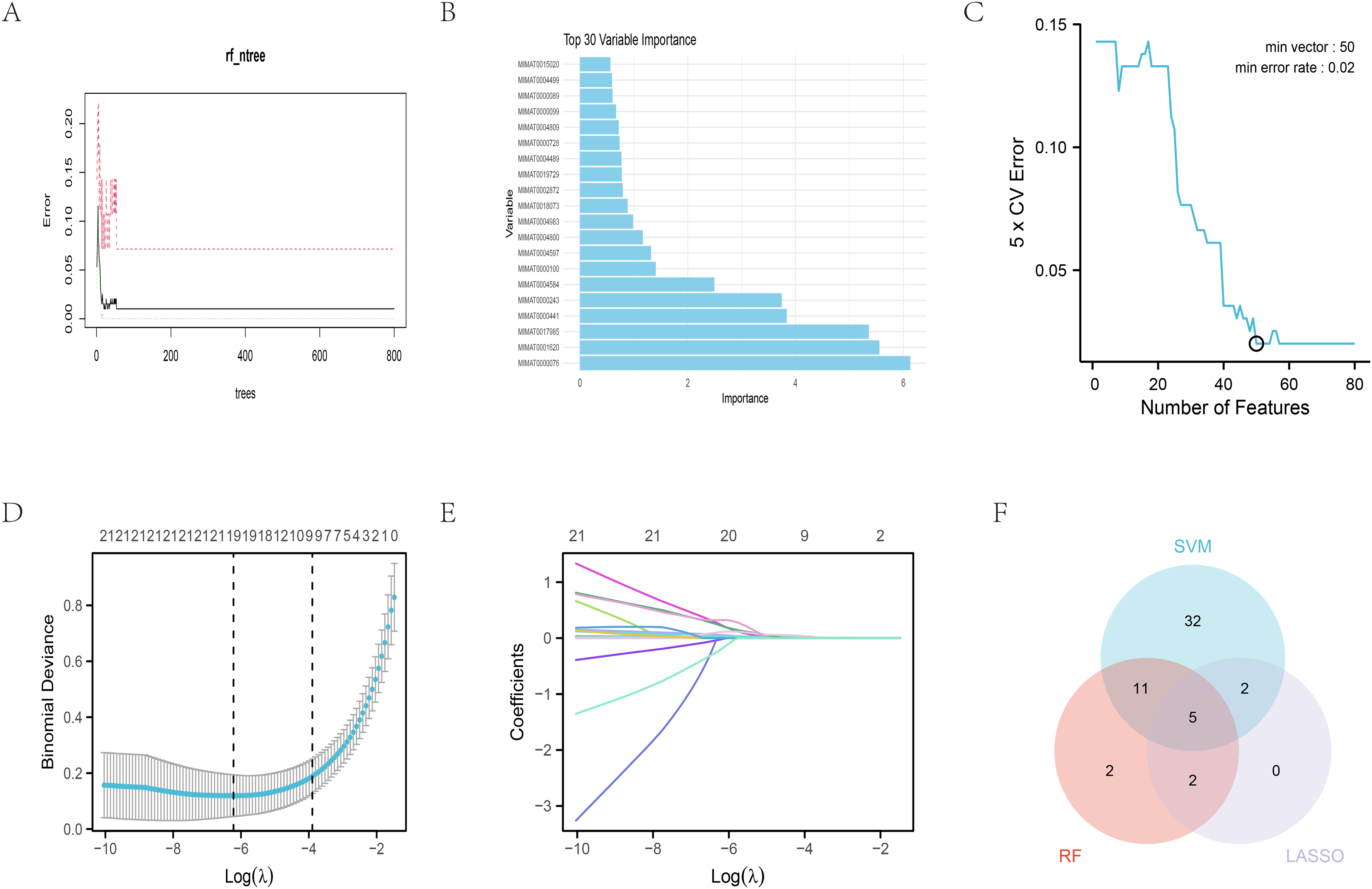
Feature selection. (A) The error rate exhibited by the random forest algorithm, plotted against the varying number of decision trees utilized. On the x-axis, the number of decision trees utilized by the random forest algorithm is depicted, while the y-axis represents the associated error rate (estimated using the out-of-bag method from 800 trees). Black, red and green lines correspond to the gross distribution, stage I lung adenocarcinoma distribution and adjacent normal tissue distribution, respectively. (B) Feature selection of miRNAs using Random Forest. Random forest analysis classifies the levels of importance of miRNAs. The X-axis illustrates the mean decrease in accuracy and Gini coefficients associated with the random forest algorithm. The Y-axis represents a ranking of variables determined by the random forest based on these measures, indicating the relative importance of each variable in predicting the target outcome. (C) The machine learning algorithm of SVM used for classification analysis. The horizontal axis represents the number of feature variables, while the vertical axis indicates the error rate associated with these feature variables. The annotated points and labels in the figure correspond to the error rate achieved with the minimum number of feature variables. (D) Lasso analysis results of miRNAs. The lower horizontal axis depicts the lambda value, while the upper horizontal axis scale indicates the number of variables in the lasso model, where the regression coefficient (x) is non-zero. (E) The trajectory of each independent variable is plotted, with the horizontal axis representing the log value of the independent variable lambda. The vertical axis denotes the coefficient of the independent variable. (F) a Venn diagram of three feature selection methods.
Construction and evaluation of six machine learning diagnostic models
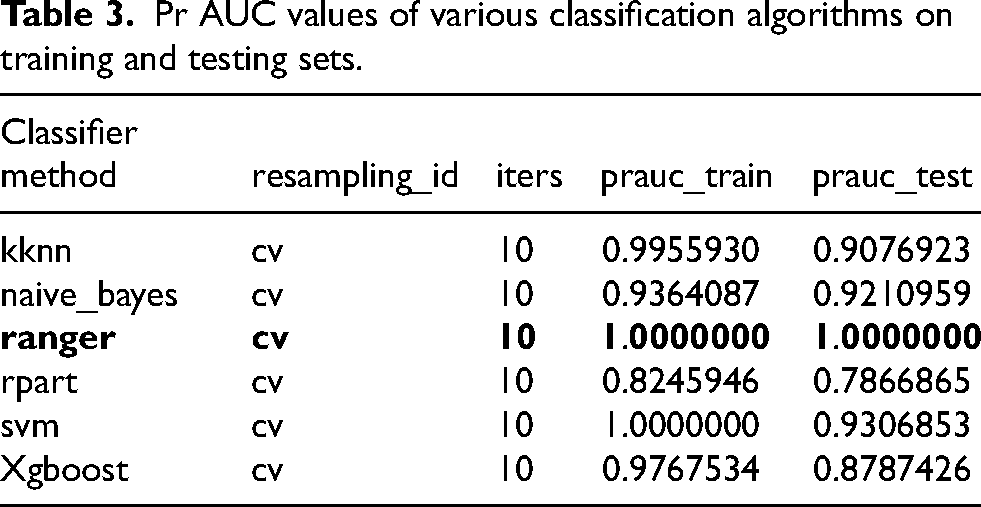
Six classification algorithms were employed for training and testing, with model performance evaluated using two metrics: AUC and PR AUC. Experimental data were based on the cross-validation method, with each algorithm run 10 iterations to ensure stability of the results. Firstly, the AUC values of each classification algorithm on both the training and testing sets were calculated. As shown in Table 2, ranger and svm achieved perfect AUC values (1.000) on the training set, followed closely by kknn and xgboost. On the testing set, classif.ranger remained in the lead with an AUC value of 1.000, while svm and naive_bayes ranked second and third, respectively, with AUC values of 0.9916667 and 0.9883333. The optimal algorithm and the AUC values for both the training and testing sets are displayed in bold font in the table. These results indicate that classif.ranger not only maintains performance on the training set but also demonstrates strong generalization ability. Additionally, the PR AUC values of each classification algorithm on both the training and testing sets were calculated. As shown in Table 3, ranger achieved perfect PR AUC values (1.000) on both sets, suggesting excellent performance in handling imbalanced datasets. The optimal algorithm and the PR AUC values for both the training and testing sets are displayed in bold font in the table. Svm and xgboost also performed relatively well on the testing set, with PR AUC values of 0.9306853 and 0.8787426, respectively. Figure 3 presents the AUC and PR AUC values for the six machine learning classification algorithms.
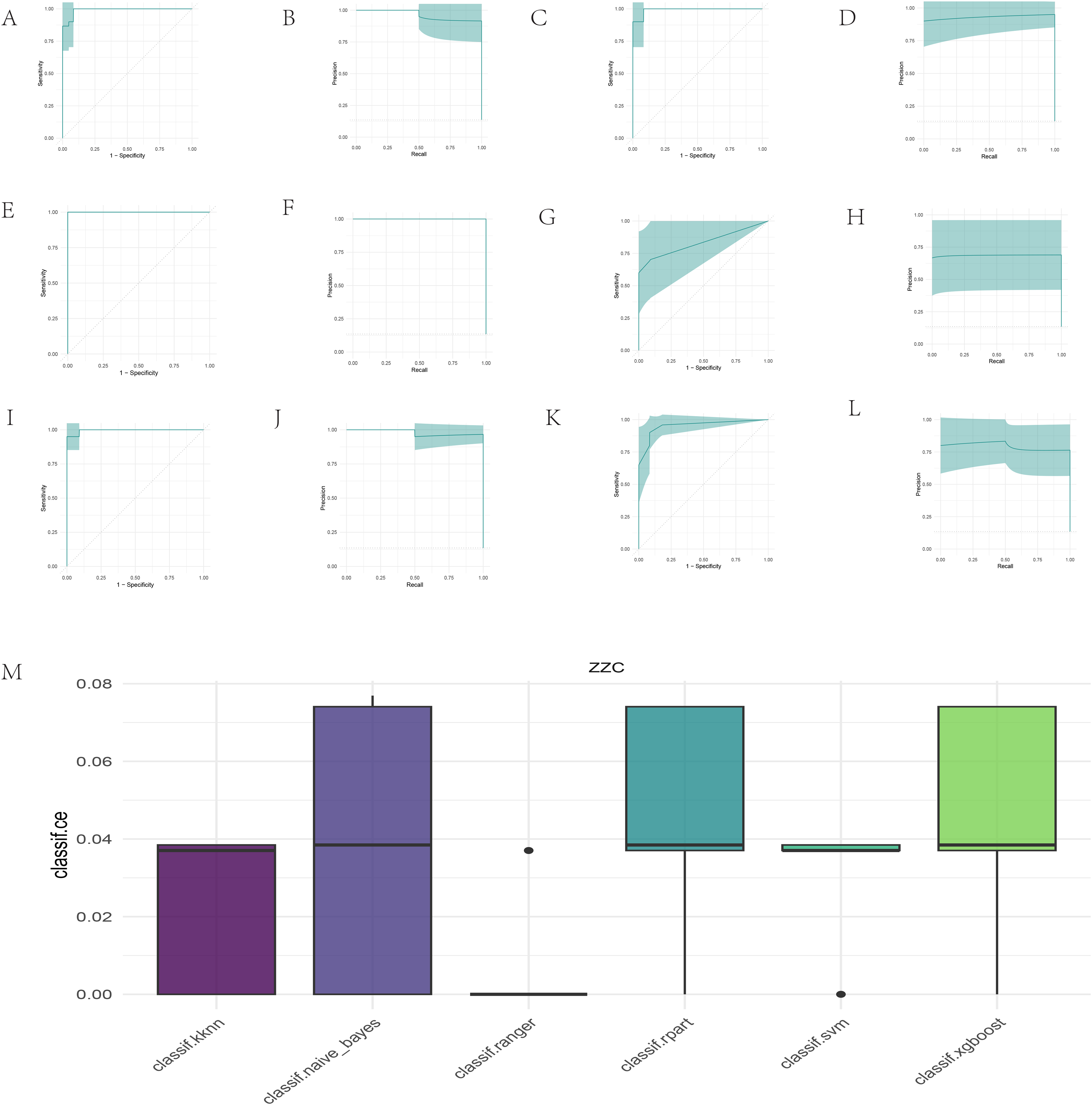
The performance of six machine learning algorithms for miRNA classification. (A) AUC of the KNN algorithm. (B) PRC AUC of the KNN algorithm. (C) AUC of the naive Bayes algorithm. (D) PRC AUC of the naive Bayes algorithm. (E) AUC of the ranger algorithm. (F) PRC AUC of the ranger algorithm. (G) AUC of the rpart algorithm. (H) PRC AUC of the rpart algorithm. (I) AUC of the SVM algorithm. (J) PRC AUC of the SVM algorithm. (K) AUC of the xgboost algorithm. (L) PRC AUC of the xgboost algorithm. (M) The error rates of the six machine learning methods in classifying miRNAs. The horizontal axis represents the classification error rates, while the vertical axis indicates the respective machine learning algorithms.
Pr AUC values of various classification algorithms on training and testing sets.
In summary, this study evaluated multiple classification algorithms using AUC and PR AUC metrics. The results demonstrate that classif.ranger maintains performance on the training set while also exhibiting strong generalization ability and the capacity to handle imbalanced datasets, making it the optimal classification algorithm in this study.
Construction prognosis models of lung adenocarcinoma
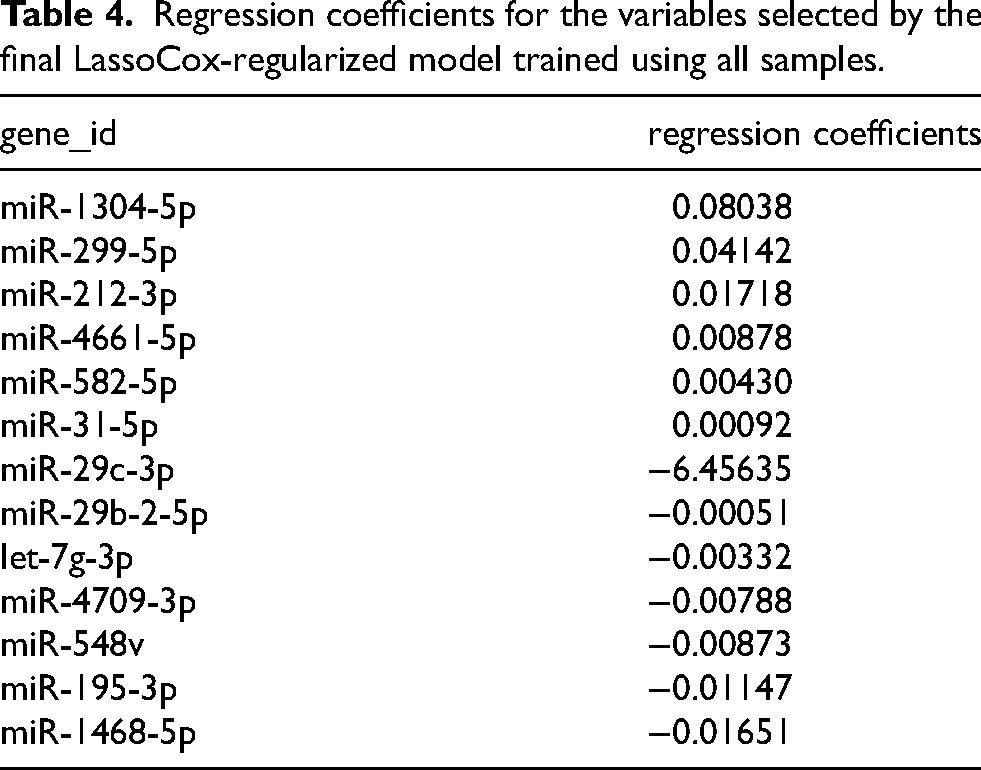
Utilizing machine learning algorithms, 81 miRNAs were identified as significant predictors of prognosis in LUAD patients (P ≤ 0.01) based on univariate Cox regression analysis. Among these, a subset of 13 miRNAs (miR-1304-5p, miR-299-5p, miR-212-3p, miR-4661-5p, miR-582-5p, miR-31-5p, miR-29c-3p, miR-29b-2-5p, let-7g-3p, miR-4709-3p, miR-548v, miR-195-3p, and miR-1468-5p) were selected through LASSO-Cox regression analysis as a prognostic model for LUAD patients. The variable coefficient selection plot is shown in Figure 4(A), and the variable trajectory plot is shown in Figure 4(B). The results of this analysis are summarized in Table 4. The predictive model is formulated as a linear combination of the expression levels of these 13 miRNAs, weighted by their respective coefficients obtained from the multivariate Cox regression. Specifically, the risk score is computed as follows: risk score = (miR-1304-5p * 0.080378882)+ (miR-299-5p * 0.041416811)+ (miR-212-3p * 0.017179747)+ (miR-4661-5p * 0.008779335)+ (miR-582-5p * 0.004307642)+ (miR-31-5p * 0.000926518)+ (miR-29c-3p * −6.45635E-06)+ (miR-29b-2-5p * −0.000512186)+ (let-7g-3p * −0.00332882)+ (miR-4709-3p * −0.007875172)+ (miR-548v * −0.008731379)+ (miR-195-3p * −0.011469174)+ (miR-1468-5p * −0.016509503). Using the median risk score as a threshold, patients in the training set were stratified into low-risk and high-risk groups based on their risk scores.
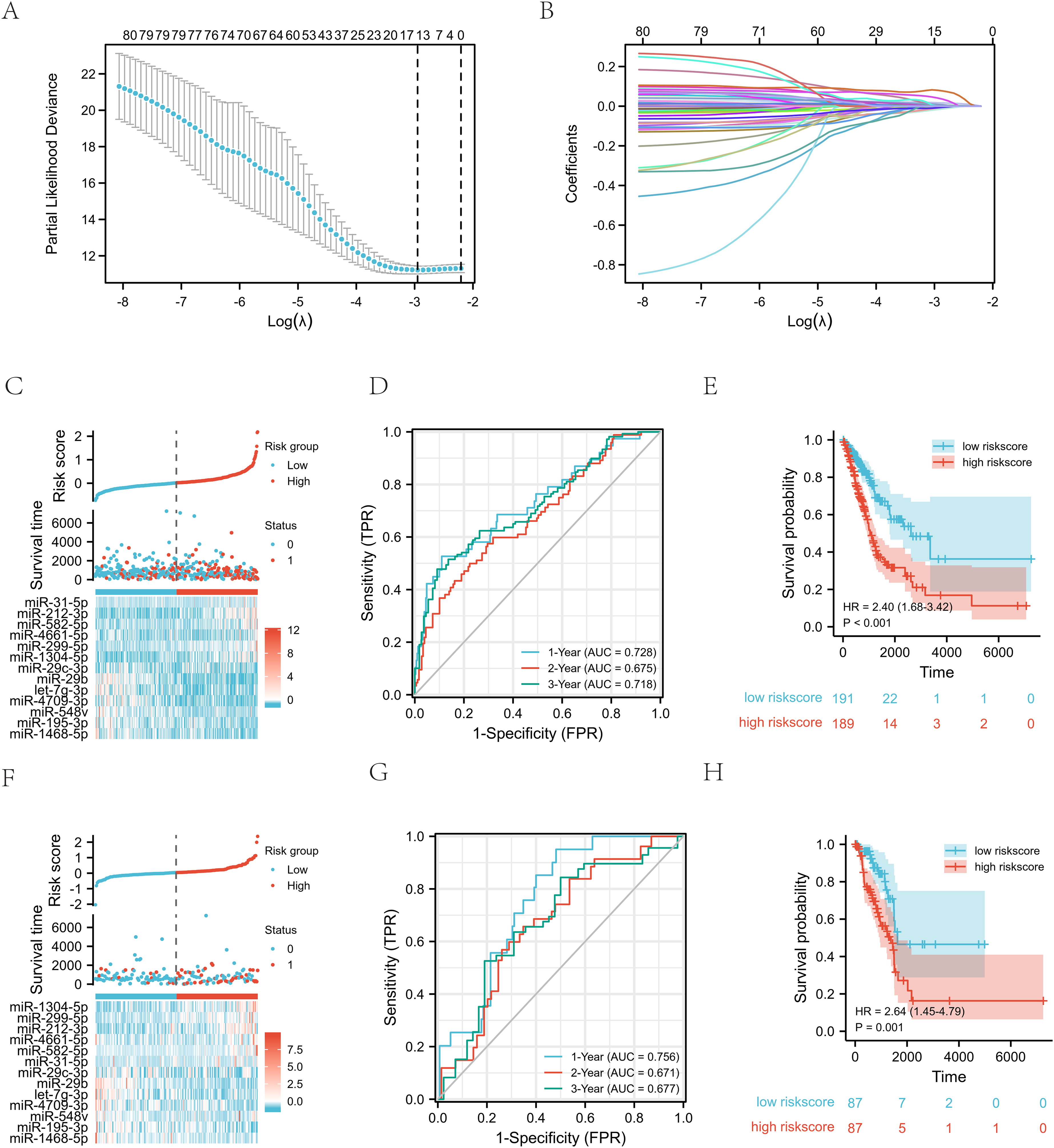
Construction and evaluation of prognosis model of lung adenocarcinoma. (A) A coefficient profile plot was generated against the log (lambda) sequence within the LASSO model. The optimal parameter (lambda) was chosen as the first black dotted line marked on the plot. (B) LASSO coefficient profiles of the 81 miRNAs. (C) A graphical representation depicting the distribution of risk scores (top panel), survival time (middle panel), and miRNA expression levels (bottom panel) within the training set is presented. The black dotted lines serve as a reference, indicating the median risk score threshold that dichotomizes patients into low- and high-risk categories. The red dots and lines highlight the patients classified as high-risk, while the blue dots and lines correspond to the low-risk group. (D) Time-varying ROC curve analyses were conducted within the training set. (E) Kaplan-Meier survival curves were generated to compare the high-risk and low-risk score groups based on the 13-miRNA signature within the training set. (F) The graphical representation displays the distribution patterns of risk scores (top panel), survival time (middle panel), and miRNA expression levels (bottom panel) in the test set. The median risk score cut-off, represented by black dotted lines, divides the patients into two distinct groups: low-risk and high-risk. The red dots and lines represent the patients categorized as high-risk, while the blue dots and lines correspond to the patients in the low-risk group. (G) The time-varying ROC curve was analyzed within the test set. (H) Kaplan-Meier survival curves were constructed to visualize the differences in survival outcomes between the high-risk and low-risk score groups, based on the 13-miRNA signature, within the test set.
Regression coefficients for the variables selected by the final LassoCox-regularized model trained using all samples.
Evaluation prognosis models of lung adenocarcinoma
Figure 4(C) shows the risk score distribution, survival status, and miRNA expression of patients. We compared gene expression between high- and low-risk groups using the 13 miRNA signature. Six miRNAs (miR-1304-5p, miR-299-5p, miR-212-3p, miR-4661-5p, miR-582-5p, and miR-31-5p) were overexpressed in the high-risk group, suggesting a positive correlation with risk. Conversely, seven miRNAs (miR-29c-3p, miR-29b-2-5p, let-7g-3p, miR-4709-3p, miR-548v, miR-195-3p, and miR-1468-5p) were underexpressed, also correlating with high risk. The prognostic model based on these miRNAs had AUC values of 0.728, 0.675, and 0.718 for one, two, and three years of OS, respectively (Figure 4(D)), indicating excellent discriminatory power. Kaplan–Meier analysis revealed superior OS for low-risk patients compared to high-risk patients (log-rank test, P < 0.001) (Figure 4(E)). These findings were consistent in the test set. Figure 4(F) displays the risk score distribution, survival status, and miRNA expression profiles of patients. A gene expression comparison between high- and low-risk groups using the 13 miRNA signature showed six miRNAs (miR-1304-5p, miR-299-5p, miR-212-3p, miR-4661-5p, miR-582-5p, and miR-31-5p) to be overexpressed in the high-risk group, correlating with higher risk. Conversely, seven miRNAs (miR-29c-3p, miR-29b-2-5p, let-7g-3p, miR-4709-3p, miR-548v, miR-195-3p, and miR-1468-5p) were underexpressed in the high-risk group. The prognostic model based on these miRNAs had AUC values of 0.756, 0.671, and 0.677 for one, two, and three years of OS, respectively (Figure 4(G)), indicating good discriminatory power. Kaplan–Meier analysis showed improved OS for low-risk patients compared to high-risk patients (log-rank test, P = 0.001) (Figure 4(H)). These findings were consistent in the test set.
The prognostic value of stage, gender, and risk score in the training set was evaluated through the use of univariate and multivariate analyses
Univariate analysis revealed that the risk score was a significant risk factor that had a significant impact on patient survival (P <0.001) (Figure 5(A)). Multivariate Cox regression analysis further confirmed that both stage and risk score served as independent prognostic factors (P <0.001) (Figure 5(B)). Additional details can be found in Table 5. These results demonstrate the strong predictive power of the risk score.
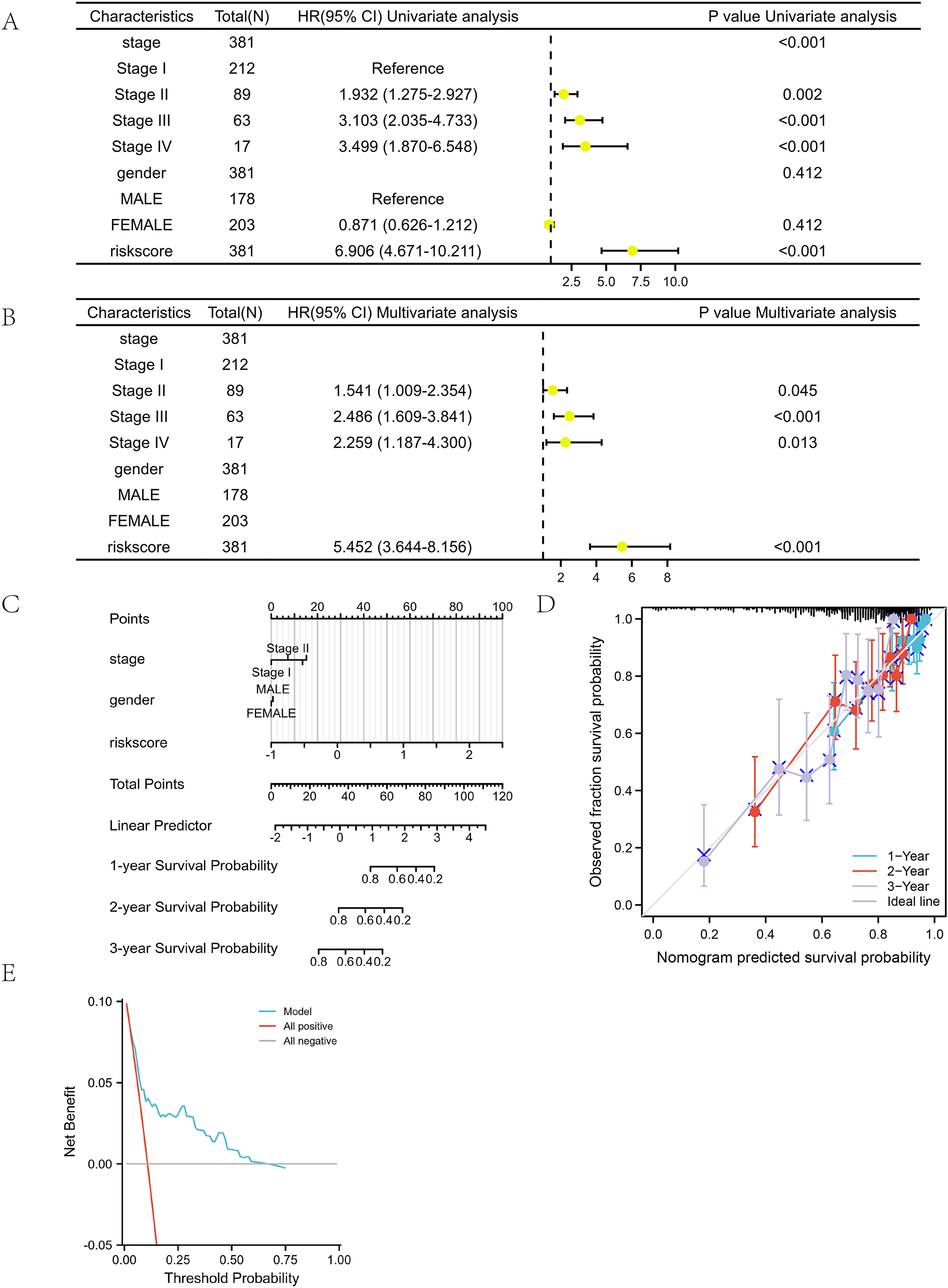
The forest plots, nomogram, calibration curve, and decision curve of the prognosis model in the training set were all compared with respect to their relationship with the risk score and clinical characteristics. (A) The results of univariate analysis demonstrated that both the stage and risk score were significant risk factors that had a significant impact on patient survival, with a P-value less than 0.001. (B) The results of multivariate Cox regression analysis indicated that the risk score might serve as an independent prognostic indicator for the overall survival of patients with lLUAD, with a statistically significant P-value less than 0.001. (C) The nomograms were established to predict the OS of patients with LUAD. When utilizing the nomogram, the value of each individual patient with LUAD is represented on each variable axis, and a line is drawn upward to ascertain the points awarded for each variable value. Subsequently, the summation of these points is located on the total point axis, and a line is traced downward to the survival axes, enabling the determination of the probabilities of 1-, 2-, and 3-year survival for OS. (D) The calibration curve, which is used to predict the 1-, 2-, and 3-year survival rates of OS in patients with LUAD within the training cohort, illustrates the actual OS rates plotted along the y-axis and the nomogram-predicted OS rates plotted along the x-axis. (E) DCA was employed to assess the potential clinical utility of the nomogram in terms of its ability to predict outcomes. The X-axis represents the probability threshold, and the Y-axis represents the net benefit.
The univariate and multivariate Cox regression analysis of risk score, gender and stage in the training set.
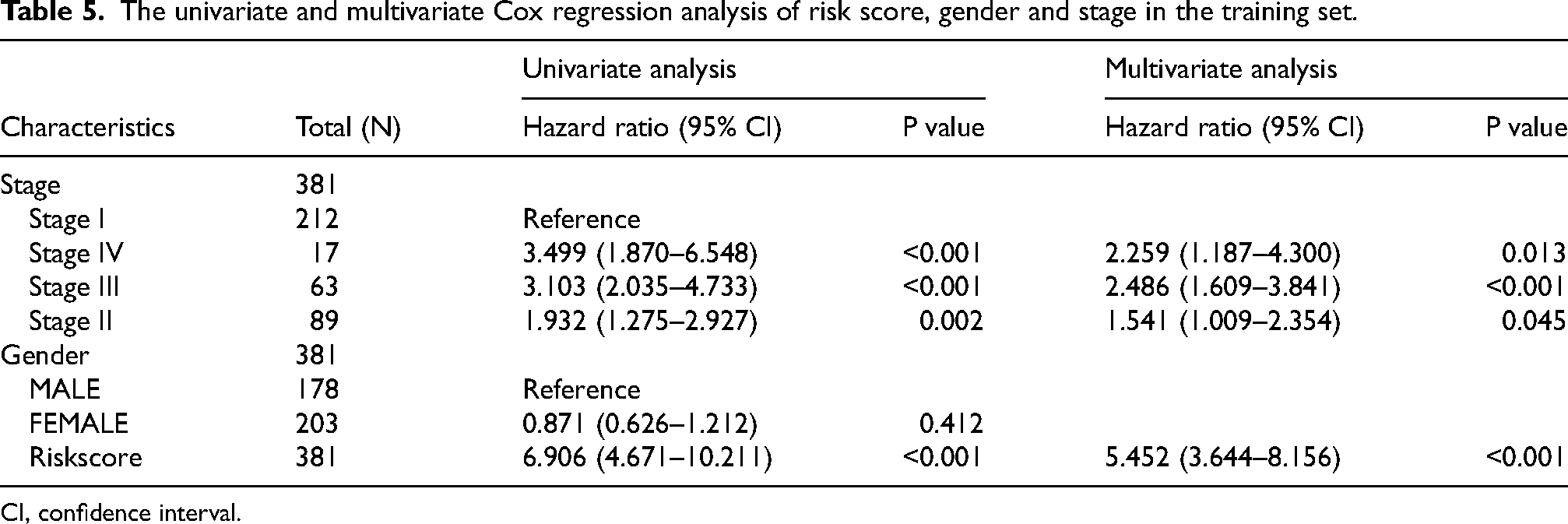
CI, confidence interval.
The prognostic nomogram was established in the training set
The optimism-corrected C-index values were specifically calculated for OS, resulting in a value of 0.702 with a 95% confidence interval ranging from 0.677 to 0.728. This finding indicated that the proposed nomograms accurately predicted the one-, two-, and three-year OS of lung adenocarcinoma patients (Figure 5(C)). The calibration curves for OS at 1-, 2-, and 3-year intervals visually demonstrated a good fit between predicted and observed survival, thereby validating the prediction accuracy of the prognostic nomograms (Figure 5(D)). Moreover, the decision curve analysis demonstrated significant benefits in the utilization of the model in clinical decision-making (Figure 5(E)). The X-axis represents the probability threshold, and the Y-axis represents the net benefit.
The prognostic value of stage, gender, and risk score in the test set was evaluated through the use of univariate and multivariate analyses
Univariate analysis demonstrated that the risk score was a significant risk factor that had a significant impact on patient survival (P = 0.002) (Figure 6(A)). In the multivariate Cox regression analysis, both stage and risk score were identified as independent prognostic factors (P = 0.004) (Figure 6(B)). Additional details can be found in Table 6. These results demonstrate the strong predictive power of the risk score.
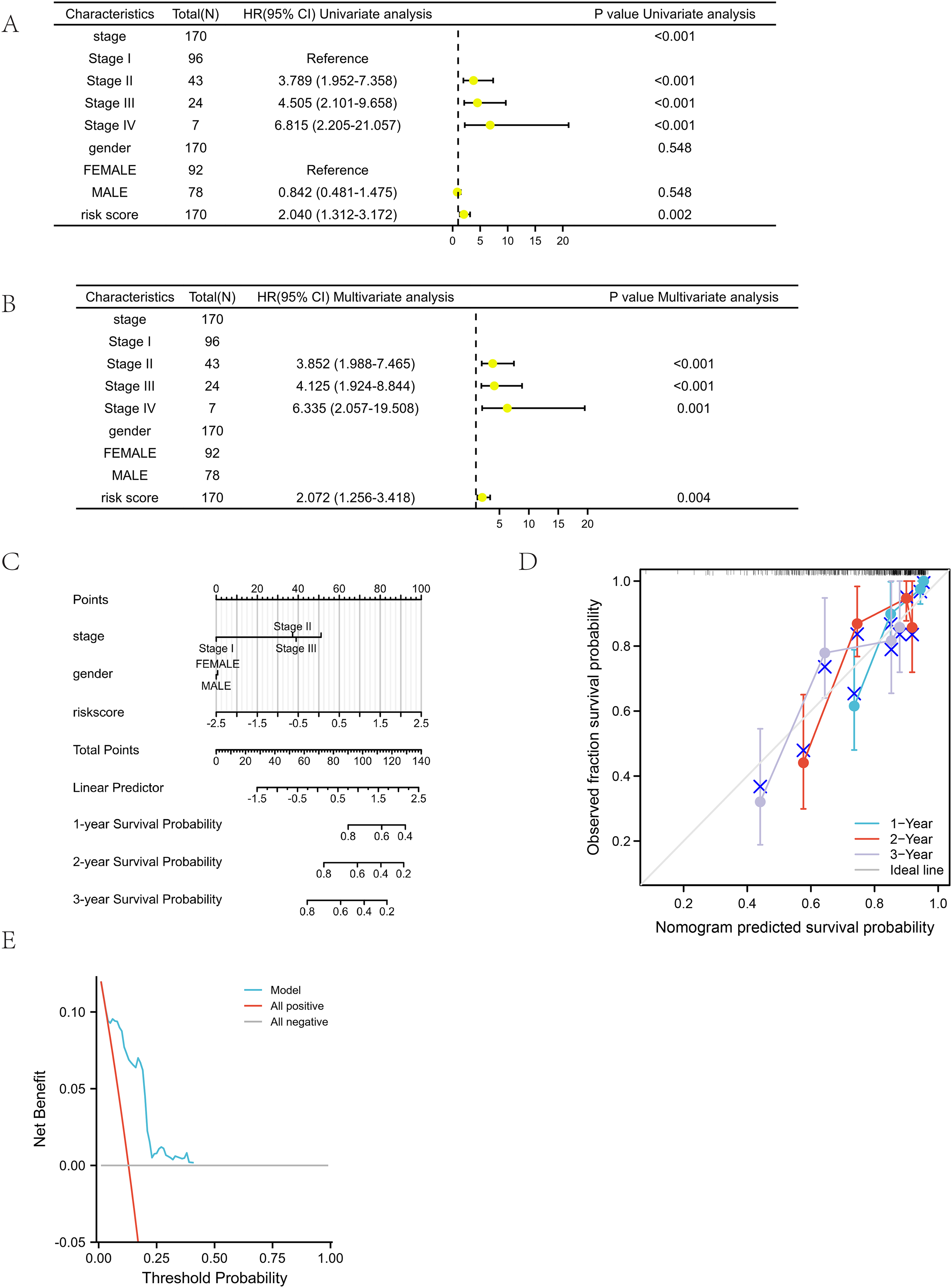
Forest plots between risk score and clinical characteristics, the nomogram,calibration curve and decision curve of prognosis model in test set. (A) The results of the univariate analysis indicated that both the stage and risk score were significant factors that influenced patient survival, with a P-value less than 0.001. (B) Based on the results of multivariate Cox regression analysis, it was suggested that the risk score might serve as an independent prognostic indicator for the OS of patients with LUAD, with a statistically significant P-value less than 0.001. (C) The nomograms were established for predicting OS in patients with LUAD. These nomograms required the input of individual patient values on each variable axis, upon which a line was drawn upward to ascertain the points awarded for each variable value. Subsequently, the summation of these points was located on the total point axis, and a line was traced downward to the survival axes, enabling the determination of the probabilities of 1-, 2-, and 3-year survival for OS. (D) The calibration curve, which is utilized for predicting the 1-, 2-, and 3-year survival rates of OS in patients with LUAD within the training cohort, depicts the actual OS rates along the y-axis and compares them with the OS rates predicted by the nomogram, which are plotted along the x-axis. (E) DCA was employed as a tool to assess the potential clinical utility of the nomogram by evaluating its capacity to inform decision-making in a clinical setting.
The univariate and multivariate cox regression analysis of risk score, gender and stage in the test set.
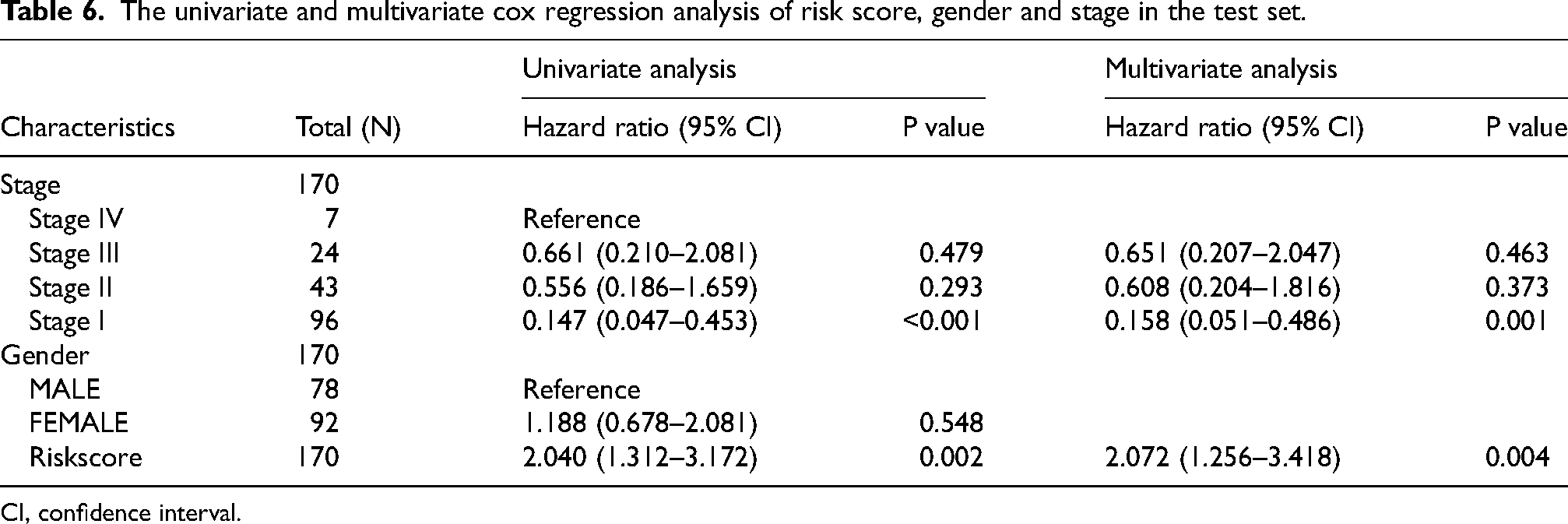
CI, confidence interval.
The prognostic nomogram was established in the test set
The optimism-corrected C-index values were specifically calculated for OS, resulting in a value of 0.701 with a 95% confidence interval ranging from 0.669 to 0.736. This finding indicated that the proposed nomograms accurately predicted the one-, two-, and three-year OS of lung adenocarcinoma patients (Figure 6(C)). The 1-, 2-, and 3-year calibration curves for OS visually demonstrated a good fit between predicted and observed survival (Figure 6(D)). Moreover, the decision curve analysis demonstrated significant benefits in the utilization of the model in clinical decision-making (Figure 6(E)). The X-axis represents the probability threshold, and the Y-axis represents the net benefit. A K-M survival analysis was conducted on LUAD patients from TCGA for 13 miRNAs, including miR-1304-5p, miR-299-5p, miR-212-3p, miR-4661-5p, miR-582-5p, miR-31-5p, miR-29c-3p, miR-29b-2-5p, let-7g-3p, miR-4709-3p, miR-548v, miR-195-3p, and miR-1468-5p. As shown in Figure 7, the analysis results indicated that high expression levels of miR-1304-5p, miR-299-5p, miR-212-3p, miR-4661-5p, miR-582-5p, and miR-31-5p were associated with poorer OS, whereas high expression levels of miR-29c-3p, miR-29b-2-5p, let-7g-3p, miR-4709-3p, miR-548v, miR-195-3p, and miR-1468-5p were associated with better OS. These findings are consistent with the heatmap results presented in Figures 4(C) and 4(F).
Gene Ontology (GO), Kyoto Encyclopedia of Genes and Genomes (KEGG), and subgroup analysis were all employed
A Venn diagram was constructed to illustrate the overlapping and unique target gene predictions for miRNAs obtained from both TargetScan and miRDB databases. A Venn diagram is presented in Figure 8(A). The enrichment analysis revealed that the correlated genes were enriched for focal adhesion, protein digestion and absorption, ECM-receptor interaction, and other related processes (Figure 8(B)). For more detailed information, please refer to the list provided in Table 7. Furthermore, K-Mcurves were plotted to investigate the relationships between the risk signature and stage. The subgroups of stage I-II and stage III-IV were based on the median risk score. The resulting K-M curves demonstrated significant differences in survival time between the subgroups, with 2.34 and 2.96 in stage I-II and stage III-IV subgroups, respectively (P < 0.01) (Figure 8(C)-(D)).
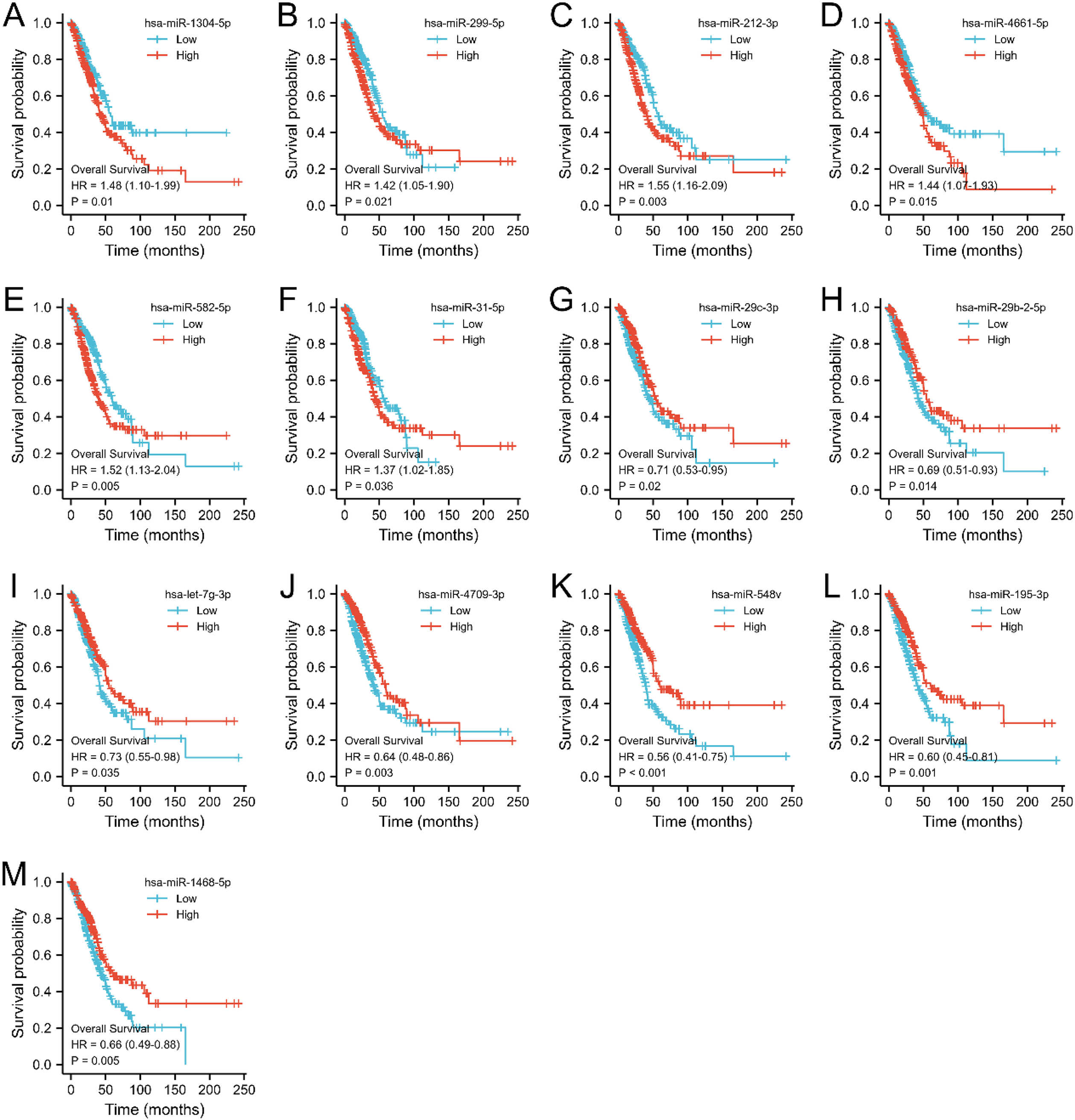
Survival analysis of TCGA dataset. (A-M) K-M survival analysis of miR-1304-5p, miR-299-5p, miR-212-3p, miR-4661-5p, miR-582-5p, miR-31-5p, miR-29c-3p, miR-29b-2-5p, let-7g-3p, miR-4709-3p, miR-548v, miR-195-3p and miR-1468-5p in TCGA-LUAD patients.
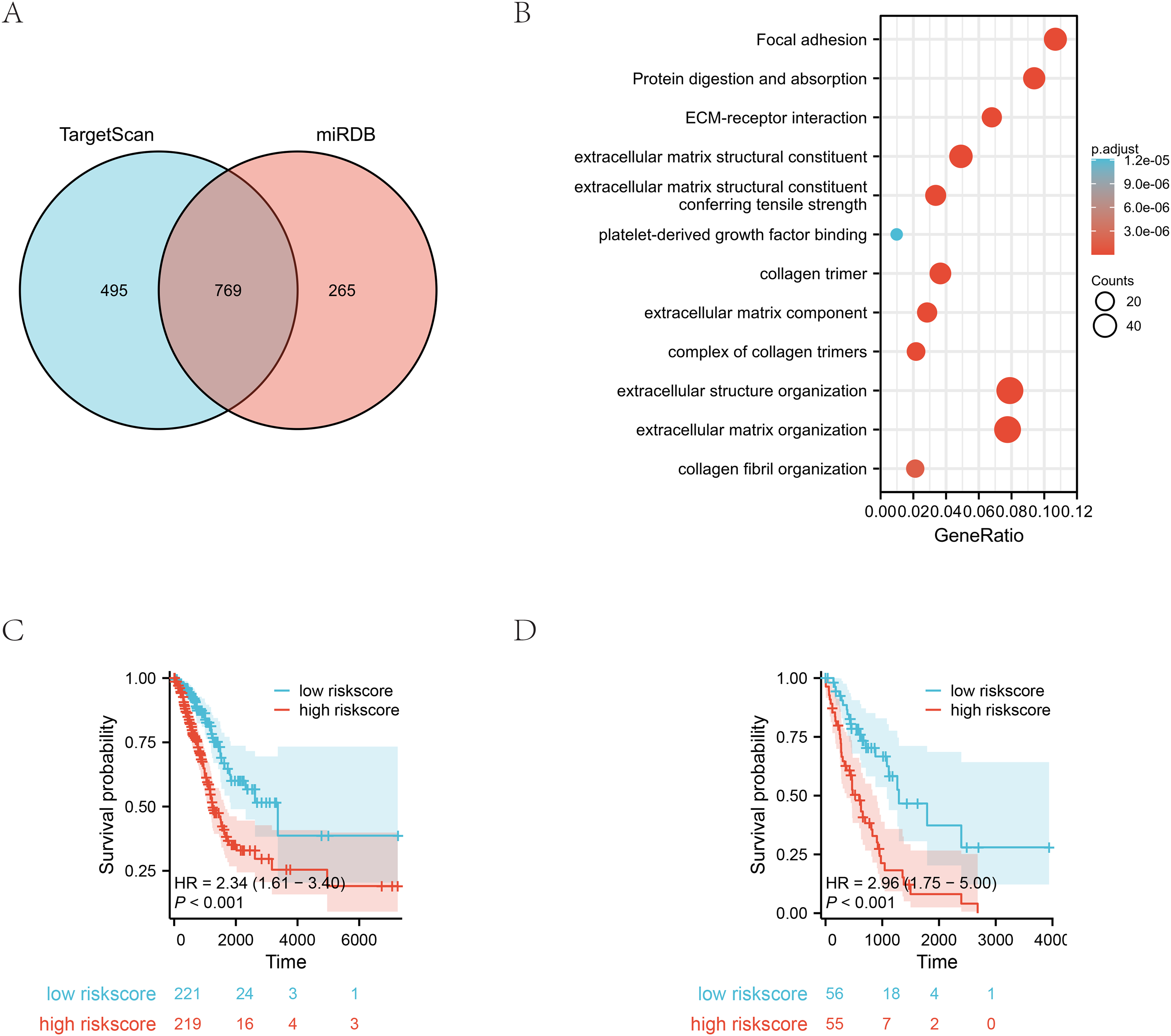
Gene Ontology (GO), Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analyses and subgroup analysis. (A)Venn diagram. (B) Gene Ontology (GO), Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analyses. (C) The K-M curves for subgroup of stage I-II (D) The KM curves for subgroup of stage III-IV.
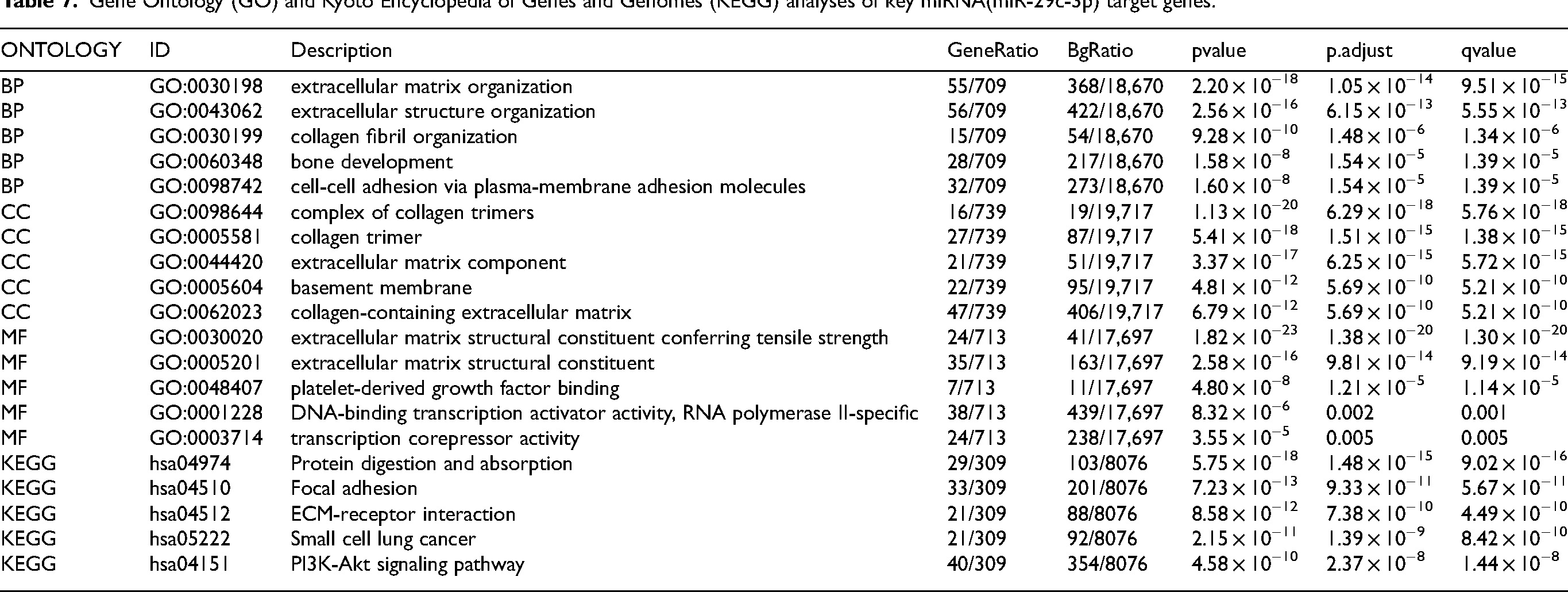
Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analyses of key miRNA(miR-29c-3p) target genes.
Discussion
LUAD is the predominant lung cancer subtype, accounting for 40% of cases. Recent gene signatures aid tumor classification and prognosis prediction, but no method accurately classifies suspicious lung nodules early. This study used TCGA Lung Adenocarcinoma data to screen stage I pulmonary nodules. Machine learning techniques were employed for dimension reduction and feature selection, ultimately identifying two optimal miRNAs. Six machine learning algorithms were trained and evaluated, exhibiting high discriminative power and strong predictive performance (AUC: 0.9337719-1; error rate: < 10%). Lasso-Cox regression identified 13 miRNAs, with low-risk patients showing significantly longer OS. The risk score, reliable for LUAD prognosis prediction (AUC max: 0.756), was an independent prognostic factor for OS. The risk score was confirmed as an independent risk factor for poor LUAD prognosis. Integrated into a nomogram, the risk score facilitated clinical prognosis predictions (C-index: 0.706). The model demonstrated excellent consistency between predicted and observed probabilities for 1-, 2-, and 3-year survival. Patients with higher risk scores had poorer prognoses across both I/II and III/IV stages.
A recent study identified miR-21-5p as a plasma miRNA with potential prognostic significance for pancreatic cancer patients. 18 A study analyzed miR-21-5p expression in lung cancer tissues and cell lines. By transfected miR-21-5p mimics, inhibitors, and negative controls into lung cancer cells, qRT-PCR and MTT assays were conducted. The findings revealed that miR-21-5p inhibitors suppressed lung cancer cell proliferation, invasion, and migration. Additionally, numerous studies have shown that miR-21-5p promotes cell proliferation by targeting TGFBI in non-small cell lung cancer cells. 19 The above findings suggest stability and credibility in our results. Our screenings imply that miR-21-5p dysregulation might be an early marker in lung adenocarcinoma development, aligning with prior research. Prior studies have shown that mir-29c-3p plays a pivotal role as a tumor suppressor gene in gastric cancer development. 20 A study showed that miR-29c-3p suppresses the epithelial-mesenchymal transition, inhibiting cervical cancer cell proliferation, invasion, and metastasis by targeting SPARC. 21 Above research results indicated that our model was reasonably accurate and acceptable.
However, our study has limitations. Firstly, while we have utilized training, cross-validation, and testing datasets, the validation of our model using an external validation set would have been ideal. Secondly, validating our findings in a larger patient cohort is crucial, and both these aspects should be prioritized in future studies. Additionally, we observed that the hazard ratio of the risk score was higher than that of staging in the training set, whereas the opposite was true in the test set, which may be influenced by sample size. Although a P-value < 0.05 and an HR > 1 indicate statistical significance for the risk score, the magnitude of the effect is closely tied to the sample size. Therefore, when interpreting these results, it is necessary to comprehensively consider factors such as sample size, effect size, and confidence intervals. Furthermore, statistical significance does not equate to causality. The association between the risk score and event risk requires additional experimental evidence and investigation of the underlying biological mechanisms for confirmation, in order to more accurately understand its role in disease and provide a reliable basis for clinical diagnosis and treatment. Despite these limitations, our research provides a theoretical reference for early diagnosis, prognosis prediction, and targeted drug development in lung adenocarcinoma.
Conclusion
Employing machine learning algorithms, we discovered a five-miRNA signature that can accurately distinguish LUAD from adjacent normal tissue. Furthermore, we developed a prognostic prediction model utilizing 13 miRNAs. Both the diagnostic and predictive models exhibit robust reliability.
Footnotes
Ethical statement
The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013).
Author contributions
i. Conception and design: Yongxia Bao and Lin Lin. ii. Collection and assembly of data: Lin Lin. iii. Data analysis and interpretation: Lin Lin. iv. Manuscript writing: Yongxia Bao and Lin Lin. v. Final approval of manuscript: Yongxia Bao and Lin Lin.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by Grants from the Natural Science Foundation of Heilongjiang Province of China (Grant No. LH2020H060).
Declaration of Conflicting Interests
All authors have completed the ICMJE uniform disclosure. The authors have no conflicts of interest to declare.
Reporting checklist
The authors have completed the TRIPOD reporting checklist.
Data sharing statement
The authors have completed the Data Sharing Statement
Permission and copyright
All figures and tables in the article are original.
