Abstract
Background
Apolipoprotein C1
Methods
Various bioinformatics analysis methods were used, including gene expression profiling interactive analysis, the University of Alabama at Birmingham cancer data analysis portal, biomarker exploration of solid tumors software, the BioPortal for Cancer Genomics, search tool for the retrieval of interacting genes/proteins, gene multiple association network integration algorithm, Metascape, transcriptional regulatory relationships unraveled by sentence-based text-mining, LinkedOmics, and genomics of drug sensitivity in cancer analysis.
Results
APOC1 and APOE expression were strongly downregulated in patients with ACC. APOC1 and APOE expression levels were lower in male patients with ACC than those in female patients. Furthermore, APOC1 and APOE expression levels affected the prognosis of patients with ACC. The main functions of APOC1 and its altered neighboring genes (ANG) were organophosphate ester transport, rRNA processing, and positive regulation of cytokine production. Cytolysis, protein ubiquitination, and histone modification were the main functions of APOE and its ANGs. The transcription factor E2F1, tumor protein p53, miR-182, miR-493, Erb-B2 receptor tyrosine kinase 2, and cyclin dependent kinase 1 were key regulatory targets of APOC1, APOE, and the ANGs. APOC1 and APOE expression in patients with ACC were positively associated with immune cell infiltration. Furthermore, anti-programmed cell death protein 1 immunotherapy strongly downregulated the expression of APOC1 in patients with ACC. Both pilaralisib and elesclomol strongly inhibited SW13 cell growth.
Conclusions
This study preliminarily clarified that APOC1 and APOE might be potential therapeutic and prognostic targets for ACC, and identified new targets and treatment strategies for ACC.
Introduction
Adrenal cortical carcinoma (ACC), is a rare endocrine adrenal malignancy that occurs in 0.7–2 cases per million per year.1,2 The biological behavior of tumors in patients with ACC is usually aggressive, and 50%–70% of these patients have hormone overdose symptoms and signs. Glucocorticoids and/or androgens are hypersecreted by two-thirds of patients.3,4 Excess hormone secretion in patients with ACC causes increased abdominal mass, weight loss, and other constitutional symptoms. 4 Females are more likely to suffer from the disease than males (1.5:1).5,6 Due to the difficult clinical presentation of ACC and the fact that most patients are diagnosed at a late metastatic stage, the prognosis for ACC is extremely poor. Patients with locoregional ACC are treated surgically; however, approximately 75% experience recurrence after surgery. 7 Furthermore, for patients with advanced metastatic disease, overall survival is 12–15 months and 15% at 5 years.8,9 Compared to patients with other tumor types, patients with ACC do not benefit from improvements in overall tumor treatment. Chemotherapy and radiation therapy are ineffective in most ACC cases. Mitotane is the only drug approved by the US Food and Drug Administration for ACC and is usually only temporarily effective and has obvious side effects.10,11 Complete tumor resection remains the only curative treatment. 12 Therefore, it is important to identify the target genes and regulatory targets that affect the prognosis of patients with ACC under circumstances of limited therapeutic drugs, poor therapeutic effects, and a high risk of disease recurrence.
Apolipoprotein C1 (APOC1) and Apoprotein E (APOE) are members of the apolipoprotein family. APOC1, the smallest apolipoprotein (6.6 kDa), is a triglyceride-rich and high-density lipoprotein closely related to inflammation, immunity, sepsis, and diabetes. 13 APOE, a 34 glycoprotein, is a high-density lipoprotein with antioxidant, anti-inflammatory, and antiatherogenic properties. 14 Recently, several studies have shown that APOC1 and APOE are involved in the development and occurrence of multiple cancers. APOC1 and APOE are diagnostic and prognostic biomarkers for many cancers. Interestingly, APOC1 and APOE expression levels were different in different tumors. Therefore, the function of APOC1 and APOE in different tumors may differ. For example, APOC1 is significantly overexpressed and its overexpression is associated with lymph node metastasis, tumor-node-metastasis stage, distant metastasis, and poor prognosis in colorectal cancer, gastric cancer, and clear cell renal cell carcinoma.15–17 The overexpression level of APOE in endometrial cancer has been correlated with histological grade, lymph node metastasis, and the International Federation of Gynecology and Obstetrics stage. 18 However, APOC1 has significantly lower expression in esophageal cancer and hepatocellular carcinoma, which is associated with reshaping the tumor immune microenvironment and inhibiting the proliferation, migration, and invasion of cancer cells.19,20 Serum APOE levels tend to decrease in patients with laryngeal squamous cell carcinoma. 21 It is currently unknown if APOC1 and APOE play a role in ACC. Hence, in this study, the expression of APOC1 and APOE in patients with ACC, as well as the gene regulatory network, prognostic value, target prediction, and potential therapeutic agents were examined. These data clarify the relationships between APOC1, APOE, and ACC.
Materials and methods
Gene expression profiling interactive analysis (GEPIA)
GEPIA (http://gepia.cancer-pku.cn/index.html) is an intuitive network application tool to assess the biological relationships between gene expression and prognostic information in patients with cancer. 22 GEPIA was used to analyze the relationships between gene expression, tumor pathological stages, and prognosis. The screening criteria were: (1) genes: APOC1 and APOE; (2) dataset: ACC; and (3) 77 patients with ACC; threshold setting conditions: P-value cutoff ≤ 0.05. Student's t-test was used to analyze the expression of APOC1 and APOE in ACC. Kaplan–Meier curves were used to analyze the prognosis of patients with ACC. 23
The university of Alabama at Birmingham CANcer data analysis portal (UALCAN)
UALCAN (http://ualcan.path.uab.edu/analysis.html) facilitates tumor subgroup gene expression and survival analyses. 22 UALCAN was used to analyze APOC1 and APOE expression in ACC. The “Expression Analysis” module of the UALCAN database was used to analyze The Cancer Genome Atlas (TCGA) gene expression data. The screening criteria were set as: (1) genes: APOC1 and APOE; (2) dataset: ACC; (3) 79 patients with ACC (31 male and 48 female); threshold setting conditions: p-value cutoff = 0.05. Student's t-test was used for comparative analyses. 23
Biomarker exploration of solid tumors (BEST)
BEST (https://rookieutopia.com/app_direct/BEST/) provides a curated database and innovative analytical pipelines to explore cancer biomarkers at high resolution. Protein expression, immune cell infiltration, candidate agents, and immunotherapy of APOC1 and APOE in ACC were analyzed using the BEST software. The “Cell infiltration,” “Immunotherapy,” and “Candidate agents” modules of the BEST database were used to analyze Gene Expression Omnibus and TCGA gene expression data. The screening criteria were set as: (1) genes: APOC1 and APOE; and (2) dataset: ACC (10 datasets with 508 patients). 23
Bioportal for cancer genomics (cBioPortal)
cBioPortal (http://cbioportal.org) is an online database used for tumor gene mutation analysis. cBioPortal was used to analyze gene alterations to APOC1, APOE, and the top 50 altered neighboring genes (ANGs), respectively. 22 A total of 75 ACC samples were analyzed and mRNA expression z-scores were obtained relative to all samples (log RNA Seq V2 RSEM) using a z-score threshold of ±2.0. 23
Search tool for the retrieval of interacting genes/proteins (STRING) and gene multiple association network integration algorithm (GeneMANIA)
STRING (https://string-db.org/cgi/input.pl) and GeneMANIA (http://www.genemania.org) are online databases used to analyze gene-protein and protein-protein interactions. 22 STRING was used to build a low-confidence level (0.150) protein-protein interaction (PPI) network to screen criteria for species defined as humans. Cytoscape software (version 3.10.2, cytoHubba plug-ins) was used to discover the core proteins in PPI network. GeneMANIA was used to explore the function of APOC1, APOE, and the top 50 ANGs, respectively. 23
Metascape
Metascape (https://metascape.org) is an online database used to analyze the functions and signaling pathways of genes and proteins. 22 In this study, Metascape was used to analyze the function and signaling pathways of APOC1, APOE, and the top 50 ANGs, respectively. 23
Transcriptional regulatory relationships unraveled by sentence-based text-mining (TRRUST)
TRRUST (https://www.grnpedia.org/trrust/) is an online database used to analyze regulatory targets of gene transcription. 22 Here, TRRUST was used to identify the key transcriptional regulators of APOC1, APOE, and the top 50 ANGs, respectively. 23
Linkedomics
LinkedOmics (http://www.linkedomics.org/) is a public online platform used to analyze correlations between differentially expressed genes related to tumor target genes and to predict microRNA (miRNA) and kinase targets. 22 LinkedOmics was used to identify kinase and miRNA targets, as well as differentially expressed genes related to APOC1 and APOE, respectively. 23
Genomics of drug sensitivity in cancer analysis
The Genomics of Drug Sensitivity in Cancer database (http://www.cancerRxgene.org) is a specialized public database to obtain information on potential anticancer drugs. 23 This database was used to identify drugs targeting APOC1 and APOE and to predict their anti-ACC activity.
Results
APOC1 and APOE expression, prognosis, and genetic alteration in ACC
As shown in Figure 1(a) to (i), APOC1 and APOE are significantly downregulated in patients with ACC (p < 0.05). A significant correlation between APOC1 expression and the pathological stage was observed in patients with ACC (p < 0.05) (Figure 1(j) and (k)). However, APOC1 and APOE expression were lower in male patients than that in female patients with ACC (p < 0.05) (Figure 1(l) and (m)). Patients with ACC who expressed low levels of APOC1 had longer overall survival times (p = 0.0083) (Figure 1(n)). Patients with ACC with low APOC1 and APOE expression had longer disease-free survival times than those with high APOC1 and APOE expression levels (p = 0.017 and p = 0.049, respectively) (Figure 1(o) and (q)). According to these results, APOC1 and APOE expression levels were altered by 9% and 15%, respectively, in patients with ACC (Figure 1(r) and (s)).
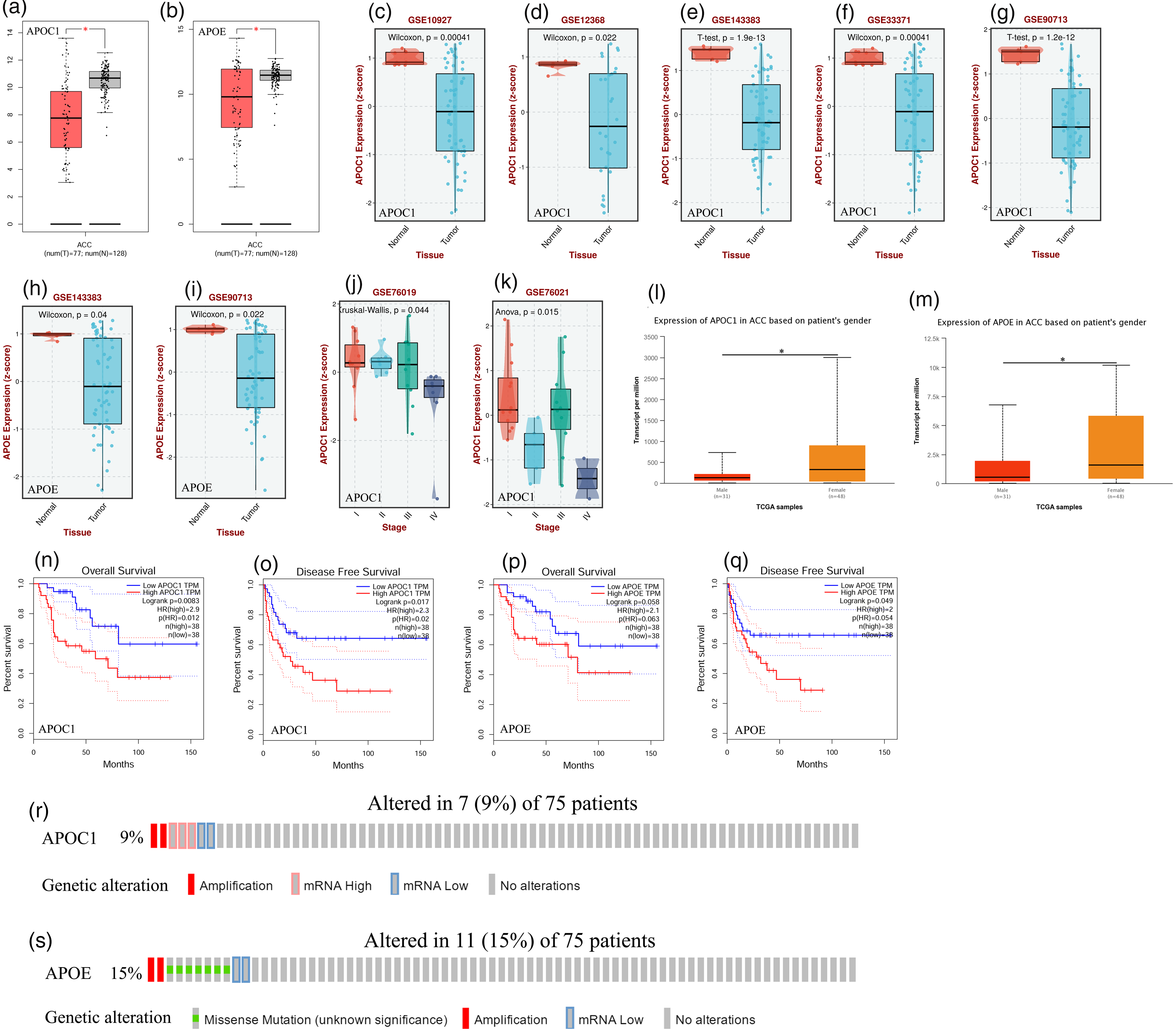
The transcription levels, prognostic value, and genetic alteration of APOC1 and APOE in ACC (GEPIA, BEST, UALCAN, and cBioPortal). (a) Boxplot showing transcription level of APOC1 in patients with ACC (GEPIA); (b) Boxplot showing transcription level of APOE in patients with ACC (GEPIA); (c-g) Boxplot showing transcription level of APOC1 in patients with ACC (BEST); (h and i) Boxplot showing transcription level of APOE in patients with ACC (BEST); (j and k) Boxplot showing correlation between the pathological stage and different expressed APOC1 in patients with ACC (BEST); (l) Boxplot showing transcription level of APOC1 in patients with ACC based on the sex (UALCAN); (m) Boxplot showing transcription level of APOE in patients with ACC based on sex (UALCAN); (n) The overall survival curve of APOC1 in patients with ACC (GEPIA); (o) The disease-free survival curve of APOC1 in patients with ACC (GEPIA); (p) The overall survival curve of APOE in patients with ACC (GEPIA); (q) The disease-free survival curve of APOE in patients with ACC (GEPIA); (r) Genetic alteration of APOC1 in patients with ACC (cBioPortal); (s) Genetic alteration of APOE in patients with ACC (cBioPortal); *p < 0.05.
Interaction network of APOC1, APOE, and ANGs in ACC
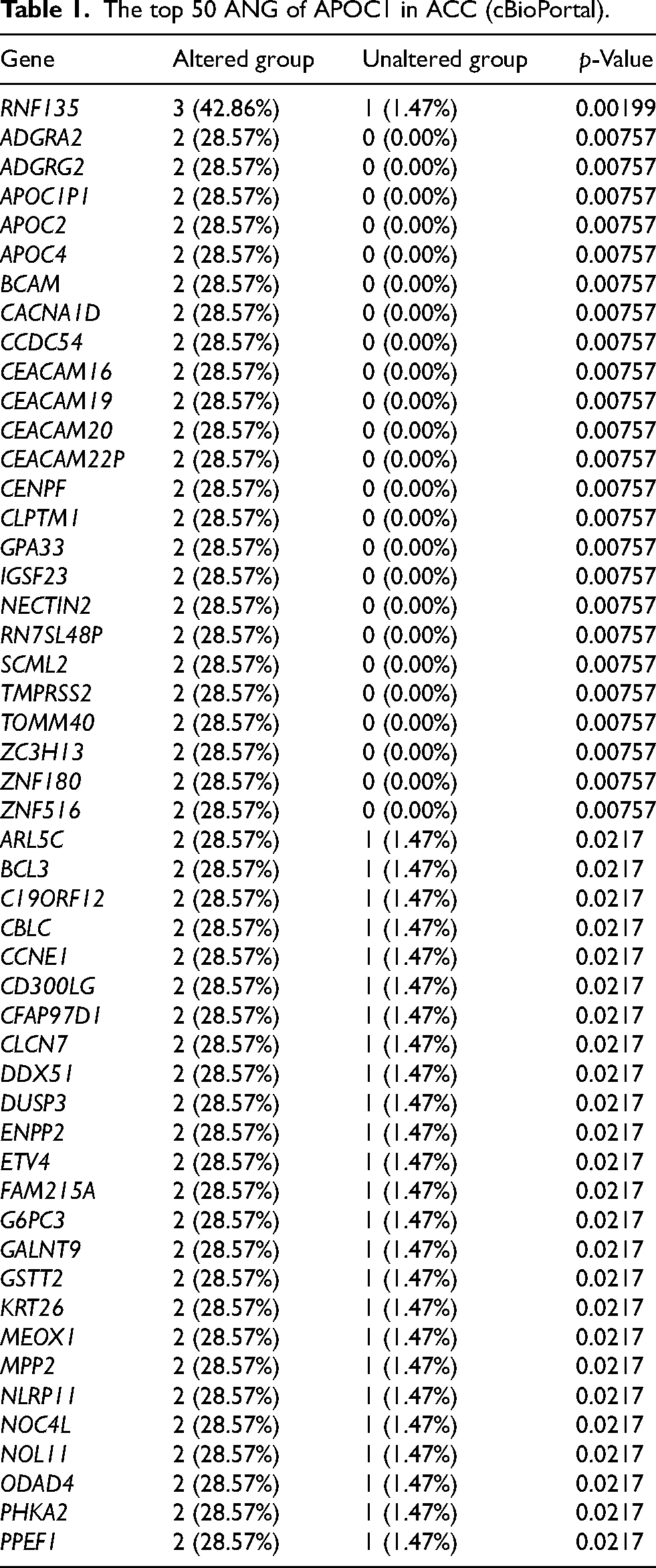
As shown in Tables 1 and 2, APOC1 and APOE ANG alteration frequencies of ≥ 28.57% and ≥ 27.27% were observed in the 50 most frequent ANGs in patients with ACC, respectively. Ring finger protein 135 (42.86%), adhesion G protein-coupled receptor A2 (28.57%), and adhesion G protein-coupled receptor G2 (28.57%) were the most frequent ANGs of APOC1 in patients with ACC (Table 1). Furthermore, the most frequent ANGs of APOE in patients with ACC were mucin 2 (MUC2) (72.73%), MUC4 (63.64%), and outer dense fiber of sperm tails 3 like 2 (36.36%) (Table 2). A total of 38 nodes and 150 edges were identified in the PPI network (Figure 2(a)). On the basis of the PPI network, the core proteins were extracted with Cytoscape software. The results showed the core protein with high scoring were as follows (Top 5): TOMM40, PVRL2, CLPTM1, APOC1, and APOC2 (Figure 2(e)). A complex interaction network was discovered between APOC1 and its ANGs by co-expression, co-localization, physical interactions, shared protein domains, and genetic interactions (Figure 2(b)). Moreover, 43 nodes and 152 edges were obtained in the PPI network (Figure 2(c)). Our results showed CDC34, ODF3L2, C2CD4C, MIER2, and TPGS1 were the core protein with high scoring (Top 5) (Figure 2((f)). An intricate network of interactions existed between APOE and its ANGs, including co-expression, physical interactions, pathways, shared protein domains, and co-localization (Figure 2(d)).
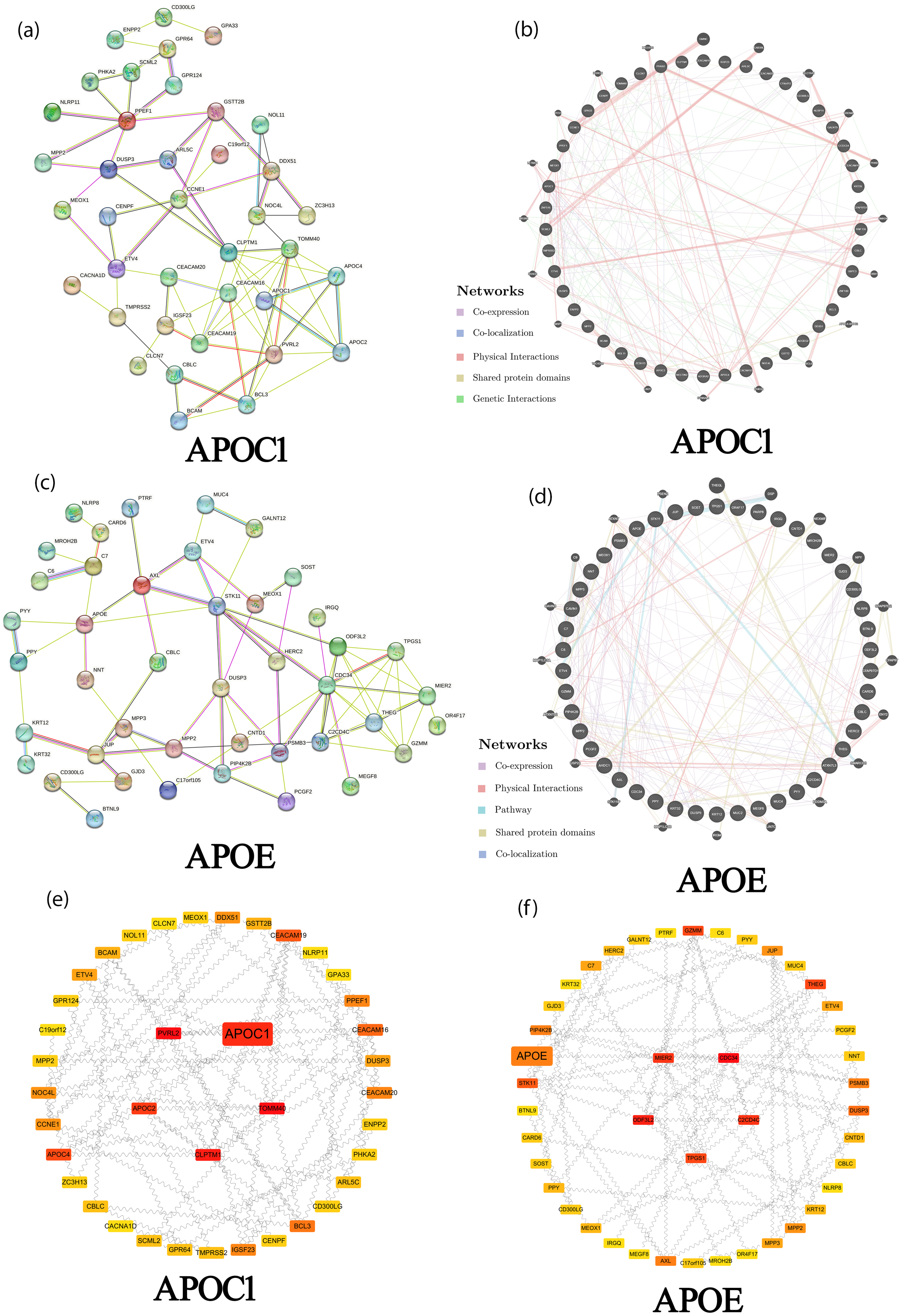
Interaction analyses of APOC1, APOE, and their ANG in ACC (STRING and GeneMANIA). (a) PPI network of APOC1 and its ANG in patients with ACC (STRING); (b) Network analyses of APOC1 and its ANG in patients with ACC (GeneMANIA); (c) PPI network of APOE and its ANG in patients with ACC (STRING); (d) Network analyses of APOE and its ANG in patients with ACC (GeneMANIA); (e) Core proteins in PPI network of APOC1 and its ANG in patients with ACC (Cytoscape); (f) Core proteins in PPI network of APOE and its ANG in patients with ACC (Cytoscape).
The top 50 ANG of APOC1 in ACC (cBioPortal).
The top 50 ANG of APOE in ACC (cBioPortal).

Gene ontology function and Kyoto encyclopedia of genes and genomes (KEGG) pathway enrichment analysis of APOC1, APOE, and ANGs in ACC
As shown in Figure 3(a), molecular functions related to APOC1 and its ANGs are mainly associated with protein tyrosine kinase binding, phosphoric ester hydrolase activity, DNA and transcription factor binding, and channel activity. Moreover, organophosphate ester transport, rRNA processing, cell surface receptor signaling pathways involved in cell-cell signaling, negative regulation of phosphate metabolic processes, and positive regulation of cytokine production were the main biological processes of APOC1 and its ANGs (Figure 3(b)). Cellular components of APOC1 and its ANGs included very-low-density lipoprotein particles, actin-based cell projections, and apical plasma membranes (Figure 3(c)). However, the molecular functions of APOE and its ANGs included protein tyrosine kinase binding, protein domain-specific binding, ubiquitin-protein transferase activity, and transcriptional co-regulator activity (Figure 3(d)). Furthermore, the main biological processes of APOE and its ANGs were cytolysis, negative regulation of kinase activity, regulation of the canonical Wnt signaling pathway, response to reactive oxygen species, protein ubiquitination, response to growth factors, cell surface receptor signaling pathways involved in cell-cell signaling, and histone modification (Figure 3(e)). The cellular components of APOE and its ANGs included intermediate filaments, cell-cell junctions, and the extracellular matrix (Figure 3(f)). KEGG pathway analysis of APOE and its ANGs revealed ubiquitin-mediated proteolysis and regulation of the actin cytoskeleton (Figure 3(g)).
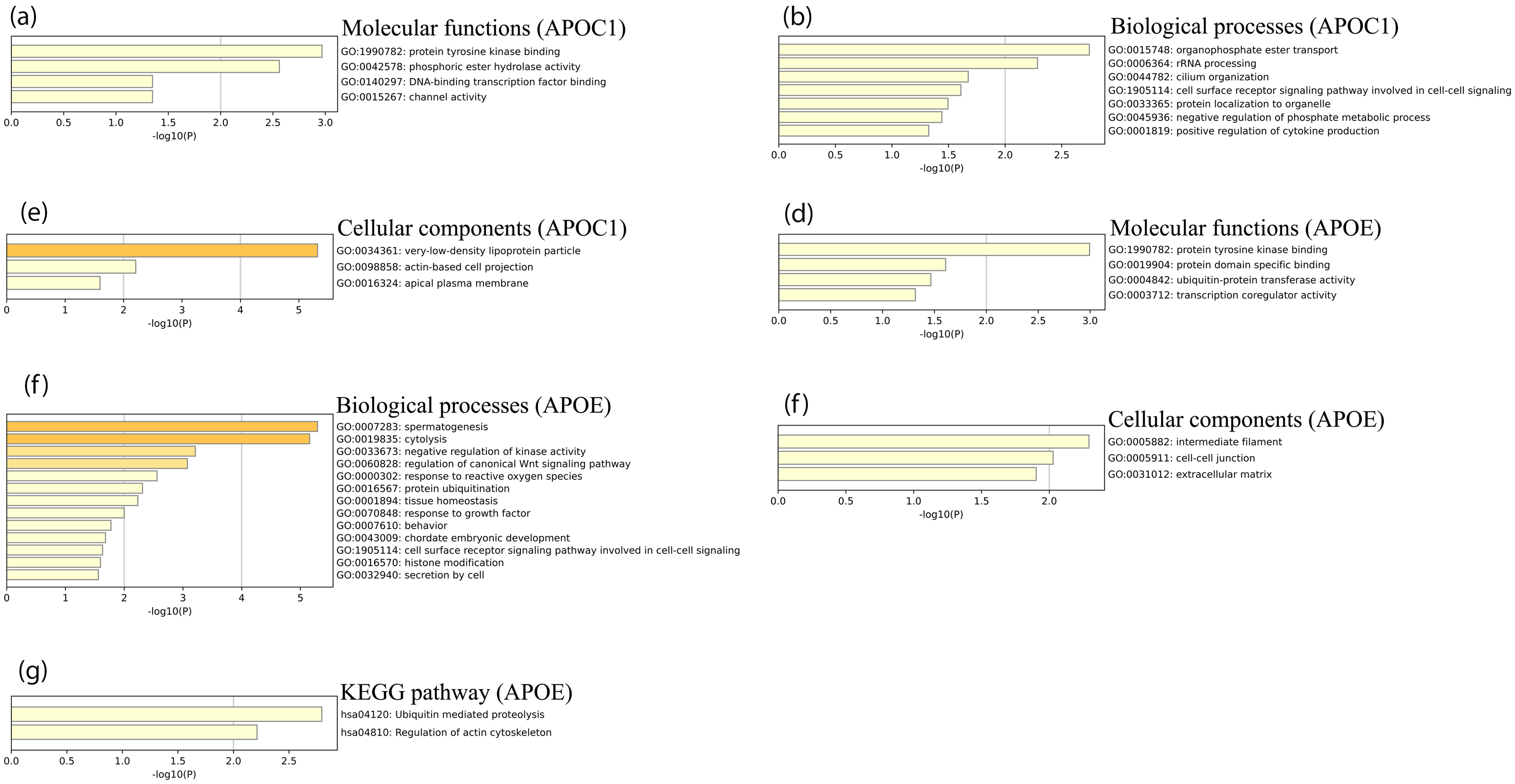
Go function and KEGG pathways enrichment analyses of APOC1, APOE, and their ANG in patients with ACC (metascape). (a) Molecular functions of APOC1 and its ANG in patients with ACC; (b) Biological processes of APOC1 and its ANG in patients with ACC; (c) Cellular components of APOC1 and its ANG in patients with ACC; (d) Molecular functions of APOE and its ANG in patients with ACC; (e) Biological processes of APOE and its ANG in patients with ACC; (f) Cellular components of APOE and its ANG in patients with ACC; (g) KEGG pathway analysis of APOE and its ANG in patients with ACC. The GO enriched terms are colored by p-value, where terms containing more genes tend to have more significant p-value.
Transcription factor, miRNA, and kinase targets of APOC1 and APOE in patients with ACC
As shown in Table 3, E2F1 is the key transcription factor involved in the network of APOC1 and its ANGs (p < 0.05). E2F1 regulates the functions of cyclin E1 and ETS variant transcription factor 4 (ETV4). Moreover, E2F1, signal transducer and activator of transcription 3 (STAT3), specificity protein 1 (SP1), and tumor protein p53 (TP53) were key transcription factors for APOE and its ANGs (p < 0.05). Among these, ETV4 and MUC4 were the main regulatory genes of E2F1. STAT3 regulated MUC4 and serine/threonine kinase 11(STK11) function and APOE
Key regulated factor of APOC1, APOE, and their top 50 ANG in ACC (TRRUST).

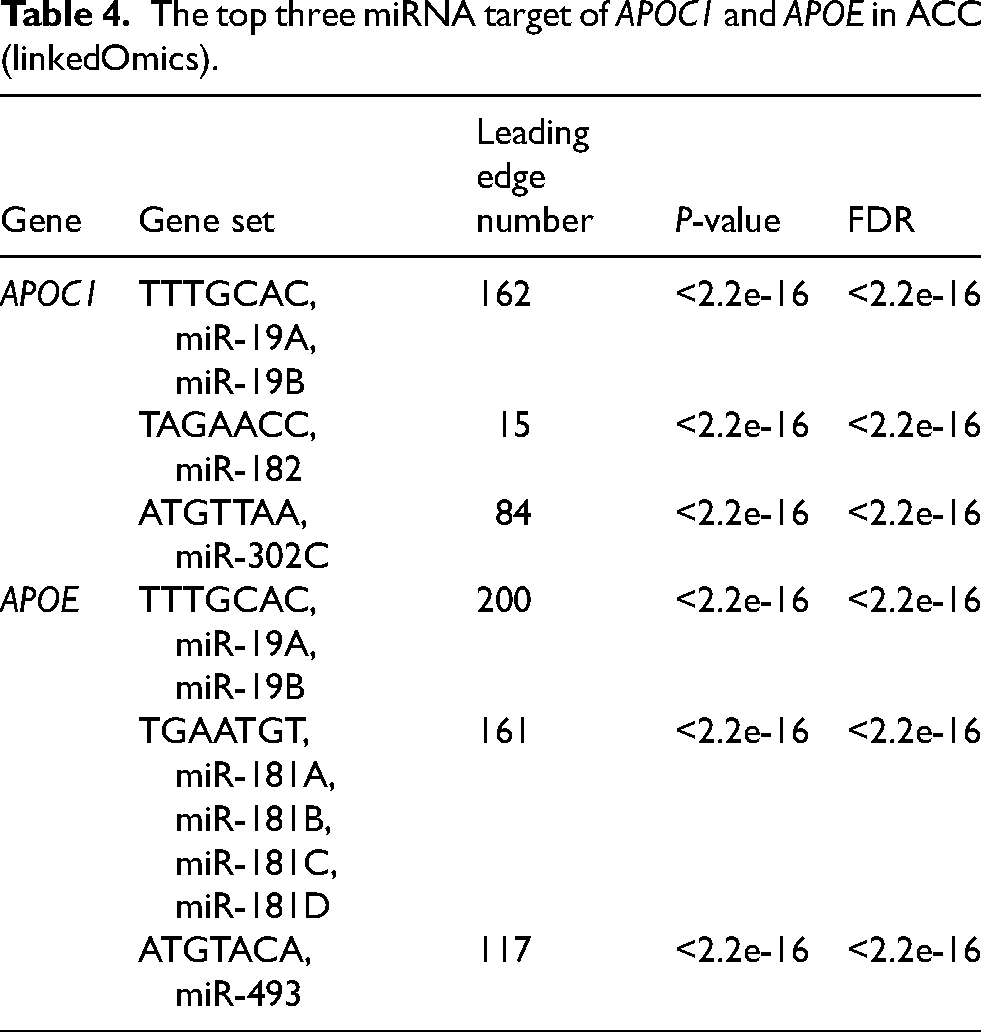
The top three miRNA target of APOC1 and APOE in ACC (linkedOmics).
The top three kinase target of APOC1 and APOE in ACC (linkedOmics).

Correlation of differentially expressed genes and expression of APOC1 and APOE in patients with ACC
As shown in Figure 4(a) and (d), 4471 and 4021 genes are closely related to APOC1 and APOE, respectively, in patients with ACC. Among them, 2115 and 1696 genes showed positive correlations whereas 2356 and 2325 genes showed negative correlations with APOC1 and APOE expression, respectively (Figure 4(a) and (d)). In addition, 50 genes were significantly positively and negatively correlated with APOC1 and APOE expression in patients with ACC (Figure 4(b), (c), (e), and (f)). Among them, APOC1 expression was strongly positively associated with APOE (Pearson correlation coefficient (PCC) = 0.7713, p = 9.06e-17) (Figure 4(g)), apolipoprotein C1 pseudogene 1 (APOC1P1) (PCC = 0.6812, p = 4.872e-12) (Figure 4(h)), and negatively associated with sparc/osteonectin, cwcv and kazal-like domains proteoglycan (testican) 1 (SPOCK1) (PCC = −0.5938, p = 8.008e-9) (Figure 4(i)). APOE expression was positively associated with APOC1 (PCC = 0.7713, p = 9.06e-17) (Figure 4(j)), family with sequence similarity 196 member B (FAM196B) (PCC = −0.5897, p = 1.076e-8) (Figure 4(k)), and calcium/calmodulin dependent protein kinase 1D (CAMK1D) (PCC = −0.5669, p = 5.122e-8) (Figure 4(l)) expression.
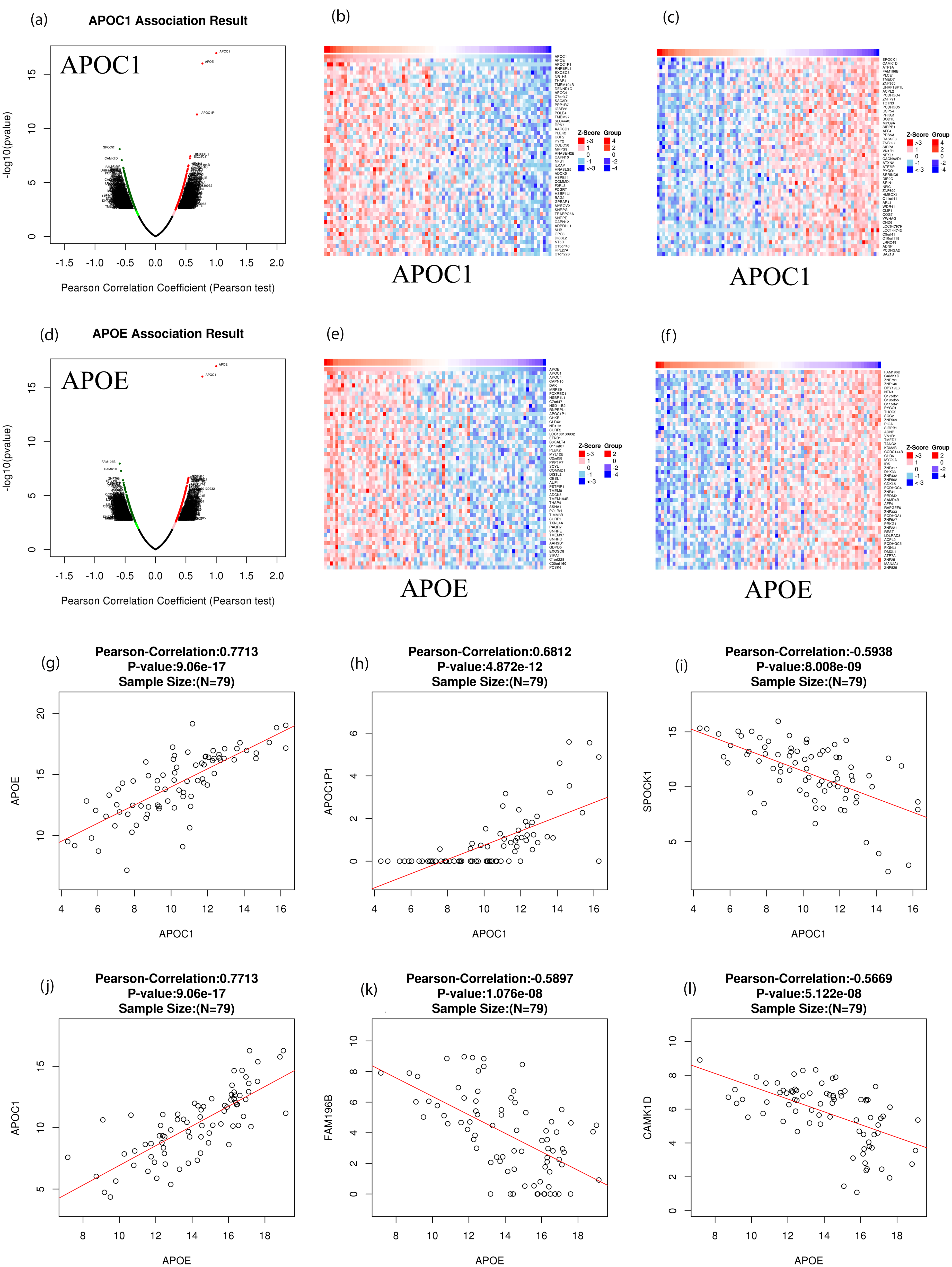
Genes differentially expressed in correlation with APOC1 and APOE expression in ACC (linkedOmics). (a and d) Pearson test was used to analyze correlations between APOC1, APOE and genes differentially expressed in ACC, respectively; (b, c, e, and f) Heat maps showing genes positively and negatively correlated with APOC1 and APOE in ACC, respectively (TOP 50 genes); The scatter plot shows Pearson correlation of APOC1 and APOE expression with expression of APOE (g), APOC1P1 (h), SPOCK1 (i), APOC1 (j), FAM196B (k), and CAMK1D (l) in ACC; Red indicates positively correlated genes and blue indicates negatively correlated genes.
Correlation of immune cell infiltration and APOC1 and APOE expression and anti-programmed cell death protein 1 (PD1)/programmed death-ligand 1 (pd-L1) immunotherapy in ACC
As shown in Figure 5(a) to (h), APOC1 and APOE expression in patients with ACC is positively associated with immune cell infiltration (p < 0.05). In addition, APOC1 was significantly downregulated in patients with ACC treated with anti-PD1 (p = 0.01) (Figure 5(i)). However, APOE expression in patients with ACC treated with anti-PD1/PD-L1 was significantly upregulated (p = 0.05) (Figure 5(j)).
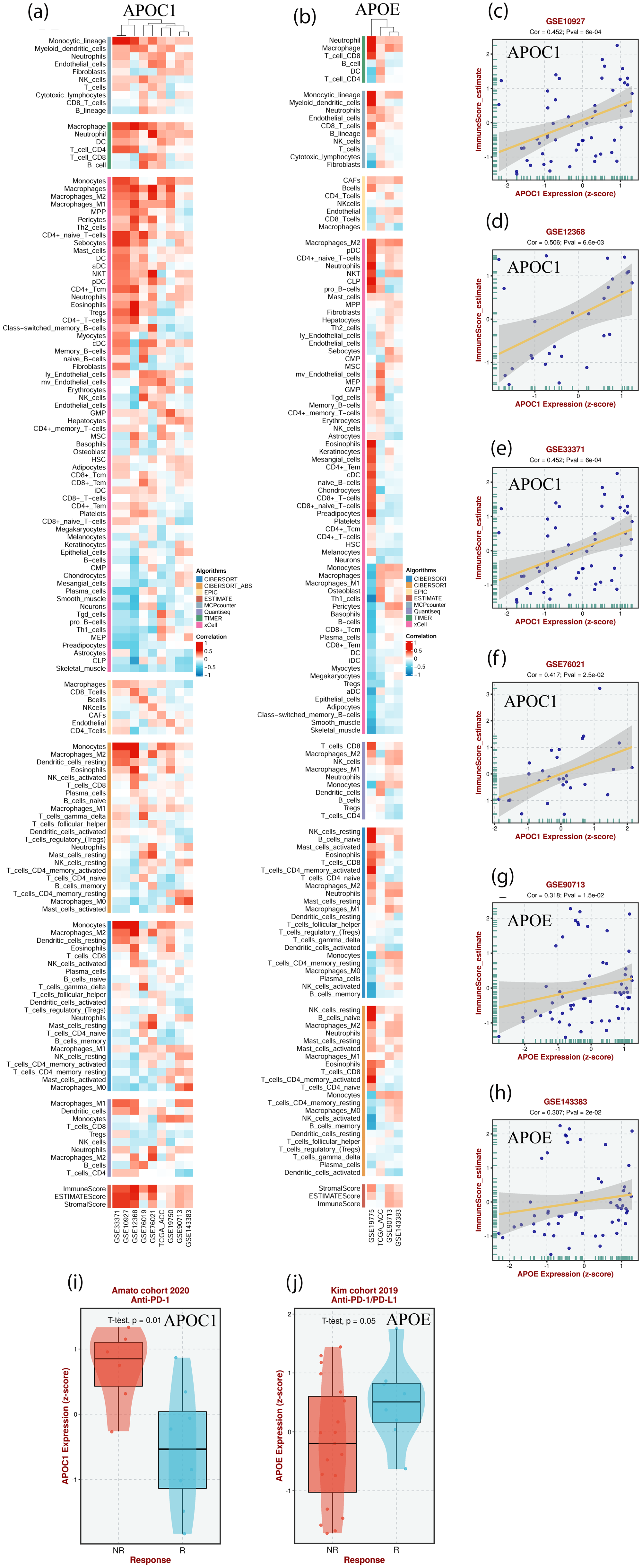
The correlation between APOC1, APOE expression and immune cell infiltration and anti-PD1/pd-L1 immunotherapy in ACC (BEST). (a and b) Heat maps showing the correlation between APOC1, APOE and immune cell infiltration in ACC, respectively; (c-h) The correlation between APOC1, APOE expression and immunescore in ACC, respectively; (i and j) Boxplot showing the correlation between APOC1, APOE expression and anti-PD1/PD-L1 immunotherapy in ACC, respectively.
Therapeutic drugs of APOC1 and APOE in ACC
The BEST database was used to evaluate APOC1 and APOE expression, where high expression indicated drug resistance; pilaralisib and elesclomol were identified as first-line drugs, respectively (Figure 6(a) and (f)). Next, the Genomics of Drug Sensitivity in Cancer database was used to evaluate the inhibitory effects of pilaralisib and elesclomol on ACC cell lines. Pilaralisib inhibited 919 cell lines with area under the curve (AUC) values greater than 0.374 (Figure 6(c)). On these cell lines, pilaralisib had a good inhibitory effect (0.573 ≤ IC50 (μM) ≤ 740) (Figure 6(b)). Moreover, pilaralisib strongly inhibited the ACC cell line SW13 (AUC values =0.857985, IC50 (μM) = 16.488967) (Figure 6(d) and (e)). In contrast, elesclomol inhibited 921 cell lines, with AUC values greater than 0.0209 (Figure 6(h)). On these cell lines, elesclomol had a good inhibitory effect (0.000231 ≤ IC50 (μM) ≤ 10.3) (Figure 6(g)). Furthermore, elesclomol also strongly inhibited SW13 (AUC values =0.421688, IC50 (μM) = 0.007628) (Figure 6(i) and (j)).
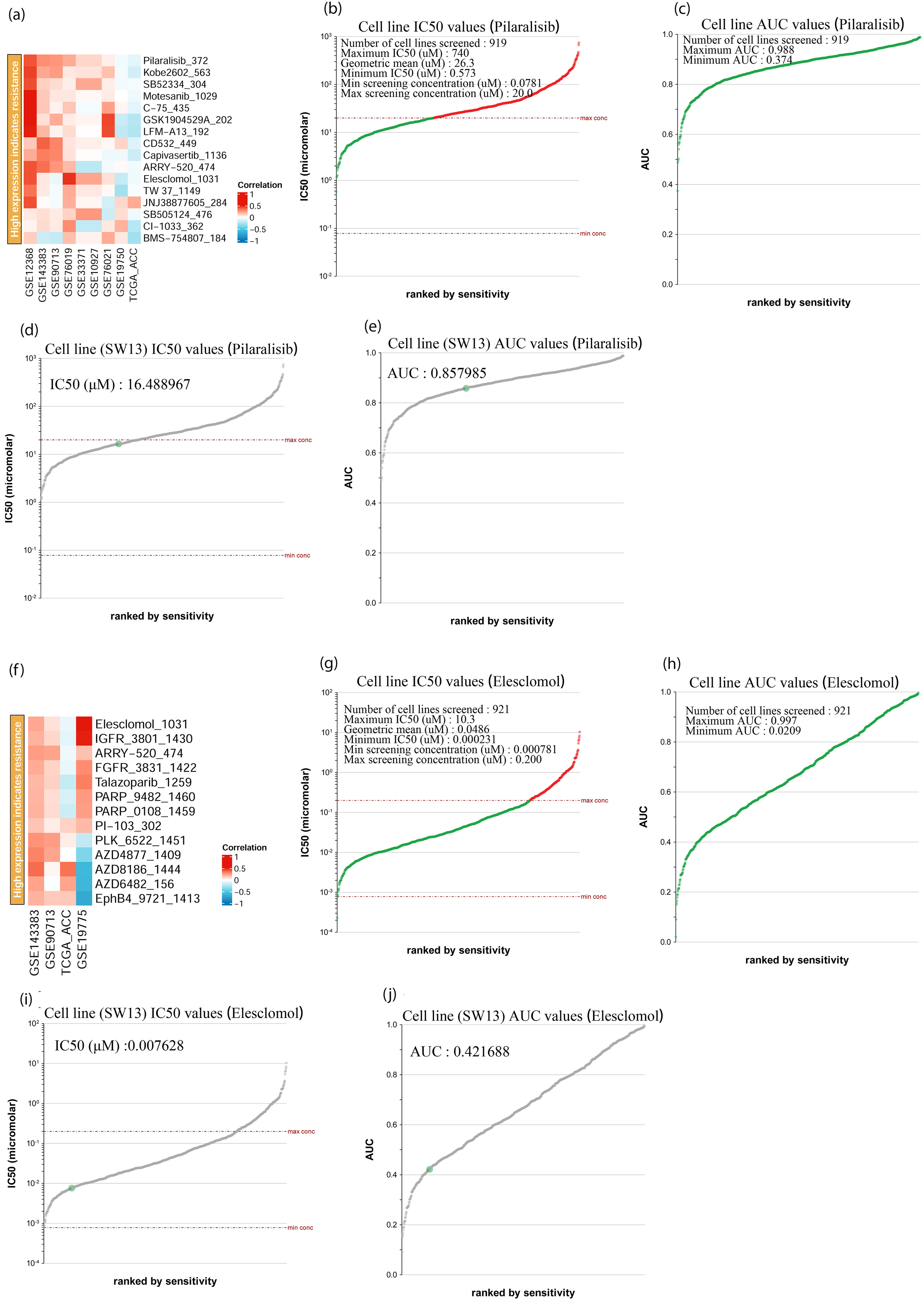
IC50 evaluation of pilaralisib and elesclomol in different tissue types of cancer (BEST and genomics of drug sensitivity in cancer). (a and f) Heat maps showing APOC1 and APOE high expression indicates resistance drugs ranking, respectively; (b) Cell line IC50 values of pilaralisib; (c) Cell line AUC values of pilaralisib; (d) SW13 cell line IC50 values of pilaralisib; (e) SW13 cell line AUC values of pilaralisib. (g) Cell line IC50 values of elesclomol; (h) Cell line AUC values of elesclomol; (i) SW13 Cell line IC50 values of elesclomol; (j) SW13 cell line AUC values of elesclomol.
Discussion
Abnormal APOC1 and APOE expression have been observed in various tumors. However, APOC1 and APOE expression in patients with ACC remains unknown. Strong downregulation of APOC1 and APOE expression has been observed in patients with ACC. APOC1 and APOE expression levels were lower in male patients with ACC than those in female patients with ACC. Low APOC1 and APOE expression levels were associated with longer survival times in patients with ACC. The number of cases in the female population generally outnumbers that of the male population (1.5:1),5,6 suggesting that sex differences in APOC1 and APOE expression may be an important factor affecting the prognosis of patients with ACC. However, this hypothesis requires further investigation. These findings suggest that APOC1 and APOE are potential targets for ACC therapy. Next, the abnormal expression of APOC1 and APOE in patients with ACC was explained through genetic alterations. These results revealed that APOC1 and APOE expression were altered by 9% and 15%, respectively, in patients with ACC, with the type of genetic alteration mainly including amplification and high and low RNA levels. In addition, genetic alterations may lead to decreased expression of APOC1 and APOE. However, further investigation is required to address this issue. Finally, the prognostic value of APOC1 and APOE expression in patients with ACC was evaluated. According to these results, patients with ACC with low APOC1 and APOE expression had longer survival times than those with high APOC1 and APOE expression. Thus, APOC1 and APOE may serve as potential prognostic markers for patients with ACC.
The potential interactions between APOC1, APOE, and the ANGs were evaluated. As a result of co-expression, co-localization, and shared protein domains, APOC1, APOE, and the ANGs were linked to a complex interaction network. The functions of APOC1, APOE, and the ANGs were also evaluated. The molecular functions related to APOC1 and its ANGs mainly included protein tyrosine kinase binding, phosphoric ester hydrolase activity, DNA and transcription factor binding, and channel activity. Protein tyrosine kinases play important roles in various cellular processes, including growth, motility, differentiation, and metabolism. Abnormal expression of protein tyrosine kinases usually leads to cell proliferation disorders and is closely related to tumor invasion, metastasis, acquired resistance, and tumor angiogenesis. 24 DNA- and RNA-binding proteins are a broad class of molecules that regulate numerous cellular processes in all living organisms, and create intricate dynamic multilevel networks to control nucleotide metabolism and gene expression. These interactions are highly regulated, and their dysregulation contributes to the development of various diseases, including cancer. 25 Targeting transcription factors and DNA-binding proteins is an important strategy in cancer treatment.26,27 The molecular functions of APOE and its ANGs included protein tyrosine kinase binding, ubiquitin-protein transferase activity, and transcriptional co-regulatory activity. Ubiquitination, one of the most important post-translational modifications, plays a versatile role in cancer-related pathways and is involved in protein metabolism, cell cycle progression, apoptosis, and transcription. 28 The regulation of ubiquitination is an important cancer treatment strategy. The current results showed that the biological processes related to APOC1 and its ANGs mainly included rRNA processing, negative regulation of phosphate metabolic processes, and positive regulation of cytokine production. The biological processes affecting tumor proliferation, invasion, metastasis, and metabolism have been reported.29–31 The biological processes related to APOE and its ANGs mainly included cytolysis, negative regulation of kinase activity, protein ubiquitination, response to growth factors, and histone modification. These biological processes affect tumor proliferation, invasion, metastasis, and angiogenesis.32–34 KEGG pathway analysis of APOE and its ANGs showed that ubiquitin-mediated proteolysis and actin cytoskeleton regulation were enriched. Actin filaments are major components of the cytoskeleton in eukaryotic cells. Mutations and abnormal expression of cytoskeletal and cytoskeleton-associated proteins play important roles in cancer cell resistance to chemotherapy and metastasis. 35 Taken together, APOC1, APOE, and the ANGs may contribute to the development and progression of ACC. Thus, ACC might be treated by regulating these genes and signaling pathways.
Transcription factors and miRNA and kinase targets of APOC1 and APOE were examined in patients with ACC. These results showed that E2F1, STAT3, SP1, and TP53 were key transcription factors of APOC1, APOE, and the ANGs in patients with ACC. E2F1 cooperates with enhancer of zeste 2 polycomb repressive complex 2 subunit to stimulate the expression of genes involved in ACC aggressiveness. 36 STAT3 promotes angiogenesis in patients with ACC, thereby making STAT3 a selective target for molecular-targeted therapy of ACC. 37 SP1 is a well-known member of the transcription factor family that plays important roles in cell growth, differentiation, apoptosis, and carcinogenesis. 38 However, its role in ACC has not yet been reported. ACC is a rare tumor type associated with TP53 mutations. Genetic susceptibility caused by mutations in TP53 is associated with approximately 50% of childhood ACC cases, but only 3%–6% of adult ACC cases. 39 Moreover, miR-19A, miR-19B, miR-182, miR-302C, miR-181A, miR-181B, miR-181C, and miR-181D were targets of APOC1 and APOE in patients with ACC. These miRNAs are involved in the proliferation, migration, invasion, and drug resistance of tumor cells, making them promising targets for cancer treatment.40–42 However, these miRNAs have not been reported to be associated with ACC. ERBB2, TGFBR1, CDK1, MAP2K7, and CDK2 were kinase targets of APOC1 and APOE in patients with ACC. CDK1 serves as a therapeutic target for ACC by regulating the epithelial-mesenchymal transition, G2/M phase transition, and PANoptosis. 43 CDK2 mRNA expression was strongly upregulated in ACC and CDK inhibitors showed dose-dependent antiproliferative effects. CDK2 mRNA expression is strongly upregulated in ACC and the CDK inhibitor flavopiridol has a dose-dependent antiproliferative effect. 44 ERBB2, TGFBR1, and MAP2K7 might be promising targets for cancer therapy. However, their relationship to ACC remains unexplored. Thus, these transcription factors, miRNAs, and kinases of APOC1, APOE, and the ANGs may be able to treat ACC.
Next, the correlation between APOC1 and APOE expression and the differentially expressed genes in patients with ACC were examined. A correlation was observed between APOC1 and APOE expression and 4471 and 4021 genes, respectively. Among them, APOC1P1 was positively and SPOCK1, FAM196B, CAMK1D were negatively correlated with the expression of APOC1 and APOE. Thus, targeting these genes may provide additional therapies for ACC. Immune infiltration of tumors plays a key role in tumor progression and outcome. 45 Cancer immunotherapy has led to significant advances in the treatment of multiple cancers. As expected, immune cell infiltration was positively correlated with APOC1 and APOE expression levels in patients with ACC. Expression levels of APOC1 and APOE in tumors are closely related to the infiltration of various immune cells. In addition, APOC1 and APOE were significantly co-expressed with immunomodulatory genes. APOC1 and APOE are immunological biomarkers that are associated with the infiltration of various immune cells.13,46 Furthermore, APOC1 and APOE were strongly downregulated and upregulated, respectively, in patients with ACC that were treated with anti-PD1/PD-L1. In patients with ACC, targeting APOC1, APOE, or the related regulatory targets may improve the immune microenvironment. Patients with ACC that are treated with anti-PD1 may have a better prognosis. Furthermore, the inhibitory effects of pilaralisib and elesclomol on ACC lines were investigated. Pilaralisib and elesclomol inhibited the expression in 919 and 921 cancer cell lines, respectively. Among them, elesclomol had a stronger inhibitory effect on the SW13 cell line than that of pilaralisib. In summary, pilaralisib and elesclomol inhibited cancer cells on a broad spectrum. The phosphoinositide 3-kinase inhibitor pilaralisib and the HSP90 inhibitor elesclomol have good antitumor effects and are safe. However, the effect of pilaralisib and elesclomol on ACC remains unclear. Thus, pilaralisib, elesclomol, and inhibitors of APOC1 and APOE or their regulatory targets, may be important strategies for the treatment of ACC. We identified the roles of APOC1 and APOE in ACC using bioinformatics methods. However, further validation through in vitro and ex vivo experiments is necessary to confirm their relationship.
In conclusion, this study preliminarily clarified that APOC1 and APOE might be potential therapeutic and prognostic targets for ACC and provided insights into the mechanism and treatment of ACC.
Footnotes
Author contributions
INTERPRETATION OR ANALYSIS OF DATA: SL and YS.
PREPARATION OF THE MANUSCRIPT: SL and SX.
REVISION FOR IMPORTANT INTELLECTUAL CONTENT: SL, SX and YS.
SUPERVISION: YS.
Funding
This research was funded by postdoctoral Foundation of Guangdong Medical University (4SG22292G).
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
