Abstract
Background
Fragile histidine triad (FHIT) is a strong tumor suppressor gene, and cells deficient in FHIT are prone to acquiring cancer-promoting mutations. Due to its location, deletions within FHIT are common in cancer. Over 50% of cancers show loss of FHIT expression. However, to date, expression levels, gene regulatory networks, prognostic value, and target prediction of FHIT in lung adenocarcinoma (LUAD) and lung squamous cell carcinoma (LUSC) have not been fully reported. Therefore, systematic analysis of FHIT expression, gene regulatory network, prognostic value, and targeted prediction in patients with LUAD and LUSC has important guiding significance, providing new therapeutic targets and strategies for clinical treatment of lung cancer to further improve the therapeutic effect of lung cancer.
Methods
Multiple free online databases were used for the abovementioned analysis in this study, including cBioPortal, TRRUST, Human Protein Atlas, GeneMANIA, GEPIA, Metascape, UALCAN, LinkedOmics, and TIMER.
Results
FHIT was upregulated in patients with LUAD, and downregulated in patients with LUSC. Genetic alterations of FHIT were found in patients with LUAD (7%), and LUSC (10%). The promoter methylation of FHIT was lower in patients with LUAD and LUSC. FHIT expression significantly correlated with LUSC pathological stages. Furthermore, patients with LUAD and LUSC having low FHIT expression levels had a longer survival than those having high FHIT expression levels. FHIT and its neighboring genes (the 50 most frequently altered neighboring genes of FHIT) functioned in the regulation of protein kinase and DNA binding in patients with LUAD, as well as cell channels and membrane potential in patients with LUSC. Gene ontology enrichment analysis revealed that the functions of FHIT and its neighboring genes are mainly related to disordered domain-specific binding, protein kinase binding, and ion gated channel activity in patients with LUAD, as well as calcium ion binding and intracellular ligand-gated ion channel activity in patients with LUSC. Transcription factor targets of FHIT and its neighboring genes in patients with lung cancer were found: USF1, SOX6, USF2, SIRT1, VHL, LEF1, EZH2, TP53, HDAC1, ESR1, EGR1, YY1, MYC, RELA, NFKB1, and E2F1 in LUAD; and HDAC1, DNMT1, and E2F1 in LUSC. We further explored the FHIT-associated kinase (PRKCQ, AURKB and ATM in LUAD as well as PLK3 in LUSC) and FHIT-associated miRNA targets (MIR-188, MIR-323, and MIR-518A-2 in LUAD). Furthermore, the following genes had the strongest correlation with FHIT expression in patients with lung cancer: NICN1, HEMK1, and BDH2 in LUAD, and ZMAT1, TTC21A, and NICN1 in LUSC. FHIT expression was positively associated with immune cell infiltration (B cell) in patients with LUAD, as well as B cell, CD8 + T, CD4 + T cells, macrophages, and dendritic cells in patients with LUSC. Nevertheless, FHIT expression was negatively associated with CD8 + T cells and neutrophils in patients with LUAD.
Conclusions
The expression, gene regulatory network, prognostic value and targeted prediction of FHIT in patients with LUAD and LUSC were systematically analyzed and revealed in this study, thereby laying a foundation for further research on the role of FHIT in LUAD and LUSC occurrence. This study provides new LUAD and LUSC therapeutic targets and prognostic biomarkers as a reference for fundamental and clinical research.
Introduction
Cancer is second only to heart disease as the leading cause of death worldwide. According to the International Agency for Research on Cancer, the number of cancer cases increased to 19.3 million and the number of deaths increased to 10 million in 2020. 1 Lung cancer (LC) is one of the most common cancers, accounting for 11.6% of all cancer cases worldwide. According to the Global Burden of Disease Study, the 5-year survival rate of LC in 2020 was only 17.8%, significantly lower than other types of cancer. 2 Moreover, LC accounts for 18.4% of all cancer deaths worldwide. Non-small cell lung cancer (NSCLC) is the most common pathological type of LC, accounting for more than 80% of LC. Lung adenocarcinoma (LUAD) and lung squamous cell carcinoma (LUSC) are two important histopathological subtypes of NSCLC. LUAD is mostly found in non-smoking women. Due to drug resistance, early diagnosis failure, and lack of effective treatment, patients with LUAD have a low survival rate. On the other hand, LUSC is one of the most common forms of NSCLC, with approximately 400,000 new cases per year, accounting for 20%–30% of NSCLC. 3 Different cancer subtypes have different origins, histology, and genetic and epigenetic changes. Recent studies suggest that LUAD and LUSC should be classified and treated as distinct cancers. 4 Therefore, systematic analysis of targeted prediction and prognostic markers in patients with different subtypes of LC is helpful to achieve accurate treatment and to improve the survival rate of patients with LC.
Fragile histidine triad (FHIT) is a tumor suppressor gene that induces apoptosis and inhibits epithelial-mesenchymal transition. 5 Due to the location of FHIT (located at 3p14. 2, which contains the FRA3B gene, which is the most common vulnerability site in the genome), 6 and loss of FHIT is common in cancer (loss of FHIT expression were shown in over 50% of cancers). 7 FHIT is considered to be the guardian of the genome, which is similar to p53. The expression of FHIT can prevent replication pressure, forks collapse, and opposing replication forks stalling, and the accumulation of double strand breaks during DNA replication. 8 FHIT-deficient cells are prone to acquire oncogenic mutations and are more easily transformed in the presence of DNA damage factors. 9 FHIT is a promising candidate biomarker for the early diagnosis of cancer because of the absence of FHIT expression observed in precancerous lesions in a variety of cancers. 10 In addition, detection of FHIT activity may contribute to precancer diagnosis and prevention. Given the differences in FHIT expression levels in different tumors, 11 either FHIT agonists or inhibitors may be promising anticancer drugs for the treatment of cancers with abnormal FHIT expression.
In this study, we used multiple free online databases to identify the expression level, gene regulation network, prognostic value, and regulation target of FHIT in patients with LUAD and LUSC. Moreover, this study aimed to further reveal the relationship between FHIT expression and LC occurrence, and to provide new insights into the target treatment and prognostic biomarkers of patients with LUAD and LUSC.
Materials and methods
UALCAN analysis
UALCAN (http://ualcan.path.uab.edu/analysis.html) is an online database used as a tool for tumor subgroup gene expression and survival analyses. 12 The “Expression Analysis” module of UALCAN database was used to analyze TCGA gene expression data, and the screening criteria were set as: (a) Gene: FHIT; (b) Dataset: LUAD and LUSC; and (c) Threshold setting conditions: P-value cutoff = 0.05. The Student’s t-test was used for comparative analysis.
Human protein atlas analysis
The Human Protein Atlas (https://www.proteinatlas.org/) is an open access resource for human protein search for specific genes/proteins. The screening criteria were as follows: (a) Gene: FHIT; (b) choose section: tissue and pathology; (c) choose tissue: lung and lung cancer (LUAD and LUSC); and (d) choose picture of tissue types: normal lung tissue, LUAD, and LUSD.
GEPIA
Gene expression profiling analysis (GEPIA) (http://gepia.cancer-pku.cn/index.html) is a free online platform that provides RNA sequencing expression data of 9736 tumors and 8587 normal samples for analyzing differential mRNA expression, promoter methylation, pathological stage, and correlative prognosis in this study. 13 The screening criteria were: (a) Gene: FHIT; (b) dataset: LUAD and LUSC; and (c) threshold setting conditions: P-value cutoff = 0.05. The Student’s t-test was used to analyze the expression of FHIT and the pathological stage of LUAD and LUSC. The Kaplan–Meier curve was used to analyze the prognosis of patients with LUAD and LUSC.
cBioPortal analysis
The cBioPortal (http://cbioportal.org) is an open online database used to visualize, study, and analyze cancer genetic data. 14 In our study, the analysis of genetic alterations of FHIT and its neighboring genes (the top 50 altered neighboring genes of FHIT) was conducted using the cBioPortal database. The screening criteria were: (a) 230, and 178 samples of LUAD and LUSC, respectively, were analyzed; (b) mRNA expression z-scores relative to all samples (log RNA Seq V2 RSEM) were obtained using a z-score threshold of ± 2.0; and (c) gene: FHIT.
STRING analysis
STRING (https://string-db.org/cgi/input.pl) is an online free database used to construct protein–protein interaction (PPI) networks between target proteins. In this study, we built the PPI network interaction by screening conditions with low confidence (0.150) and defined species as Homo sapiens.
GeneMANIA analysis
GeneMANIA (http://www.genemania.org) is an analysis tool for building PPIs, generating hypotheses about gene function, analyzing gene lists, and sequencing genes for function determination. In this study, we constructed interaction networks for exploring the role of FHIT and the top 50 altered neighboring genes.
Metascape analysis
Metascape (https://metascape.org) is a simple and powerful gene function annotation and analysis tool that can help users apply the currently popular bioinformatics analysis methods for the analysis of batch genes and proteins to understand gene or protein function. 15 In our study, gene ontology (GO) function and the Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis of FHIT and its altered neighboring genes in LUAD and LUSC were investigated using Metascape.
TRRUST analysis
TRRUST (https://www.grnpedia.org/trrust/) is a manually curated database of human transcriptional regulatory networks that contain 8444 and 6552 transcription factor-target regulatory relationships of 800 human transcription factors. 16 In this study, we attempted to identify the key factor regulating the expression of FHIT and its altered neighboring genes in patients with LUAD and LUSC using the TRRUST database.
LinkedOmics analysis
LinkedOmics (http://www.linkedomics.org/) is a public online platform that includes multi-omics data from all 32 The Cancer Genome Atlas (TCGA) cancer types. 17 It provides methods for analyzing and comparing cancer multi-omics data within and across tumor types. In our study, kinase target enrichment, miRNA target enrichment, and genes differentially expressed in correlation with FHIT were conducted using the “LinkInterpreter” module of LinkedOmics. The screening criteria were set as follows: (a) a minimum number of three genes (size); (b) cancer type: LUAD and LUSC; (c) a simulation of 500; (d) search attribution: FHIT and the top 50 altered neighboring genes; and (e) target dataset: RNA-seq (data type).
Timer analysis
TIMER (https://cistrome.shinyapps.io/timer/) is a comprehensive resource for the systematical analysis of tumor-infiltrating immune cells, including B cells, CD4 + T cells, CD8 + T cells, neutrophils, macrophages, and dendritic cells. 18 In our study, the correlation between FHIT expression level and the infiltration of immune cells was assessed by the “Gene module” of TIMER.
Results
FHIT expression in patients with LUAD and LUSC
We compared the expression levels of FHIT in patients with LUAD and LUSC having normal human tissues, and found that the transcriptional levels of FHIT stratified by sample type, sex, and pathological stage were significantly upregulated in patients with LUAD (P < 0.05, P < 0.01, or P < 0.001) (Figure 1(a) to (c)). In addition, immunohistochemical results showed that the expression level of FHIT protein in patients with LUAD was significantly increased compared with healthy humans (Supplementary Figure 1(a) and (b). Inversely, the expression level of FHIT in LUSC tissues was decreased based on sample type, sex, and pathological stage (P < 0.001) (Figure 1(d) to (f)). Immunohistochemical results revealed that the expression level of FHIT protein was significantly decreased (Supplementary Figure 1(a) and (c)).

The transcription levels of FHIT in lung cancer, stratified based on gender and stages (UALCAN database). (a) Boxplot showing relative expression of FHIT in healthy and LUAD samples. (b) Boxplot showing relative expression of FHIT of either gender in healthy individuals and LUAD patients. (c) Boxplot showing relative expression of FHIT of stages in healthy individuals and LUAD patients. (d) Boxplot showing relative expression of FHIT in healthy and LUSC samples. (e) Boxplot showing relative expression of FHIT of either gender in healthy individuals and LUSC patients. (f) Boxplot showing relative expression of FHIT of stages in healthy individuals and LUSC patients. Data are mean ± SE. *P < 0.05; **P < 0.01; ***P < 0.001.
Furthermore, we assessed the correlation between the differential expression of FHIT and the pathological stage in patients with LUAD and LUSC. Our results showed a significant correlation between the expression of FHIT and the pathological stage in patients with LUSC (P = 0.00494) (Figure 2(b). Moreover, we used GEPIA to evaluate the prognostic value of FHIT expression in patients with LUAD and LUSC. The overall survival was longer in patients with LUSC having lower FHIT expression levels (P = 0.011) (Figure 3(b)). The percentage survival was higher in patients with LUAD having lower FHIT expression levels before 100 months (P = 0.046) (Figure 3(c)).
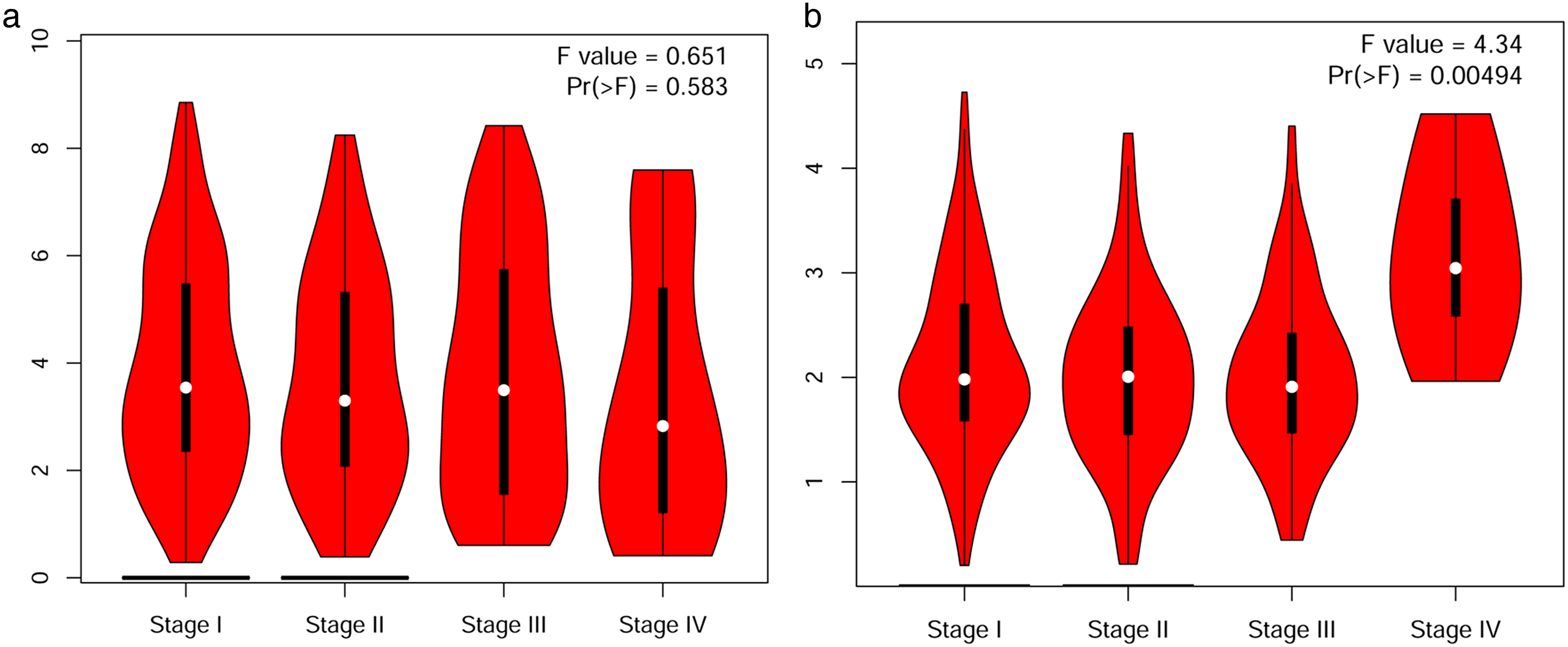
Correlation between the expression level of FHIT and the pathological stage in lung cancer (GEPIA database). (a) LUAD; (b) LUSC.

The prognostic value of FHIT in lung cancer (GEPIA database). (a) The overall survival curve of FHIT in LUAD. (b) The overall survival curve of FHIT in LUSC. (c) The disease-free survival cure of FHIT in LUAD. (d) The disease-free survival cure of FHIT in LUSC.
Genetic alteration and promoter methylation level of FHIT expression in patients with LUAD and LUSC
Genetic alterations of FHIT in patients with LUAD and LUSC were evaluated using TCGA. Our results showed that FHIT expression was altered by 7% in patients with LUAD (Supplementary Figure 2(a)). However, the promoter methylation level of FHIT expression stratified by sample type, sex, and pathological stage were lower in patients with LUAD than in healthy humans (P < 0.001) (Supplementary Figure 2(b) to (d)). Moreover, we found a 10% FHIT expression level alteration in patients with LUSC (Supplementary Figure 2(e)). Similarly, the promoter methylation level of FHIT expression stratified by sample type, sex, and pathological stage were lower in patients with LUSC than in healthy humans (P < 0.05, P < 0.01, or P < 0.001) (Supplementary Figure 2(f) to (h)).
Neighboring gene alteration and FHIT interaction network in patients with LUAD and LUSC
The neighboring gene alteration of FHIT expression in patients with LUAD and LUSC was analyzed using cBioPortal software. We found FHIT neighboring gene alteration frequencies ≥18.75%, and ≥23.53% (the 50 most frequently altered neighboring genes) in patients with LUAD and LUSC, respectively (Supplementary Table 1 and Table 2). The most frequently altered FHIT neighboring genes in patients with LUAD were TP53 (56.25%), and TTN (50.00%) (Table 1). In addition, TP53 (82.35%), RYR2 (70.59%), and TTN (70.59%) were the most frequently altered FHIT neighboring genes in patients with LUSC (Supplementary Table 2).
Key regulated factor of FHIT and the top 50 neighbor-altered genes in lung cancer (TRRUST database).
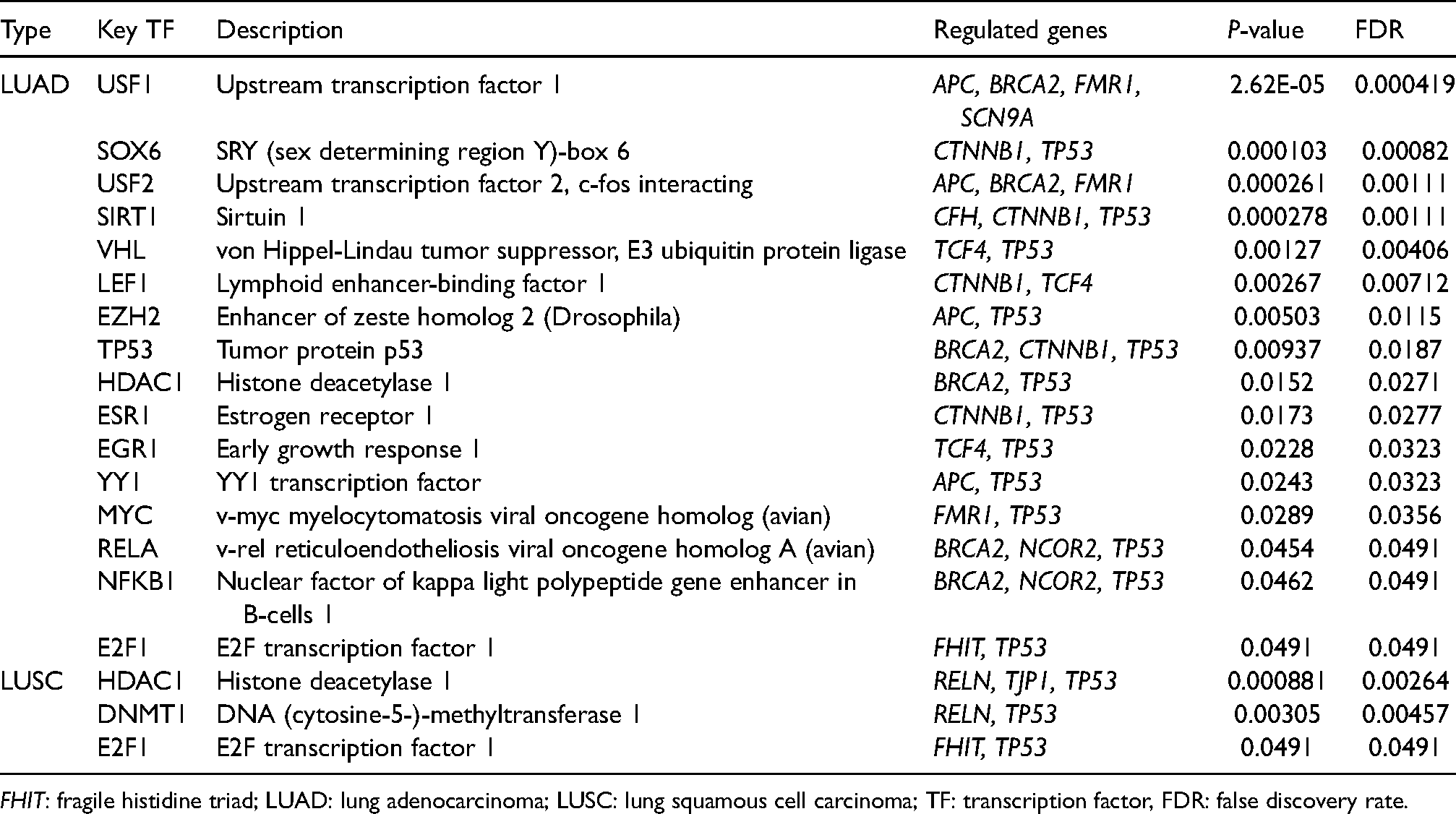
FHIT: fragile histidine triad; LUAD: lung adenocarcinoma; LUSC: lung squamous cell carcinoma; TF: transcription factor, FDR: false discovery rate.
We then explored the potential interactions between FHIT and its neighboring genes. The PPI network was constructed using STRING software. We obtained 48 nodes and 226 edges in the PPI networks in patients with LUAD (Supplementary Figure 3(a)). Moreover, protein kinase binding, regulatory region DNA binding, and regulatory region nucleic acid binding were the primary functions of FHIT and its neighboring genes in patients with LUAD (Supplementary Figure 3(b)). We also obtained 47 nodes and 258 edges in the PPI networks in patients with LUSC (Supplementary Figure 3(c)). FHIT and its neighboring genes had the following functions in patients with LUSC: intracellular ligand-gated ion channel activity, sarcoplasmic reticulum calcium ion transport, and regulation of membrane potential (Supplementary Figure 3(d)). FHIT is connected to its neighboring genes in a complex interaction network by co-expression, physical interactions, predicted shared protein domains, and co-localization (Supplementary Figure 3).
GO and KEGG pathway enrichment analysis
The GO function and KEGG pathway enrichment analysis of FHIT and the top 50 altered neighboring genes in patients with LUAD and LUSC were explored using Metascape software. Our results showed that the cellular components related to FHIT and its neighboring genes in patients with LUAD were mainly involved in Z disc, ion channel complex, photoreceptor cell cilium, and the microtubule organizing center (Supplementary Figure 4(a)). Moreover, cellular response to nitrogen compound, heart morphogenesis, histone modification, and apoptotic signaling pathway was the main biological processes of the expression of FHIT and its altered neighboring genes in patients with LUAD (Supplementary Figure 4(b)). The molecular functions of FHIT and its top 50 altered neighboring genes in patients with LUAD mainly included disordered domain-specific binding, protein kinase binding, and ion gated channel activity (Supplementary Figure 4(c)). The KEGG pathway of FHIT and its neighboring genes in LUAD were mainly involved in the signaling pathway of breast cancer, Cushing syndrome, and arrhythmogenic right ventricular cardiomyopathy (Supplementary Figure 4(d)). The Z disc, catenin complex, and sarcolemma were cellular components related to the expression of FHIT and its neighboring genes in patients with LUSC (Supplementary Figure 4(e)). In addition, the main biological processes involving the expression of FHIT and its top 50 altered neighboring genes in patients with LUSC were cell–cell junction organization, heart development, and muscle structure development (Supplementary Figure 4(f)). The molecular functions of FHIT and its neighboring genes in patients with LUSC were calcium ion binding, intracellular ligand-gated ion channel activity, and actin binding (Supplementary Figure 4(g)). Extra-cellular matrix (ECM)-receptor interaction, hypertrophic cardiomyopathy, and small cell lung cancer were involved in the KEGG pathway analysis of FHIT and its top 50 altered neighboring genes in patients with LUSC (Supplementary Figure 4(h)).
Transcription factor targets of FHIT in patients with LUAD and LUSC
Potential transcription factor of FHIT in patients with LUAD and LUSC were acquired using TRRUST software (Table 1). Upstream transcription factor 1 (USF1), SRY (sex determining region Y)-box 6 (SOX6), upstream transcription factor 2, c-fos interaction (USF2), sirtuin 1 (SIRT1), von Hippel-Lindau tumor suppressor, E3 ubiquitin protein ligase (VHL), lymphoid enhancer-binding factor 1 (LEF1), enhancer of zeste homolog 2 (Drosophila) (EZH2), tumor protein p53 (TP53), histone deacetylase 1 (HDAC1), estrogen receptor 1 (ESR1), early growth response 1 (EGR1), YY1 transcription factor (YY1), v-myc myelocytomatosis viral oncogene homolog (avian) (MYC), v-rel reticuloendotheliosis viral oncogene homolog A (avian) (RELA), nuclear factor of kappa light polypeptide gene enhancer in B-cells 1 (NFKB1), and E2F transcription factor 1Hypoxia inducible factor 1 (E2F1) were the key transcription factors involved in the network of FHIT and its neighboring genes in patients with LUAD (P < 0.05) (Table 1). APC, BRCA2, FMR1, and SCN9A were the regulated genes of USF1. In addition, SOX6 regulated the functions of CTNNB1 and TP53. Furthermore, APC, BRCA2, and FMR1 were the regulated genes of USF2. CFH, CTNNB1, and TP53 were the regulated genes of SIRT1. VHL regulated the functions of TCF4 and TP53. The regulated genes of LEF1 were CTNNB1 and TCF4. EZH2 regulated the functions of APC and TP53. BRCA2, CTNNB1, and TP53 were regulated genes of TP53. The regulated genes of HDAC1 were BRCA2 and TP53. ESR1 regulated the functions of CTNNB1 and TP53. TCF4 and TP53 were regulated genes of EGR1. The regulated genes of YY1 were APC and TP53. MYC regulated the functions of FMR1 and TP53. BRCA2, NCOR2, and TP53 were regulated genes of RELA and NFKB1. The regulated genes of E2F1 were FHIT and TP53 (Table 1). To a greater degree, the key transcription factor targets involved in the network of FHIT and its neighboring genes were histone deacetylase 1 (HDAC1), DNA (cytosine-5-)-methyltransferase 1 (DNMT1), and E2F transcription factor 1 (E2F1) in patients with LUSC (P < 0.05) (Table 1). The regulated genes of HDAC1 were RELN, TJP1, and TP53. In addition, DNMT1 regulated the functions of RELN and TP53. Furthermore, FHIT and TP53 were the regulated genes of E2F1 (Table 1).
Kinase and miRNA targets of FHIT in patients with LC
We also obtained the top three kinase targets and miRNA targets of the FHIT network with LinkedOmics (Table 2). Kinase PRKCQ, kinase AURKB, and kinase ATM were the kinase targets of FHIT in patients with LUAD (P < 0.01). Moreover, kinase targets of FHIT were kinase PLK3 in patients with LUSC (P < 0.05). FHIT miRNA-target network in LUAD was associated with (AAGGGAT) MIR-188, (TAATGTG) MIR-323, and (TTTGCAG) MIR-518A-2 (P < 0.01).
The kinase target and miRNA target of FHIT in different type of lung cancer (LinkedOmics database).
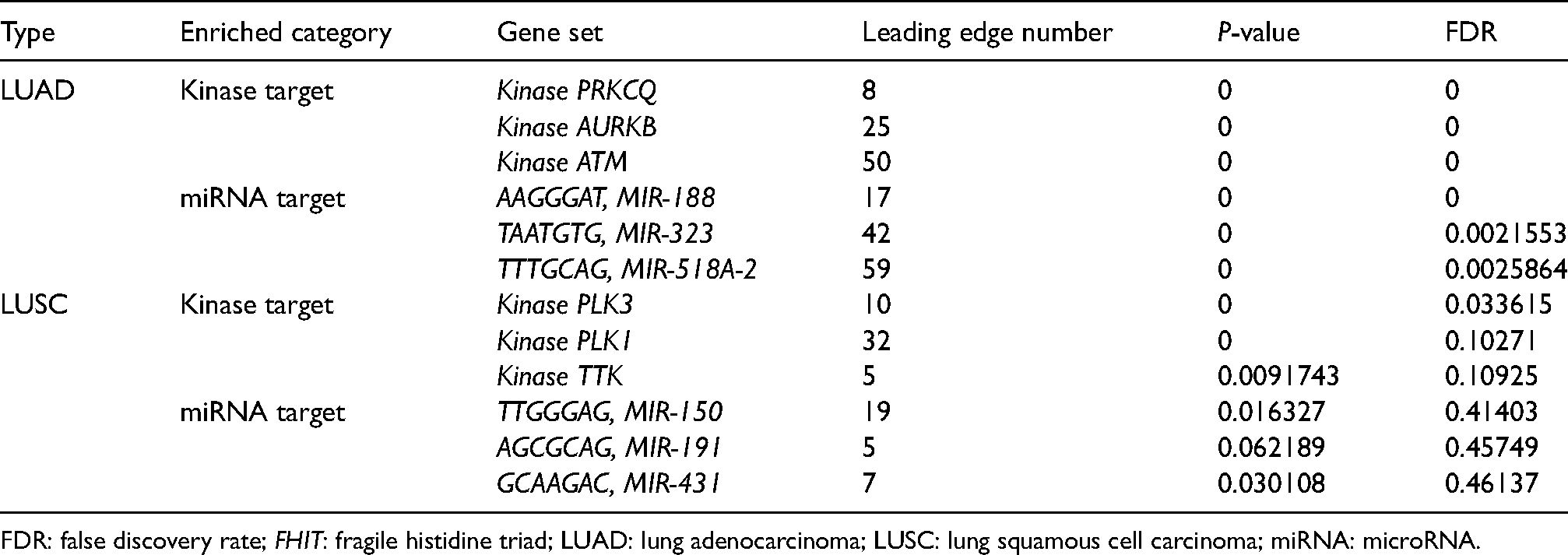
FDR: false discovery rate; FHIT: fragile histidine triad; LUAD: lung adenocarcinoma; LUSC: lung squamous cell carcinoma; miRNA: microRNA.
Correlation of differentially expressed genes and FHIT expression in patients with LUAD and LUSC
We obtained mRNA sequencing data of 515 patients with LUAD in the TCGA database of LinkedOmics. As shown in Supplementary Figure 5(a), 19,988 genes were closely related to FHIT. Among them, 11,442 and 8546 genes showed positive and negative correlations, respectively, with FHIT expression. In addition, 50 significant genes were positively and negatively correlated with FHIT expression in patients with LUAD (P < 0.05) (Supplementary Figure 5(b) and (c)). Moreover, the expression of FHIT had a strong positive association with NICN1 (Pearson correlation coefficient = 0.4955, P = 2.891e–33) (Supplementary Figure 6(a)), HEMK1 (Pearson correlation coefficient = 0.4943, P = 4.348e–33) (Supplementary Figure 6(b)), and BDH2 (Pearson correlation coefficient = 0.4934, P = 5.912e–33) (Supplementary Figure 6(c)) expressions. Furthermore, we analyzed mRNA sequencing data of 501 patients with LUSC in the TCGA database, and found that 20,103 genes were closely related to FHIT expression. Among them, 13,645 and 6458 genes showed positive and negative correlations, respectively, with FHIT expression (Supplementary Figure 5(d)). We also identified 50 significant genes that were positively and negatively correlated with FHIT expression in patients with LUSC (Supplementary Figure 5(e) and (f)). In addition, FHIT expression was positively associated with ZMAT1 (Pearson correlation = 0.5412, P = 1.799e–39) (Supplementary Figure 6(d)), TTC21A (Pearson correlation = 0.5036, P = 1.4446e–33) (Supplementary Figure 6(e)), and NICN1(Pearson correlation = 0.4991, P = 6.495e–33) (Supplementary Figure 6(f)) expressions.
Immune cell infiltration and FHIT expression in patients with LUAD and LUSC
To further reveal the relationship between immune cell infiltration and FHIT expression in patients with LUAD and LUSC, we examined the correlation between immune cell infiltration and FHIT expression using TIMER software. The expression level of FHIT in patients with LUAD was positively associated with the infiltration of B cells (Cor = 0.16, P = 4.09e-4) (Supplementary Figure 7(a)). However, CD8 + T cells (Cor = −0.119, P = 8.75e − 3) and neutrophil infiltrations (Cor = −0.163, P = 3.23e − 4) were negatively associated with FHIT expression (Supplementary Figure 7(a)). Furthermore, we analyzed the correlation between immune cell infiltration and FHIT expression in patients with LUSC. The infiltration of B cells (Cor = 0.197, P = 1.57e-5), CD8 + T (Cor = 0.144, P = 1.64e-3), CD4 + T cells (Cor = 0.239, P = 1.41e-7), macrophages (Cor = 0.194, P = 2.04e-5), and dendritic cells (Cor = 0.172, P = 1.71e-4), were positively associated with the expression of FHIT (Supplementary Figure 7(b)).
Discussion
FHIT has been well demonstrated as a tumor suppressor gene and has been reported in LC through deletion, reduced expression, or promoter methylation inactivation. 19 However, the expression level and promoter methylation of FHIT in patients with LUAD and LUSC remain controversial.20–22 In LC cell lines, restoration of FHIT expression leads to the induction of apoptosis and tumorigenic inhibition, so the gene is considered a potential drug for targeted gene therapy. Due to a variety of reasons, there are currently no reports on anti-cancer drugs targeting FHIT. Whether it is an FHIT agonist or an inhibitor, it may be a promising anticancer agent for the treatment of cancers with an abnormal expression of FHIT. Therefore, we sought to reveal the expression, gene regulatory network, prognostic value, and targeted prediction of FHIT in patients with NSCLC subtypes (LUAD and LUSC).
Moreover, we explored the expression level of FHIT and the correlation between the differential expression of FHIT and pathological stage in patients with LUAD and LUSC. FHIT expression was upregulated in patients with LUAD than in healthy humans. Nevertheless, patients with LUSC had a downregulated FHIT expression. These findings were similar to previous study findings in patients with LUSC, and contrary to patients with LUAD. 20 Our results also showed that in different cancer stages, the expression level of FHIT is inconsistent. From stage 1 to stage 4, the expression level of FHIT in the patients with LUAD gradually decreases to the level of normal people. We found that the expression level of FHIT in males and females was also different, and the expression level of FHIT in males was significantly higher than in females. Although the expression level of FHIT in patients with LUAD is in conflict with the results of previous studies, the gender difference in FHIT expression is consistent with the results of previous studies. 20 Studies had found that FHIT is involved in abnormal cell proliferation by disrupting laryngeal G0–G1 arrest and apoptosis, and the down-regulation of FHIT levels may be a clinically useful prognostic indicator in laryngeal squamous cell carcinoma. 23 In addition, overexpressed FHIT protein did not (a) alter tumor cell morphology; (b) inhibit tumor cell proliferation; and (c) change tumor cell cycle dynamics. 11 Other studies have revealed that LUSC show more frequent FHIT loss than LUAD (87% vs. 57%), 24 and loss of FHIT expression is a relatively late event in this development of LUAD. 25 A pilot study was conducted on 50 BALf samples: 25 from LUSC patients and 25 from healthy donors. Analysis of RQ values revealed down-regulation of FHIT in LUSC patients than healthy donors (0.75 vs. 1.01). 22 Unfortunately, there is still no relevant experimental data to support the high expression of FHIT in patients with LUAD. In brief, as a tumor suppressor gene, there are differences in the expression level and function of FHIT in different tumors and different stages, and the reasons still need to be further studied. It should be noted that due to the limitations of the UALCAN platform, there may be some errors in the direct Student’s t-test without the normality test of the data in Figure 1. But from the box plots observation, it basically conformed to the normal distribution. Next, we attempted to explain the conflicting results through gene alteration and promoter methylation in patients with LUAD and LUSC. We found that genetic alterations occurred in 7% and 10% of patients with LUAD and LUSC, respectively. Moreover, interestingly, the promoter methylation level of FHIT was lower in patients with LUAD and LUSC than in healthy individuals, which is not consistent with previous findings.26,27 We hypothesized that there were differences in methylation expression in different NSCLC subtypes at different stages. Moreover, we assume that the different genetic alteration and demethylation of FHIT may be the leading cause of different FHIT expressions in patients with LUAD and LUSC. The disadvantage is that the differences in FHIT expression, methylation, and mutation levels were obtained from different databases, which may have a certain bias in the inference of this causal relationship.
In addition, we found a significant correlation between FHIT expression and LUSC pathological stages. Furthermore, the survival of patients with LUSC having low FHIT expression levels was longer than that of patients with LUSC having high FHIT expression levels. Notably, we obtained same results before 100 months in patients with LUAD based on disease-free survival curves. Thus, the FHIT expression level may be a prognostic indicator in patients with LUAD and LUSC. Some studies have shown that the loss of FHIT expression is significantly negatively correlated with patient survival. It may be an independent prognostic indicator of clinical outcome in patients with NSCLC. 28
Moreover, the 50 most frequent FHIT neighboring genes were altered in patients with LC according to the following frequencies: ≥18.75% (LUAD) and ≥23.53% (LUSC). We then explored the potential interactions and functions of FHIT and its neighboring genes, which were found to have complex and tight connection networks. In patients with LUAD, these genes were mainly involved in regulation of protein kinase, region DNA binding, and nucleic acid. However, in patients with LUSC, FHIT neighboring genes functioned in ion transport and membrane potential. It has been confirmed that various protein kinases are involved in the development of LC. For example, phosphorylation of MAP2Ks by MAP3K19 protein kinase activates ERK and JNK kinases and increases the viability of KRAS mutant LC cells. 29 In addition, affecting the function and damage of DNA and nucleic acids can affect cell proliferation and differentiation, which can cause cancer. 30 Finally, some studies have confirmed that there are extensive changes in ion homeostasis and membrane potential during the development of cancer, which play a key role in the occurrence and development of malignant tumors. 31 The above evidence reveals that FHIT and neighboring genes might impact the occurrence and progression of LUAD and LUSC.
Furthermore, GO enrichment analysis revealed that the functions of FHIT and its neighboring genes in patients with LCAD are mainly related to disordered domain-specific binding, protein kinase binding, and ion gated channel activity. However, our results showed that calcium ion binding, intracellular ligand-gated ion channel activity, and actin binding were the molecular functions of FHIT and its neighboring genes in patients with LUSC. The consistency of the function of FHIT and its neighboring genes was verified again. We found that the signaling pathway of breast cancer and Cushing syndrome were mainly involved in the KEGG pathway analysis of FHIT and its neighboring genes in patients with LUAD. Small cell lung cancer is a neuroendocrine tumor that has the potential to secrete various peptides or hormones that may cause ectopic Cushing syndrome. 32 Cushing syndrome cardiomyopathy is an unusual manifestation of small cell lung cancer. Studies have shown that breast cancer patients are more likely than the general population to develop LC; a potential link between them. 33 Furthermore, KEGG pathway analysis revealed that the functions of FHIT and its neighboring genes in patients with LCSC are mainly related to the signaling pathway of ECM-receptor interaction and small cell lung cancer. An increasing number of studies have shown that ECM-receptor interaction is involved in tumor proliferation, differentiation, and metastasis. 34 Thus, regulating these signaling pathways may be a potential treatment strategy of LUAD and LUSC.
We also analyzed the targets and regulators of FHIT in patients with LUAD and LUSC. We first explored the transcription factor targets of FHIT and its neighboring genes in patients with LUAD and LUSC, and found that USF1, SOX6, USF2, SIRT1, VHL, LEF1, EZH2, TP53, HDAC1, ESR1, EGR1, YY1, MYC, RELA, NFKB1, and E2F1 were crucial regulatory factors in patients with LUAD. USF1 is a commonly expressed transcription factor, increasing evidence indicates that USF1 up-regulation can inhibit lung metastasis and delay the progression of LC, suggesting that USF1 overexpression can be used as a biomarker of LC. 35 In addition, studies have shown that USF1 and USF2 mRNA levels are reduced in LUAD compared with non-tumor tissues. 36 SOX6 is related to tumorigenesis and plays a crucial role in cell proliferation and differentiation. Low expression of SOX6 may accelerate the occurrence and progression of tumors. MIR-181b may reduce the proliferation and metastasis of LC cells by targeting SOX6. 37 Evidence revealed that SIRT1 is both a tumor suppressor gene and an oncogene. SIRT1 deacetylates and reduces the activity of the p53 protein. SIRT1 activators (resveratrol and SRT1720) enhance the deacetylation of p53, while SIRT1 inhibitors (tenovin-6 and sirtinol) activate p53 by inhibiting deacetylation. Thus, targeting at SIRT1 may be a good way for regulating the occurrence and development of LC. 38 VHL expression may be an important biological indicator of the occurrence, development, clinical biological behavior, and prognosis of LC. 39 Studies have found that LEF1 overexpression leads to enhanced LC cell proliferation and invasion. 40 EZH2, a key molecule of PRC2 complex, plays an important role in epigenetic regulation and is overexpressed in a variety of tumors. EZH2 inhibitors (GSK343 and DZNep) in combination with gefitinib effectively inhibit LC cell proliferation and migration. 41 HDAC1 is an important epigenetic factor that is thought to be involved in the progression and prognosis of cancer. HDAC1 can be used as a good diagnostic and prognostic marker of LC. 42 Similarly, ESR1, EGR1, YY1, MYC, RELA, NFKB1, and E2F1 are closely related to the occurrence and development of LC. We also found that HDAC1, DNMT1, and E2F1 were significantly regulated factors in patients with LUSC. DNMT1 is a maintenance DNA methyltransferase, and its role in cancer has been widely revealed in recent years. DNMT1 expression was upregulated in NSCLC cell line A549, DNMT1 inhibitor procaine can induce apoptosis of human gastric cancer cells by re-expression of the methylated silenced tumor suppressor gene. 43 These results suggest that finding transcription factor target-related inhibitors or activators of FHIT and its neighboring genes is an important strategy in the treatment of LUAD and LUSC.
We further explored the FHIT-associated kinase targets (PRKCQ, AURKB, and ATM in LUAD, as well as PLK3 in LUSC) and FHIT-associated miRNA targets (MIR-188, MIR-323, and MIR-518A-2 in LUAD). Over the past decade, many studies have shown that protein kinase disorders or mutations play a causal role in cancer occurrence. Cancer research has demonstrated the critical role of many protein kinases in human tumorigenesis and cancer progression, making these molecules effective candidates for new targeted therapies. Kinase inhibitors have revolutionized the treatment of certain types of malignant tumors. Recent studies have found that inhibition of ATM, AURKB, and PLK3 can inhibit the invasion, metastasis, and drug resistance of LC.44–46 However, the role of PRKCQ in LU has not been found. miRNAs are a class of non-coding RNAs that play an essential role in the regulation of gene expression. miRNA inhibitors reduce the expression of target genes by interacting with the 3′ untranslated region of target genes. Their important role in regulating the expression of tumor suppressor genes and oncogene expression enhances their role in tumorigenesis. The discovery of miRNA targets may eventually clarify the mechanisms underlying tumorigenesis and anticancer drug development. Studies have shown that MIR-188 and MIR-323 are involved in regulating the proliferation and apoptosis of LC cells.47,48 However, the relationship between MIR-518A-2 and LC has not been reported. This study provides a reference and highlights the importance of further analyses of regulated factors, kinases, and miRNAs as potential therapeutic targets for LUAD and LUSC treatment.
We explored the correlation between differentially expressed genes and FHIT expression in patients with LUAD and LUSC. More than 19,000 genes were positively and negatively correlated with FHIT expression in patients with LC. Among them, the genes with the highest correlation with FHIT were found in patients with LC: NICN1, HEMK, and BDH2 in LUAD; and ZMAT1, TTC21A, and NICN1 in LUSC. However, targeting these cancer-related genes may provide an adjuvant therapy for LUAD and LUSC. Immunoinfiltration of LC is closely related to clinical prognosis. 49 As expected, we found that FHIT expression in patients with LUAD was positively associated with infiltrations of B cells, and negatively associated with CD8 + T cells and neutrophil infiltrations. Furthermore, the infiltration of B cells, CD8 + T, CD4 + T cells, macrophages, and dendritic cells were positively associated with the expression of FHIT. We hope to improve the immune cell infiltration of LUAD and LUSC by developing drugs acting on FHIT- or FHIT-related regulatory targets.
In conclusion, by exploring the expression levels, gene regulatory networks, prognostic value, and target prediction of FHIT in patients with LUAD and LUSC, we provided more insights into the study and treatment of LC. Furthermore, we identified new therapeutic targets and prognostic biomarkers for the accurate prediction of LC patient survival.
Footnotes
Acknowledgments
This work was supported by the project of financial fund science and technology special competitive allocation of Zhanjiang (Zhanke[2010]174).
Author contributions
Zheng Shao and Qinying Xu performed data analysis work and aided in writing the manuscript. Yongli Situ designed the study and assisted in writing the manuscript. Ruxiu Gao, Lei Lei, Li Deng edited the manuscript. All authors read and approved the final manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the the project of financial fund science and technology special competitive allocation of Zhanjiang, (grant number Zhanke[2010]174).
Supplementary material
The Supplementary material for this article can be found online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
