Abstract
Background
Circular RNA hsa_circ_0002346 has been implicated in the progression of various tumors, yet the functional role in breast cancer remains poorly understood. This study aimed to investigate the significance of hsa_circ_0002346 in breast cancer (BC).
Methods
Quantitative reverse transcriptase polymerase chain reaction assays were performed to detect hsa_circ_0002346 expression in BC cell lins and 27 patients with BC. Then, siRNAs were used to knock down hsa_circ_0002346. And detecting function of downregulated hsa_circ_0002346 by proliferation colony formation, apoptosis assays, wound-healing assays and Transwell assays. Finally, we assess the levels of the EMT-associated proteins.
Results
Our findings suggest that hsa_circ_0002346 levels were significantly downregulated in breast cancer and may play a crucial role in regulating key cellular processes associated with cancer progression. The expression level of hsa_circ_0002346 was correlated with lymph node metastasis. The knockdown of hsa_circ_0002346 resulted in increased tumor cell proliferation, invasion, migration and decreased apoptosis. Additionally, alterations in the expression of EMT-associated proteins further support the hypothesis that hsa_circ_0002346 is implicated in the metastatic processes of breast cancer.
Conclusion
hsa_circ_0002346 emerges as a promising biomarker for breast cancer and a potential therapeutic target for future treatment strategies.
Introduction
Breast cancer (BC), according to the International Agency for Research on Cancer, is now the most prevalent malignancy in the world, having overtaken lung cancer. Despite recent advances in BC treatment, tumor invasion and metastasis are still the main causes of death in patients with BC. Therefore, it is necessary to identify new biomarkers to further our understanding of BC progression and overcome poor prognosis.
Circular RNAs (circRNAs) lack coding ability and have attracted much recent attention. CircRNAs are highly stable due to their covalent closed-loop structures. 1 There is growing evidence for the involvement of circRNAs in a variety of biological processes, including cancer progression, and they may also sponge microRNAs (miRNAs), modulating gene expression. In breast cancers, the circRNAs circGNB1, circKIF4A, circTADA2As, and circRAD18 have been observed to sponge miRNAs to modulate the progression of the disease.2–5 circRNA-UBAP2 has been found to regulate ovarian carcinoma through the miR-382-5p/ pre-mRNA-processing-splicing factor 8 (PRPF8) axis, 6 while circNDUFB2 modulates protein ubiquitination, degradation, and cellular immune responses via destabilization of insulin-like growth factor-2 mRNA-binding proteins (IGF2BPs) to inhibit the progression of non-small cell lung cancer. 7 However, there are few studies on the circRNAs associated with BC.
Hsa_circ_0002346 has been observed to reduce invasion and metastasis in lung adenocarcinoma cells and has been suggested to be a promising biomarker in non-small cell lung cancer8,9 However, the role of hsa_circ_0002346 in BC remains largely unknown. The findings indicated that hsa_circ_0002346 has significant associations with the proliferation, migration, invasion and apoptosis of BC cells, suggesting that it could be served as a potential biomarker for BC progression.
Materials and methods
CircRNA identification
The reported involvement of hsa_circ_0002346 in several cancer types led to the speculation that it may also affect BC progression. KangChen Bio-tech completed the Arraystar Human circRNA Array V2 analysis of the samples that we submitted. Total RNA from each sample was quantified using the NanoDrop ND-1000. The sample preparation and microarray hybridization were performed based on the Arraystar's standard protocols. Briefly, total RNAs were digested with Rnase R (Epicentre, Inc.) to remove linear RNAs and enrich circular RNAs. Then, the enriched circular RNAs were amplified and transcribed into fluorescent cRNA utilizing a random priming method (Arraystar Super RNA Labeling Kit; Arraystar). The labeled cRNAs were hybridized onto the Arraystar Human circRNA Array V2 (8 × 15 K, Arraystar). After having washed the slides, the arrays were scanned by the Agilent Scanner G2505C. Small-sample size CircRNA microarray screening showed that the expression of hsa_circ_0002346 in breast cancer tissues was significantly lower than that in normal tissues. In addition, the levels of hsa_circ_0002346 were initially evaluated in BC cells, and the results indicated significant differential expression between BC and normal breast epithelial cells. Thus, hsa_circ_0002346 was used for further investigation.
Cell culture
Six human cell lines (BT -549, MDA-MB-231, SKBR-3, MCF-7, BT-474, and MCF-10A) were used. The BT-549 and MDA-MB-231 cell lines are derived from triple-negative BC tissues, while BT-474 represents luminal-B BC, SKBR-3 HER-2 BC, and MCF-7 luminal-A type BC. These cell lines encompass all subtypes of breast cancer cells. The SKBR-3, BT-549, and MDA-MB-231 lines, as well as the normal mammary epithelial cell line MCF-10A were purchased from MICRO-G BIOTECH, (Zhejiang, China), MCF-7 and BT-474 cells were purchased from Taizhou Hospital Enze Laboratory (Zhejiang, China). MDA-MB-231, SKBR-3, MCF-7, and BT-474 cells were grown in DMEM, Gibco, Waltham, MA, USA) and BT-549 cells in RPMI 1640 medium. All media contained 10% fetal bovine serum, 100 U/mL penicillin, and 100 mg/mL streptomycin. MCF-10A cells were cultured in MEpiCM (ScienCell, USA). Cells were grown in a humidified incubator at 37 °C with 5% CO2.
Specimens
All specimens were obtained from Shaoxing People's Hospital between April 2022 and June 2023. The tissues were immediately frozen in liquid nitrogen after the surgical excision. A total of 27 pairs of tissue specimens, all from female patients, were collected. The median age of the patients was 59 years, with an age range from 27 to 77 years. None of the patients had received prior radiation therapy, chemotherapy, immunotherapy, or targeted therapy. The histopathological diagnosis of the tumor was determined by the consensus of two experienced pathologists. This study was approved by the Ethics Committee of Shaoxing People's Hospital, and informed consent was obtained from all patients.
qRT-PCR
All primers used in this study were purchased from RiboBio (Guangzhou, China) and their sequences were as follows: hsa_circ_0002346 forward 5′-AACACCATTCGAACCTGCAG-3′ and reverse 5′-TTGGCAAAGTACAGCAACCA-3′; GAPDH forward 5′-GAACGGGAAGCTCACTGG-3′ and reverse 5′-GCCTGCTTCACCACCTTCT-3′. Total RNA was extracted from tissue samples and cultured breast cells with the RNA-Quick Purification Kit (Yishan Biotechnology, Shanghai, China). The Ribo qRT-PCR Starter Kit (Ribo, China) was used for qRT-PCR. The final reaction volume was 20 µl, including 2 µl cDNA, 2 µl each primer, 10 µl SYBR Green, and 4 µl ddH20. Thermal cycling was as follows: 95°C for 5 min, then 40 cycles at 95°C for 5 s and 60°C for 1 min. Data were analyzed using the 2-ΔΔCq method. 10
CCK-8 assay
Cells (2X103 per well) were plated in 96-well plates and 10 µl CCK-8 solution (Beyotime, China) was added to each well and incubated for 2 h at 37 °C. Absorbances at 450 nm were read in a microplate reader. Experiments were performed in triplicate.
Cell transfection
The cells were cultured in 6-well plates. Five groups of cells were set up, namely, si 1#, si 2#, si 3#, si-NC, and Mock. Each group contained three replicates with consistent numbers of cells in all. Transfection was performed when the cells were approximately 50% confluent. Three siRNAs purchased from RiboBio (Guangzhou, China) were designed to knock down hsa_circ_0002346 expression. Their sequences were as follows: si-hsa_circ_0002346-1# 5′-TTTGCCAACAAGATGAGAA-3′; si-hsa_circ_0002346-2# 5′-TGCCAACAAGATGAGAACT-3′; si-hsa_circ_0002346-3# 5′-CAACAAGATGAGAACTGGA-3′. MDA-MB-231 and BT-549 cells were transfected with 50 nM siRNAs or negative control (si-NC) with siRNA Transfection Kit from RiboBio (Guangzhou, China). After incubating at 37°C for 48 h, the knockdown of circRNA was confirmed via qRT-PCR. Follow-up experiments were performed 24 h after transfection.
Wound healing
Forty-eight hours after transfection, cells were plated in 6-well plates and a scratch was made with a pipette tip (200 µl) in the center of the cell monolayer. The plates were washed twice in PBS before adding serum-free medium and measuring the wound widths at 0 and 24 h. The wounds were examined and imaged by microscopy and the images were analyzed using ImageJ software (NIH, USA).
Colony formation
BC cells (2X103 per well) were seeded in 6-well plates and incubated at 37 °C for 1 week. The cells were then fixed with 4% paraformaldehyde, stained with 0.1% crystal violet, and photographed with a camera.
Transwell assays
Transfected cells (1 × 104) in 200 μL medium with 10% FBS were added to the Matrigel-coated upper chamber while the lower chamber contained 600 µl medium with 20% FBS. After 24 h, the migrated cells were fixed with 4% paraformaldehyde, stained in 0.1% crystal violet, and counted and imaged.
Measurement of apoptosis
Apoptosis was measured using an Annexin V-FITC cell apoptosis kit (Beyotime, China) and cells were examined by fluorescence microscopy and flow cytometry.
Western blotting
Total protein was extracted in RIPA lysis buffer (Beyotime, China), separated on SDS-PAGE, and electroblotted onto PVDF membranes (Millipore, USA). After blocking (room temperature, 2 h) using Western confining liquid (Biosharp, USA), the blots were incubated with primary antibodies anti-Vimentin (1:1000, #ab92547, Abcam, Cambridge, UK), anti-GAPDH (1:1000, #ab8245, Abcam, Cambridge, UK), anti-MMP2 (1:1000, #ab92536, Abcam, Cambridge, UK), anti-MMP9(1:1000, #ab76003, Abcam, Cambridge, UK) and a secondary antibody (Abcam, Cambridge, UK), and visualized by multifunctional gel-imaging.
Statistical Analysis
SPSS 25.0 and GraphPad Prism 8.0 statistical software were used for analysis of the experimental results. Quantitative data are presented as mean ± standard deviation. Normally distributed data were analyzed with t-tests for comparisons between two groups and one-way ANOVA for comparisons between multiple groups. Dunnett post hoc test was applied after ANOVA. P < 0.05 (*) was considered statistically significant; *: P < 0.05, ** : P < 0.01, *** : P < 0.001, **** : P < 0.0001, ns: P > 0.05.
Results
hsa_circ_0002346 levels are reduced in BC cell lines
qRT-PCR was used to measure hsa_circ_0002346 expression in BC cell lines. The results showed that hsa_circ_0002346 levels were significantly downregulated in BC cell lines, compared with normal mammary epithelial cells, particularly in the MDA-MB-231 and BT-549 (Figure 1(a)).
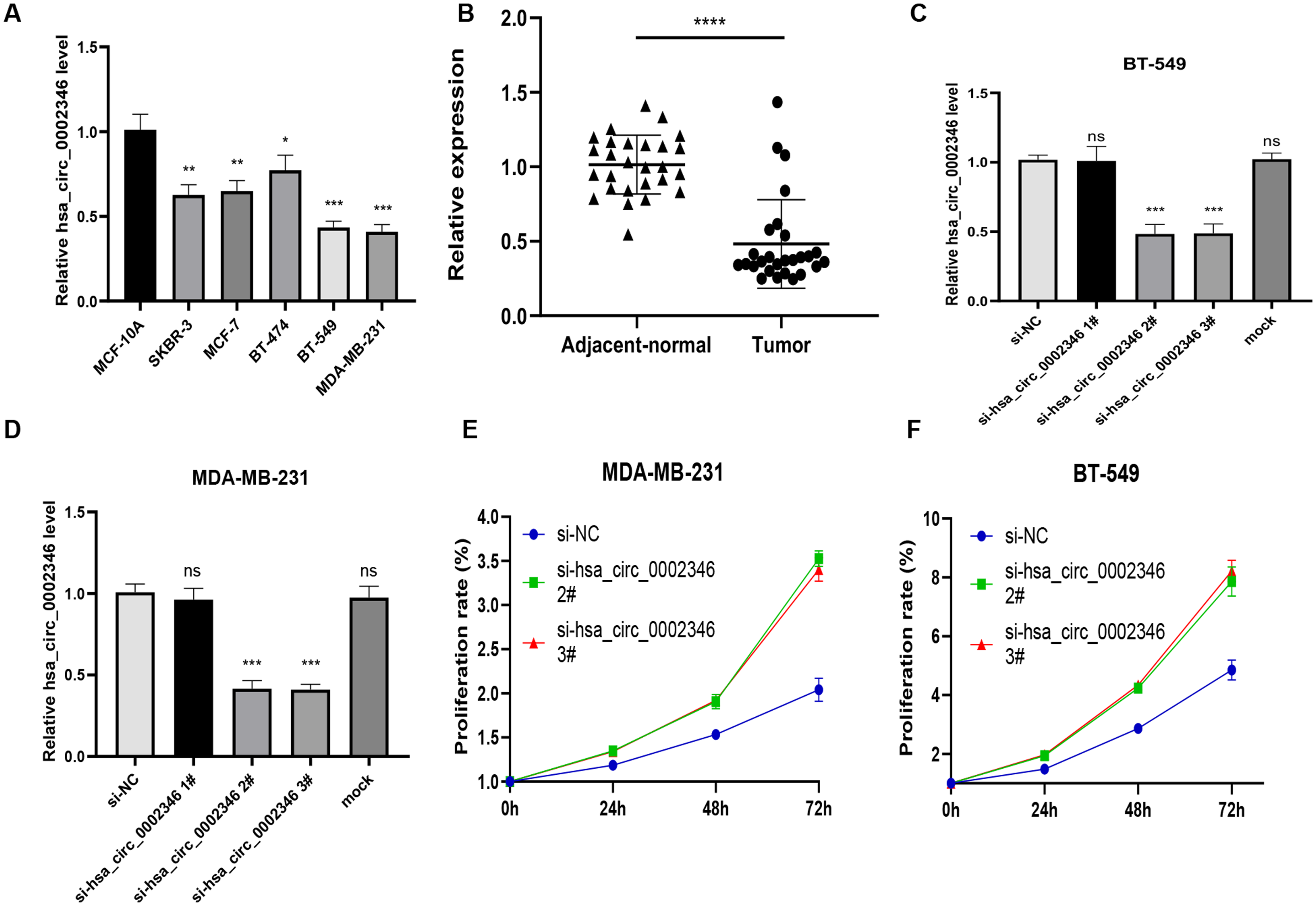
(a) Expression of hsa_circ_0002346 in the normal mammary cell line MCF-10A and breast cancer cell lines; (b) Expression of hsa_circ_0002346 in tumor and adjacent normal tissues. (c)-(d) Reduction of hsa_circ_0002346 expression after knockdown; (e) Expression of hsa_circ_0002346 in tumor and adjacent normal tissues. (e)-(f) Cell viability measured by CCK-8 assays in MDA-MB-231 and BT-549 cells transfected with control or siRNAs; (g) Quantification of apoptosis in treated MDA-MB-231 and BT-549 cells by flow cytometry. T-tests and one-way ANOVA were used for statistical analysis. *P < 0.05, **P < 0.01, ***P < 0.001, nspd > 0.05.
hsa_circ_0002346 levels are reduced in BC tissues
The expression of hsa_circ_0002346 in the tissue samples was determined by qRT-PCR. The results revealed a significant downregulation of hsa_circ_0002346 expression in BC tissues compared with the matched adjacent non-cancerous tissues (Figure 1(b)). The expression pattern of hsa_circ_0002346 in BC tissues was consistent with that in the cell lines. Therefore, based on these findings, we conducted an in-depth exploration of its relationship with BC progression.
hsa_circ_0002346 levels are associated with lymph node metastasis status in patients
In our investigation of the relationship between hsa_circ_0002346 and the clinicopathological data of the patients, we observed notable differences in hsa_circ_0002346 expression levels between patients with lymph node metastasis and those without lymph node metastasis, as presented in Table 1. The expression level of hsa_circ_0002346 showed no statistically significant differences across different ages of patients, tumor sizes, WHO grades, or TNM stages.
Associations between hsa_circ_0002346 expression levels and clinical pathological characteristics of patients with BC.

ΔΔCT = ΔCT(hsa_circ_0002346) -ΔCT(GAPDH); TNM, Tumor Node Metastasis. WHO, World Health Organization.
Downregulation of hsa_circ_0002346 promotes proliferation and colony formation while suppressing apoptosis in BC cells
The expression level of hsa_circ_0002346 in BT-549 and MDA-MB-231 cells was significantly different from that in the control group. These two types of cells are the most representative and have certain persuasiveness for the experimental results, so we chose these two cell lines to study the effect of hsa_circ_0002346. Three siRNAs targeting hsa_circ_0002346 (si-hsa_circ_0002346#1 si-hsa_circ_0002346#2, and si-hsa_circ_0002346#3) were used to knock down hsa_circ_0002346, with their effectiveness assessed by qRT-PCR.
Although si-hsa_circ_0002346#1 had little effect on hsa_circ_0002346 levels, the other two si-RNAs were more effective, as seen in marked reductions in expression of hsa_circ_0002346 (Figure 1(c) and (d)). Thus, these two siRNAs were chosen for subsequent experiments. CCK-8 assays showed that the transfected cells proliferated faster than those in the control groups, indicating that reduced levels of hsa_circ_0002346 promoted proliferation in BC cells (Figure 1(e) and (f)). Then we then determined the effects of hsa_circ_0002346 silencing on apoptosis. MDA-MB-231 cells were transfected for 72 h before staining with Annexin V-FITC/propidium iodide. Downregulation of hsa_circ_0002346 was found to inhibit apoptosis (Figure 2(a)–(c)). Moreover, colony formation assays showed that the transfected cells formed greater numbers of visible colonies, indicating that hsa_circ_0002346 reduction stimulated colony formation (Figure 2(d)–(g)). These findings demonstrated that hsa_circ_0002346 inhibited both proliferation and colony formation while promoting apoptosis in BC cells.
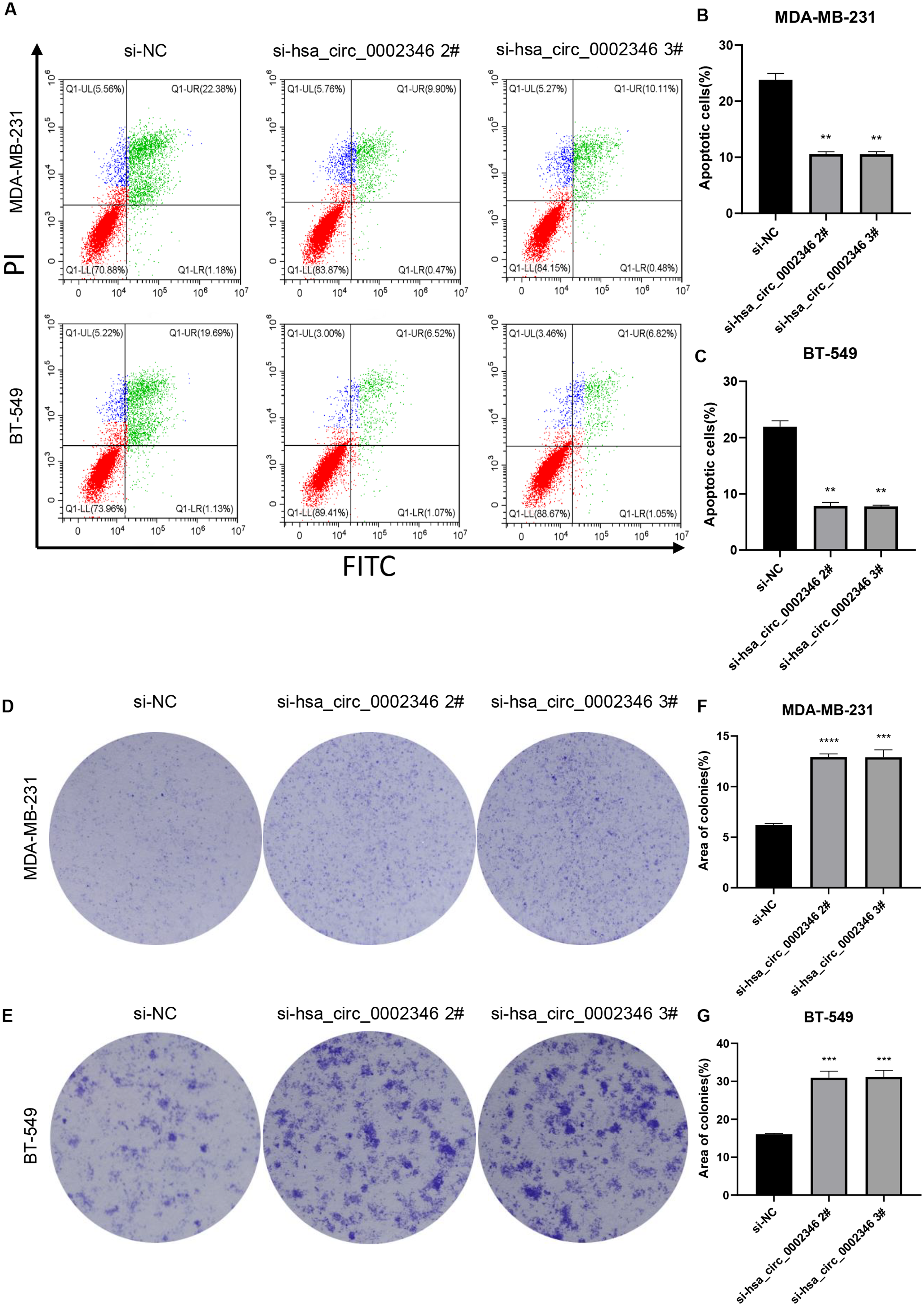
(a)-(b) Colony formation in MDA-MB-231, and BT-549 cells transfected with control or hsa_circ_0002346 siRNAs, shown by the colony-formation assays; (c)-(d) Quantification of colony areas in BC cell lines. one-way ANOVA were used for statistical analysis.*P < 0.05, **P < 0.01, ***P < 0.001.
Silencing of hsa_circ_0002346 promotes BC migration and invasion
The wound-healing assays demonstrated that, compared with non-transfected cells, the wounds in the transfected cells healed much faster, indicative of increased migratory ability (Figure 3(a)–(d)). Similarly, the Transwell assay showed that migration was enhanced in the silenced cells (Figure 3(e)–(g)) while hsa_circ_0002346 knockdown enhanced invasion in both BC cell lines (Figure 3(h)–(j)). As the epithelial-mesenchymal transition (EMT) was linked to cancer cell metastasis, western blotting was used to assess the levels of the EMT-associated proteins vimentin, N-cadherin, MMP-2, and MMP-9, observing increased levels of vimentin, MMP-2, and, especially, MMP-9 in the silenced cells (Figure 4(a)–(c)). These findings indicated that reduced hsa_circ_0002346 expression promoted migration and invasion, possibly related to the activation of the EMT in BC cells.
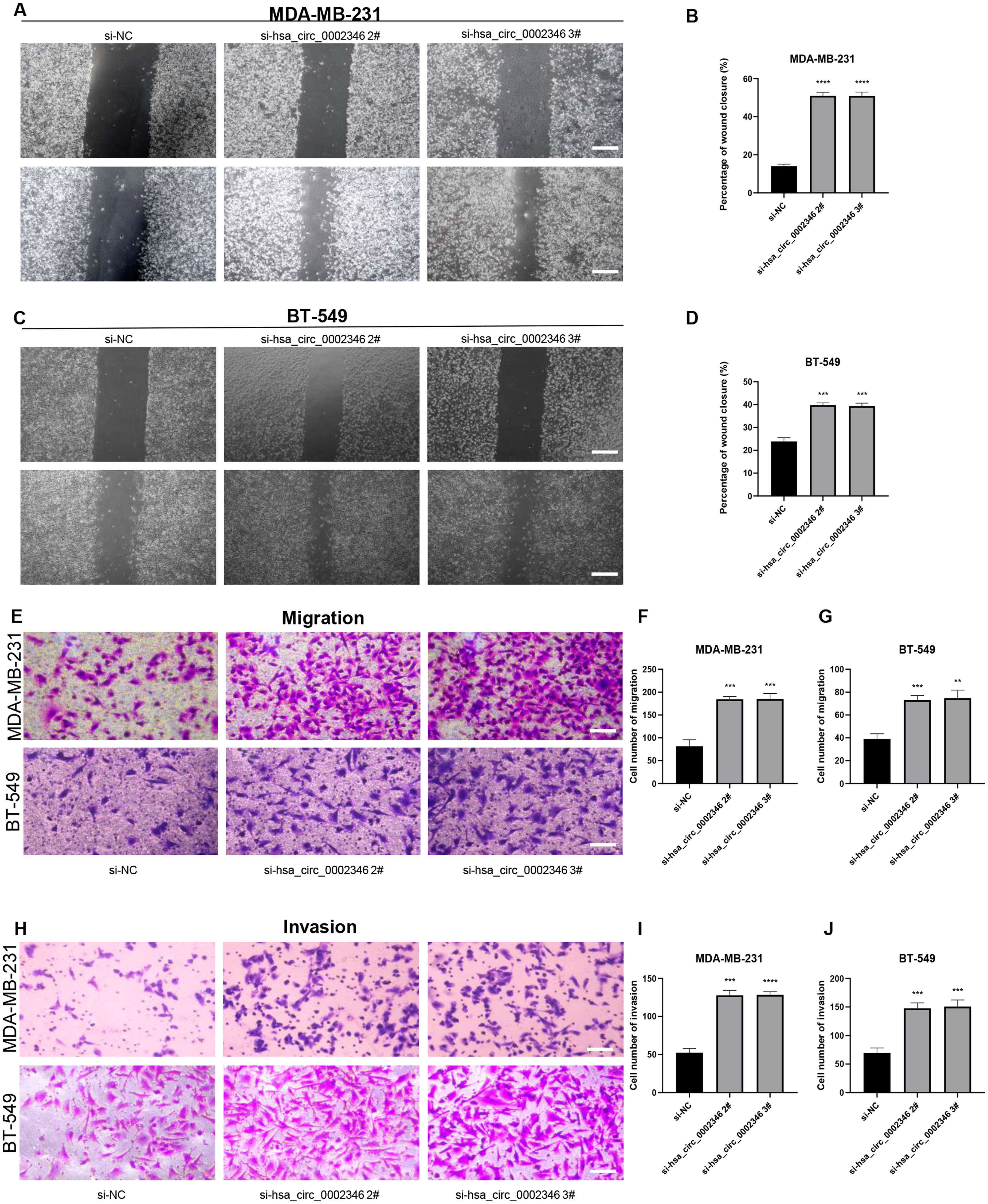
(a)-(d) The effects of hsa_circ_0002346 on cell migration, measured by wound-healing assays. Scale bar, 200 μm; (e)-(g) The effects of hsa_circ_0002346 on cell migration, measured by Transwell assays. Scale bar, 100 μm. (h)-(j) The effects of hsa_circ_0002346 on cell invasion, measured by Transwell assays. Scale bar, 100 μm. one-way ANOVA were used for statistical analysis. one-way ANOVA were used for statistical analysis.*P < 0.05, **P < 0.01, ***P < 0.001.

(a) Expression of EMT-related proteins, as shown by western blotting; (b)-(g) Quantification of the expression of EMT-related proteins. one-way ANOVA were used for statistical analysis.*P < 0.05, **P < 0.01, ***P < 0.001.
Discussion
Recent advances in high-throughput techniques and bioinformatics have led to an increased understanding of the significant roles of circRNAs in tumor processes. This is especially relevant in relation to the functions of competing endogenous RNAs (ceRNAs). 11 Specifically, circRNAs have been found to function as oncogenes in BC when upregulated, for example, circ-TFCP2L1, 12 circANKS1B, 13 circ-UBAP2, 14 and circGFRA1, 15 while others, including circFBXW7, 16 circTADA2A-E6, 4 and circAHNAK1, 17 have anti-cancer effects.
Wang. et al proposed hsa_circ_0002346 as a potential therapeutic target in lung adenocarcinoma. 8 In our study, hsa_circ_0002346 levels were reduced in BC cell lines. In the cell lines examined, there appeared to be inherent differences among BC subtypes. However, there was only one cell line of each subtype (except TNBC cell lines), it does not meet the statistical criteria to perfectly explain the differences in expression between different subtypes. Perhaps in the future, by assessing the expression levels of hsa_circ_0002346 across a greater number of breast cancer cell subtypes, a more refined explanation for this phenomenon could be provided. Additionally, our data indicated that the expression level of hsa_circ_0002346 correlated with the lymph node metastasis status of the patients. This suggested that hsa_circ_0002346 may also possess significant clinical predictive value in the future. However, analysis of clinical pathological data from patients revealed that the expression level of hsa_circ_0002346 did not exhibit significant correlations with tumor size greater than 2 cm, WHO grading, or TNM staging. This could potentially be attributed to the limited sample size analyzed and the variability in breast cancer types among different patients, leading to data biases. In future studies, we plan to increase our sample size, with a specific focus on diverse subtypes of breast cancer, to discern any variations in hsa_circ_0002346 expression levels across different patient groups. Furthermore, we observed that when hsa_circ_0002346 expression level changed, the proliferation, migration, invasion and apoptosis of BC cells changed as well. Interestingly, the knockdown of hsa_circ_0002346 significantly promoted both cancer invasion and migration, accompanied by altered expression levels of EMT-related proteins, particularly MMP9. MMP-9 is known to be involved in metastasis and angiogenesis through its ability to degrade the extracellular matrix (ECM), MMPs, and plasminogen to induce the EMT. 18 MMP-9 is able to promote carcinogenesis by binding to the surfaces of cancer cells. 19 In addition, it has also been associated with unfavorable prognosis in BC. 20 This provides a new direction for exploring the EMT-related molecular signaling pathways that Circular RNA hsa_circ_0002346 affected in the future.
Here, we found that has_circ_0002346 levels were reduced in BC. Hsa_circ_0002346 was found to reduce the proliferation, invasion, and migration of BC cells while promoting apoptosis. The mechanisms responsible are currently being investigated by our group. We will address the underlying molecular mechanisms in the future studies using overexpression, analysis of clinical samples, and in vivo animal experiments. Our study postulates that hsa_circ_0002346 may modulate the functional role of BC cells in regulating cancer progression by regulation of the EMT. However, further experimental investigations are required to substantiate this hypothesis.
Footnotes
Acknowledgements
The present study was financially supported by National Natural Science Foundation of China (grant no. 82173080) and Provincial Natural Science Foundation of Zhejiang (grant no. GF21H160088).
Author contributions
Conception: Feilin Cao and Liming Huang. Interpretation or analysis of data:Yixin Zhou. Preparation of the manuscript: Yixin Zhou, Linyi Wang and Wangjin Xu. Revision for important intellectual content:Jiaxiang An and Guoming Hu. Supervision: Feilin Cao and Liming Huang.
Funding
The present study was financially supported by National Natural Science Foundation of China (grant no. 82173080) and Provincial Natural Science Foundation of Zhejiang (grant no. GF21H160088).
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
