Abstract
Background:
Cathepsins are pivotal regulators of critical physiological processes implicated in cancer, rheumatic disorders, and inflammatory conditions.
Objectives:
This study employed Mendelian randomization (MR) to evaluate causal relationships between cathepsins and ankylosing spondylitis (AS).
Design:
A retrospective study.
Methods:
Single nucleotide polymorphism data of cathepsins were obtained from the INTERVAL study, and AS data were obtained from the FinnGen database. Inverse variance weighting was used as the primary method to assess the causal relationship described above. Cochran’s Q test, MR Egger intercept test, MR-PRESSO, and leave-one-out method were used to analyze study sensitivity, heterogeneity, and pleiotropy.
Results:
In the forward MR analysis, inverse variance weighted results indicated that higher cathepsin S might be associated with an increased risk of AS (inverse-variance weighting, odds ratio = 1.08, 95% confidence interval = 1.00–1.16, p = 0.047). MR-Egger intercept test and Cochran’s Q test did not detect significant heterogeneity or horizontal pleiotropy of instrumental variables. The leave-one-out method confirmed the reliability of causality. The reverse MR analysis found no significant causal relationship between cathepsins and AS.
Conclusion:
Our MR analysis results suggest a potential causal relationship between cathepsin S and AS. Further studies on the pathogenesis of cathepsin-mediated AS may provide new insights into the prevention and treatment of AS.
Plain language summary
This study employed Mendelian randomization to investigate the causal relationship between cathepsins and ankylosing spondylitis. Our analysis suggests that genetically determined higher levels of cathepsin S may modestly increase the risk of developing ankylosing spondylitis. The reliability of this finding is supported by a series of subsequent rigorous sensitivity analyses, and reverse causality was ruled out by reverse MR analysis. It should be noted that the observed association is weak and requires further confirmation in future studies. If validated, this finding would provide new insights into the etiology of the disease and may open avenues for novel preventive and therapeutic strategies.
Keywords
Introduction
Rheumatic diseases are a group of inflammatory and autoimmune disorders that can affect any part of the body across all age groups. These conditions most commonly target joints, ligaments, bones, and muscles. Ankylosing spondylitis (AS), a chronic inflammatory and autoimmune disease primarily affecting the spinal column, is characterized by chronic back pain and progressive spinal stiffness. 1 Typically presenting with early onset and slow progression, AS often leads to delayed clinical diagnosis, sometimes by several years. The etiology of AS remains complex and poorly understood. While HLA-B27 genetic susceptibility and Th17-related inflammatory signaling pathways are known contributors, current research emphasizes the role of autoimmune responses and bone stromal cells in its pathogenesis. 2 HLA-B27 is a highly sensitive genetic marker; however, its utility is constrained by differences in ethnicity, geography, and other population-related factors, which limit its application in predicting disease severity and prognosis.3,4 Consequently, identifying novel biomarkers or therapeutic targets for AS holds significant promise for improving disease management and treatment outcomes.
Cathepsin is among the most critical proteolytic enzymes in mammals. Under physiological conditions, cathepsin primarily localizes to lysosomes and is released into the cytoplasm upon specific signaling stimuli. To date, 15 categories of cathepsins have been identified in humans, classified into serine proteases (CTSA, CTSG), aspartic proteases (CTSD, CTSE), and cysteine proteases (CatB, C, F, H, K, L, O, S, V, W, and Z) according to their catalytic mechanisms. 5 Cysteine proteases constitute the predominant subclass within the cathepsin family. Cathepsins participate in diverse physiological and pathological processes, including autophagy regulation, apoptosis modulation, antigen presentation, extracellular matrix (ECM) homeostasis maintenance, infection response, and inflammatory pathways.6–8 Elevated expression levels and enhanced enzymatic activity of cathepsins are consistently observed during disease pathogenesis and progression, garnering increasing attention for their diagnostic and therapeutic potential.
In recent years, advances in understanding cathepsin in both physiological and pathological contexts have highlighted their growing significance in AS. Cathepsin K plays a critical role in bone resorption, and assessment of its active form may provide essential biological insights for metabolic bone disorders such as AS. 9 Neidhart et al. 10 demonstrated marked overexpression of cathepsin K in distinct spinal regions of AS patients, indicating persistent bone-destructive activity in chronic AS. A study by Skjøt-Arkil et al. 11 demonstrated that the cathepsin-mediated degradation products of C-reactive protein can more effectively reflect the inflammatory severity in patients with AS than other conventional inflammatory markers. However, the functional contributions of individual cathepsin in rheumatic diseases may differ, and specific proteases remain uncharacterized in AS. Further investigation is therefore warranted to elucidate causal relationships between specific cathepsin subtypes and AS pathogenesis.
Advances in high-throughput sequencing, proteomics, and metabolomics have provided increasing evidence for the involvement of genetic factors in rheumatic diseases. 12 Mendelian randomization (MR) employs genetic variants as instrumental variables (IVs), utilizing genome-wide association study (GWAS) data to infer causal relationships between exposures and outcomes. This study leveraged GWAS summary statistics for nine cathepsins and AS to conduct two-sample MR analyses, followed by reverse MR to account for the pleiotropic effects of genetic instruments and potential confounders. These analyses aimed to evaluate putative genetic causal associations between cathepsins and AS, offering theoretical foundations for AS prevention and therapeutic strategies.
Methods
Study design
Using two-sample MR, we analyzed pooled summary statistics from publicly available GWAS to evaluate potential causal relationships between cathepsins and AS. This study first assessed whether cathepsins influence AS incidence by designating cathepsins as exposure variables and AS as the outcome variable. MR analyses must satisfy three core assumptions: (1) Relevance assumption: genetic instruments must exhibit significant associations with exposure factors; (2) Independence assumption: genetic variants must remain independent of confounders affecting both exposures and outcomes; (3) Exclusion restriction assumption: genetic instruments influence outcomes exclusively through the specified exposure, without alternative causal pathways (i.e., absence of horizontal pleiotropy). 13 Figure 1 provided a schematic representation of these assumptions. This study was reported in accordance with the Strengthening the Reporting of Observational Studies in Epidemiology for Mendelian Randomization (STROBE-MR) statement. 14
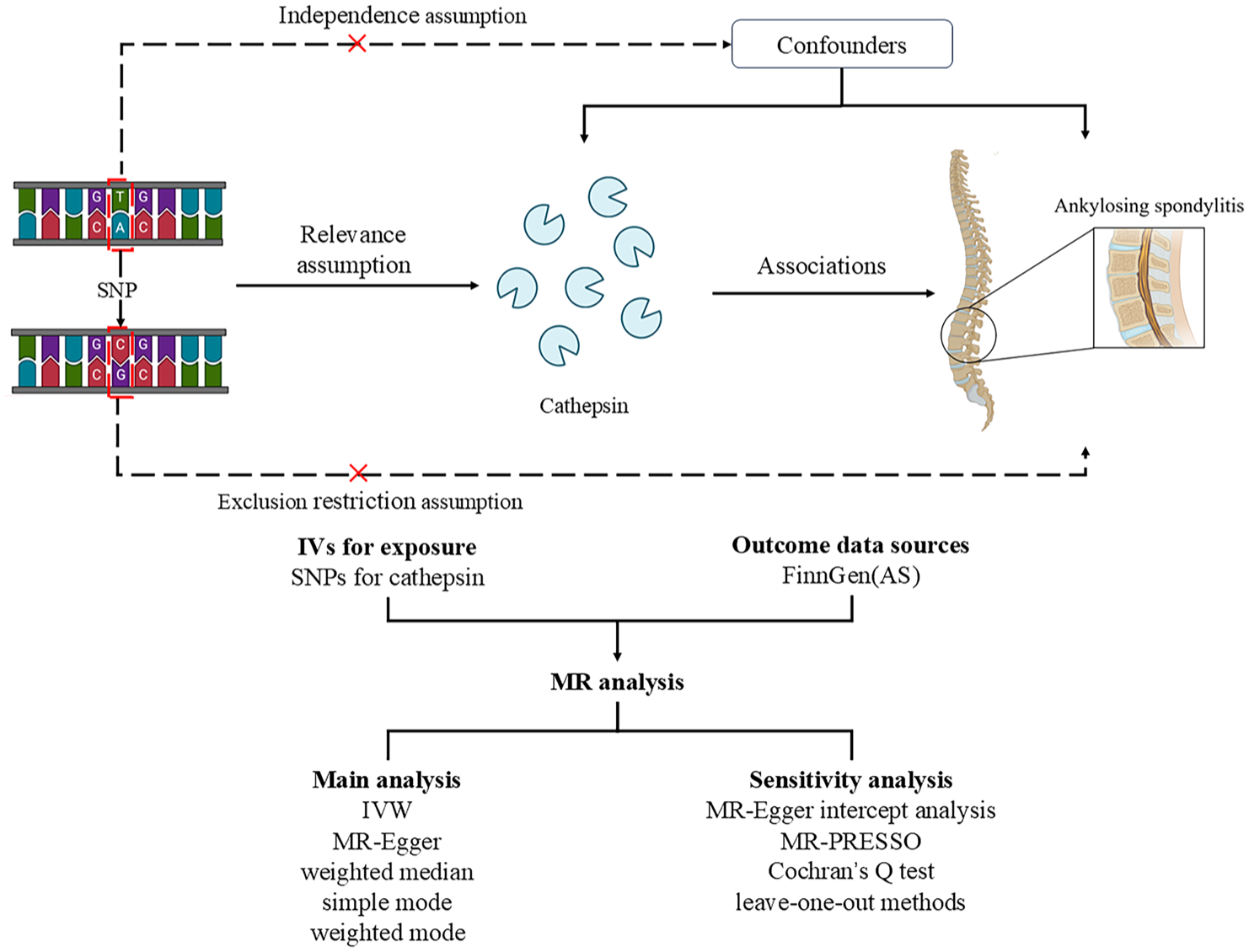
The design and analysis process of MR.
Exposure data sources
Cathepsin-related genetic IVs were derived from the INTERVAL study (https://gwas.mrcieu.ac.uk), focusing on a European ancestry subset comprising 3301 individuals. 15
Outcome data sources
AS summary statistics were obtained from the FinnGen Consortium R9 version (https://r9.risteys.finngen.fi/), including 2860 AS cases.
Instrument variables selection
Single nucleotide polymorphisms (SNPs) with genomewide significant association strength with cathepsins were selected as IVs, and the following inclusion criteria were established: (1) SNPs highly associated with exposure factors were included in the analysis, and a significance threshold of p < 1 × 10−5 was adopted.16,17 This threshold aimed to increase the availability of analytical tools, enhance statistical power, and facilitate comprehensive causal exploration; (2) To reduce the influence of linkage disequilibrium, independence between IVs was ensured, defined as a linkage distance of 10,000 kb and R2 < 0.001 18 ; (3) Use the PhenoScanner database (http://www.phenoscanner.medschl.cam.ac.uk/phenoscanner) to detect and eliminate confounding factors related to IV to meet the MR independence assumption for the analysis of principle. (4) Strength was evaluated via the F-statistic (F > 10 indicated minimal weak instrument bias), calculated as F = [(NK − 1)/K]×[R2/(1 − R2)], where N is the number of protease GWAS samples, K is the number of SNPs, and R2 is the variance of each SNP explaining the exposure. The specific R2 was calculated as follows: R2 = 2 × β 2 × (1 − EAF) × EAF, where β 2 is the estimated genetic effect of the SNP on protease, and EAF is the frequency of the effect allele. 19
Statistical analysis
A two-sample MR approach was employed to assess causal relationships between cathepsins and AS risk using five methods: inverse-variance weighting (IVW), 20 MR-Egger, 21 weighted median, 22 simple mode, 23 and weighted mode. 24 Odds ratios (ORs) with 95% confidence intervals (CIs) were calculated to quantify causal effects.
IVW is the main method for estimating the overall effect size, weighted according to the effect of each variant on the risk of the disease under study, and finally, a random effects inverse variance meta-analysis is used to obtain the overall value. MR-Egger intercept analysis and MR-PRESSO were used to detect horizontal pleiotropy. MR-Egger intercept analysis was used to assess the relationship between IV and other potential confounders, and a statistically insignificant difference would indicate the absence of horizontal pleiotropy. If horizontal pleiotropy was suggested by MR-Egger intercept analysis (p < 0.05), horizontal pleiotropy was corrected with the MR-PRESSO outlier test by deleting or decreasing the outlier value. 15 ORs < 1 suggested protective effects of exposure on outcomes, while ORs > 1 indicated risk-enhancing effects.
Heterogeneity was assessed using Cochran’s Q statistic, with p < 0.05 denoting significant heterogeneity.25 Sensitivity analyses via leave-one-out methods evaluated whether individual SNPs disproportionately influenced the outcome. False alarm rates were assessed using false discovery rate (FDR) correction. A suggestive association between cathepsins and AS was considered to exist when p < 0.05 and q < 0.1.
Reverse MR analysis investigated the potential causal effects of AS on cathepsins identified in forward MR analyses, following identical analytical procedures. All analyses were implemented in R software (v4.3.1) using the TwoSampleMR and MRPRESSO packages.
Results
Genetic causality of cathepsins and AS
In accordance with the screening criteria, our study incorporated nine cathepsins (cathepsin B, E, F, G, H, O, S, L2, and Z) as IVs. All IVs demonstrated F-statistic values exceeding 10, effectively mitigating weak instrument bias. Detailed characteristics of the selected SNPs are provided in Table S1.
MR analyses were performed to assess the relationship between each cathepsin and AS, with the IVW method serving as the primary analytical approach. As shown in Table 1, cathepsin S (OR = 1.08, 95% confidence interval (CI) = 1.00–1.16, p = 0.047, IVW) exhibited a potential causal association with AS, suggesting that elevated cathepsin S levels may contribute to AS pathogenesis. When the FDR adjustment was carried out, however, these correlations failed to be significant (q > 0.1). Specifically, an OR greater than 1 indicates that higher cathepsin S levels are associated with an increased risk of AS. The scatterplot in Figure 2(A), depicting the effect distribution of individual SNPs across the five MR methods, demonstrated consistent directional estimates with slopes greater than 1, further supporting the positive association between elevated cathepsin S levels and AS risk.
Full result of MR estimates for the association between cathepsin and AS.

AS, ankylosing spondylitis; CI, confidence interval; MR, Mendelian randomization; OR, odds ratio; SNP, Single nucleotide polymorphism.
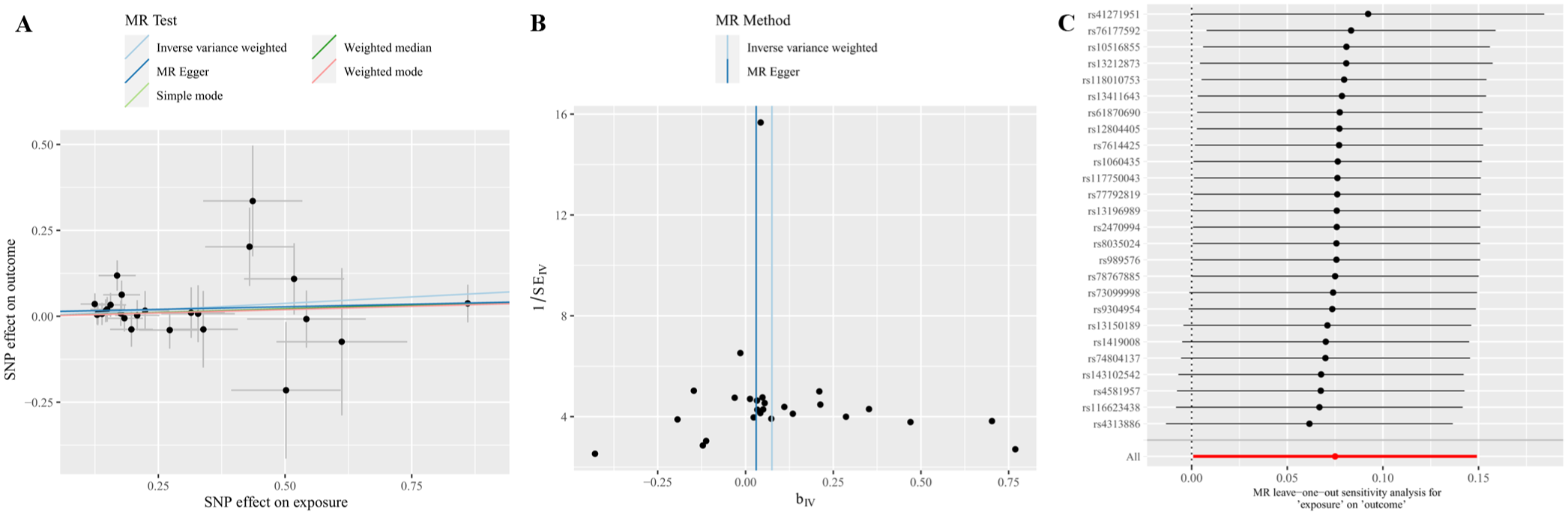
Causal effects of cathepsin S on AS. (A) Scatter plot of the causal relationship between 26 SNPs of cathepsin S and AS by five Mendelian randomization methods (the slope of the line shows the causal association of each method, with a positive slope indicating that cathepsin S increases the risk of AS). (B) MR analysis of the effect of 26 SNPs of cathepsin S on AS Funnel plot. (Each point denotes an individual SNP’s IV estimate (bIV) against its standard error (SEIV). Lines indicate MR Fit results from inverse variance weighting and MR Egger regression analysis.) (c) Leave-one-out sensitivity analysis for the causal relationship between cathepsin S and AS. (Red dots depict pooled effect estimates from all 26 SNPs; black dots show estimates after sequentially excluding individual SNPs. Error bars clustered to the right of the null line (0) confirm the result robustness.)
Sensitivity analyses
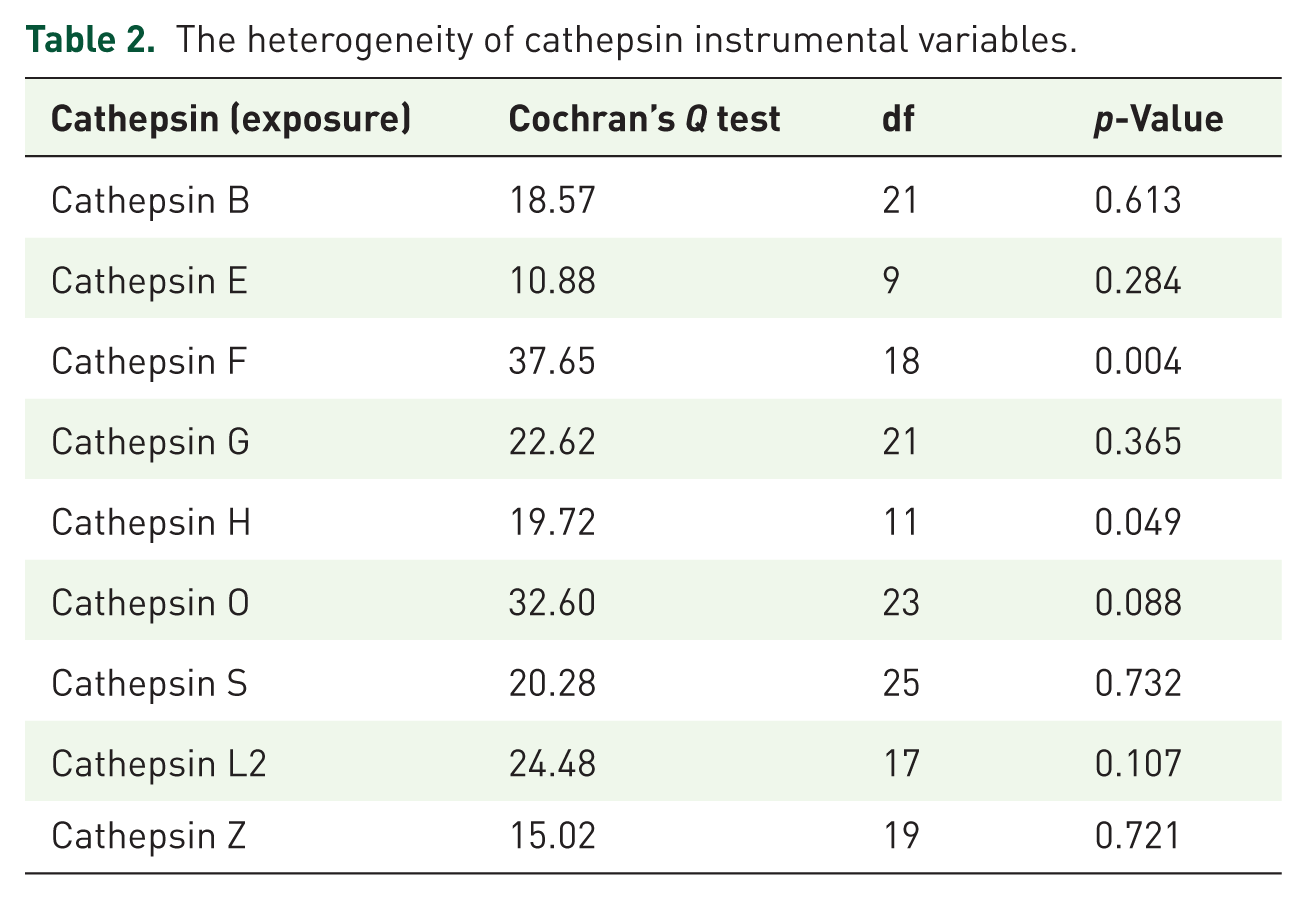
Heterogeneity and horizontal pleiotropy were evaluated using Cochran’s IVW Q-test and MR-Egger regression intercept analysis, respectively. Cochran’s IVW Q-test revealed no significant heterogeneity in the IVs for cathepsin S (Q = 20.28, p = 0.73; Table 2). Figure 2(B) further demonstrates no substantial outliers across SNP estimates. MR-Egger intercept analysis also indicated no evidence of horizontal pleiotropy for cathepsin S (p = 0.42; Table 3). In addition, MR PRESSO was used to ensure overall horizontal pleiotropy and improve study reliability (Table S2). As illustrated in Figure 2(C), leave-one-out sensitivity analysis evaluated the influence of individual SNPs on the overall MR estimates. Consistent directional effects were observed, with error bars clustered on one side of the null line upon sequential exclusion of each SNP, confirming that no single SNP disproportionately drove the association. The result supports the reliability of the causal inference between cathepsin S and AS.
The heterogeneity of cathepsin instrumental variables.
Directional horizontal pleiotropy assessed by the intercept term in MR Egger regression of the association between cathepsin and AS.

AS, ankylosing spondylitis; MR, Mendelian randomization.
Reverse MR analysis
To investigate potential reverse causation, reverse MR analyses were conducted with AS as the exposure and cathepsins as the outcomes. Results revealed no significant causal effect of AS on cathepsin S (OR = 0.97, 95% CI = 0.93–1.01, p = 0.17, IVW), as detailed in Table S3.
Discussion
This study employs a two-sample MR analysis to evaluate the causal relationship between cathepsins and AS. Among the MR methods utilized, the random-effects IVW method demonstrated stable performance and robust statistical power, serving as the primary analytical approach in this research. The findings reveal a causal association between cathepsin S and AS, while no reverse causal relationship from AS to cathepsin S was observed. While the association between Cathepsin S and AS was relatively weak (p = 0.047), it remained suggestive. To the best of our knowledge, this is the first study to report a causal link between cathepsin S and AS.
Cathepsin S, a member of the cysteine protease family, has garnered significant attention not only for its proteolytic functions but also due to its unique biochemical properties. Notably, cathepsin S retains catalytic activity beyond the acidic lysosomal environment, and its inactive proenzyme form undergoes autocatalytic activation at neutral pH. 26 Furthermore, cathepsin S is predominantly expressed in immune cells, including antigen-presenting cells, B cells, dendritic cells, and macrophages, 27 underscoring its pivotal role in immune regulation. Cathepsin S is implicated in the pathophysiology of diverse diseases, with its expression modulated in respiratory disorders, cardiovascular diseases, autoimmune conditions, neurological pathologies, metabolic disorders (e.g., obesity/diabetes), immune modulation, cancer, and inflammatory processes.28–30
Cathepsin S may contribute to tissue damage and disease progression by modulating inflammatory cytokines such as IL-6 through signaling pathways involving PAR2 and NF-κB.31,32 Liu et al. 33 identified cathepsin S as a significant risk factor enhancing rheumatoid arthritis (RA) susceptibility, underscoring its critical involvement in RA pathogenesis. In AS-related studies, cathepsin K has been proposed as a biomarker for assessing disease presence and severity. 10 Sloan et al. 34 demonstrated that cathepsin F induces ECM remodeling through degradation of ECM components (e.g., collagen and elastin), potentially contributing to structural alterations in spinal and joint tissues associated with AS-related stiffness and pain. Cathepsin S and K exhibit high structural homology and tissue-specific distribution, suggesting functional synergy or overlap, such as their shared role in promoting cytokine overexpression and secretion.31,35 This study provides the first evidence of cathepsin S as a risk factor for AS development; however, its precise mechanistic contributions to AS pathogenesis remain to be elucidated. Further preclinical and clinical investigations are warranted to explore this association.
Current management of AS combines pharmacotherapy and physical therapy to alleviate symptoms and decelerate disease progression. Standard pharmacological interventions include nonsteroidal anti-inflammatory drugs (etoricoxib, diclofenac), conventional synthetic disease-modifying antirheumatic drugs (DMARDs; sulfasalazine, methotrexate, leflunomide), biologic DMARDs (TNF inhibitors or IL-17 inhibitors), targeted synthetic DMARDs (JAK inhibitors), and corticosteroids. 36 Targeted therapy is a key future prospect for AS. B cells have garnered significant attention due to their crucial role in the pathogenesis of AS, and rituximab, which targets CD20 on B cells, has shown effective clinical responses in AS patients.37,38 Furthermore, cathepsin-specific treatments have begun to be explored; cathepsin K inhibitors, for example, have been the most extensively studied in osteoporosis, laying groundwork for their potential repurposing in AS treatment. Cathepsin S, the second most researched cathepsin to date, has been intensively investigated as a therapeutic target for inflammatory and autoimmune disorders. 27 This study identifies cathepsin S as a novel therapeutic target for AS prevention and treatment. Furthermore, the enhanced structural stability of cathepsin S compared to other cathepsins provides a robust rationale for further mechanistic and translational exploration.
In summary, this study reveals a putative genetic causal relationship between cathepsin S and AS. This study partially addresses the research gap in this field, and it opens up more possibilities for targeted diagnosis and therapy of AS via cathepsin in the future. Of course, our study has certain limitations: First, given that most participants in the pooled data of our study were of European ancestry and the sample size was small, our findings may have limited generalizability to other populations. Thus, future research involving non-European cohorts is needed to examine the broader applicability of the observed cathepsin-AS association. Second, we used a significance level of p < 1×10−5 for SNPs to obtain additional IVs, which meant that the SNPs used in the analysis in this study did not meet the standard GWAS significance threshold (p < 5 × 10−8). To mitigate weak instrument bias, we rigorously evaluated the F-statistic for each SNP. Finally, the precise pathophysiological interplay between cathepsins and AS remains incompletely understood, necessitating further mechanistic investigation. These limitations may affect the reliability of the results of this study, and more cross-sectional studies can be carried out in the future to explore the relationship between cathepsins and AS in real-life settings.
Conclusion
Our study identifies cathepsin S as a risk factor for AS. This work provides the first evidence of a causal relationship between cathepsins and AS, highlighting the critical role of cathepsin S in AS pathogenesis and proposing it as a novel therapeutic target for AS prevention and treatment.
Supplemental Material
sj-docx-1-tab-10.1177_1759720X251408873 – Supplemental material for The causal relationship between cathepsins and ankylosing spondylitis: a two-sample Mendelian randomization study
Supplemental material, sj-docx-1-tab-10.1177_1759720X251408873 for The causal relationship between cathepsins and ankylosing spondylitis: a two-sample Mendelian randomization study by Baofeng Wu, Ru Li, Jian Li, Qin An, Shuqing Jin, Qinhao Liu, Yi Zhang and Yunfeng Liu in Therapeutic Advances in Musculoskeletal Disease
Supplemental Material
sj-xlsx-2-tab-10.1177_1759720X251408873 – Supplemental material for The causal relationship between cathepsins and ankylosing spondylitis: a two-sample Mendelian randomization study
Supplemental material, sj-xlsx-2-tab-10.1177_1759720X251408873 for The causal relationship between cathepsins and ankylosing spondylitis: a two-sample Mendelian randomization study by Baofeng Wu, Ru Li, Jian Li, Qin An, Shuqing Jin, Qinhao Liu, Yi Zhang and Yunfeng Liu in Therapeutic Advances in Musculoskeletal Disease
Supplemental Material
sj-xlsx-3-tab-10.1177_1759720X251408873 – Supplemental material for The causal relationship between cathepsins and ankylosing spondylitis: a two-sample Mendelian randomization study
Supplemental material, sj-xlsx-3-tab-10.1177_1759720X251408873 for The causal relationship between cathepsins and ankylosing spondylitis: a two-sample Mendelian randomization study by Baofeng Wu, Ru Li, Jian Li, Qin An, Shuqing Jin, Qinhao Liu, Yi Zhang and Yunfeng Liu in Therapeutic Advances in Musculoskeletal Disease
Supplemental Material
sj-xlsx-4-tab-10.1177_1759720X251408873 – Supplemental material for The causal relationship between cathepsins and ankylosing spondylitis: a two-sample Mendelian randomization study
Supplemental material, sj-xlsx-4-tab-10.1177_1759720X251408873 for The causal relationship between cathepsins and ankylosing spondylitis: a two-sample Mendelian randomization study by Baofeng Wu, Ru Li, Jian Li, Qin An, Shuqing Jin, Qinhao Liu, Yi Zhang and Yunfeng Liu in Therapeutic Advances in Musculoskeletal Disease
Footnotes
Acknowledgements
The authors would like to thank the IEU Open GWAS project for providing the GWAS summary data, as well as the participants and researchers in the FinnGen project who provided AS data.
Declarations
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
