Abstract
Axial spondyloarthritis (axSpA) is a complex disease characterized by a diverse range of clinical presentations. The primary manifestation is inflammatory lower back pain, often accompanied by other clinical manifestations such as peripheral arthritis, enthesitis, uveitis, psoriasis, and inflammatory bowel disease. However, the presentation of axSpA can vary widely among patients. Despite extensive research, the precise pathogenesis of axSpA remains largely unknown. The lack of complete understanding poses challenges in subgrouping the disease, developing specific treatment approaches, and predicting treatment response. In this review, we will explore the limitations in diagnosing and treating axSpA. In addition, we will examine the current knowledge and potential opportunities provided by various omics and technological advancements in enhancing the diagnosis and personalized treatment of axSpA.
Plain language summary
The precise pathogenesis of axSpA remains unknown and is likely to be complex. Further efforts are needed to understand the disease mechanism to improve patient classification. Precision diagnosis integrates genetic data, environmental factors, and clinical characteristics to define subcategories. With the rapid advancement of technology, conducting more studies on the mechanism of SpA using multi-omics technology may yield new insights into the disease. It is also important to strike a balance between early treatment and avoiding overtreatment. Future studies should aim to combine multi-omic data, allowing the development of a more precise and individualized treatment strategy for SpA patients.
Axial spondyloarthritis (axSpA) is an inflammatory rheumatic disease characterized by inflammatory lower back pain and sacroiliitis on imaging. Some patients may also develop new bone formation (syndesmophytes) in the spine, leading to pain and limited spinal mobility. In addition, patients with axSpA can exhibit various articular and extra-musculoskeletal manifestations, including peripheral arthritis, enthesitis, dactylitis, acute anterior uveitis, psoriasis, and inflammatory bowel disease (IBD). Most patients with axSpA are carriers of human leukocyte antigen (HLA)-B27. 1 Conventionally, X-rays have been used to detect sacroiliitis. Over the past two decades, magnetic resonance imaging (MRI) has helped detect sacroiliitis earlier in axSpA patients, even before it becomes apparent on X-ray. This provides earlier treatment opportunities for these patients before structural damage occurs. However, there are some limitations in the use of MRI; therefore, new imaging modalities have been explored to assess sacroiliitis.
Currently, non-steroidal anti-inflammatory drugs (NSAIDs) are the first-line treatment for patients with axial involvement. For those patients who do not respond to NSAIDs, the emergence of biologics such as anti-tumor necrosis factor (anti-TNF), anti-interleukin (IL) 17, and janus kinase inhibitors (JAKi) have significantly improved patients’ quality of life.2–4 Despite these treatment options, a proportion of patients continue to experience persistent pain and disease progression, highlighting the need for a better understanding of the underlying pathogenesis and the identification of biomarkers to guide personalized treatment approaches.
The pathogenesis of axSpA is believed to be influenced by a combination of genetic and environmental factors, contributing to its complex etiology. Over the years, numerous studies have explored different aspects of axSpA, including pathogenesis, diagnosis, and treatment, using various omics technologies. In the era of artificial intelligence (AI), integrating multiple omics data, such as genomics, transcriptomics, proteomics, and metabolomics, may offer valuable insights into characterizing the axSpA patients and identifying potential biomarkers for early diagnosis and personalized treatment. This review aims to discuss the challenges associated with axSpA, provide an overview of the current understanding of different omics in axSpA, and propose conceptual advancements in this field.
Challenges in axSpA diagnosis
AxSpA is a disease that presents with a heterogeneous range of symptoms. Rheumatologists often incorporate both clinical assessment and the Assessment of Spondyloarthritis International Society (ASAS) classification criteria for diagnosing axSpA.5,6 The classification criteria were mainly developed to facilitate clinical trials by classifying patients with radiographic axSpA (r-axSpA) and non-radiographic axSpA (nr-axSpA). Therefore, clinical experts play an important role in making the diagnosis. According to these criteria, patients with lower back pain lasting for 3 months or more, and an onset age of less than 45 years, can be classified as having axSpA if they exhibit sacroiliitis on imaging (X-ray or MRI) or if they have the HLA-B27 allele along with two clinical features of SpA. AxSpA patients can be further divided into two subtypes: r-axSpA versus nr-axSpA, based on the presence or absence of sacroiliitis on X-ray and MRI. Patients with definite sacroiliitis on X-ray pelvis are defined as r-axSpA, while those without sacroiliitis on X-ray but with evidence of bone marrow edema (BME) on MRI are considered to have nr-axSpA. The use of MRI allows for the early detection of inflammation at the sacroiliac joint (SIJ), even before sacroiliitis becomes apparent on X-ray, enabling early diagnosis. According to the ASAS/OMERACT group consensus, the presence of BME in SIJ on at least two consecutive slices or more than one BME on a single slice, located in the typical anatomical areas (subchondrally and periarticularly) and highly suggestive of sacroiliitis associated with SpA, is regarded as positive MRI for sacroiliitis in the ASAS classification criteria for axSpA.6–8
Classification criteria are important for classifying patients with SpA for research purposes. However, there are several issues with this classification criteria. First, the “clinical arm” of the criteria showed only moderate specificity of 77%, while the “imaging arm” showed excellent specificity of 97%, and therefore the “clinical arm” lowered the overall specificity of the classification criteria. Second, the definition of active sacroiliitis on MRI used in these criteria is nonspecific as BME can be present in other conditions such as healthy individuals, athletes, or post-partum women.9–11 The variability in interpreting MRI results among different raters also affects the accuracy of the diagnosis. Third, a study showed that positive family history was not an independent factor for axSpA diagnosis in patients with HLA-B27 positivity. Yet, both HLA-B27 and positive family history are included as separate factors in the classification criteria, which increases the risk of over-classification. 12 Therefore, further research is necessary to enhance our understanding of axSpA patients, by revising the classification criteria for greater specificity. This will enable us to provide optimal treatment to the appropriate patients.
Challenges in axSpA treatment
AxSpA patients display diverse disease phenotypes, including various extra-musculoskeletal manifestations, and their disease courses and progression can differ significantly. Some patients may experience periods of inactive disease after several years, while others may have fluctuating symptoms. In addition, they exhibit varying responses to biological treatments. There is no one-size-fits-all approach for SpA. In addition to the gaps in our understanding of the disease’s pathogenesis, it remains uncertain which patients benefit most from biologic treatments and which biologic is most effective for specific patient subtypes.
The disease presentation and progression can vary significantly from person to person, even among those with r-axSpA. While syndesmophyte formation may not occur in all patients, some may experience early formation. Current studies have demonstrated that syndesmophyte formation and progression can be predicted by male sex, elevated C-reactive protein (CRP), and radiographic sacroiliitis.13,14
Radiographic sacroiliitis and elevated CRP levels might predict response to anti-TNF therapy. 14 Some studies have shown that patients with nr-axSpA and r-axSpA have a similar response to biologic treatment,15,16 while others have shown that the response to treatment in nr-axSpA patients differs from that in r-axSpA, especially in those with normal inflammatory markers. 17 Currently, the choice of biological treatment is based on patients’ comorbidities. For example, anti-IL17 therapy is preferred in patients at higher risk of tuberculosis while it should be avoided in those with symptoms of IBD. Therefore, it is essential to identify biomarkers that can assist in selecting the most appropriate biological treatment for patients, ensuring they receive treatment that yields the best response.
In addition, there are a certain number of patients with very difficult-to-treat (D2T) axSpA. Currently, there is no consensus on the definition of D2T axSpA. Fakih et al. 18 reviewed the French national healthcare database and found that one-fifth of the axSpA patients suffered from D2T SpA, based on the definition of failure of three or more biologics or targeted synthetic disease-modifying antirheumatic drugs (DMARDs) or two or more biologics or targeted synthetic DMARDs with a different mode of actions. Another French group extrapolated the EULAR definition of D2T for rheumatoid arthritis into axSpA patients and conducted a study to review the clinical characteristics of D2T axSpA patients. Those who have failed to respond to two or more biologics or targeted synthetic DMARDs with different mechanisms of action are regarded as D2T axSpA. They further classified those who failed two or more biologics or targeted synthetic DMARDs in less than 2 years as having very D2T axSpA. This study included 88 D2T axSpA patients and compared to 223 non-D2T axSpA, finding that the D2T axSpA patients had a higher prevalence of peripheral involvement, higher baseline Bath Axial Spondyloarthritis Disease Activity Index (BASDAI), and a higher prevalence of fibromyalgia. High baseline CRP and the presence of IBD at baseline were common in those with very D2T axSpA. 19 Therefore, a deeper understanding of the pathogenesis of D2T axSpA is crucial, to identify more effective treatment approaches for this challenging subset of patients.
Precision medicine in the diagnosis of axSpA
Genetics
Family history is a strong predictor of axSpA development. Twin studies showed that the disease concordance between monozygotic twin pairs was much higher than that between dizygotic twin pairs.20,21 Individuals with a first-degree relative who has HLA-B27-positive axSpA have up to 20-fold higher risk of developing axSpA.22,23 The risk of axSpA recurrence within the same family significantly decreases as the degree of relationship with the proband increases.24,25 A prospective inception cohort of 123 axSpA patients showed that seven (6%) of the seemingly healthy first-degree relatives developed axSpA after 1 year of follow-up. 26
The Class I Major Histocompatibility Complex (MHC) molecule HLA-B27 is the most important allele associated with the pathogenesis of axSpA. 1 The prevalence of HLA-B27 in axSpA varies among different ethnicities, with the Northern Norway populations having a prevalence as high as over 90%. 27 However, HLA-B27 is responsible for only about 25% of the total heritability. 28 Genome-wide association studies (GWAS) have discovered over 100 single nucleotide polymorphisms (SNPs) associated with the predisposition of r-axSpA, including IL23R, ERAP1, ERAP2, and IL1R2. 29 These SNPs are involved in the Th17-mediated immunity and peptide presentation. Another study showed polymorphism in combined homozygous and heterozygous variant genotypes of TNFRSF1A-609 G > T (TNF-alpha pathway) and homozygous variant genotype of TLR1 743 T > C (rs4833095) (NFkB pathway) was associated with a higher risk of r-axSpA, 30 whereas polymorphism in the combined homozygous and the heterozygous variant genotypes of TNF -308 G > A (rs1800629) (TNF- alpha pathway) was associated with a lower risk of r-axSpA. 30 Nonetheless, besides HLA-B27 and other MHC genes, the risk of axSpA associated with other non-MHC genes is relatively low. 31 Recently, a transcriptome-wide association study was conducted, revealing the identification of 499 susceptibility genes associated with SpA, 137 were identified from whole blood, while 390 genes were identified in skeletal muscle. 32
Given the potential contribution of multiple SNPs in increasing the risk of axSpA, utilizing a polygenic risk score could be beneficial. This approach involves assigning weights to each SNP and summing them to create a predictive tool that offers improved accuracy in predicting the likelihood of developing axSpA. A polygenic risk score has demonstrated greater accuracy in differentiating axSpA patients from individuals with lower back pain compared to HLA-B27 status, CRP, or MRI of the SIJ alone. 33 However, in terms of predicting the development of axSpA, another polygenic risk score has shown inferior diagnostic performance compared to individuals who meet the ASAS imaging criteria. 34 Further studies are needed to validate these risk scores.
MicroRNA (miRNA) is a non-coding RNA that plays a crucial role in regulating gene expression. It controls the translation of messenger RNA (mRNA), which acts as a messenger to relay genetic information through protein synthesis. With the advancement in proteomic technologies and bioinformatic analysis, a recent study investigated the exosomal microRNA profile in patients with axSpA. Compared to healthy controls, it was found that axSpA patients demonstrated a distinct miRNA signature in their circulating exosomes, with 22 miRNAs being upregulated and 2 miRNAs being downregulated. Some of these differentially expressed miRNAs are involved in the process of bone remodeling, such as miR-140-3p, miR-30d-5p, and miR-29a.35,36 These miRNAs are likely involved in the new bone formation observed in axSpA. Furthermore, the specific exosomal miRNA profile may serve as a potential biomarker for the diagnosis of axSpA.
The role of genetic factors in the development of axSpA is undoubtedly crucial. However, studies of twins revealed that not all monozygotic twins develop ankylosing spondylitis.20,21 This suggests that, in addition to genetic predisposition, other factors may also significantly contribute to triggering the onset of the disease.
Imaging
Sacroiliitis is a crucial component in diagnosing axSpA, so it is important to accurately define active sacroiliitis for proper diagnosis. For decades, X-rays have been used for the assessment of sacroiliitis. However, the sensitivity and specificity of X-rays in diagnosing sacroiliitis are inconsistent, even among trained rheumatologists and radiologists. 37 Moreover, X-rays are not capable of detecting the early stages of sacroiliitis. As a result, MRI has been utilized for the detection of sacroiliitis. The ASAS Classification criteria defines a positive MRI for sacroiliitis as the presence of BME at typical anatomical sites that are highly suggestive of spondyloarthropathies, which appears as a hyperintense signal on short tau inversion recovery (STIR) imaging or T2-weighted images, and a hypointense signal on T1-weighted images, located peri-articularly. 8 Structural damage such as sclerosis or erosion may also be present. However, BME is a nonspecific finding and can also be observed in other conditions. Recently, the ASAS MRI group proposed a new definition of active lesions specific to axSpA with high positive prediction values. 38 Two reading exercises, involving 169 cases and 91 cases, respectively, were conducted by seven and eight experienced readers to assess active and chronic inflammation and structural lesions and to determine whether these MRI findings were suggestive of axSpA. Based on these results, they proposed a data-driven MRI definition for axSpA, in which an active lesion typical of axSpA requires BME at four or more quadrants of the SIJ at any location, or the same location in at least three consecutive slices. 38 The ASAS MRI group has also proposed a data-driven definition of the structural lesions typical of SpA which include erosions in three or more quadrants of the SIJ, or fat lesions in five or more quadrants of the SIJ. In addition, erosion or fat lesions at the same location in ⩾2 or ⩾3 consecutive slices respectively, or the presence of a deep fat lesion, are also considered structural lesions. 38 However, these definitions need further validation.
Besides conventional MRI sequences, other imaging modalities have also been explored in assessing for sacroiliitis. Low-dose computer tomography (CT) may be a good alternative to X-ray in detecting sacroiliitis as it has better sensitivity in detecting structural lesions including erosions, sclerosis, and ankylosis, and lower inter-rater variability. 39 Low-dose CT (ldCT) has also shown higher specificity in detecting sacroiliitis compared to MRI, particularly in patients with nr-axSpA as BME could be observed in other conditions, potentially resulting in false-positive findings on MRI. 39 In patients with nr-axSpA who present with inconclusive findings and do not show sacroiliitis on X-ray and MRI, the presence of structural lesions on ldCT may aid in the diagnosis of SpA-related sacroiliitis (see Figures 1 to 3). 40 A study that employed data-driven analysis revealed that ankylosis and erosions of the middle and dorsal joint portion of the SIJ exhibit excellent specificity in the diagnosis of sacroiliitis. 41 LdCT has been used as a standard reference for the detection of structural lesions in some studies. 40 However, ldCT cannot differentiate active sacroiliitis and poses a radiation risk. 40 Alternatively, other novel MRI sequences have been investigated for the assessment of structural lesions in the SIJ. The MRI sequence, known as volumetric interpolated breath-hold examination (MR-VIBE), has shown higher sensitivity in detecting erosions in the SIJ compared to T1-weighted MRI, using ldCT as the standard reference. 42 MR-VIBE also allowed the detection of smaller erosions compared to T1-weighted MRI or ldCT, likely due to the higher contrast between bone and cartilage. The diagnostic confidence was highest when using MR-VIBE, followed by ldCT and T1-weighted MRI in descending order. In addition, zero echo time MRI and gradient echo “black bone” MRI were new MRI sequences that demonstrated superiority over T1-weighted MRI in detecting cortical bone lesions, with a sensitivity of around 80% and a specificity of 99% compared to ldCT.43,44 Other novel imaging modalities such as diffusion-weighted MRI (DWI) and dual-energy CT (DECT) have also demonstrated good specificity in detecting inflammatory sacroiliitis BME.45,46 However, the STIR sequence appeared to be more reliable than DWI for identifying active sacroiliitis. 47 While DECT has the limitation of being unable to differentiate between sclerosis and BME in some circumstances. 40 As the field of imaging continues to rapidly evolve, it is important to determine the most sensitive and specific imaging technique for diagnosing SpA-related sacroiliitis. This is crucial for ensuring early diagnosis while also avoiding over-diagnosis.

Low-dose CT revealed definite evidence of moderate ankylosis in the left sacroiliac joint.
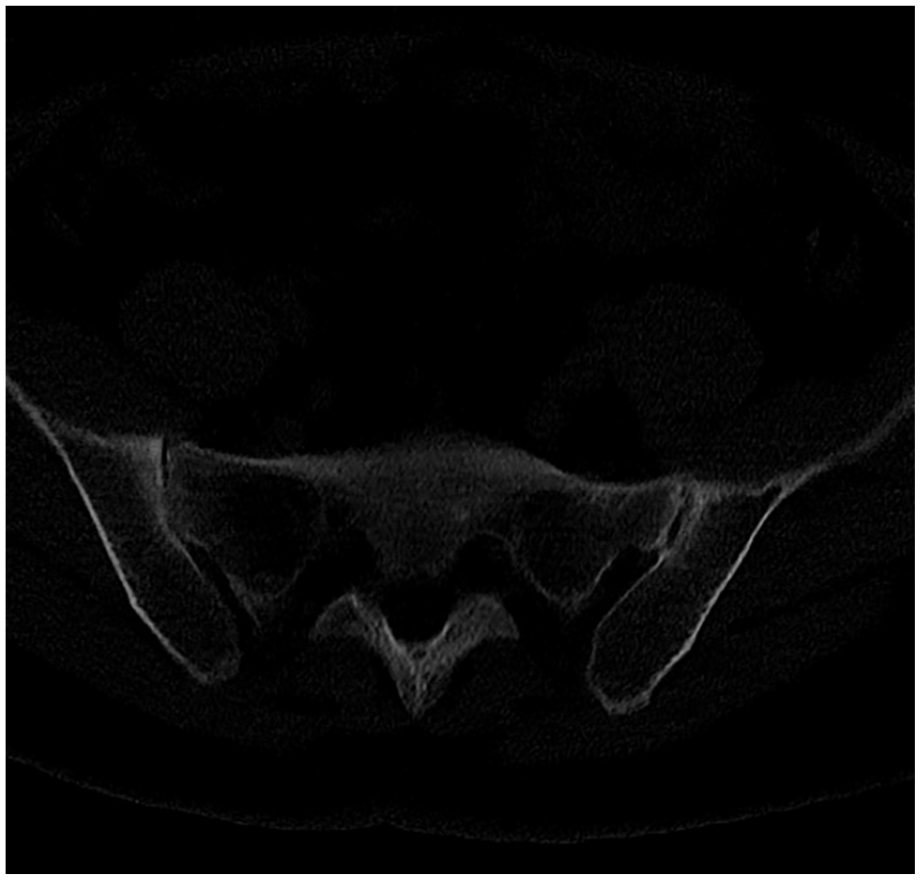
Low-dose CT showed moderate narrowing of the right sacroiliac joint with small erosions.

Low-dose CT showed moderate narrowing of the right sacroiliac joint with mild ankylosis.
Accurate interpretation of imaging studies poses a challenge and requires extensive training. The application of AI in reading imaging, including X-ray, MRI, and CT films, has shown promise in increasing accuracy and reducing labor. X-ray pelvis has been used to assess for radiographic sacroiliitis; however, the interrater agreement was suboptimal among both trained readers and local rheumatologists and radiologists, with a Kappa value of around 0.5. 48 Deep artificial neural networks, in turn, have facilitated more consistent detection of definite sacroiliitis on X-ray and CT.49,50 Bressem et al. 49 showed good performance between the reference agreement and the neural network assessment of sacroiliitis on X-ray, with a good kappa value of 0.79 and 0.72 for the test and validation cohorts, respectively. Another study led by Belgian and Canadian groups developed a deep neural network that could automatically perform SIJ segmentation as well as detect erosions and ankylosis on the SIJ CT with high accuracy. 50 Automated machine-learning systems have also been developed for reading BME, which saves time, enhances objectivity, and potentially improves accuracy.51,52 A study using a deep learning model to assess SIJ inflammation on STIR sequence in MRI showed excellent performance, which was comparable to the assessment by radiologists. 52 Another team further developed a fully automated algorithm that could detect and segment the SIJ, followed by quadrant extraction and identification of BME in the SIJ. The algorithm demonstrated good performance in the detection of inflammation in the validation cohort, as assessed by the area under the curve, balanced accuracy, and F1 score. 51 This is an important step toward the fully automated assessment of SIJ. In the foreseeable future, with the aid of AI, we can hope for an easier, accurate, and consistent diagnosis of sacroiliitis.
Gut microbiome and metabolomic profiling
Growing evidence from animal and clinical studies supports the role of the gut–joint axis in the development of SpA. 53 Dysbiosis, characterized by an imbalance in gut microbiome composition, is believed to be a key factor in triggering autoimmune diseases. A systemic review revealed distinct microbiome profiles in terminal ileum biopsies and stool samples of patients with axSpA. 54 Among the consistent findings across different studies, there was a reduced abundance of Bacteroidales and Parasutterella, as well as an increased abundance of Actinobateria (Phylum), Dialister, Streptococcus, and Clostridium bolteae in the gut microbiome of these patients compared to healthy individuals. However, further research is required to ascertain whether patients with axSpA exhibit a distinct gut microbiome profile that can distinguish them from individuals with other forms of SpA or inflammatory arthritis, aiding in the diagnostic process. Moreover, it is important to note that the applicability of these results is limited by the heterogenicity of disease characteristics observed in patients across different studies, as well as the assessment methods employed. Most studies utilized the 16S rRNA gene sequencing method, which may restrict the identification of specific genes. In the future, it is recommended to conduct more studies utilizing metagenomic sequencing methods to identify additional disease-specific bacteria and facilitate the discovery of microbial markers.
Metabolites are the products that arise from the interaction between genes and the environment. Few metabolomics studies showed potential biomarkers of axSpA using plasma, urine, and ligament samples.55,56 However, metabolome could be influenced by many factors such as sex, age, and body build, and its level can fluctuate rapidly. This variability makes it challenging to standardize and interpret the data. Therefore, additional research is required to validate the panel of metabolites that have the potential to be specific for diagnosing axSpA and ensure that the findings can be replicated reliably.
Prediction of disease progression
It is important to identify patients at high risk of disease progression to prevent irreversible new bone formation that can lead to limited spinal mobility, debilitating pain, and reduced quality of life. The Modified Stoke Ankylosing Spondylitis Spine Score (mSASSS) has been identified as the most important predictor of radiographic progression. 57 High disease activity, as defined by the Axial Spondyloarthritis Disease Activity Score (ASDAS), is associated with greater radiographic damage measured by mSASSS. 58 HLA-B27 positivity is also linked to more severe radiographic changes and increased syndesmophytes formation. 59 Male gender, age, smoking, obesity, and the presence of syndesmophytes at baseline are all associated with radiographic progression.59–61 Furthermore, there are sex-specific predictors of spinal radiographic progression. A study showed that exposure to bisphosphonate in women was associated with new syndesmophyte formation and spinal radiographic progression, while current smokers were associated with new syndesmophyte formation in men. 61 In young axSpA patients, low bone mineral density may predict the formation of new syndesmophytes. 60
MRI can detect early bone changes, and studies have shown that inflammatory lesions on the MRI spine may resolve without permanent damage. However, chronic inflammatory lesions are associated with new syndesmophyte formation. 62 Also, the presence of fatty lesions, indicating an early stage of the bone remodeling process, is also linked to new syndesmophyte formation. 62 Therefore, early aggressive treatment should be considered for patients, before the occurrence of these lesions that may lead to irreversible damage.
High levels of CRP and erythrocyte sedimentation rate (ESR) are widely recognized as risk factors for radiographic progression in axSpA. 31 In addition, various other biomarkers, including microRNA; calprotectin (inflammation marker); visfatin and leptin (adipokines); and sclerostin and serum C-terminal telopeptide fragments of type I collagen (bone turnover marker), have been investigated for their potential in predicting radiographic progression in axSpA patients.63,64 However, the results regarding these novel biomarkers are inconsistent. 31 Bin et al. attempted to develop algorithms to identify risk factors associated with radiographic spinal progression, defined by dichotomous discrimination, in axSpA patients. They tested various known risk factors using seven different machine-learning models. However, the presence of baseline syndesmophytes was the only consistent and significantly important risk factor across the models. 65 Recently, a retrospective study utilized machine-learning methods, including linear regression models and artificial neural networks to develop a composite model for predicting spinal progression in SpA patients in terms of mSASSS. The model incorporated known and potential risk factors, such as baseline clinical characteristics and treatment. 57 These predictive models demonstrated excellent performance in predicting radiographic progression, which may help identify patients at high risk of spinal progression and formulate more individualized treatment plans (see Figure 4).

This figure outlines current precision medicine approaches for diagnosing and treating axSpA, as well as predicting disease progression. Key areas covered include genome analysis, gut microbiome, and imaging. Future research should focus on multi-omic studies and developing risk score models, integrating technological advancements to enable more personalized treatment strategies for axSpA.
Precision medicine in treatment prediction
NSAIDs are considered the first-line treatment for axSpA patients. A study demonstrated that BME improved after 6 weeks of NSAID use, despite no significant clinical improvement. 66 Also, a significant proportion of patients were unable to tolerate the full dose of NSAIDs. Biologic agents have shown effectiveness in many patients including anti-TNF, anti-IL17, and JAK inhibitors. 67 In terms of mSASSS progression, users of all three classes of biologics, anti-TNF, anti-IL17, and JAK inhibitors, had a slower rate and minimal spinal disease progression.67–69 Similarly, the above three classes of biologics users also showed improvement in spinal inflammation on MRI.67,70 Although two network meta-analyses showed that anti-TNFα treatment outperformed anti-IL17 and JAK inhibitors in terms of disease activity improvement, as indicated by ASAS5/6, ASAS20, and ASAS40 responses.71,72 However, careful interpretation of these studies is needed as most studies included in the meta-analyses were randomized controlled trials or cohort studies, and head-to-head studies on different classes of biologics in axSpA were limited. A study on secukinumab (an anti-IL17) used adalimumab (an anti-TNF) as an active reference group and showed both drugs performed better than placebo in terms of achieving ASAS40 response. 73 A recent head-to-head study comparing tofacifitib (a JAK inhibitor) with adalimumab showed both drugs achieved similar clinical improvement at 6 months. 74
Several factors have been identified as predictors of good response to biological treatment. AxSpA patients with younger age responded better to anti-TNFα than older patients, regardless of disease duration. 75 In patients with r-axSpA, active inflammation on the MRI spine, a disease duration of less than 10 years, high clinical disease activity, and elevated CRP levels are predictive of a good clinical response to anti-TNF therapy. 76 For patients with nr-axSpA, inflammation in SIJ, positive HLA-B27 status, and high CRP levels at baseline may also serve as predictors of good response to anti-TNF treatment. 77 Similarly, higher CRP levels, the presence of inflammation on MRI at baseline, and male sex were also shown to predict better response to anti-IL17 (secukinumab and ixekizumab) compared to placebo in both radiographic and nr-axSpA patients.78–80 Elevated CRP levels and inflammation on MRI of SIJ at baseline also predicted a favorable response to JAK inhibitors (tofacitinib and upadacitinib) in axSpA patients compared to placebo.81,82 Consistently, axSpA patients with both elevated CRP and sacroiliitis on MRI had better responses to all three classes of biological therapies: anti-TNF, anti-IL17, and JAK inhibitors. 83 There are a few research agendas that could be focused on in the future (see Table 1).
Future research agenda.

axSpA, axial spondyloarthritis.
Genetics
Pharmacogenetic testing is an exciting field aiming to use genetics to predict treatment response. Previous studies have aimed to identify genetic variants that can predict treatment response in patients with SpA. However, the relationship between treatment response and genes encoding TNF or molecules involved in the TNF receptor pathway remains inconsistent. 84 In addition to genes involved in the TNF pathway, the IL-17 pathway has also been studied. A study of Polish axSpA patients showed IL-17F rs763780 polymorphism may predict response to anti-TNF treatment, with patients homozygous for A allele responding better than heterozygotes. However, further studies are needed for external validation. Recently, another study reviewed the role of genetics in predicting treatment response to secukinumab, an anti-IL17 agent, in axSpA patients, yielded negative results. 85
Potential serum biomarkers
Many cytokines have been studied to predict treatment response to anti-TNF. High baseline IL-17A level and lower baseline level of Jun N-terminal kinase pathway-associated phosphatase (JKAP) were found to be associated with better response to adalimumab, an anti-TNF therapy, with similar predictive value as CRP.86,87 Another study showed that serum type 1 interferon activity, interferon-α level, and interferon-γ level were associated with better response to adalimumab.87,88 For IL-6, there were inconsistent findings regarding the association of treatment response to anti-TNFα and baseline IL-6 level with different studies.86,89 A study showed that a lower baseline level of TNFα-producing CD8 cells was associated with a positive response to anti-TNFα treatment and the level of TNFα-producing CD8 cells is likely influenced by age. 75 On the other hand, although patients with axSpA exhibit significantly elevated serum levels of matrix metalloproteinase 3 (MMP3), these levels failed to predict the response to etanercept, an anti-TNF therapy. 90 By contrast, a study has demonstrated that a change in serum calprotectin levels during the first month of treatment may serve as a predictive factor for a favorable treatment response. 91 A recently published study showed that higher serum zoulin levels and lower serum haptoglobulin levels were associated with poorer response to biological treatment. 92 All these potential biomarkers are novel, and studies have not compared their sensitivity and specificity to CRP/ESR. Further studies are required for validation and to explore their applicability in clinical practice.
Microbiome
Efforts have been made to identify any biomarker for predicting treatment response. A study identified seven operational taxonomic units (OTUs) that could potentially predict treatment response. Among them, the prevalence of OTU0112, in the genus Sutterella, could predict non-responders to anti-TNF treatment, based on ASDAS criteria. 93 Another pilot study that included 19 SpA patients showed that a higher abundance of Burkholderiales at baseline may predict good anti-TNF response in SpA patients. 94 A study involving 30 axSpA patients revealed a tendency to predict non-responsiveness to adalimumab in patients who had a higher abundance of Comamonmas in their stool before treatment. 95 The available data on the use of gut microbiome in predicting treatment outcomes in patients with SpA are currently very limited. Larger-scale studies are needed to confirm these findings.
Conclusion
The precise pathogenesis of axSpA remains unknown and is likely to be complex. Further efforts are needed to understand the disease mechanism to improve patient classification. Precision diagnosis integrates genetic data, environmental factors, and clinical characteristics to define subcategories. With the rapid advancement of technology, conducting more studies on the mechanism of SpA using multi-omics technology may yield new insights into the disease. It is also important to strike a balance between early treatment and avoiding overtreatment. Future studies should aim to combine multi-omic data, allowing the development of a more precise and individualized treatment strategy for SpA patients.
