Abstract
Amyotrophic lateral sclerosis (ALS) is a fatal neurodegenerative disease characterized by upper and lower motor neuron (MN) degeneration. Astrocytes surrounding MNs are known to modulate ALS progression. When cocultured with astrocytes overexpressing the ALS-linked mutant Cu/Zn superoxide dismutase (SOD1G93A) or when cultured with conditioned medium from SOD1G93A astrocytes, MN survival is reduced. The exact mechanism of this neurotoxic effect is unknown. Astrocytes secrete extracellular vesicles (EVs) that transport protein, mRNA, and microRNA species from one cell to another. The size and protein markers characteristic of exosomes were observed in the EVs obtained from cultured astrocytes, indicating their abundance in exosomes. Here, we analyzed the microRNA content of the exosomes derived from SOD1G93A astrocytes and evaluated their role in MN survival. Purified MNs exposed to SOD1G93A astrocyte-derived exosomes showed reduced survival and neurite length compared to those exposed to exosomes derived from non-transgenic (non-Tg) astrocytes. Analysis of the miRNA content of the exosomes revealed that miR-155-5p and miR-582-3p are differentially expressed in SOD1G93A exosomes compared with exosomes from non-Tg astrocytes. Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis indicates that miR-155-5p and miR-582-3p predicted targets are enriched in the neurotrophin signaling pathway. Importantly, when levels of miR-155-5p were reduced by incubation with a specific antagomir, SOD1G93A exosomes did not affect MN survival or neurite length. These results demonstrate that SOD1G93A-derived exosomes are sufficient to induce MN death, and miRNA-155-5p contributes to this effect. miRNA-155-5p may offer a new therapeutic target to modulate disease progression in ALS.
Introduction
Amyotrophic lateral sclerosis (ALS) is a neurodegenerative condition that selectively damages the motor neurons (MNs) in the cortex, brainstem, and spinal cord (Brown & Al-Chalabi, 2017). This induces progressive paralysis that leads to death within 3–5 years postdiagnosis due to respiratory failure (Brown & Al-Chalabi, 2017). Most ALS cases are sporadic, but 5–10% have a familiar history and mendelian inheritance with more than 13 associated genes, including Cu/Zn superoxide dismutase (SOD1), TAR DNA binding protein-43 (TDP-43), fused in sarcoma (FUS), and chromosome 9 open reading frame 72 (C9Orf72), (Kiernan et al., 2011). Overexpression of mutant human SOD1 gene in rodents leads to progressive MN degeneration resembling many aspects of ALS in human patients and has contributed to elucidating various aspects of ALS pathogenesis (Gurney et al., 1994; Howland et al., 2002).
Although ALS is characterized by MN degeneration, glial cells play a key role in the development of the disease (Clement et al., 2003). Several neuroglia cell types, such as astrocytes, microglia, and oligodendrocytes, are involved in ALS progression (Barbeito et al., 2004; Ilieva et al., 2009). Experiments using mouse models with the ALS-associated human SOD1 with glycine to alanine substitution in position 93 (SOD1G93A) transgene deleted in a specific cell type manner using the Cre-Lox system revealed that removal of mutant SOD1 in astrocytes or microglial cells significantly slowed ALS progression and extended life span in mice (Boillée et al., 2006; Wang et al., 2011a; 2009; Yamanaka et al., 2008b). Astrocytes are the most abundant glial cells in the central nervous system and have a major neurotrophic function for MNs. They release trophic factors and limit MN excitotoxicity by clearing glutamate from the synaptic cleft (Verkhratsky et al., 2019). ALS astrocytes change their gene expression pattern and adopt an activated state that contributes to the progression of the disease (Liddelow & Barres, 2017). Data obtained from our group and others using isolated astrocytes in coculture with MNs have demonstrated that astrocytes expressing SOD1 mutations reduce MN survival compared to non-transgenic (non-Tg) ones (Di Giorgio et al., 2007; Marchetto et al., 2008; Nagai et al., 2007; Vargas et al., 2004; Yamanaka et al., 2008a). In agreement with this, it was demonstrated that the addition of SOD1 astrocyte-conditioned medium to MNs cultures results in neurotoxicity (Fritz et al., 2013; Madill et al., 2017; Nagai et al., 2007). However, the agent responsible for astrocyte toxicity remains unknown.
Like other cell types, astrocytes modulate different biological processes by the secretion of extracellular vesicles (EVs) (Basso et al., 2013; Ceruti et al., 2011; Falchi et al., 2013; Hajrasouliha et al., 2013; Proia et al., 2008; Sbai et al., 2010; Taylor et al., 2007; Wang et al., 2011b). EVs include microvesicles and exosomes; while microvesicles are directly shed from the plasma membrane and have a heterogeneous size ranging from a dozen nanometers to a few micrometers, exosomes are generated by inward budding from the limiting membrane of endosomes forming the multivesicular bodies (MVB), which upon fusion with the plasma membrane cause exosome release (Raposo & Stoorvogel, 2013). An alteration in exosome astrocytic production and composition has been previously reported in ALS (Gharbi et al., 2020). In particular, astrocytes that express SOD1G93A show an increase in exosome release carrying SOD1 protein, which efficiently transfers mutant SOD1 to spinal neurons and induces selective MN death (Basso et al., 2013; Silverman et al., 2019).
Exosomes transport mRNA and microRNAs (miRNAs) from one cell to another (Ratajczak et al., 2006; Valadi et al., 2007). This is especially important in ALS, whereby RNA metabolism is altered and has been proposed to initiate cellular pathology (Arnold et al., 2013; Bilsland et al., 2010; Da Cruz & Cleveland, 2011; De Vos & Hafezparast, 2017; Murakami et al., 2002; Perlson et al., 2009; Rotem et al., 2017). In particular, the miRNA pathway is dysregulated in ALS and has been suggested as an interphase of stress and disease (Emde et al., 2015). miRNAs are RNAs of about 18–25 nucleotides long that act by binding to their target mRNAs resulting in translational repression and/or mRNA degradation (Ambros, 2004; Bartel, 2004). Two ALS-linked genes, FUS and TDP-43, are essential for miRNA biogenesis and pre-miRNA processing (Lagier-Tourenne et al., 2010). Mutations in these genes correlate with dysregulated RNA processing and metabolism in ALS cells or tissues (Le Gall et al., 2020). miRNA dysregulation has been implicated in the disease, with numerous studies supporting a role for miRNAs in ALS (Campos-Melo et al., 2013; De Felice et al., 2012; Emde et al., 2015; Hoye et al., 2017; Koval et al., 2013; Zhou et al., 2013).
Here, we further evaluated the contribution of the exosomes from SOD1G93A astrocytes conditioned media (CM) in their ability to influence MN survival and neurite integrity. In addition, we focus on the microRNA content of exosomes that may contribute to these effects.
Materials and Methods
Materials
Culture media and serum were obtained from Gibco (Thermo Fisher Scientific, Waltham, MA, USA). Unless otherwise specified, all other reagents were from Sigma Chemical Co. (Saint Louis, MO, USA).
Ethics Statement
Procedures using laboratory animals were conducted following international guidelines and were approved by the Institutional Animal Committee (Comisión honoraria de experimentación animal de la Universidad de la República http://www.chea.csic.edu.uy; CHEA), protocol no. 070153-000729-18.
Animals
Rats (Rattus norvegicus) were maintained in a controlled environment (12-h light-dark cycle; 20 ± 1°C), fed with ad libitum access to food and water, and housed in a cage with five female or male rats. Male hemizygous NTac: SD-Tg (SOD1G93A) L26H rats (RRID: IMSR_TAC:2148) were obtained from Taconic (Hudson, NY, USA) and were mated with outbred Sprague–Dawley background. Progeny pups were genotyped with polymerase chain reaction (PCR), as previously described (Pehar et al., 2004; Vargas et al., 2006). In our colony, the onset of symptomatic disease (160–170 days), as well as lifespan (180–195 days), was considerably delayed from the original report (Howland et al., 2002).
Astrocyte Cultures
Cortical astrocyte cultures were prepared from 1-day transgenic SOD1G93A or nontransgenic rat pups genotyped by PCR as previously described (Pehar et al., 2004; Vargas et al., 2006). Briefly, the brain cortex was dissected, meninges were carefully removed, and tissue was chopped and dissociated with 0.25% trypsin-EDTA for 25 min at 37°C. Trypsinization was stopped with Dulbecco's modified Eagle's medium supplemented with 10% fetal bovine serum (s-DMEM) in the presence of 50 μg/ml DNase I. Subsequently, the tissue was mechanically disaggregated by repeated pipetting, and the suspension was passed through an 80-μm mesh and spun 10 min at 300 × g. The pellet was resuspended in s-DMEM medium and plated at a density of 1.5 × 106 cells per 25-cm2 tissue culture flask. When confluent, cultures were shaken for 48 h at 250 rpm and incubated for another 48 h with 10 μM cytosine arabinoside. Next, cells were plated at a density of 2 × 104 cells/cm2 in a 75 cm2 Nunc flask (Nunc, Thermo Fisher Scientific). Astrocytes were maintained in s-DMEM, 4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid (HEPES, 3.6 g/L), penicillin (100 IU/mL), and streptomycin (100 mg/mL). Astrocyte monolayers were 98% pure as determined by glial fibrillary acidic protein (GFAP) immunoreactivity.
MN Cultures
MN cultures were prepared from embryonic day 15 (E15) rat spinal cord by a combination of OptiPrep™ (Merck, Rahway, NJ USA, 1:10 in L15 medium) gradient centrifugation and immunopanning with the monoclonal antibody 192-IgG hybridoma supernatant (Chandler et al., 1984) against p75 low-affinity neurotrophin receptor as previously described with minor modifications (Henderson et al., 1993). The remaining cell population after immunopanning consisted of over 95% neurons, expressing the MN markers p75 and ISLet1. Purified MNs were seeded at a density of 300 cells/cm2 onto confluent glial feeder layers (FL) in s-Leibovitz's L15 medium or on a polyornithine–laminin substrate and maintained in s-Neurobasal™ (NB) medium. MN cultures were supplemented with 1 ng/ml glial cell-derived neurotrophic factor (GDNF) as previously described (Cassina et al., 2002; Díaz-Amarilla et al., 2011) and subjected to different treatments described later. For RNA extraction, when high amounts of MNs were required, immunopanning was avoided.
Exosome-Enriched Fraction
Exosomes were isolated from a conditioned medium harvested from a confluent monolayer of cultured astrocytes from two 75 cm2 flasks (6 × 106 astrocytes on 30 ml final volume). Before CM extraction, astrocytes were washed twice with phosphate-buffered saline (PBS, NaCl2 137 mM, 2.7 mM KCl, 4.3 mM Na2HPO4·7H2O, 1.5 mM KH2PO4) and incubated with DMEM, supplemented with HEPES (3.6 g/L), penicillin (100 IU/ml), and streptomycin (100 mg/ml). After 24 h incubation, exosomes were prepared from astrocyte culture mediums. Exosomes were purified as described previously with minor modifications (Thery et al., 1999) by three successive centrifugations at 300 × g (10 min), 2,000 × g (10 min), and 10,000 × g (30 min) to eliminate cells and debris followed by a 100,000 × g centrifugation for 80 min. The exosome pellet was washed once in 6 ml PBS, centrifuged 100,000 × g (1 h), and resuspended in 30 µl of PBS or 200 µl TRIzol® (Thermo Fisher Scientific) or protein lysis buffer for western blotting. To evaluate exosome integrity or exosome membrane protein in survival or neuron morphometric parameter experiments, fractions were subjected to incubation with 0.25% trypsin or Milli-Q water for 10 min at 37°C, as described previously (Lopez-Verrilli et al., 2013). After treatment, fractions were washed in PBS, ultracentrifuged, and resuspended in PBS. The exosomes obtained were analyzed by Microfluidic Resistive Pulse Sensing (MRPS, Spectradyne, Torrance, CA) to determine the size and concentration of the exosomes (Arab et al., 2021). A mean concentration of 12.8 ± 6.3 × 1010 particles/ml was obtained, corresponding to approximately 4,000 particles per astrocyte.
MNs Treatment and Counting
After being cultured for 24 h, MNs were subjected to the following treatments: astrocytic conditioned medium (CM), exosome-free astrocyte CM (at a 1:10 dilution), or exosomes at a concentration of 6.4 × 108 particles/ml. This concentration is equivalent to approximately 5 × 105 particles/MN, which matches the final concentration present in a similar volume of astrocyte CM. Additionally, MNs were treated with either a 0.01 μM 155-5p miRNA mimetic (MIMAT0030409 Ambion, Thermo Fisher Scientific) or an inhibitor (MIMAT0030409, Ambion, Thermo Fisher Scientific).
After 24–48 h posttreatment, cells were processed for immunofluorescence and MN counting. MN survival was assessed by directly counting all intact neurites longer than 4 somas in diameter in a prefixed area of the dishes. Counts were performed in an area of 0.90 cm2 along a diagonal in 24-well plates.
Western Blot Analysis
Proteins were extracted from exosomes enriched fraction in 1% SDS supplemented with 2 mM sodium orthovanadate and a complete protease inhibitor cocktail (Roche, Basel, Switzerland). Lysates were resolved by electrophoresis on 12% SDS–polyacrylamide gels and transferred to a polyvinylidene fluoride membrane (PVDF; Thermo Fisher Scientific). The membrane was blocked for 1 h at room temperature in 5% skimmed milk in TBS-T (Tris-buffered saline with 0.1% Tween). The membrane was then probed overnight with primary antibodies in 1% skimmed milk in TBS-T at 4°C. Primary antibodies include rabbit monoclonal anti-TSG101, 1:1,000 (Abcam, Cambridge, UK, Cat# ab125011, RRID: AB_10974262); mouse monoclonal anti-flotillin-1, 1:500 (BD Biosciences, Franklin Lakes, NJ, USA, Cat# 610820, RRID: AB_398139); rabbit polyclonal anti-human SOD1 antibody kindly provided by Prof. Mónica Marin from Facultad de Ciencias, UdelaR, 1:500 (Palacios et al., 2010); and rabbit polyclonal anti-BIP, 1:1,000 (Abcam, ab21685, RRID: AB_2119834). Afterward, the membranes were washed in TBS-T and incubated for 60 min at room temperature with IRDye 680RD-conjugated goat anti-mouse IgG and IRDye 800CW-conjugated goat anti-rabbit IgG secondary antibodies (1:15,000 in PBS each, LI-COR Biosciences, Cat# 926-68070, RRID: AB_10956588 and Cat# 925-68071, RRID: AB_2721181, respectively). The Odyssey system (LI-COR Biosciences) was used to detect the bands.
Electron Microscopy
Exosomes were processed for transmission electron microscopy (TEM) by negative staining with 2% uranyl acetate at the Electron Microscopy Unit (Facultad de Ciencias, Universidad de la República, Montevideo, Uruguay). Briefly, astrocyte exosome-enriched fractions were prepared as mentioned and resuspended in 30 µl PBS. 10 µl exosome suspension was added on top of carbon film-coated electron microscopy grids (Electron Microscopy Sciences) and incubated for 15 min, and excess liquid was removed with filter paper. Samples were incubated with 2% uranyl acetate for 5 min, with excess liquid removed as before, and let dry completely before imaging. Exosome preparations were examined using a Jeol JEM1010 TEM operated at 100 kV and equipped with 4,000 AM DVC and a HAMAMATSU C-4742-95 digital camera under the control software AMT ADVANTAGE. The exosome diameter was measured in electron micrographs using Fiji (ImageJ) software (NIH; RRID: SCR_002285).
Immunofluorescence
MN cultures were fixed for 15 min on ice with 4% paraformaldehyde plus 0.1% glutaraldehyde in PBS. The cultures were permeabilized with 0.1% Triton X-100 for 15 min and incubated for 2 h at room temperature (25°C) in blocking solution [0.1% Triton X-100, 2% bovine serum albumin (BSA), and 10% goat serum in PBS]. Primary antibodies diluted in the blocking solution were incubated overnight at 4°C. After washing with PBS, the cultures were further incubated for 1 h at room temperature with the secondary antibodies diluted in the blocking solution. The slides were then washed with PBS, rinsed with distilled water, and mounted with Prolong Antifade mounting kit (Thermo Fisher Scientific). The primary antibody used was mouse anti-beta III tubulin (1:3,000, Abcam, Cambridge, UK, Cat# ab41489, RRID: AB_727049), and the secondary was Alexa Fluor 488-conjugated goat anti-mouse IgG secondary antibody (1:1,000, Thermo Fisher Scientific, # A-11029, RRID: AB_2534088). Nuclei were stained with DAPI. Images were captured with a DC290 Zoom Kodak Digital Camera coupled to a Nikon fluorescence microscope.
RNA Extraction and Quantitative RT-PCR
Astrocyte cells and exosomes were lysed with TRIzol reagent (Thermo Fisher Scientific, USA) to extract the required RNA, as the manufacturer's instructions recommended. To increase the RNA yields, minor modifications were applied at the precipitation step of exosome RNA purification. Briefly, RNA precipitation was performed with two volumes of isopropyl alcohol; 5 µg linear acrylamide was added, and the solution was incubated for 20 min at −20°C. After 30 min centrifugation at 12,000g, 4°C supernatant was discarded, and RNA was resuspended in 20 µl nuclease-free water (Thermo Fisher Scientific) and 40 U RNase inhibitor (Thermo Fisher Scientific). Possible DNA contaminations were eliminated with DNase treatment using the DNase-free Kit (Thermo Fisher Scientific). RNA quality was evaluated by agarose gel electrophoresis followed by ethidium bromide staining or by using Agilent 2100 BioAnalyzer (Agilent, Santa Clara, CA, USA). RNA quantification was performed by 260 nm absorption using a nanodrop spectrophotometer (Thermo Fisher Scientific).
Mature miRNA qPCR was done by Stem Loop RT-qPCR as described previously (Kramer, 2011) and mRNA by SYBR Green-based RT-qPCR. Briefly, before starting Stem Loop RT-qPCR, a stock solution of 100 μM SL primer was done and after was folded by heating to 95°C for 10 min. Heat was slowly reduced to 75°C, and then, the temperature was held at 75, 68, 65, and 62°C for an hour each and held at 60°C for 4 h, and the solution was stored at −20°C. Stem Loop RT-qPCR was done by reverse transcription of 5 ng of total RNA with 1 μl SL primer (1 μM), 0.5 μl dNTPs (10 μM), 5xRT buffer, 2 µl DTT, and 1 µl M-MLV-reverse transcriptase 200 U/µl (Thermo Fisher Scientific) in a final volume of 20 µl. Retrotranscription was done in a thermal cycler by incubation for 30 min at 16°C, followed by pulsed RT of 60 cycles at 30°C for 30 s, 42°C for 30 s, and 50°C for 1 s. After that, the reaction was incubated at 85°C for 5 min. For real-time PCR, 1 µl of the cDNA was used in Biotools Quantimix Easy master mix (Biotools, South San Francisco, CA, USA) with 0.5 μl primer mix (Universal RV and Fw miR specific primers, Fw:Rev, 5 μM:1 μM) in a final volume of 10 μl. Samples were incubated at 95°C for 10 min, followed by cycles of 95°C for 15 s, 60°C for 20 s, and 92°C for 0.5 seg using a Rotor-Gene 6000 System (Corbett, Life Science), and data were analyzed using Rotor-Gene 6000 software (Corbett Life Science; RRID: SCR_017552). The ΔΔCt method was performed using MNs without treatment with exosomes as a negative control and GAPDH mRNA as an exogenous reference control.
For conventional real-time PCR, 100 ng of this total RNA was reverse transcribed using the MLV-reverse transcriptase, 200 U/µl, and 1 µl of the resulting cDNA was diluted in Biotools Quantimix Easy master mix and amplified by PCR over 40 cycles using the Corbett System. Collected data were analyzed using Rotor-Gene software. The ΔΔCt method was performed using MNs without treatment with exosomes as a negative control and GAPDH mRNA as an exogenous reference control. GAPDH was used as an endogenous reference gene because of its well-documented stability and its prior utilization in exosomal miRNA studies (Chen et al., 2022; Gorji-Bahri et al., 2021; Shi et al., 2015; Yu et al., 2020). Moreover, PCR analysis confirmed the presence of GAPDH mRNA in every sample, providing additional evidence to support its suitability as a normalization control.
All reactions were done in 10 μl volume and triplicate in strip tubes (Axygen® Brand Products, San Francisco, CA, USA), using specific forward and reverse primers (Table 1).
List of Primer Sequences of Primers Employed for Real-Time RT-PCR.
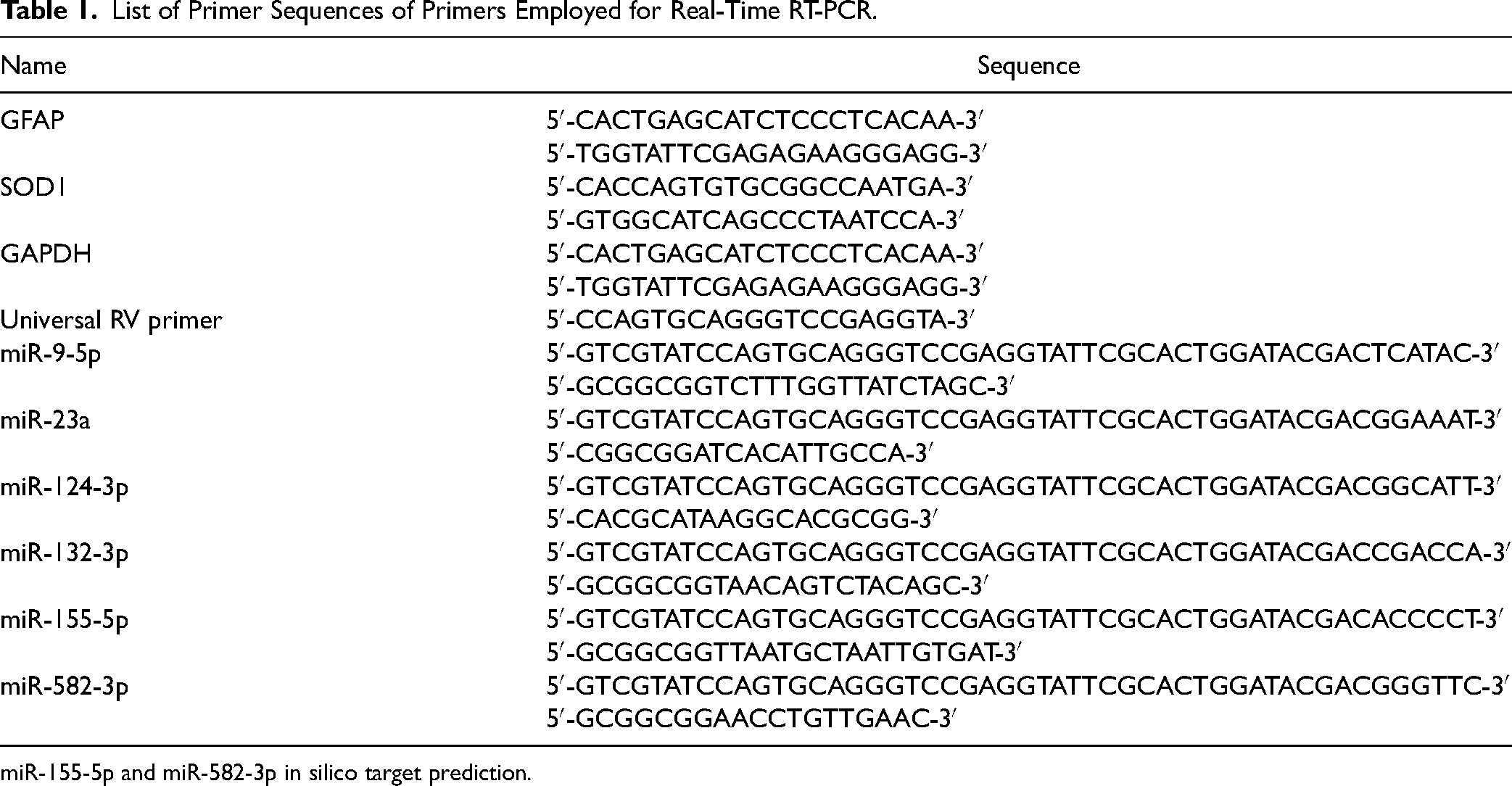
miR-155-5p and miR-582-3p in silico target prediction.
In all cases, a uniform amplification of the products was rechecked by analyzing the melting curves of the amplified products (dissociation graphs), and gel electrophoresis was performed to confirm the correct size of the amplification and the absence of nonspecific bands.
To predict messengers targeted by miR-155-5p or miR-582-3p, we used TargetScan (Agarwal et al., 2015) and DIANA-microT-CDS (Paraskevopoulou et al., 2013; Vlachos et al., 2015). Because we wished to extrapolate our results to humans for its greater accuracy than Rattus norvegicus, we used the Homo sapiens database to predict miRNA target genes. miRNAs predicted targets that were predicted with both algorithms were used in an enrichment analysis with DAVIDv6.8 (Huang et al., 2009; Sherman et al., 2022) online software to identify overrepresented Kyoto Encyclopedia of Genes and Genomes (KEGG) signaling pathways and Gene Ontology (GO) categories (biological process, molecular function, and cellular component). The terms with modified Fisher exact p-value (EASE score) ≤ .05 and false discovery rate (FDR) ≤ 0.05 were considered significantly enriched.
Statistics
All culture assays were performed in duplicate, and each experiment was repeated at least three times. Quantitative data were expressed as mean ± standard error of the mean (SEM). Paired one- or two-tailed t-tests were performed to compare data with two experimental groups. A two-way analysis of variance (ANOVA) followed by Fisher's LSD test for multiple comparisons or a one-way ANOVA followed by Tukey's multiple comparisons test was performed as indicated using GraphPad Prism version 8.0.0 for Windows, GraphPad Software, San Diego, CA, USA. A value of p ≤ .05 was considered statistically significant.
Results
Exosomes are Obtained From Astrocyte CM
After successive centrifugations of CM from astrocyte primary cultures, we obtained a pellet fraction from 100,000 × g ultracentrifugation (p100). This fraction was analyzed by TEM following negative staining. The micrographs showed several arrays of round-shaped vesicles. Around 77% of the total vesicles measured were between 60 and 100 nm in diameter with a median of 100.2 nm, which is in the range reported for exosome size (Raposo & Stoorvogel, 2013) (Figure 1(a) and (b)). To further characterize this fraction, the presence of the exosome-specific proteins flotillin-1 (Thery et al., 1999) and the component of the endosomal sorting complex responsible for transport 1 (ESCRT-I): tumor susceptibility gene 1 (TSG101) (De Gassart et al., 2003) were assayed by western blotting. Flotillin-1 and TSG101 were detected in all p100 fractions analyzed (Figure 1(c)). Moreover, the exosomes were negative for BiP, which is a protein found in the endoplasmic reticulum (Supplemental Figure 1). These findings suggest that the samples are enriched in exosomes and do not contain other subcellular components. In addition, the presence of the human SOD1 was analyzed in the same fraction as had been previously shown (Basso et al., 2013). As expected, human SOD1 was present exclusively in SOD1G93A – but not in the non-Tg-derived exosomes.
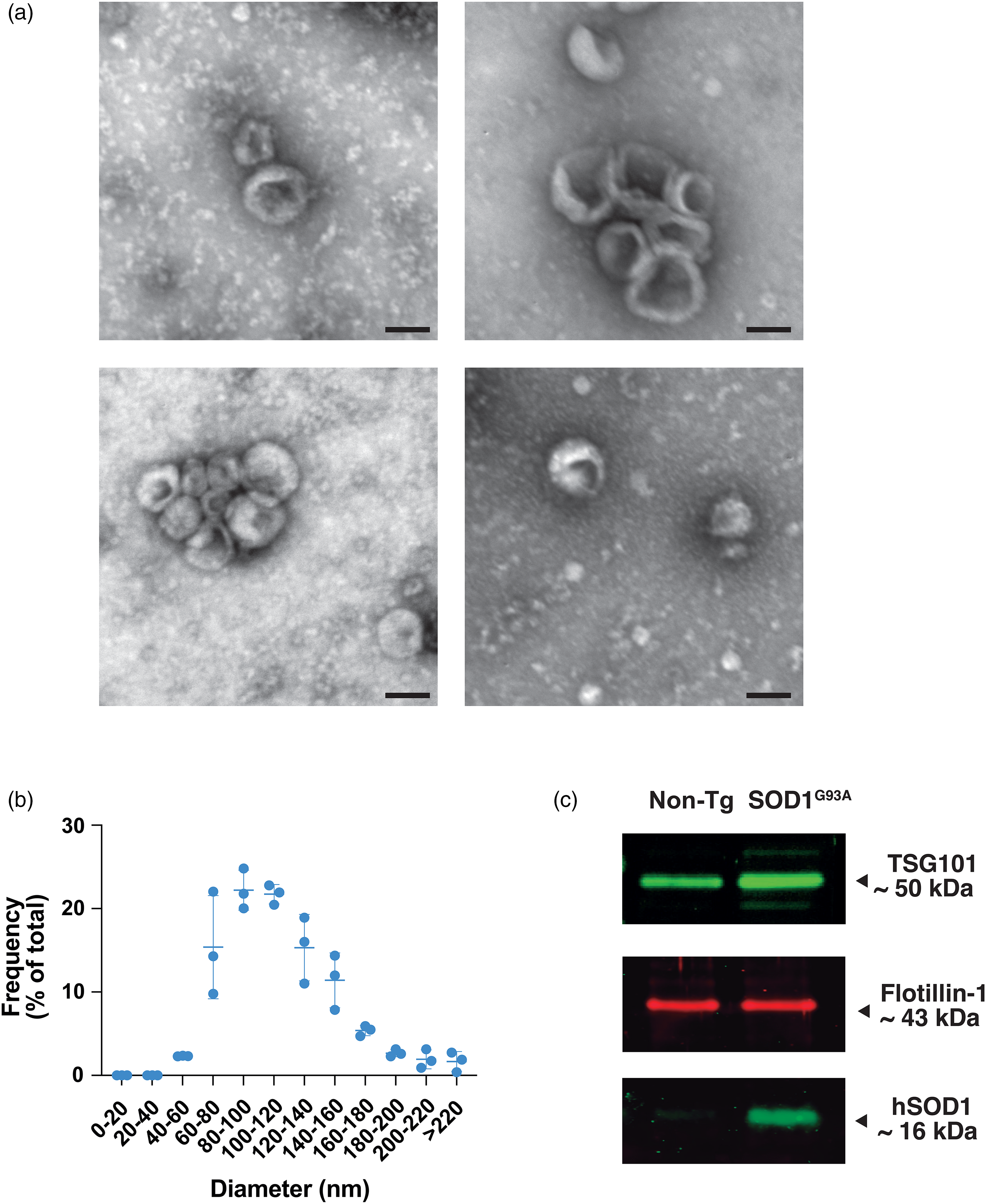
Morphological characterization and size distribution of extracellular vesicles present in astrocyte CM. (a) TEM micrographs (negative staining) from extracellular vesicles isolated from astrocyte CM. Bars: 100 nm. (b) Frequency distribution of EV diameter as a percentage of the total number of vesicles from the three independent experiments. (c) Representative western blot of the exosomes from non-Tg or SOD1G93A astrocyte CM using antibodies against TSG101, flotillin-1, and human SOD1.
SOD1G93A Astrocytes-Derived Exosomes Exhibit the Characteristic RNA Size Profile and No Detectable Levels of Human SOD1G93A mRNA
Previous reports indicate that exosomes contain substantial amounts of RNA (Valadi et al., 2007). The electropherogram profile of total RNA from the astrocyte-derived exosomes was compared to the total RNA extracted from cultured astrocytes and analyzed (Figure 2(a)). As described, the RNA extracted from exosomes included a broad range of RNA sizes (30–500 nt), with undetectable levels of 18S and 28S ribosomal RNAs (rRNAs). However, these ribosomal RNAs were present in RNA extracted from cultured astrocytes. The presence of human SOD1 (hSOD1) mRNA in exosomes of SOD1G93A astrocytes was analyzed by RT-PCR. Although SOD1G93A protein was detected in the exosomes of SOD1G93A astrocytes, hSOD1 mRNA was not identified in any of the analyzed samples (Figure 2(b)).
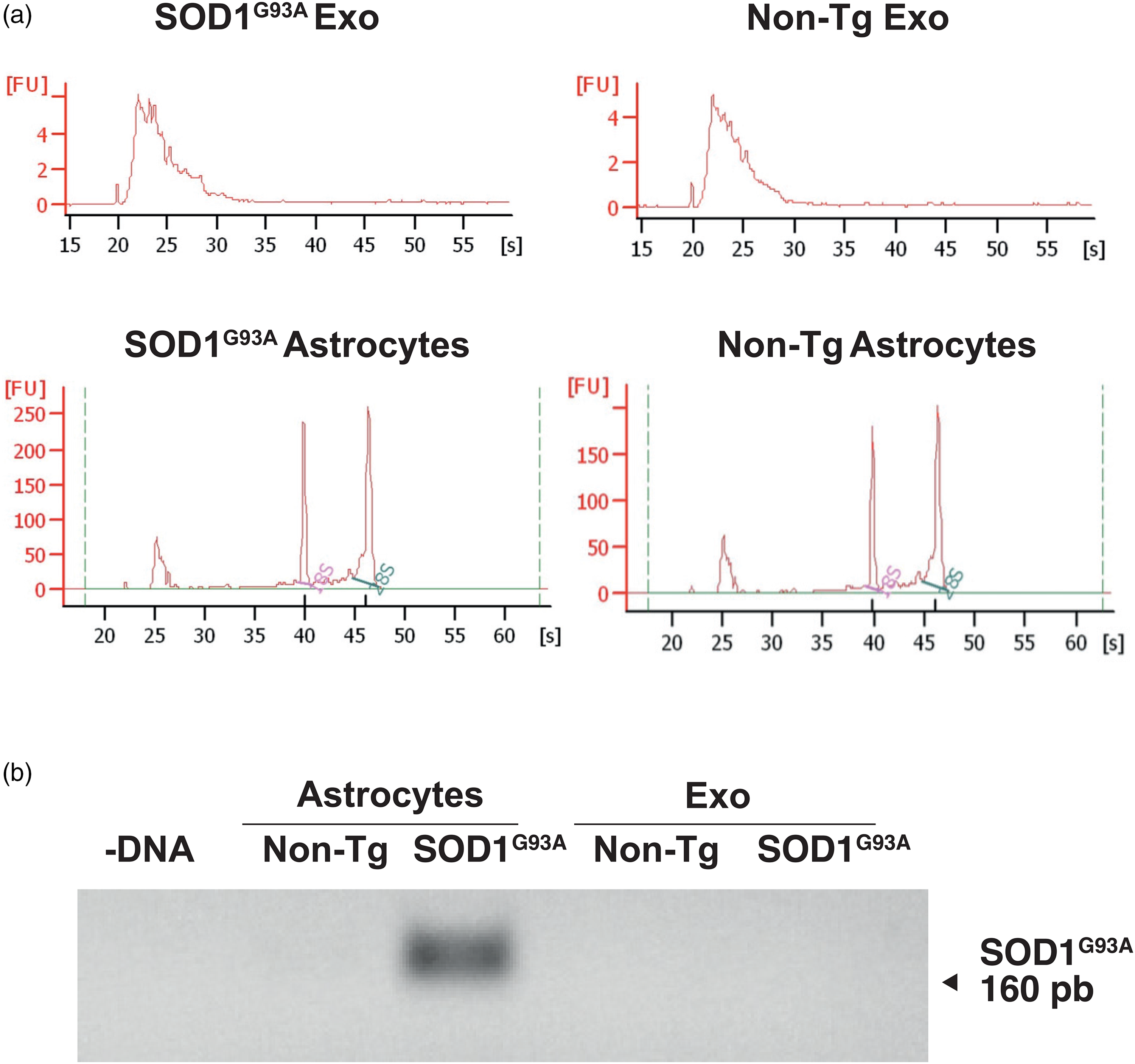
Human SOD1G93A mRNA is undetectable in exosomes of SOD1G93A astrocytes. (a) Electropherogram results of total RNA extracted with TRIzol. (b) SOD1G93A mRNA expression in non-Tg and SOD1G93A astrocytes and astrocyte-derived exosomes, as assessed by RT-PCR.
SOD1G93A Astrocyte-Derived Exosomes are Sufficient to Reduce the Survival of MNs and the Length of Neurites
To address the role of SOD1G93A astrocyte-derived exosomes on the ability of astrocytes to promote the survival of MNs, we compared the effects of non-Tg or SOD1G93A astrocytes CM versus exosomes on purified non-Tg MN cultures exposed to GDNF (Figure 3). CM from SOD1G93A astrocytes (1:10 dilution) significantly reduced the survival of MNs exposed to GDNF, compared to CM from non-Tg astrocytes, as previously reported (Figure 3(a) and (b)) (Fritz et al., 2013; Madill et al., 2017; Nagai et al., 2007). The incubation of pure MN cultures with SOD1G93A astrocyte exosomes induced a significant reduction in MN survival in the presence of GDNF (25.6% ± 10.6), while no such effect was detected when non-Tg exosomes were applied to MN cultures (Figure 3(a) and (b)). The reduced ability of SOD1G93A astrocyte CM to support MN survival was not present when EVs were depleted from SOD1G93A astrocytes CM (Figure 3(b)).
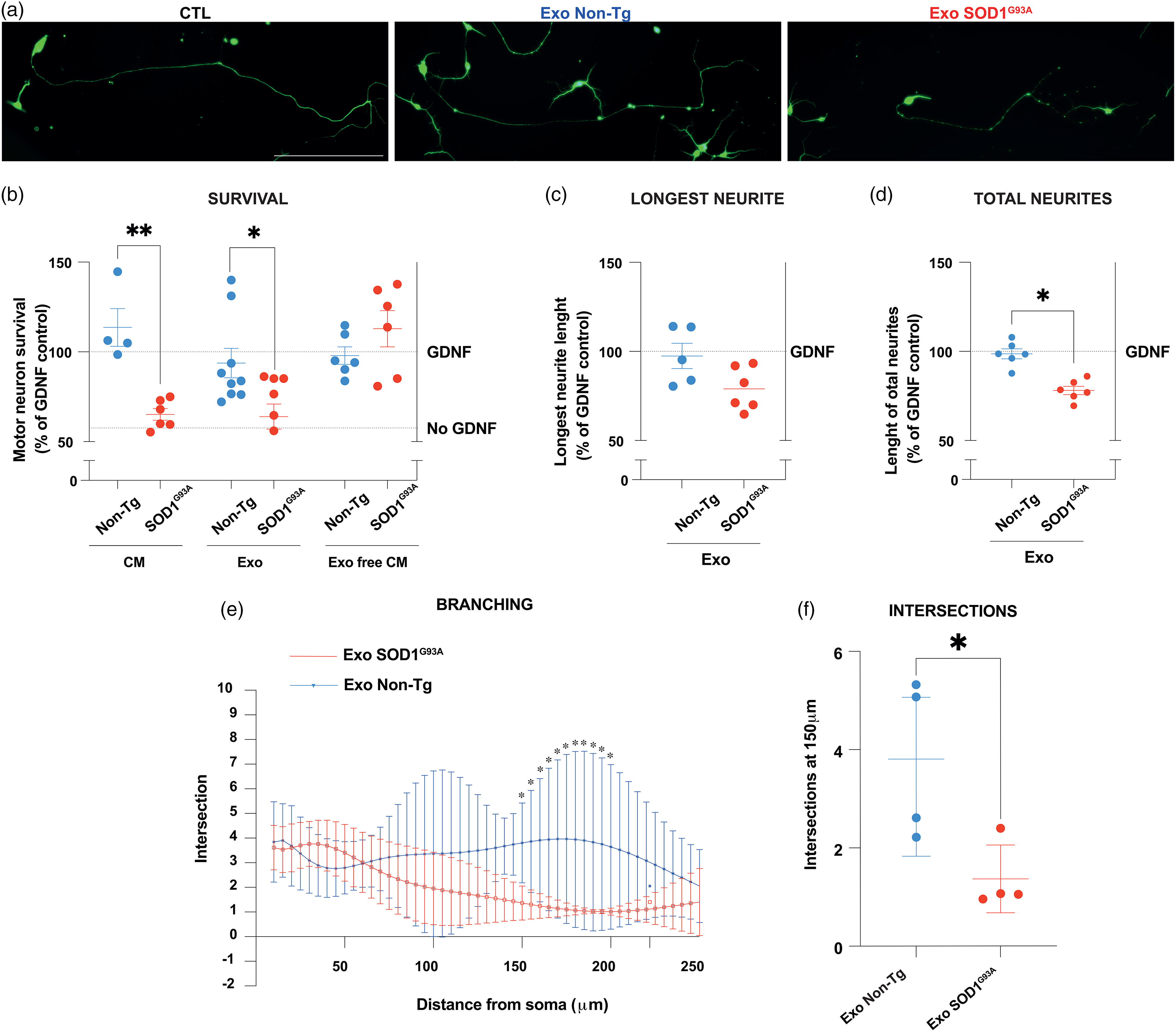
SOD1G93A astrocyte-derived exosomes reduce MN survival, neurite length, and neurite branching. (a) Representative images of primary cultures of MNs treated with GDNF (1 ng/ml, CTL) or with GDNF and exosomes from non-Tg or SOD1G93A astrocytes. MNs were immunostained with beta III tubulin, scale bar = 250 μm. (b) MN survival after treatment with GDNF and following the addition of the indicated CM (non-Tg and SOD1G93A), exosomes (Exo; non-Tg and SOD1G93A) or CM depleted of exosomes (Exo free CM; non-Tg and SOD1G93A). Data are expressed as a percentage of survival after GDNF addition (100%, upper dotted line) compared to MN survival with no addition of trophic factor (lower dotted line). Unpaired t-tests were carried out, and the levels of significance were represented as *p < .05, **p < .01. (c) and (d) Longest neurite length and total neurite length plot. MNs were treated with GDNF (control) and with GDNF and exosomes from nontransgenic and SOD1G93A astrocytes (Exo; non-Tg and SOD1G93A). At least seven neurites per treatment were quantified with the Fiji program. Data are expressed as the mean ± SEM from five to six independent experiments (GDNF, Exo; non-Tg and SOD1G93A). Unpaired t-tests were performed, and significance levels were denoted as unpaired t-tests, *p < .05. (e) Representative images of Sholl graphs of MNs treated with exosomes. Mean ± SEM from at least three independent experiments. A two-way ANOVA followed by Fisher's LSD test for multiple comparisons was performed. p < .05 was used to determine statistical significance. (f) Intersections at 150 μm from the soma. An unpaired t-test was used to analyze the significant branching of neurites at 150 μm. An unpaired t-test was executed, and the significance levels were indicated as follows: *p < .05. Experiments performed with non-Tg astrocyte fractions are depicted in blue, and those from SOD1G93A ones are in red.
In addition, the effect of the exosomes on the length and branching of MN neurites was assessed. Neurite length was significantly decreased in MNs treated with SOD1G93A astrocyte-derived exosomes compared to non-Tg ones following GDNF exposure (Figure 3(c) and (d)). There was a 21 ± 12% and a 22 ± 6% decrease in the longest neurite (Figure 3(c)) and total neurite length (Figure 3(d)), respectively. The morphology of motoneuron arborizations, as revealed by the Sholl analysis, is characterized by few neurites emerging from the soma and usually the longer one branches near the end. The length of the longest neurite varies among cells. Neurite branching showed an overall reduced complexity in MNs treated with exosomes from SOD1G93A astrocytes significant at 150 μm from the soma (Figure 3(e)). SOD1G93A astrocyte exosome-mediated effects were abolished when exosome membrane integrity was disrupted or membrane proteins were digested. MNs incubated with exosome preparations that were subjected to osmotic shock or trypsin digestion induced no significant changes in the longest neurite or total neurite length (data not shown).
miR-155-5p and miR-582-3p are Dysregulated in SOD1G93A Astrocyte Exosomes
Because exosomes can transport miRNAs that are functionally transferred to other cells (Lopez-Verrilli et al., 2013; Record et al., 2014) and miRNA profile is altered in ALS, we assessed whether miRNAs that have been reported to show modified expression in ALS (Paez-Colasante et al., 2015) were differentially expressed in SOD1G93A astrocyte exosomes (Table 2).
Analyzed miRNAs in Astrocytes or Astrocyte-Derived Exosomes.
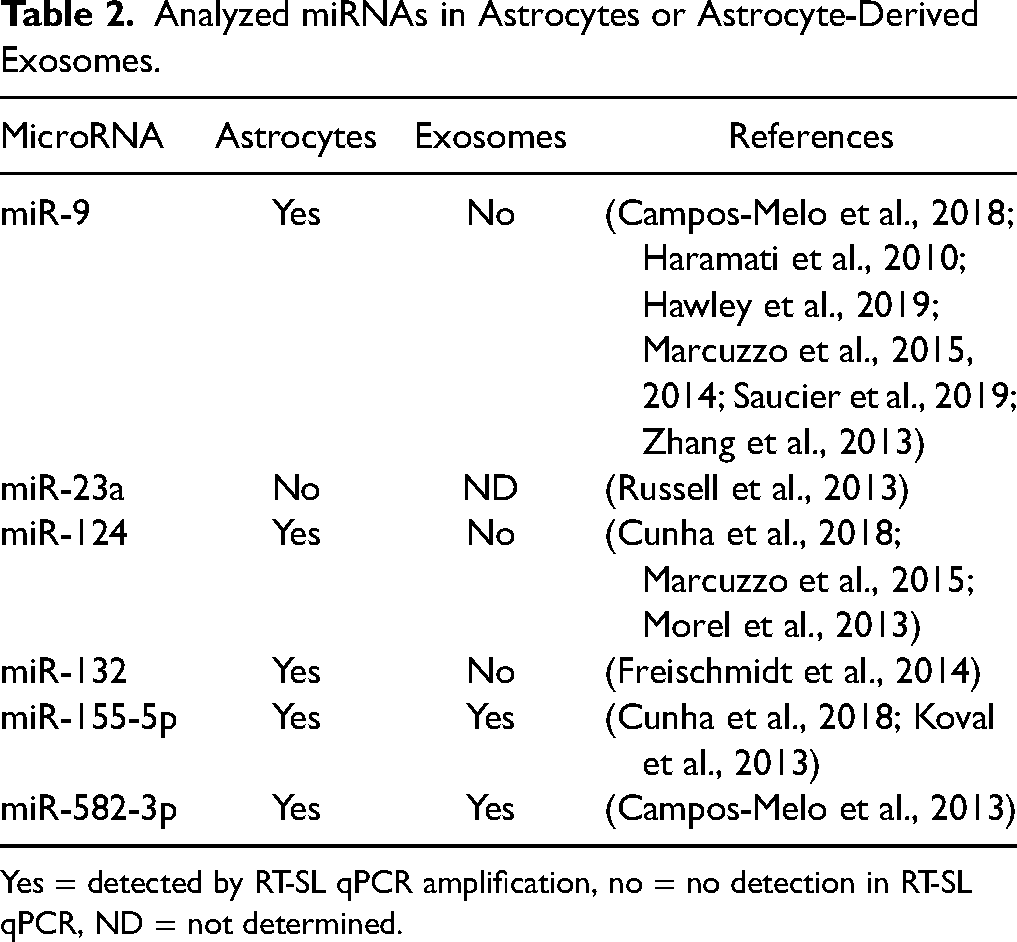
Yes = detected by RT-SL qPCR amplification, no = no detection in RT-SL qPCR, ND = not determined.
Most of the analyzed miRNAs were expressed in cultured whole astrocytes, but only miR-155-5p and miR-582-3p were also detected in the exosomes (Table 2). Notably, there were significant differences in the gene expression levels of miR-155-5p and miR-582-3p between SOD1G93A astrocyte exosomes and non-Tg exosomes (Figure 4). Specifically, miR-155-5p was upregulated, while miR-582-3p was downregulated in SOD1G93A astrocyte exosomes (Figure 4(b) and (d)). Interestingly, the expression levels of miR-155-5p were significantly lower in whole SOD1G93A astrocytes than in non-Tg ones (Figure 4(a)).
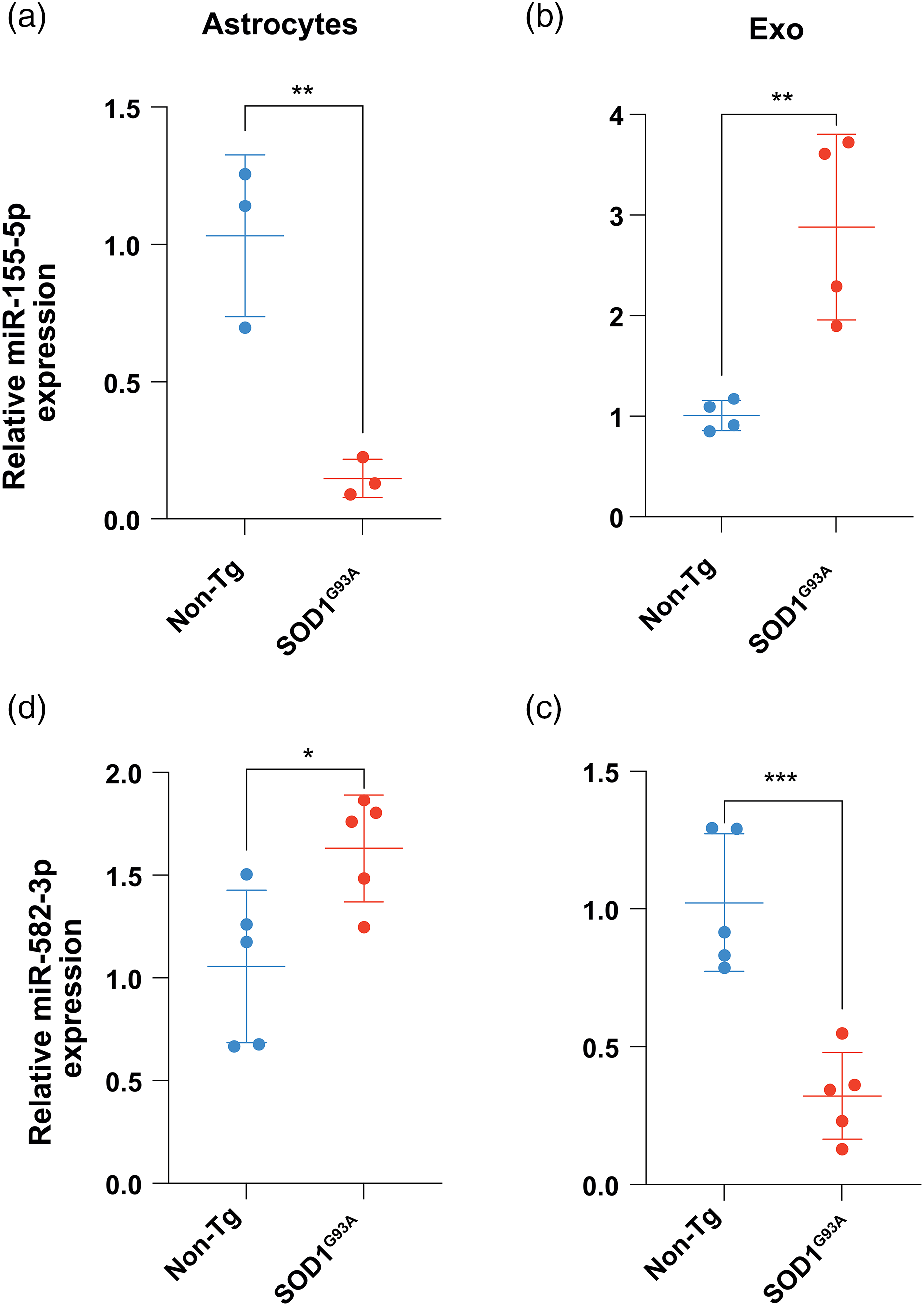
miR-155-5p and miR-582-3p show different expression levels in SOD1G93A and non-Tg astrocyte exosomes. (a) and (c) Relative expression levels of miR-155-5p and miR-582-3p in astrocytes. (b) and (d) Relative expression levels of miR-155-5p and miR-582-3p in astrocyte exosomes. qRT-PCR quantification was performed with the ΔΔCt method using non-Tg samples as negative control and GAPDH mRNA as an endogenous reference control. Data are the mean ± SEM from at least three independent experiments. Unpaired t-tests were performed, and significance levels were denoted as *p < .05, **p < .01, ***p < .005.
KEGG Analysis Shows That miR-155-5p and miR-582-3p Predicted Targets are Enriched in Neurotrophin Signaling Pathway Genes
To gain insights into miR-155-5p and miR-582-3p miRNA regulatory mechanisms, we searched for potential gene targets and associated signaling pathways using TargetScan (Agarwal et al., 2015) and DIANA-microT-CDS (Paraskevopoulou et al., 2013; Vlachos et al., 2015) bioinformatic platforms (Figure 5).
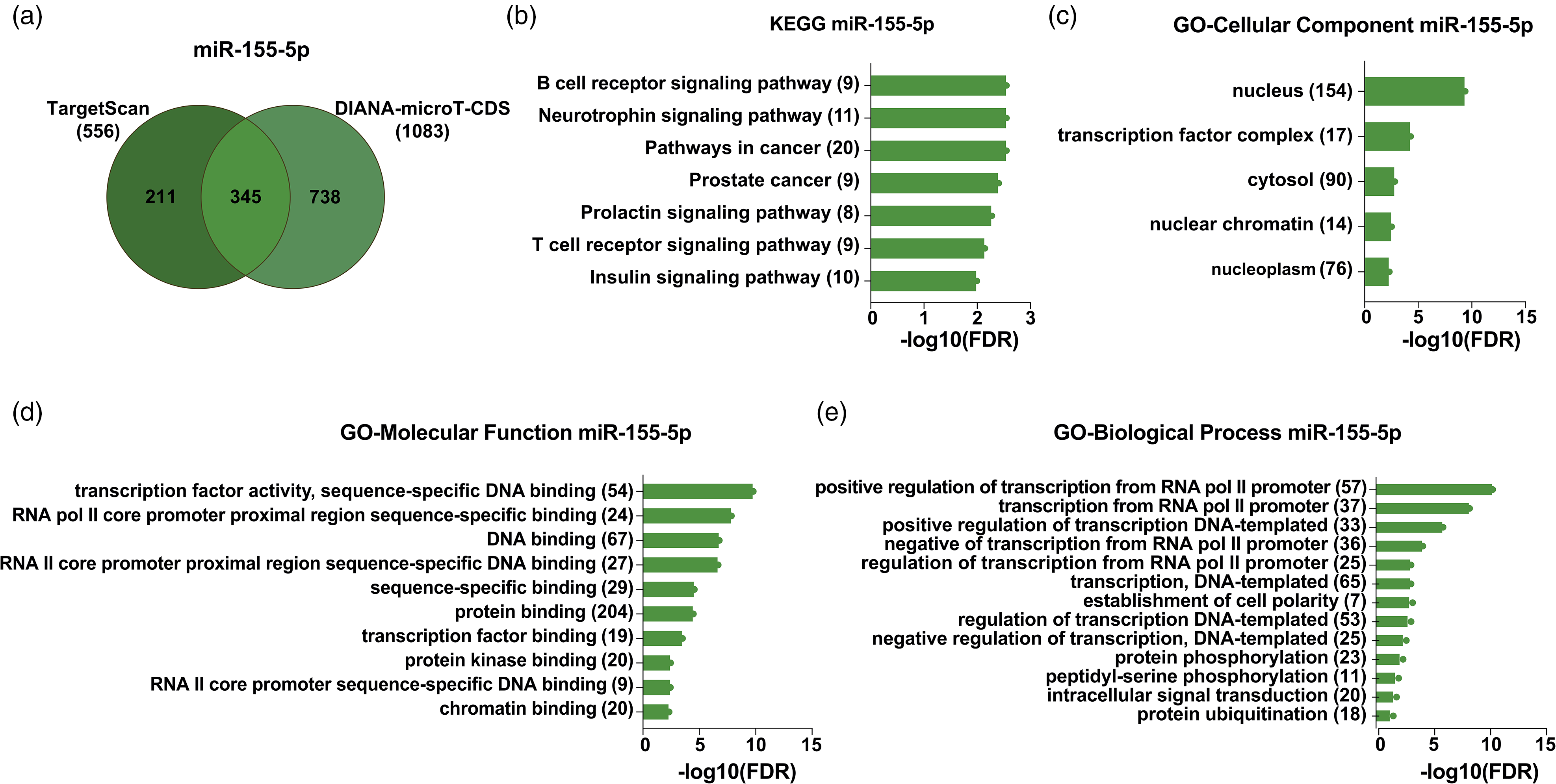
Enrichment analysis of miR-155-5p predicted targets. (a) Venn diagram of miR-155-5p predicted targets with TargetScan and DIANA-microT-CDS. (b) Enrichment analysis of KEGG pathways (123), (c) cellular component (322), (d) molecular function (309), and (e) biological process (309) in terms of miR-155-5p predicted targets. The x-axis represents the enrichment value as a −10 logarithm of the adjusted p-value (FDR). The number of genes identified in each pathway is reported inside parentheses. All represented KEGG and GO terms are significantly enriched, with an adjusted p-value < .05 and FDR < 0.05. KEGG = Kyoto Encyclopedia of Genes and Genomes.
For miR-155-5p, 556 predicted target genes were found with TargetScan and 1,082 with DIANA-T-CDS. In total, 345 targets were predicted at the intersection of both algorithms (Figure 5(a)). Out of these, 341 were identified in the DAVID database and used in GO enrichment analyses (Figure 5(a) to (d)). The biological processes category exhibited a noteworthy enrichment of genes primarily associated with regulating transcription, intracellular signal transduction, and protein ubiquitination (Figure 5(e)). Genes within the molecular function category exhibited enrichment in DNA binding, protein binding, and chromatin binding, as indicated in Figure 5(d). In the cellular component enrichment analysis, most of the target genes were associated with the nuclear compartment except for a reduced but significant set that localized to the cytosol (Figure 5(c)). Additionally, 123 genes were detected in KEGG pathway enrichment analysis, most of them related to signaling pathways. Neurotrophin signaling (KEGG: 04722, with a p-value of 0.00005 and FDR of 0.003) was among the top five enriched pathways (Figure 5(b)). Furthermore, we found that key factors such as PI3 K, Ras, or NFκB of neurotrophin signaling were putatively regulated by miR-155-5p (Supplemental Figure 2), which could be the effect observed on MN survival and neurite length.
On the other hand, miR-582-3p predicted target gene analyses identified 730 putative targets with TargetScan and 3,310 with DIANA-microT-CDS software, of which 628 were predicted with both algorithms (Figure 6(a)). The enrichment analysis showed no enrichment in the biological process category, only one term in molecular function, protein binding (FDR = 0.00003), and several enriched terms in the cellular component category. KEGG pathway enrichment analysis only identified the neurotrophin pathway as a pathway enriched in miR-582-3p target genes (Figure 6(b) to (d)).
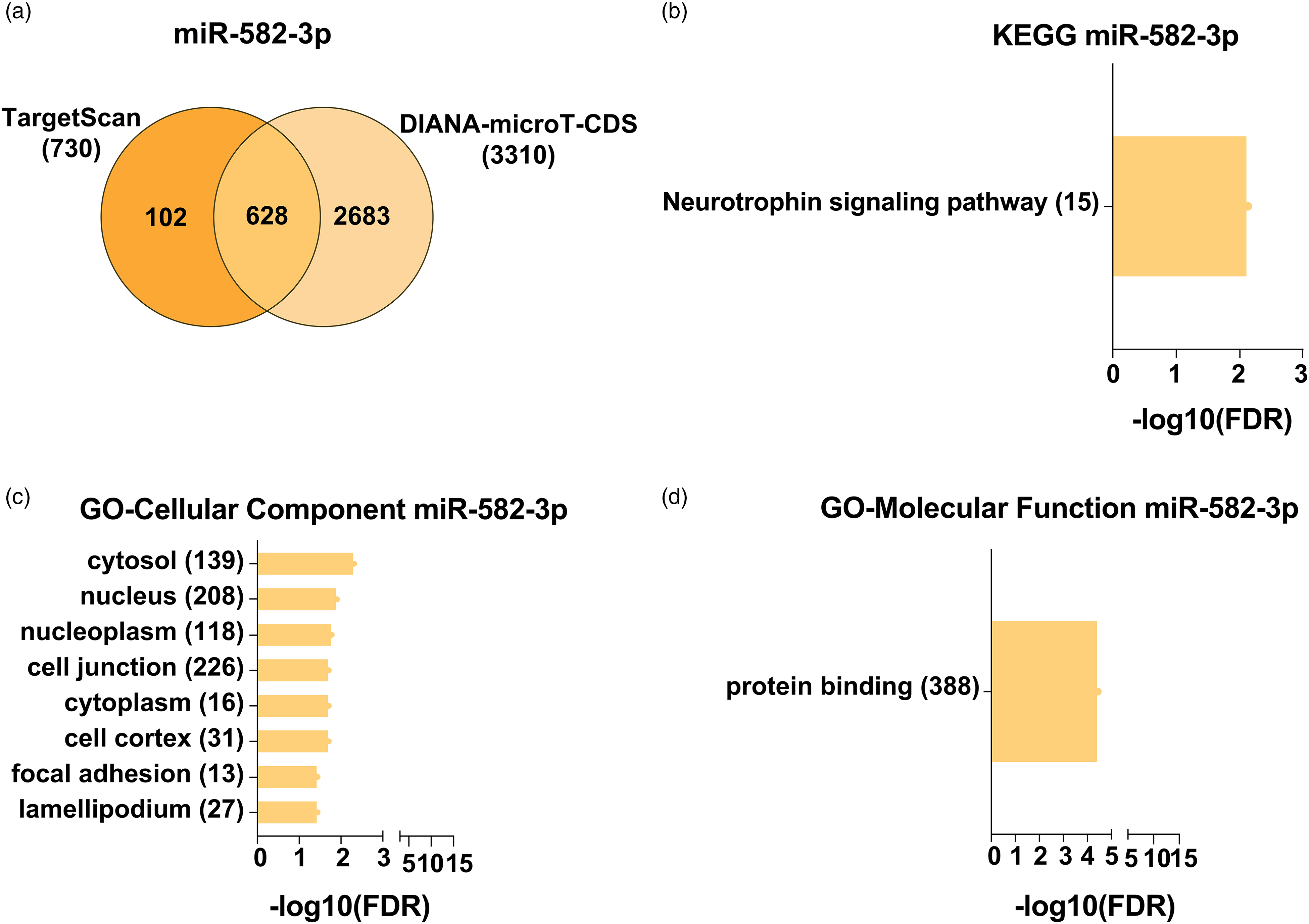
Enrichment analysis of miR-582-3p predicted targets. (a) Venn diagram of miR-582-3p predicted targets with TargetScan and DIANA-microT-CDS. (b) Enrichment analysis of KEGG pathways (199), (c) cellular component (538), and (d) molecular function (526), in terms of miR-582-3p predicted targets. The x-axis represents the enrichment value as the −10 logarithm of the adjusted p-value (FDR). The number of genes identified in each pathway is reported inside parentheses. All represented KEGG and GO terms are significantly enriched, with an adjusted p-value < .05, and FDR< 0.05. KEGG = Kyoto Encyclopedia of Genes and Genomes.
SOD1G93A Astrocyte-Derived Exosome-Mediated MN Death Was Reduced by Depletion of miR-155-5p
Due to the overexpression of miR-155-5p in SOD1G93A exosomes and its involvement in neurotrophin signaling pathways with more putative targets compared to miR-582-3p, we conducted further analysis to investigate the role of miR-155-5p in MN death induced by SOD1G93A astrocyte exosomes. Primary MNs were incubated with astrocyte exosomes (non-Tg or SOD1G93A) together with miR-155-5p mimics or miR-155-5p inhibitors, and cell survival and neurite branching were analyzed (Figure 7(a)). Interestingly, when MNs were treated with non-Tg astrocyte-derived exosomes in the presence of a miR155-5p mimetic, a reduction in MN survival was observed (Exo-non-Tg-miR, 64.16 ± 20.07%). Importantly, when MNs were treated with exosomes of SOD1G93A astrocytes and transfected with oligonucleotides that inhibit miR-155-5p, MN death induced by SOD1G93A astrocyte exosomes was not detected (Exo-SOD1G93A-αmiR, 50.51 ± 19.42%; Exo-SOD1G93A 104.45 ± 21.61%; Figure 7(a)). Conversely, when MNs were treated with exosomes of non-Tg astrocytes in the presence of miR155-5p inhibitors or SOD1G93A astrocytes in the presence of miR-155-5p mimetics, no differences were found compared to those with the respective exosomes alone (Exo-non-Tg-αmiR 116.98 ± 2.86%; Exo-SOD1G93A-miR 16.31 ± 5.50%). In addition, no significant effect on MN survival was detected when adding either the miRNA or the antisense miRNA alone (miR, 70.24 ± 32.06%; αmiR 155-5p, 92.74 ± 80.97% compared to MNs treated with non-Tg astrocyte exosomes). Interestingly, MN branching was increased when MNs were treated with SOD1G93A astrocyte-derived exosomes in the presence of miR-155-5p inhibitors compared to the one developed when treated with the SOD1G93A astrocyte-derived exosomes alone (Figure 7(b) and (c)).
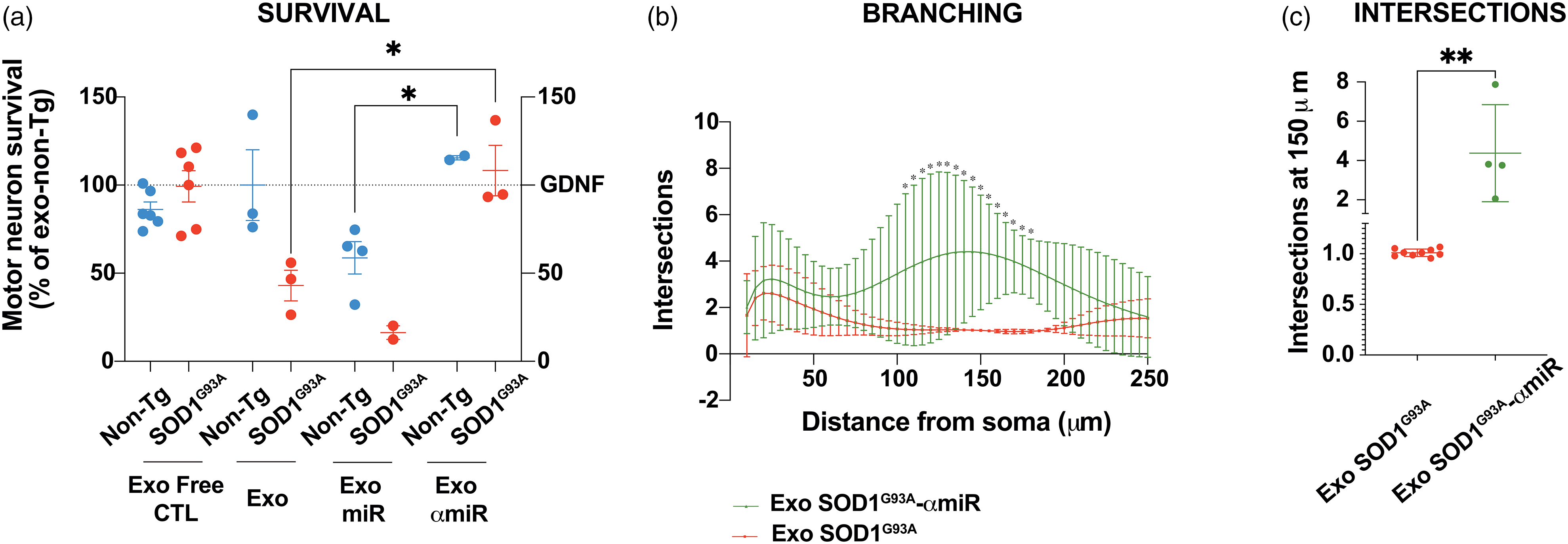
Inhibition of miR-155-5p rescue MN death induced by SOD1G93A astrocyte exosomes. MN survival 24 h after incubation with SOD1G93A (red) or non-Tg (blue) exosomes plus the miR155-5p mimic or inhibitor as indicated. Data show the percentage of MNs compared to control (MNs treated with non-Tg astrocyte exosomes). 24 h primary MN cultures were treated with non-Tg CM depleted of exosomes from non-Tg and SOD1G93A astrocytes (Exo free CM; non-Tg, and SOD1G93A, respectively), or with exosomes of non-Tg and SOD1G93A astrocytes (Exo; non-Tg and SOD1G93A, respectively), cotransfected with miR-155-5p mimetics (Exo miR; non-Tg or SOD1G93A) or miR-155-5p inhibitors (α-miR; non-Tg and SOD1G93A). Mean ± SEM from at least three independent experiments. One-way ANOVA results: *p < .05, from non-Tg treated control. (b) Representative images of Sholl graphs of MNs treated with exosomes from SOD1G93A astrocytes (Exo-SOD1G93A) in the absence or presence of miR-155-5p inhibitors (Exo-SOD1G93A-αmiR). Mean ± SEM from at least three independent experiments. A two-way ANOVA followed by Fisher's LSD test for multiple comparisons was performed. p < .05 was employed to determine statistical significance. (c) Intersections at 150 μm from the soma. An unpaired t-test was used to analyze the significant branching of neurites at 150 μm. Experiments performed with Exo-SOD1G93A astrocytes are depicted in red, and those from Exo-SOD1G93A-αmiR ones are in green. An unpaired t-test was conducted, and the results were denoted as *p < .05, **p < .01, and ***p < .005.
Discussion
In the present work, we demonstrate that SOD1G93A astrocyte-derived exosomes are sufficient to reduce neurite outgrowth and survival of MNs in the presence of GDNF to the level exhibited upon trophic factor deprivation. In addition, we provide evidence that miRNA 155-5p contributes to SOD1G93A exosome-mediated MN death.
The fraction isolated from CM of primary SOD1G93A and non-Tg astrocyte cultures contained round-shaped vesicles of 60–100 nm diameter and carried flotillin-1 and ESCRT-I complex component, TSG101, consistent with previously reported exosome features (De Gassart et al., 2003; Thery et al., 1999). Following prior findings, the exosomes derived from SOD1G93A astrocytes contained human SOD1 protein (Basso et al., 2013; Silverman et al., 2019). Exosomes containing mutated or misfolded SOD1 are reported to be secreted by MNs, astrocytes, and microglia (Basso et al., 2013; Gomes et al., 2007; Massenzio et al., 2018; Vaz et al., 2019). Mutated or misfolded SOD1 decorates the surface of exosomes and can be transferred to healthy cells (Silverman et al., 2019). In addition, proteins associated with ALS such as VCP, TDP43, FUS, other RNA-binding proteins, and dipeptide repeats (DPRs) resulting from C9orf72 expansions have been described to be present in exosomes derived from cells overexpressing these proteins (Basso et al., 2013; Iguchi et al., 2016; Nonaka et al., 2013). While some researchers have suggested that this could be a mechanism of maintaining cell survival by conveying toxic materials including misfolded proteins out of the cell (Desdín-Micó & Mittelbrunn, 2017), others have suggested that exosomes may propagate toxicity in ALS (Basso et al., 2013; Ding et al., 2015; Grad & Cashman, 2014; Westergard et al., 2016). This work shows that the toxic effects of SOD1G93A astrocyte-derived exosomes on MNs could be related to miRNA cargo.
Earlier studies have indicated that astrocytes derived from mutant SOD1 mice (Nagai et al., 2007) and rats (Vargas et al., 2006) can induce MN death in vitro by releasing soluble specific factors present in CM. Numerous agents have been suggested to participate in this toxicity, including nitric oxide, ATP (Almad et al., 2016; Gandelman et al., 2010), lipids (Guttenplan et al., 2021), or proteins such as pro–nerve growth factor (proNGF) (Pehar et al., 2017) and amyloid precursor protein (Mishra et al., 2020). However, the exact mechanism of MN death and the nature of these toxic factors remain elusive and subject to ongoing debate (Van Harten et al., 2021). In this context, SOD1G93A and non-Tg astrocyte-derived exosomes are taken up by cultured MNs and induce MN death in vitro when applied to non-Tg spinal neuron-astrocyte cocultures (Basso et al., 2013). Moreover, SOD1 astrocyte-derived exosomes have been described to induce MN death in vitro when applied to non-Tg spinal neuron–astrocyte cocultures (Basso et al., 2013), and EVs from astrocytes derived from fibroblasts of C9orf72 ALS patients are also toxic to MNs (Varcianna et al., 2019). Here, we show that the SOD1G93A astrocyte-derived exosomes are sufficient to induce MN death and reduce neurite outgrowth in purified MN cultures, suggesting that these exosomes could contribute to astrocyte-mediated MN toxicity.
Basso et al. showed that the SOD1G93A protein carried by exosomes is transferred to spinal neurons while the SOD1G93A present in exosome-free astrocyte CM is undetectable inside spinal neurons (Basso et al., 2013). Besides protein content, exosomes are reported to carry important RNA species where they exhibit increased stability and protection from RNase (Ratajczak et al., 2006; Valadi et al., 2007) and may also affect neuronal survival. The analysis of exosomal RNA cargo revealed a broad range of RNA sizes in non-Tg and SOD1G93A astrocyte-derived exosomes. As previously described for exosomal RNAs (Valadi et al., 2007), the sizes of 18S and 28S ribosomal RNAs in astrocyte exosomes were undetectable, but they were present in astrocyte cells. We focused on miRNAs because of previous reports suggesting their involvement in ALS (Campos-Melo et al., 2013; De Felice et al., 2012; Emde et al., 2015; Hoye et al., 2017; Koval et al., 2013; Zhou et al., 2013) and its essential role in gene expression regulation. Moreover, some miRNAs were reported to be altered in ALS such as miR-9, miR-23a, miR-124, miR-132, miR-155-5p, and miR-582-3p (Campos-Melo et al., 2013, 2018; Cunha et al., 2018; Freischmidt et al., 2014; Haramati et al., 2010; Hawley et al., 2019; Koval et al., 2013; Marcuzzo et al., 2015, 2014; Morel et al., 2013; Russell et al., 2013; Saucier et al., 2019; Zhang et al., 2013).
These miRNA species, except for miR-23a, were identified in cultured astrocytes. In addition, previous studies have reported that these miRNAs, except for miR-132, are deregulated in the spinal cords of ALS patients with mutations in SOD1, TARDBP, FUS/TLS, expanded repeats on C9ORF72 (Butovsky et al., 2012; Campos-Melo et al., 2018; Freischmidt et al., 2014; Hawley et al., 2019; Koval et al., 2013; Zhang et al., 2013), or SOD1G93A murine models (Cunha et al., 2018; Marcuzzo et al., 2015, 2014; Morel et al., 2013). miR-132, on the other hand, is downregulated in cerebrospinal fluid and serum of sporadic ALS cases with mutations in TARDBP, FUS, and C9ORF72, but not SOD1 mutant patients (Freischmidt et al., 2014). Interestingly, only miR-155-5p and miR-582-3p were observed in astrocyte-derived exosomes, and quantitative PCR of miR-155-5p and miR-582 showed different miRNA expression levels in SOD1G93A astrocytes relative to non-Tg cells. Moreover, miRNA levels of miR-155-5p and miR-582-3p in astrocyte-derived exosomes are different than in astrocytes. This agrees with data reported in the literature showing that exosome miRNA expression profiles are functionally different from those of the parent cells (Zhang et al., 2015). Particularly, significant differences were observed in miRNA profiles between mouse astrocyte-derived exosomes compared to whole-cell astrocytes (Jovicic & Gitler, 2017).
Concerning both miRNAs present in astrocyte exosomes, this is the first report of miR-582-3p levels in astrocytes or astrocyte EVs. Notably, it is downregulated in the spinal cord of ALS patients with mutations in SOD1, TARDBP, FUS/TLS, or expanded repeats on C9ORF72 (Campos-Melo et al., 2013). The downregulation of miR-582-3p found in the exosomes of SOD1G93A astrocytes could be relevant in pathological spreading in ALS due to its regulatory role in physiologically relevant mRNA expression (Campos-Melo et al., 2013; Qin et al., 2018). On the other hand, miR-155 has been observed to be overexpressed in the spinal cord of familial and sporadic ALS patients as well as in the spinal cord of SOD1G93A mice at presymptomatic and symptomatic stages and has been suggested as an early marker of disease (Cunha et al., 2018). However, no differences were reported in miR-155-5p expression levels at the cerebral cortex between SOD1G93A and non-Tg mice (Gomes et al., 2019). In addition, miR-155-5p was detected in small EVs derived from astrocytes (Gomes et al., 2020). In this study, we observed a downregulation of miR-155-5p in cultured SOD1G93A astrocytes, while there was an upregulation of miR-155-5p in the SOD1G93A astrocyte exosomes. miR-155-5p has numerous suggested and validated targets (Bayraktar et al., 2015). Its synthesis is stimulated by proinflammatory cytokines such as TNF-α, which activates NF-κB transcription factor that promotes miR-155-5p biogenesis (Lee et al., 2014; Kim et al., 2017). In astrocytes, it is upregulated after IL-1 or TNFα cytokine stimulation, leading to a neurotoxic astrocyte phenotype (Tarassishin et al., 2011).
In silico target analysis identified 341 putative targets for miR-155-5p and 628 for miR-582-3p. In both cases, pathways of targeted mRNAs identified by KEGG analysis revealed an increment in neurotrophin signaling pathway-related genes. Given that miR-155-5p showed a higher number of putative targets than miR-582-3p, our study focused on exploring the implications of miR-155-5p. As for miR-582-3p, further investigations will be carried out to elucidate its involvement in astrocyte-mediated MN death.
Our study reveals that inhibiting miR-155 effectively decreased the death of MNs induced by SOD1G93A astrocyte-derived exosomes, resulting in similar survival rates to MNs treated with exosomes from nontransgenic (non-Tg) astrocytes. Additionally, we observed a significant enhancement in the growth and branching of MN neurites. This result is consistent with the KEGG analysis, which identified an increase in neurotrophin signaling pathways. Neurotrophins are trophic factors that promote neuronal survival and development and significantly regulate axonal and dendritic growth (Park & Poo, 2013). Furthermore, these findings align with previous research showing that primary cultures of dorsal root neurons from miR-155 knockout (KO) mice displayed an approximate 40% increase in neurite length, indicating enhanced arborization (Gaudet et al., 2016). Following peripheral injury, miR-155-5p KO mice show higher regenerative levels than those observed in control mice. These results are associated with an improvement in motor skills after nerve contusion (Gaudet et al., 2016). In addition, inhibition of miR-155-5p by intraventricular administration of anti-miR-155 in SOD1G93A mice significantly extended survival (Koval et al., 2013). Our results provide further evidence supporting the toxic activity of miR-155-5p toward MNs. Nonetheless, additional research is required to determine whether neurotrophin signaling pathway-related genes undergo modifications in recipient MNs when EVs derived from SOD1G93A astrocytes are applied.
In summary, our results show that miR-155-5p expression is increased in SOD1G93A astrocyte-derived exosomes and that these exosomes critically influence MN survival in culture, describing a new mechanism by which astrocytes may contribute to MN death in ALS. Moreover, these data suggest a potential new therapeutic target to modify disease outcomes in ALS consistent with previous reports (Koval et al., 2013).
Conclusion
The exosomes derived from astrocytes overexpressing the ALS-linked mutant Cu/Zn superoxide dismutase (SOD1G93A) are enriched with miRNA-155-5p. In addition, miR-155-5p reduces the ability of exosomes derived from non-Tg astrocytes to support MN survival.
Supplemental Material
sj-pdf-1-asn-10.1177_17590914231197527 - Supplemental material for SOD1G93A Astrocyte-Derived Extracellular Vesicles Induce Motor Neuron Death by a miRNA-155-5p-Mediated Mechanism
Supplemental material, sj-pdf-1-asn-10.1177_17590914231197527 for SOD1G93A Astrocyte-Derived Extracellular Vesicles Induce Motor Neuron Death by a miRNA-155-5p-Mediated Mechanism by Soledad Marton, Ernesto Miquel, Joaquín Acosta-Rodríguez, Santiago Fontenla, Gabriela Libisch and Patricia Cassina in ASN Neuro
Footnotes
Abbreviations
Acknowledgments
The data presented herein were obtained using instrumentation in the Microscopy Unit of Facultad de Medicina, Universidad de la República, Montevideo, with the support of Core staff. The facility is funded through user fees and financial support. We would like to thank Dr. Luciana Negro from the School of Biomedical Engineering, Sciences and Health Systems, Drexel University, Philadelphia; Dr. Zachary Fitzpatrick from Spark Therapeutics, Philadelphia, for kindly reading the manuscript and comments; Dr. José Tort from Facultad de Medicina, Universidad de la República, Montevideo, for bioinformatic assistance; Dr. Juan Pablo Tosar and MSc Pablo Fagúndez from Facultad de Ciencias, Universidad de la República, Montevideo, for the analysis and quantification of exosomes; and Dr. Carlos Robello from Institut Pasteur, Montevideo, Uruguay, for support in the analysis of exosomal RNA.
Author Contribution
Project conceptualization and design were developed by S.M., E.M., and P.C. Experiments and analyses were performed by S.M., E.M., J.A., and S.F. The manuscript was written and edited by S.M. and P.C.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by a research fellowship award from Agencia Nacional de Investigación e Innovación (ANII) (PD NAC 2014,1,102084) to S.M. and by ANII grant [FCE_1_2014_1_104239] to P.C.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
