Abstract
As the resident immune cells of the healthy nervous system, homeostatic microglia can rapidly become activated in response to injury/disease. Dysregulated microglia activation is a hallmark of nervous system disorders including neurodegenerative diseases such as amyotrophic lateral sclerosis (ALS) and Alzheimer's disease. The elucidation of the biological and pathological roles of microglia has recently benefitted from the development of microglia-like cells using human induced pluripotent stem cell (iPSC)-based approaches. The success of iPSC-derived microglia preparations as a disease-relevant model system depends on their representation of the in vivo spatial and temporal heterogeneity of microglia under pathological conditions. Little is currently known about the potential of human iPSC-derived microglia generated using different methods for the study of neurodegenerative diseases. We compared the transcriptomes of human iPSC-derived microglia generated using two frequently used in vitro differentiation methods to determine whether separate strategies can generate microglia with distinct transcriptional signatures in vitro. We show that microglia derived using different differentiation methods display distinct maturation characteristics after equivalent times in culture. We also reveal that iPSC-derived microglia preparations generated using these two methods are composed of different subpopulations with transcriptomic signatures resembling those of in vivo regionally distinct microglia subtypes, specifically white-matter and gray-matter microglia. These findings highlight the need to better characterize the subtype composition of each microglia preparation prior to its use to model neurodegenerative diseases.
Introduction
Microglia are the resident myeloid cells of the adult nervous system. They mediate innate immunity by sensing and controlling changes in the microenvironment caused by either external pathogens or local pathologies. In healthy conditions, microglia patrol neural tissues in a homeostatic (“resting”) state, which can rapidly change to an “activated” state in response to injury/disease. Activated microglia are the primary phagocytic cells in the nervous system. They can engulf external pathogens and cellular debris resulting from injury or disease. Activated microglia also release inflammatory cytokines that can promote astrocyte activation and be toxic to neighboring neurons. The balanced contributions of resting and activated microglia mediate both beneficial (protective) and detrimental (inflammatory) effects on the surrounding cells, depending on the specific context (reviewed by Mendes & Majewska, 2021; Miron & Priller, 2020; Prinz et al., 2019; Spiteri et al., 2022).
The presence of activated microglia is a common event in virtually all neurological diseases and disorders. Taking amyotrophic lateral sclerosis (ALS) as an example, hallmarks of microglia activation are a feature of postmortem tissues from ALS patients (Henkel et al., 2004; Lederer et al., 2007; Malaspina & de Belleroche, 2004; Yasojima et al., 2001), as well as spinal cord and cranial motor nuclei of ALS transgenic mouse models (Fendrick et al., 2007; Graber et al., 2010; Henkel et al., 2009; Kassa et al., 2009). Intercellular communication between neurons and microglia is proposed to play important and dynamic roles during the degeneration of neural circuits in ALS, as well as other neurodegenerative diseases, by mediating beneficial or detrimental effects depending on the disease stage (reviewed by Beers & Appel, 2019; Clarke & Patani, 2020; Geloso et al., 2017; McCauley & Baloh, 2019).
The study of the mechanisms underlying the roles of microglia in neurodegenerative pathophysiologies seen in ALS, Parkinson's disease, and Alzheimer's disease, has benefitted from the availability of animal experimental model systems (reviewed by Beers & Appel, 2019; Karanfilian et al., 2020; Liu et al., 2021; Thonhoff et al., 2018). However, numerous species-specific differences in gene expression exist between murine and human microglia. Moreover, several microglial genes exhibiting differences among species are often implicated in human neurodegenerative diseases (Healy et al., 2020; Masuda et al., 2019; Patir et al., 2019; Szulzewsky et al., 2016). These observations highlight the need to combine studies in animal models with investigations using human microglia-based experimental systems. Recent advances in induced pluripotent stem cell (iPSC)-based approaches have addressed this need by providing efficient and reliable approaches to generate physiologically relevant human microglia-like cells (e.g., Abud et al., 2017; Douvaras et al., 2017; McQuade et al., 2018; Muffat et al., 2016; Pandya et al., 2017; to cite a few). This remarkable progress offers previously unavailable experimental opportunities to study human microglia biology and model microglia pathophysiological mechanisms in vitro.
Recent studies have provided growing evidence that microglia represent a heterogeneous group of cells comprising region-specific subpopulations with distinct biological characteristics (DePaula-Silva et al., 2019; Grabert et al., 2016; Li et al., 2019; Masuda et al., 2019; Masuda et al., 2020; Stratoulias et al., 2019; van der Poel et al., 2019). This cellular heterogeneity poses a challenge to research strategies involving microglia-like cells derived from human iPSCs (hereafter referred to as “human iPSC-derived microglia”). It is reasonable to postulate that the success of these investigations will depend on whether or not human iPSC-derived microglia preparations will comprise those specific subtypes that are more relevant to the biological questions under study. A case in point: neuroinflammatory responses mediated by microglia in ALS are heterogeneous, resulting in the presence of distinct subsets of activated microglia that are believed to interact with disease-specific motor neurons in regionally defined patterns (Cipollina et al., 2020; Dachet et al., 2019; Liu et al., 2021; Maniatis et al., 2019). More generally, the field of nervous system disease modeling using human iPSCs is increasingly recognizing the importance of including the in vivo heterogeneity of the induced cells among the factors that need to be considered when designing ideal, disease-relevant cellular assays (Giacomelli et al., 2022; Hedegaard et al., 2020; Stifani, 2021).
In this study, we provide evidence that human iPSC-derived microglia preparations generated using different methods comprise microglia with distinct transcriptional signatures in vitro resembling gene profiles of different microglia subpopulations in vivo. These observations suggest that the harmonized use of multiple human microglia derivation methods should be implemented when modeling neurodegenerative diseases, in order to generate the most disease-relevant repertoire of microglia subgroups needed to model pathophysiological mechanisms involving regionally specialized microglia subtypes.
Materials and Methods
Human-Induced Pluripotent Stem Cells
Three separate human iPSC lines were used in these studies to ensure that the successful generation of microglia could be reproduced using multiple iPSCs. Human iPSC line NCRM-1 was obtained from the National Institutes of Health Stem Cell Resource (Bethesda, MD, USA). Human iPSC lines CS52iALS-C9n6.ISOxx (referred to as “CS52” hereafter for sake of brevity) and CS29iALS-C9n1.ISOxx (“CS29” hereafter), corresponding to corrected, isogenic lines derived from two distinct parental lines generated from ALS patients with C9orf72 gene mutations, were obtained from Cedars-Sinai (Los Angeles, CA, USA) (Thiry et al., 2022). Undifferentiated state of human iPSCs was assessed by testing for expression of the stem cell markers NANOG and OCT4 using rabbit anti-NANOG (1/1,000; Abcam; Cambridge, UK; Cat. No. ab21624) and rabbit anti-OCT4 (1 µg/ml; Abcam; Cat. No. ab19857) or goat anti-OCT3/4 (1/500; Santa Cruz Biotechnology; Dallas, TX, USA, Cat. No. sc-8628) antibodies, as described (Chen et al., 2021).
Derivation of Microglia-Like Cells from Human iPSCs
Derivation of microglia from human iPSCs was performed as described in two separate protocols published by Fossati and coworkers (Douvaras et al., 2017) and Blurton-Jones and colleagues (McQuade et al., 2018). The following modifications were made to the Douvaras et al. method. iPSCs were harvested using StemProTM AccutaseTM Cell Dissociation Reagent (ThermoFisher Scientific; Waltham, MA, USA; Cat No. A1110501) and seeded at a density of 15,000 cells per cm2 onto 6-well dishes coated with Matrigel (ThermoFisher Scientific; Cat. No. 08-774-552) in mTeSRTM1 medium (STEMCELL Technologies; Vancouver, BC, Canada; Cat. No. 85850) containing 10 µM ROCK inhibitor (compound Y-27632 2HCl; Selleck Chemicals; Cat. No. S1049) (this was considered as day 0 in vitro). Starting on the next day, the culture medium was changed to Essential 6TM Medium (ThermoFisher Scientific; Cat. No. A1516401) containing 80 ng/ml BMP4 (R&D Systems; Minneapolis, MN, USA; Cat No. 314-BPE), and cells were cultured for 2–4 days until they reached near confluence. All subsequent steps were as described in the original protocol (Douvaras et al., 2017), except that microglial progenitors were routinely collected as floating cells, without the need for fluorescence-activated cell sorting or magnetic bead separation, followed by culture for further differentiation in SF-Microglia Medium (RPMI + IL-34 + GM-CSF). When using the McQuade et al. method, iPSCs were harvested using ReLeSR™ reagent (STEMCELL Technologies; Cat. No. 05872) and seeded as small aggregates of ∼100 cells each, at 10–20 aggregates per cm2 in mTeSR™1 medium containing 10 µM ROCK inhibitor. The following day (considered as day 0), those wells containing 4–10 adhered ∼100-cell colonies per cm2 were switched from mTeSR™1 medium to STEMdiff™ Hematopoietic kit (STEMCELL Technologies: Cat. No. 05310) (2 ml of medium per well using 6-well dishes) to start the differentiation process. Multiple (n ≧ 10) cultures of either type of microglia preparations were generated and validated through morphological analysis and immunocytochemistry.
Characterization of Human iPSC-Derived Microglia by Immunocytochemistry
Induced human microglia were characterized by immunocytochemistry using the following primary antibodies: rabbit anti-IBA1 (1/1,000; FUJIFILM Wako Chemicals; Richmond, VA, USA; Cat. No. 019-19741) and rabbit anti-P2Y12R (1/150; Alomone Labs; Jerusalem, Israel; Cat. No. APR-020). The secondary antibody was a donkey antirabbit IgG conjugated to Alexa Fluor 594 (1/1,000; Invitrogen; Burlington, Ontario, Canada; Cat No. A-21207; RRID, AB_141637). More specifically, cells were rinsed two times in phosphate-buffered saline (PBS) at room temperature (RT). They were then fixed with 4% paraformaldehyde in PBS for 10 min at RT, followed by several quick rinses and three consecutive washes in PBS for 15 min each time. Cells were then permeabilized for 10 min in PBS containing 0.1% Triton-X100, followed by several quick rinses and three consecutive washes in PBS for 15 min each time. After these steps, cells were incubated for one hour at RT in PBS containing 0.05% Triton-X100 and 1% bovine serum albumin, followed by the addition of primary antibodies and incubation at 4°C for 16 h. After this step, cells were rinsed/washed as described above, followed by incubation with secondary antibodies for 1 h at RT, rinsing/washing as described, counterstaining with DAPI for 3 min at RT, and final rinses/washes in PBS. Images were acquired using a Zeiss Axio Imager M1 microscope connected to an AxioCam 503 camera, using ZEN software (Zeiss Canada Ltd., Toronto, Ontario, Canada) (Methot et al., 2018). Quantification of the proportion of cells expressing a specific marker was performed by counting the number of cells presenting positive labeling of the marker and dividing it by the total number of Hoechst-positive nuclei (Soubannier et al., 2022).
RNA Sequencing and Acquisition of Read Counts
Total RNA was isolated for RNAseq from cell pellets at different differentiation/maturation timepoints via sequential treatment with TRIzol Reagent (ThermoFisher Scientific; Cat. No. 15596026) and PureLink RNA Micro Scale Kit (ThermoFisher Scientific; Cat. No. 12183-016) following the instructions provided by the manufacturer. According to the Douvaras protocol, induced microglia begin to acquire a mature phenotype starting on day 39 in vitro (Douvaras et al., 2017); therefore, we chose a series of timepoints for RNA isolation of CS52 iPSC-derived microglia under the Douvaras protocol, specifically days in vitro 45, 48, 58, and 66. Under the McQuade protocol's conditions, induced microglial cells are considered to become mature as of day 38 in vitro (McQuade et al., 2018); therefore, we conducted RNA isolation from CS52 iPSC-derived microglia under the McQuade protocol on days in vitro 42, 45, and 46. All seven RNA sample replicates collected at different time points from microglia generated in separate derivation experiments were submitted for Illumina next-generation sequencing to the Genomics platform at the Institute for Research in Immunology and Cancer, Montreal, Quebec, Canada (https://www.iric.ca/en/research/platforms-and-infrastructures/genomics). Adaptor sequences and low-quality bases in the resulting FASTQ files were trimmed using Trimmomatic version 0·35 (Bolger et al., 2014), and genome alignments were conducted using STAR version 2·5 1b (Dobin et al., 2013). The sequences were aligned to the human genome version GRCh38, with gene annotations from Gencode v29 based on Ensemble release 94. As part of quality control, the sequences were aligned to several different genomes to verify that there was no sample contamination. Raw read-counts were obtained directly from STAR, and reads in transcripts per million (TPM) format were computed using RSEM (Li & Dewey, 2011).
In Silico Analysis
Heatmaps were used to visualize the expression of specific gene sets across multiple different samples. To generate heatmaps, read counts were converted into log2-counts-per-million (logCPM) values with the “cpm” function of the edgeR package in R. This function adds a low “prior count” value to avoid taking the log of zero and also normalizes that data by the different library sizes of the samples. Accordingly, negative values represent very low gene expression values, usually below five. Hierarchical clustering of rows was applied to the heatmaps in order to group genes with correlated expression patterns. Since logCPM values are displayed directly without additional Z-score scaling by row, color comparisons can be made across samples for each gene as well as between genes for any given sample.
Differential gene expression analysis was conducted on the raw read-count matrix using an edgeR (version 4·0) pipeline as previously described (Chen et al., 2016). Briefly, we first filtered the genes such that only those with a robust expression level were retained. Next, the differing RNA composition/library size of each sample was normalized by a scaling factor of the trimmed mean of M-values (TMM) between each pair of samples. The known sources of variability in the studied samples were the microglia differentiation protocol and batch, both of which were incorporated into the experimental design used for differential expression analysis. In order to avoid detecting transcriptomic differences resulting from different levels of in vitro developmental maturation, we determined the maturity stage of CS52 iPSC-derived microglia samples by visualizing their comparative expression of known microglia marker genes (Butovsky et al., 2014; Hickman et al., 2013; Muffat et al., 2016; Zhang et al., 2014) and applied only a subset of samples to downstream differential gene expression analysis. From among the different timepoints of microglia, we chose the most mature samples in each group, namely days in vitro 58 and 66 of microglia generated under the Douvaras protocol, and days in vitro 42, 45, and 46 of microglia generated under the McQuade protocol. The “glmQLFit” and “glmQLFTest” functions of edgeR were then implemented on this sample subset to obtain differentially expressed genes (DEGs) using the quasi-likelihood (QL) method. Genes were considered statistically significant if the adjusted p-value (false discovery rate; FDR) was <0.05, with no fold-change cutoff applied to restrict the number of DEGs identified.
The GeneOverlap package in R was used to reveal statistically significant overlaps between gene sets in comparison to the genomic background. We used the DEGs upregulated in microglia derived using either protocol and compared them with previously published gene signatures for different microglial subtypes. All the gene sets used in this analysis are listed in Supplementary Table 1 available in Supplementary Materials. Fisher's exact test was used to generate the overlapping p-value, with the null hypothesis that the odds ratio ≤1. An odds ratio is a statistic that quantifies the strength of association between two events (in this case, gene sets); an odds ratio ≥ 1 means there is a correlation between the two gene sets.
Gene set enrichment analysis (GSEA) was conducted on the DEGs using the Functional Annotation Tool of DAVID Bioinformatics Resources 6·8 (Huang da et al., 2009a; 2009b). Gene Ontology (GO) terms and Kyoto Encyclopaedia of Genes and Genomes (KEGG) pathways enriched in the DEGs were computed separately for upregulated and downregulated genes, as this increases the statistical power to identify pathways/terms pertinent to the phenotype of interest (Hong et al., 2013). For each GO category (e.g., Biological Process or Molecular Function), the “direct” function in DAVID was used to restrict the results to those directly annotated by the source database and to avoid repetitive and generic parent terms. For the KEGG pathways, only the non-disease-specific pathways are shown.
Open Source Data
RNAseq data (raw read counts) for iPSC lines were obtained from the LINCS Data Portal (LSC-1002 and LSC-1004 from Dataset LDS-1355) (http://lincsportal.ccs.miami.edu/datasets/). LSC-1002 and LSC-1004 correspond to iPSC lines CS14i-CTR-n6 and CS25iCTR-18n2, whose source provider was Cedar Sinai Stem Cell Core Laboratory. RNAseq data for the microglia line C20 were obtained from the Digital Expression Explorer 2 (DEE2) repository (Accession# SRR3546647) (http://dee2.io/). All these raw data were combined with the RNAseq data from the present microglia preparations and analyzed using a single pipeline. Additional metadata for all open-source RNAseq data are provided in Supplementary Table 2.
Results
Characterization of Human iPSC-Derived Microglia Generated Using Different Protocols
The in vivo heterogeneity of microglia is seldom considered when human iPSCs are used to derive microglial cells to study their roles in (patho)biological mechanisms in health and disease. This omission has repercussions when human iPSC-derived microglia with no defined regional characteristics are used to model neurodegenerative diseases that affect specific neuronal cell types in specific regions of the central nervous system. It is unlikely that the numerous published microglia derivation protocols starting from human iPSCs would give rise to equivalent preparations, since the experimental conditions used by different methods vary considerably (e.g., Abud et al., 2017; Douvaras et al., 2017; McQuade et al., 2018; Muffat et al., 2016; Pandya et al., 2017). We, therefore, sought to compare the gene expression signatures of human iPSC-derived microglia generated using different in vitro differentiation methods to begin to understand whether different methods have the potential to give rise to transcriptionally distinct microglia populations.
Motivated by our long-term interest in using human iPSC-derived microglia for the study of ALS pathophysiology, we chose to compare the microglia derivation methods described by Douvaras and colleagues (Douvaras et al., 2017) and McQuade and coworkers (McQuade et al., 2018), because either one or the other of these widely used methods have been utilized recently to model the involvement of human microglia in ALS pathophysiology (Allison et al., 2022; Kerk et al., 2022). Both of these derivation protocols achieve the robust generation of microglia under serum-free conditions and in the presence of defined cytokines (Douvaras et al., 2017; McQuade et al., 2018). In the context of the present study, we shall hereafter operationally refer to microglia generated using the Douvaras et al. protocol as “Douvaras-microglia,” and microglia generated using the McQuade et al. method as “McQuade-microglia.”
Both protocols generated induced cultures highly enriched in cells expressing the purinergic receptor P2RY12, whose expression distinguishes microglia from peripheral macrophages (Butovsky et al., 2014) (fractions of P2RY12+ cells: McQuade-microglia, 95.2% ± 1.2%; Douvaras-microglia, 96.3% ± 3.1%) (Figure 1A). In our hands, McQuade microglia displayed more frequently a compact ramified morphology (“type 1” morphology in Figure 1A(a); higher-magnification depicted in panel A(a)-1), or a slender elongated morphology (“type 2” morphology in Figure 1A(a); higher magnification depicted in panel A(a)-2). Less frequently, we observed P2RY12+ microglia with larger cell bodies (“type 3” morphology in Figure 1A(a); higher magnification depicted in panel A(a)-3). Douvaras-microglia cultures typically contained few cells with a type-1 morphology, and most cells had either a type-2 or type-3 morphology (Figure 1A(b) and panels A(b)-1-3). Similar morphologies were observed when cultures were stained with an antibody against the microglia/macrophage protein IBA1 (Figure 1B(a), see insets (a)-1 and (a)-2; and Figure 1B(b), see insets (b)-2 and (b)-3). Similar robust yields, and similar morphologies, were observed when microglia were derived from two other iPSC lines, namely NCRM-1 (Figure 1C and D) and CS29 (Figure 1E). In all cases examined, staining for both P2RY12 and IBA1 expression revealed that Douvaras-microglia were generally larger in size than McQuade microglia in our preparations (Figure 1A–E).
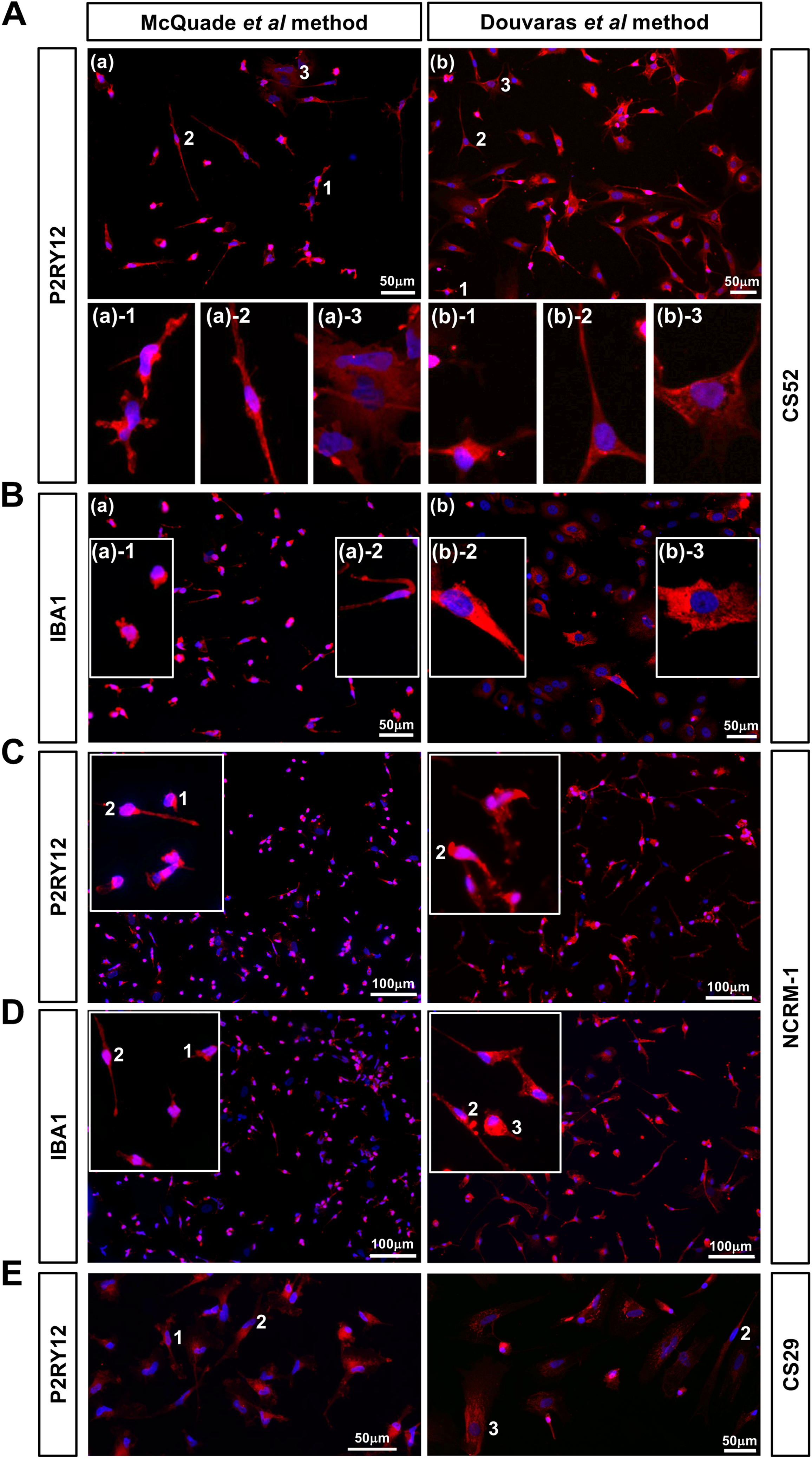
Characterization of human iPSC-derived microglia by immunocytochemistry. (A, B) Immunocytochemical analysis of either P2RY12 (A) or IBA1 (B) expression in microglia generated from iPSC line CS52. Cells were examined at differentiation day 43 (A(a), B(a)) when using the McQuade et al. protocol, and at differentiation days 51 (A(b)) or 49 (B(b)) in the case of microglia derived using the Douvaras et al. protocol. Scale bars are indicated. McQuade-microglia display mainly type-1 and type-2 morphologies (marked by numbers 1 or 2 next to representative cells in A(a), and shown at higher magnification in A(a)-1, A(a)-2), B(a)-1, and B(a)-2). Less frequently, McQuade microglia exhibit a type-3 morphology (shown at higher magnification in A(a)-3). Douvaras-microglia display mainly type-2 and type-3 morphologies (shown at higher magnification in A(b)-2, A(b)-3), B(b)-2, and B(b)-3), and less frequently type-1 morphology (shown at higher magnification in A(b)-1). (C, D) Immunocytochemical analysis of either P2RY12 (C) or IBA1 (D) expression in microglia generated from iPSC line NCRM-1. Cells were examined at either differentiation day 42 (McQuade et al. protocol) or differentiation day 48 (Douvaras et al. protocol). Scale bars are indicated. Insets show higher magnification views of type-1 and type-2 cell morphologies most prevalent in McQuade microglia, or type-2 and type-3 morphologies most common in Douvaras microglia, respectively. (E) Immunocytochemical analysis of P2RY12 expression in microglia generated from iPSC line CS29. Cells were examined at either differentiation day 51 (McQuade et al. protocol) or differentiation day 45 (Douvaras et al. protocol). Cell morphologies are marked by numbers 1, 2, or 3 next to representative cells. Scale bars are indicated.
To further characterize the properties of human iPSC-derived microglial cell preparations generated using these two methods, we performed bulk RNAseq studies on microglia derived from iPSC line CS52. We examined a series of timepoints in order to characterize cells at different stages of in vitro developmental maturation. Specifically, RNAseq analysis was conducted on samples obtained from Douvaras-microglia collected at day 45, 48, 58, and 66, and RNA samples obtained from McQuade-microglia collected at day 42, 45, and 46 (Figure 2A). Based on RNAseq data, we used the expression of a panel of 38 known microglia marker genes (Butovsky et al., 2014; Hickman et al., 2013; Muffat et al., 2016; Zhang et al., 2014) to compare Douvaras-microglia and McQuade-microglia at different timepoints. This analysis suggested that McQuade-microglia acquired a robust microglia gene-expression profile 42 days after the start of the differentiation process; in comparison, Douvaras-microglia required an additional 15–20 days to exhibit a transcriptional signature similar to differentiation day 42–46 McQuade-microglia (Figure 2A).
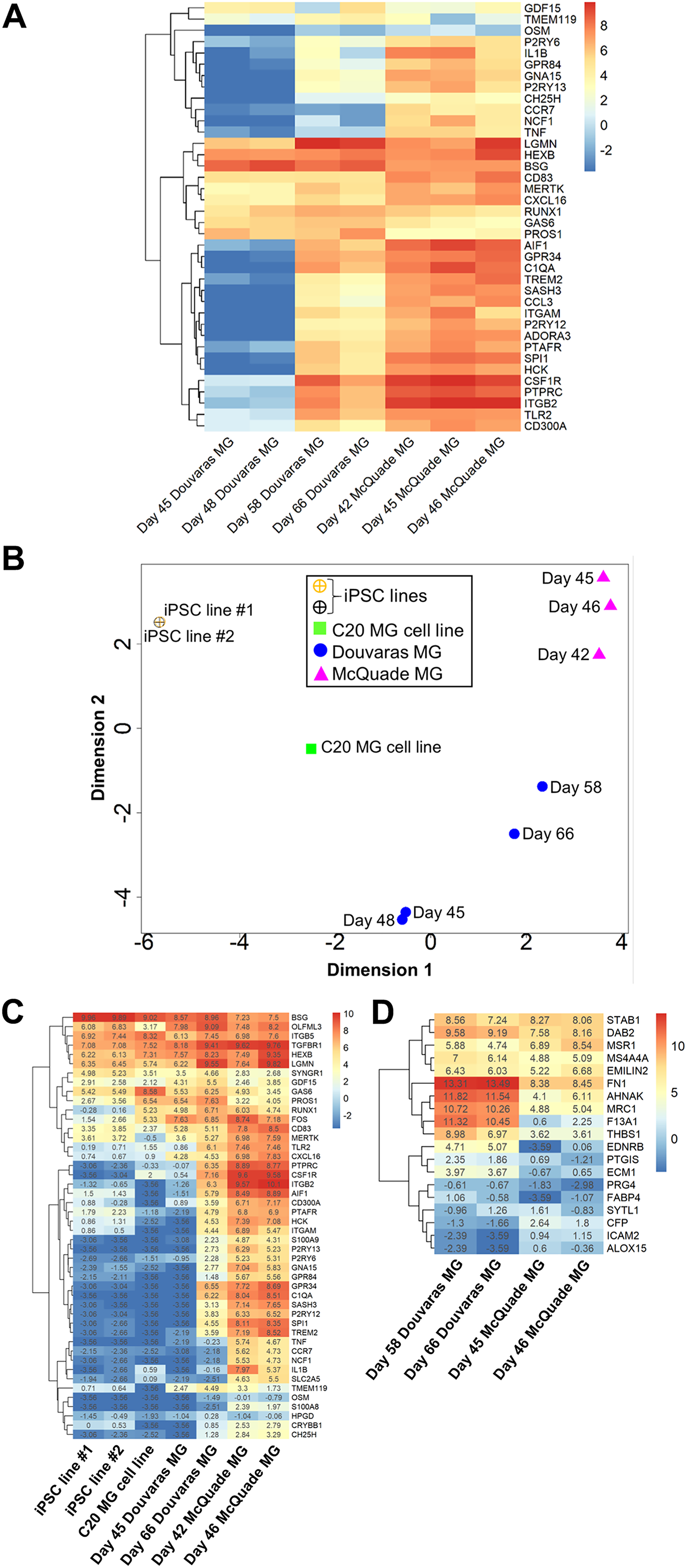
Characterization of human iPSC-derived microglia by RNA sequencing. (A, B) Heatmap depicting the expression of known microglia genes in human CS52 iPSC-derived microglia (MG) preparations generated with either the Douvaras or McQuade protocol. Each heatmap column represents one biological replicate cultured with the indicated differentiation protocol and examined at the indicated differentiation day. Cool colors indicate low expression and warm colors indicate increased/high expression, as shown in the color scale. Subgroups of genes with correlated expression patterns are portrayed by the dendrogram on the left side of the heatmap. (B) Principle component analysis plot showing the relationship between the transcriptomic profiles of CS52 iPSC-derived microglia (MG) generated with either the Douvaras or McQuade protocols (at the indicated differentiation days) compared with the transcriptomic profiles of two undifferentiated iPSC cell lines (data from two different lines from LINCS Data Portal open-source dataset LDS-1355) and the immortalized human microglia cell line C20 (data from DEE2 repository open-source data). Each plotted data point represents one biological replicate cultured with the indicated differentiation protocol and examined on the indicated differentiation day. (C) Heatmap depicting the expression of known microglia genes in two undifferentiated human iPSC lines, immortalized human microglia cell line C20, and CS52 iPSC-derived microglia generated with either the Douvaras or McQuade protocol (the least and most mature samples sequenced under each differentiation protocol). (D) Heatmap depicting the expression of a macrophage-enriched gene signature in CS52 iPSC-derived microglia generated with either the Douvaras or McQuade protocol (the two most mature samples sequenced under each protocol). (C, D) Each heatmap column of original data (columns 4–7 in panel C and all columns in panel D) represents one biological replicate cultured with the indicated differentiation protocol and examined on the indicated differentiation day. Cool colors indicate low expression and warm colors indicate increased/high expression, as shown in the color scale. The logCPM value (corresponding to the color scale) for each data point is listed in the heatmap. Subgroups of genes with correlated expression patterns are portrayed by the dendrogram on the left side of the heatmaps.
To further compare microglia generated using the two methods, we next performed a principle component analysis, which showed that both Douvaras-microglia and McQuade-microglia grouped together as iPSC-derived microglia along one dimension, clustering away from two undifferentiated iPSC cell lines and from the immortalized human microglial cell line C20 (Garcia-Mesa et al., 2017) (Figure 2B). However, the iPSC-derived microglia generated under the two different protocols were distinct enough in their transcriptomic profiles to still be separated into different clusters along another dimension of the PCA plot (Figure 2B). Using transcriptomic data, we defined the expression level of a list of 46 microglial cell marker genes (Butovsky et al., 2014; Hickman et al., 2013; Muffat et al., 2016; Zhang et al., 2014) at our earliest timepoint (day 45 for Douvaras-microglia and day 42 for McQuade-microglia) and latest timepoint (day 66 for Douvaras-microglia and day 46 for McQuade-microglia) of both microglia preparations. The majority of these 46 microglial genes were upregulated in both Douvaras-microglia and McQuade-microglia when compared with two undifferentiated human iPSC lines and the microglial cell line C20 (Figure 2C). In agreement with data depicted in Figure 2A, McQuade-microglia acquired a robust microglia gene-expression profile, compared to C20 microglia, 40–45 days after the start of the differentiation process. In comparison, Douvaras-microglia were transcriptionally similar to C20 microglia at in vitro differentiation day 45 and required an additional 15–20 days to exhibit a transcriptional signature similar to differentiation day 42–46 McQuade-microglia (Figure 2C). Both microglia preparations displayed generally low expression of a previously described macrophage-specific gene signature (Hickman et al., 2013), as displayed in the heatmap of the most mature microglia samples from each protocol (Figure 2D). The few genes in exception, such as DAB2, MSR1, and MRC1, are known to be expressed in both macrophages and microglia (Kobashi et al., 2020; Zeiner et al., 2019), which might point to the presence of common precursors of microglia and macrophages in those cultures (Figure 2D). Together, these findings show that both microglial preparations display transcriptional signatures of physiological microglia; moreover, they suggest that McQuade-microglia undergo a more rapid developmental maturation in vitro than Douvaras-microglia.
Microglia-Subtype Composition of Different Human iPSC-Derived Microglia Preparations
To further compare the characteristics of Douvaras-microglia and McQuade-microglia, we next performed DEG analysis of RNAseq data collected from cells generated with either methods. In order to minimize the possibility of detecting differences resulting from different levels of in vitro developmental maturation, we compared cells displaying similar levels of microglial maturity, as determined based on data depicted in Figure 2A–C. Two separate Douvaras-microglia preparations (RNA collected at days in vitro 58 and 66) and three McQuade microglia preparations (RNA collected at days in vitro 42, 45, and 46) were compared using an EdgeR analysis pipeline for DEG analysis (Chen et al., 2016). Data from these five separate RNAseq experiments revealed 1943 genes that were significantly upregulated in McQuade-microglia compared to Douvaras-microglia, while 2836 genes were upregulated in the latter (adjusted p < 0.05 with no fold-change cutoff). In silico functional analyses (GO and KEGG analyses combined) of the DEGs between McQuade-microglia and Douvaras-microglia revealed that GO and KEGG terms involving biological functions typically expected from mature microglia (such as immune response, chemotaxis, motility, and migration) were overrepresented in McQuade-microglia, suggesting further that these cells were more developmentally mature than Douvaras-microglia (Figure 3A). This analysis also showed that a number of biological processes/pathways suggestive of a more activated microglia phenotype were overrepresented in McQuade-microglia, including phagocytosis, apoptosis, NF-κB signaling pathway, and others (Figure 3B).
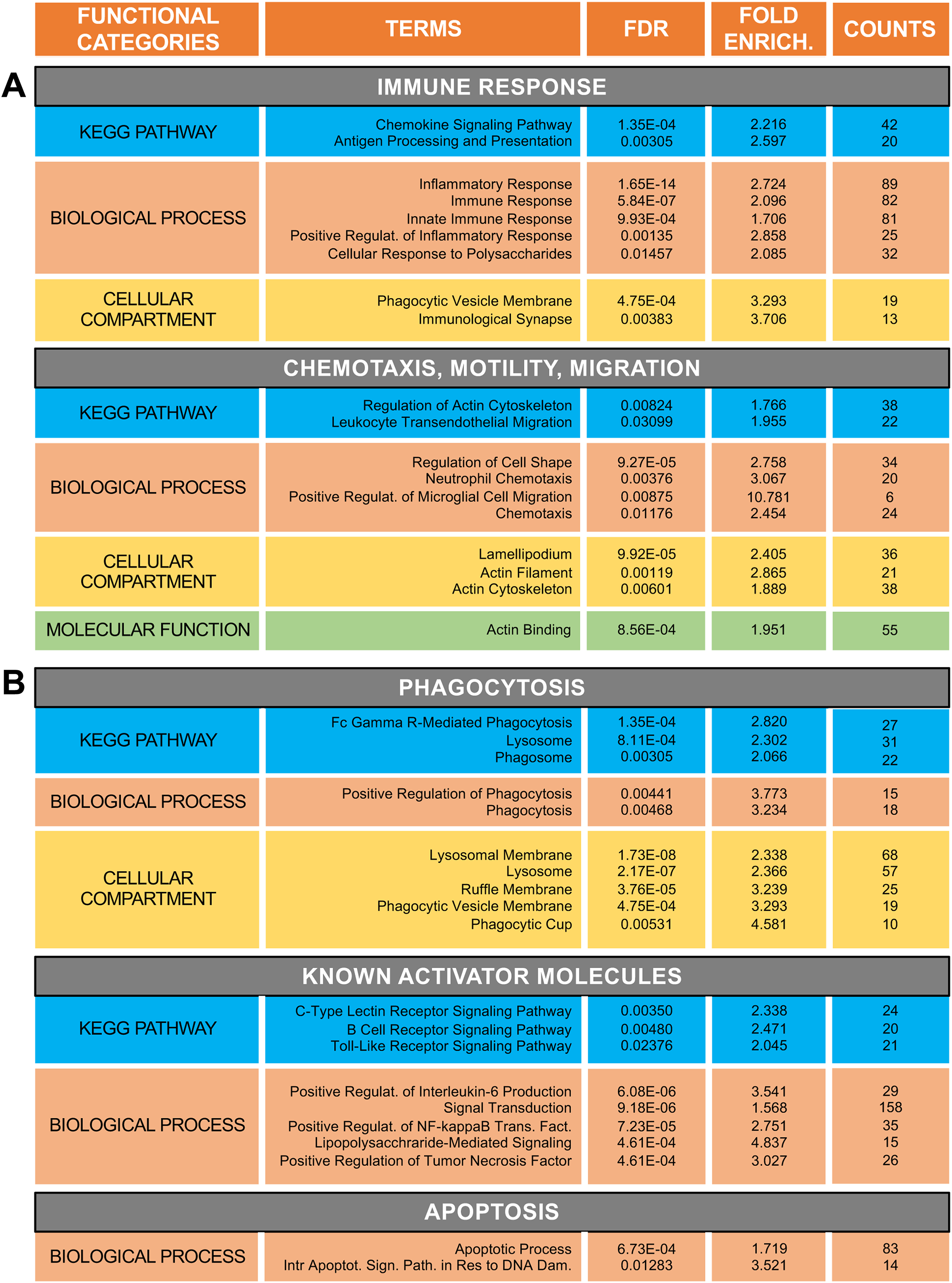
Human iPSC-derived microglia generated with different protocols exhibit distinct transcriptomic profiles. (A, B) Functional analysis of DEGs between Douvaras-microglia (n = 2; RNA collected at days in vitro 58 and 66) and McQuade-microglia (n = 3; RNA collected at days in vitro 42, 45, and 46). Shown are functional categories (KEGG Pathway, Biological Process, Cellular Compartment, Molecular Function), Terms within each Functional Category, false discovery rate (FDR), Fold Enrichment (defined here and in succeeding figures as the enrichment of each term in the studied samples compared to its enrichment in the background population (number of DEGs in term/total number of DEGs) / (number of human genes in term/total number of genes in the human genome), and number of Counts. (A) Gene ontology terms related to functions of more developmentally mature microglia (e.g., immune response, chemotaxis, motility, and migration) that are significantly overrepresented in McQuade-microglia compared to Douvaras-microglia. (B) Gene ontology terms associated with microglia activation (categorized under “phagocytosis,” “known activator molecules,” and “apoptosis”) that are significantly overrepresented in McQuade-microglia compared to Douvaras-microglia. Only non-disease-specific KEGG terms are included. Here and in succeeding figures, all microglia were derived from iPSC line CS52.
We next investigated whether the transcriptomic divergence observed between Douvaras-microglia and McQuade-microglia might also be due, at least in part, to a different microglia-subtype composition in these preparations. The DEGs significantly upregulated in each preparation of microglia were compared to previously described gene signatures of regionally distinct microglia subtypes in vivo. At first, we compared the transcriptional profiles of Douvaras-microglia and McQuade-microglia to those of three distinct mouse microglia subtypes described by Grabert and colleagues (Grabert et al., 2016). These investigators isolated microglia from different regions of the adult mouse brain (cortex, hippocampus, striatum, and cerebellum) and identified three major subtypes based on their transcriptomic profiles. We observed that one of these three microglia subtypes was significantly enriched in Douvaras-microglia (Fisher's exact test; p = 0.002), whereas a separate subtype was significantly enriched in McQuade-microglia (Fisher's exact test; p = 6.6 × 10−6) (Figure 4A). Importantly, the study by Grabert and coworkers had shown that the particular microglia gene signature that we observed to be enriched in Douvaras-microglia (Figure 4A) exhibited high expression in the mouse cerebellum and hippocampus, but low expression in the cortex and striatum. In contrast, the gene signature that we found to be enriched in McQuade microglia (Figure 4A) had reported high expression in the cortex and striatum, but low expression in the cerebellum and hippocampus (Grabert et al., 2016). These combined observations suggest that Douvaras-microglia and McQuade-microglia comprise regionally distinct microglial cell populations.
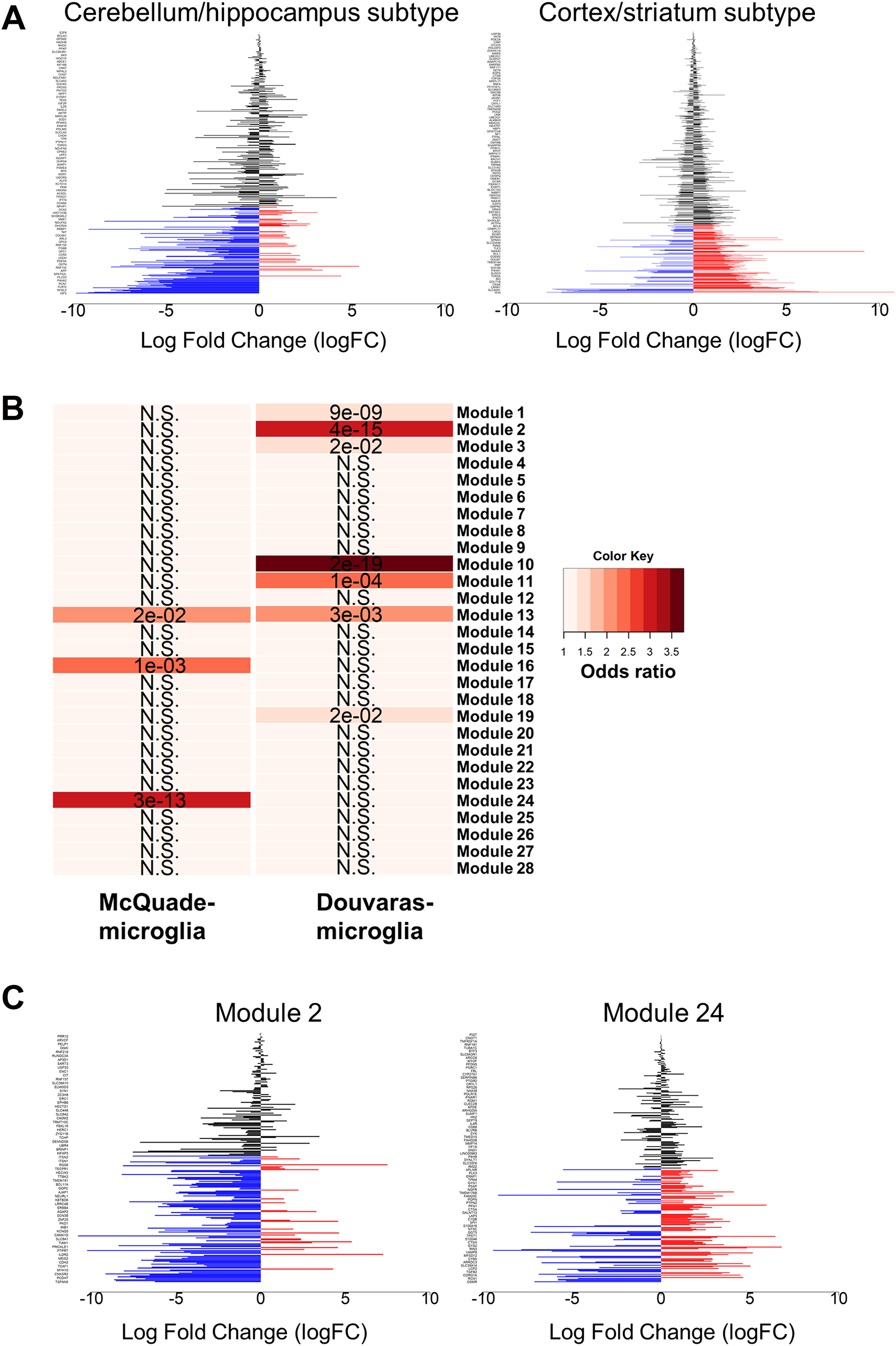
Overlap of regionally distinct microglia gene signatures with iPSC-derived microglia generated using different methods. (A) Barplots depicting the gene expression signatures of regionally distinct mouse microglia populations previously reported to preferentially localize to either the cerebellum and hippocampus or to the cortex and striatum (Grabert et al., 2016) in Douvaras-microglia and McQuade-microglia, respectively. Blue bars correspond to genes with significantly higher expression in Douvaras-microglia, whereas red bars correspond to genes with significantly higher expression in McQuade-microglia. Black bars are genes not significantly increased in microglia derived from either protocol. Log fold change in expression between the two protocols is displayed on the X-axis. Genes listed on the Y-axis can be found in Supplementary Table 1 or in the source publication (Grabert et al., 2016), where the “cortex/striatum subtype” and “cerebellum/hippocampus subtype” are, respectively, named “Cluster 1” and “Cluster 2.” (B) Heatmap depicting the enrichment distribution of regionally distinct cell “modules” described in post mortem spinal cord samples from ALS patients (Maniatis et al., 2019), in Douvaras-microglia and McQuade-microglia, by pairwise gene-overlap analysis. The color gradient represents the degree of association between the gene sets (odds ratio), and the overlapping P-value (Bonferroni-corrected) is listed in each cell. NS = not statistically significant. C) Barplots depicting the expression of Module 2 and Module 24 gene signatures deconvoluted from ALS patient-derived spinal cord tissue, in the microglia derived using the two different protocols. Blue bars are genes with significantly higher expression in Douvaras-microglia, whereas red bars are genes with significantly higher expression in McQuade-microglia. Black bars are genes not significantly increased in microglia derived using either protocol. Log fold change in expression between the two protocols is displayed on the X-axis. Genes listed on the Y-axis can be found in Supplementary Table 1, or in the source publication (Maniatis et al., 2019), where “Module 2” and “Module 24” are correspondingly named “human module 2” and “human module 24.”
To test this possibility further, we next cross-examined transcriptomic data from Douvaras-microglia and McQuade-microglia with previous results by Maniatis and colleagues, who conducted a spatial transcriptomic analysis of postmortem human spinal cord tissues from ALS patients (Maniatis et al., 2019). These authors identified 28 regionally distinct cell “modules” with different gene signatures (including but not restricted to microglia), among which we observed 10 gene signatures that significantly overlapped with upregulated genes in human iPSC-derived microglia generated using either tested protocol. Importantly, six of the seven modules that were enriched in Douvaras-microglia did not overlap with the three modules we found enriched in McQuade-microglia (Figure 4B). Based on the spatial analysis conducted by Maniatis and coworkers (Maniatis et al., 2019), we noticed that six of seven modules enriched in Douvaras-microglia showed strong preferential expression for the gray matter of human spinal cord samples from ALS patients, while two of three modules enriched in McQuade-microglia were preferentially expressed in the white matter of ALS spinal cords (Figure 4B). Two representative gene signatures enriched in Douvaras-microglia and McQuade-microglia, namely Module 2 (Fisher's exact test; p-value for overlap with Douvaras-microglia = 3.9 × 10−15) and Module 24 (Fisher's exact test; p-value for overlap with McQuade-microglia = 5 × 10−13), are plotted in Figure 4C. These observations suggest further that microglia generated using the two in vitro differentiation methods under study are not equivalent and raise the possibility that Douvaras-microglia and McQuade-microglia resemble gray-matter or white-matter microglia, respectively.
To examine the latter possibility more, gene signatures of both types of in vitro generated microglia were compared to a previously described transcriptomic characterization of mouse white-matter microglia (Safaiyan et al., 2021). We observed that white-matter-associated genes displayed significantly increased expression in McQuade-microglia compared to Douvaras-microglia (Fisher's exact test; overlapping p-value = 2 × 10−3) (Figure 5A). In agreement with this finding, GO and KEGG analysis showed that, compared to Douvaras-microglia, McQuade-microglia displayed an overrepresentation of biological processes/pathways associated with white-matter microglia, such as phagocytosis, lysosome, antigen processing and presentation, and others (Amor et al., 2022; Lee et al., 2019) (Figure 5B). In contrast, Douvaras-microglia exhibited overrepresented functions characteristic of gray-matter microglia, such as synaptic interaction, neuron interaction, and others (Figure 5C).
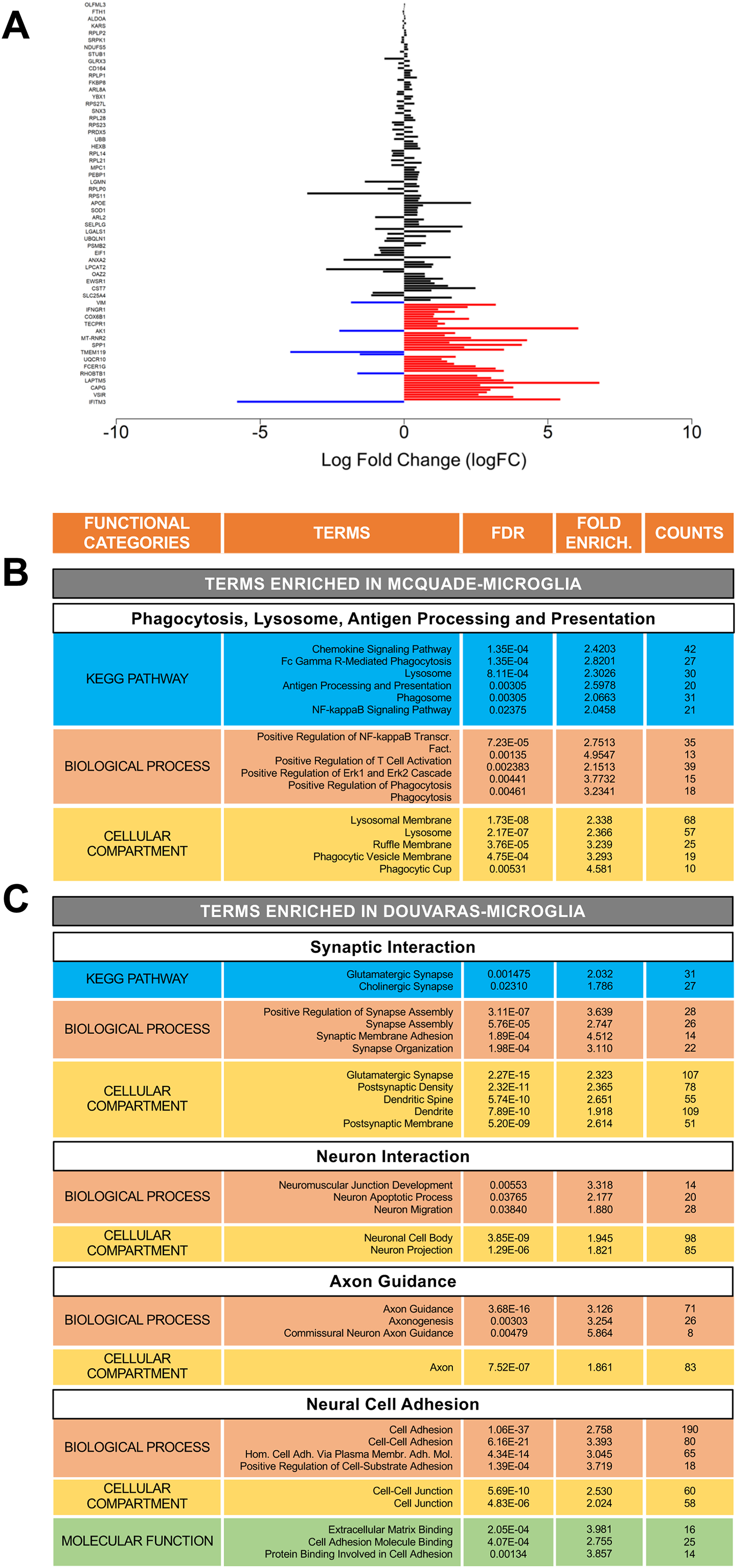
Gene signatures of iPSC-derived microglia generated with different protocols. (A) Barplot depicting the expression of white-matter-associated microglia (WAM) gene signature (Safaiyan et al., 2021) in microglia derived using the two methods under study. Blue bars correspond to genes with significantly increased expression in Douvaras-microglia, while red bars correspond to genes with significantly increased expression in McQuade-microglia. Black bars are genes not significantly increased in microglia derived from either protocol. Log fold change in expression between the two protocols is displayed on the X-axis. Genes listed on the Y-axis can be found in Supplementary Table 1 or in the source publication. (B, C) Shown are either the gene ontology terms related to functions of white-matter microglia (such as phagocytosis, lysosome, antigen processing, and presentation) that are significantly overrepresented in McQuade-microglia compared to Douvaras-microglia (B) or the gene ontology terms related to functions of gray-matter microglia (such as synaptic interaction, neuron interaction, axon guidance, and neural cell adhesion) that are significantly overrepresented in Douvaras-microglia compared to McQuade-microglia (C). Columns depict Functional Categories, Terms, False Discovery Rate (FDR), Fold Enrichment, and number of Counts. All functional categories with relevant terms are included, namely KEGG Pathway, Biological Process, Molecular Function, and Cellular Compartment. Only non-disease-specific KEGG terms are included.
Taken together, these results provide evidence that the two different human iPSC-derived microglia preparations under study contain cell populations with distinct gene expression profiles correlating with regionally distinct microglial subpopulations identified in vivo. More specifically, Douvaras-microglia and McQuade-microglia exhibit gene signatures that overlap, at least in part, with microglia subpopulations present in the gray matter or white matter, respectively.
Discussion
The elucidation of the roles of microglia during the progression of ALS, and other neurodegenerative diseases such as Alzheimer's disease and Parkinson's disease, has benefitted from the recent development of robust and reliable methods to generate microglia from human iPSCs, including iPSCs derived from patients affected by familial forms of these diseases and their matching genome-edited isogenic iPSCs. To date, the published studies using human iPSC-derived microglia to model ALS pathophysiology, which have used either one or the other derivation method investigated in the present study (Allison et al., 2022; Kerk et al., 2022), have not provided information on the subcellular composition of the microglia cultures that were examined. There is a growing need to gather more information on the diversity of different in vitro microglia experimental systems, stemming from the demonstrated spatial heterogeneity of microglia in vivo (DePaula-Silva et al., 2019; Grabert et al., 2016; Li et al., 2019; Masuda et al., 2019; Masuda et al., 2020; Stratoulias et al., 2019; van der Poel et al., 2019). Lack of this information can delay progress in the field of ALS research because it is likely that different subtypes of microglia interact with the variety of regionally distinct motor neurons that are affected in ALS. Local microglia heterogeneity is also relevant in the context of other neurodegenerative diseases with region-specific neuronal pathologies along the rostrocaudal and/or dorsoventral axes of the nervous system. The future success of studies utilizing human iPSC-derived microglia to model ALS, and other neurodegenerative diseases, will depend in large part on the ability to generate, and study, those specific microglial subtypes that are relevant to the pathobiological questions under investigation. In this work, we have provided evidence suggesting that by comparing, and integrating, different human iPSC-based microglia derivation protocols, it will be possible to gain insight into how to develop enhanced experimental systems that will take microglia heterogeneity, as well as other factors such as activation state, into consideration when modeling non-cell autonomous mechanisms of neuronal degeneration using human iPSC-derived microglia.
Specific human iPSC-based neuronal or glial cell derivation methods are often chosen over other protocols because of length and easiness considerations. As important as these factors are, they carry the risk that the fastest or most amenable protocols may give rise to cultures enriched in only a few particular types of induced cells, which may be appropriate to study certain biological mechanisms but not others. The results of the present study suggest that it will be beneficial to the success of disease-focused investigations utilizing human iPSC-derived microglia to conduct preliminary studies to select the most appropriate derivation methods on the basis of multiple criteria. The latter should include not only the rapidity and easiness of the method but also the cellular composition of the induced microglial cultures that would be ideal for the biological questions of interest (e.g., gray-matter versus white-matter microglia). The activation state of the induced microglia should also be considered, since intrinsically activated preparations may be less useful when studying dysregulated activation mechanisms in microglia derived from patient iPSCs, compared to isogenic control cells.
The present work provides an example of studies that can be performed to address the abovementioned questions, before certain (or combinations of) microglia derivation methods are selected. Specifically, our transcriptomic data suggest that human iPSC-derived microglia generated using the method described by McQuade and colleagues (McQuade et al., 2018) are enriched in cells resembling white-matter microglia. In contrast, microglial cultures generated as described by Douvaras and coworkers (Douvaras et al., 2017) are enriched in cells exhibiting gene signatures similar to those of gray-matter microglia. This finding provides support to the important notion that human iPSC-based approaches have the potential to give rise to distinct microglia subpopulations preferably suited to model specific mechanisms of disease pathophysiology. We have also shown that microglia preparations generated using the McQuade method develop more rapidly and display more robust signs of activation than microglia derived using the Douvaras method at in vitro differentiation time points when these cells would usually be experimentally investigated. Thus, different derivation methods may offer both advantages and disadvantages in different contexts, further emphasizing the need to carefully consider the planned experimental strategies. It is reasonable to predict that when an increased number of transcriptomic studies of different human iPSC-derived microglia become available, including the results of single-cell RNAseq, it will become increasingly recognized that certain methods would be preferable for certain studies. For instance, preparations enriched in white-matter microglia would be better suited to address mechanisms of interaction of microglia with axons along myelinated fibers than gray-matter microglia. In the long run, a harmonized use of multiple methods might be the recommended approach to provide disease-relevant experimental systems, as this would generate a larger repertoire of microglia with optimal potential to reflect the heterogeneity present in vivo.
In conclusion, selecting the most informative human iPSC-based microglia derivation strategies is expected to become a key step in the development of disease-relevant experimental systems to model the involvement of microglia in pathophysiological mechanisms. This goal will be facilitated by the growing understanding of microglia diversity in vivo and the mechanisms underlying the generation of this diversity. Combining this increased biological knowledge with a deeper characterization of the properties of in vitro-generated microglia will offer opportunities to optimize the development of advanced cellular systems to elucidate the specific roles of microglia in the pathophysiology of ALS, as well as other neurodegenerative diseases.
Supplemental Material
sj-xlsx-1-asn-10.1177_17590914221145105 - Supplemental material for Comparing the Characteristics of Microglia Preparations Generated Using Different Human iPSC-Based Differentiation Methods to Model Neurodegenerative Diseases
Supplemental material, sj-xlsx-1-asn-10.1177_17590914221145105 for Comparing the Characteristics of Microglia Preparations Generated Using Different Human iPSC-Based Differentiation Methods to Model Neurodegenerative Diseases by Ye Man Tang, Nisha S. Pulimood and Stefano Stifani in ASN Neuro
Supplemental Material
sj-xlsx-2-asn-10.1177_17590914221145105 - Supplemental material for Comparing the Characteristics of Microglia Preparations Generated Using Different Human iPSC-Based Differentiation Methods to Model Neurodegenerative Diseases
Supplemental material, sj-xlsx-2-asn-10.1177_17590914221145105 for Comparing the Characteristics of Microglia Preparations Generated Using Different Human iPSC-Based Differentiation Methods to Model Neurodegenerative Diseases by Ye Man Tang, Nisha S. Pulimood and Stefano Stifani in ASN Neuro
Footnotes
Acknowledgments
The authors thank Panagiotis Douvaras and Valentina Fossati for invaluable advice on microglia derivation, and Thomas Durcan and Jean-Francois Cloutier for providing access to scientific equipment. We also thank Rita Lo, Louise Thiry, Vincent Soubannier, Gilles Maussion, and Carol Chen for assistance, discussions, and advice.
Author Contributions
Y.T. performed all tissue culture and microscopy studies. N.S.P. performed all bioinformatics studies. S.S. and N.S.P. wrote the manuscript. S.S. supervised the study. All authors have read and agreed to the published version of the manuscript.
Data Sharing Statement
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: These studies were funded by the ALS Society of Canada, and Canadian Institutes for Health Research and Fonds de la recherche en Santé-Quebec under the frame of E-Rare-3, the ERA-Net for Research on Rare Diseases (SS). S.S is a Distinguished James McGill Professor of McGill University.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
