Abstract
Human induced pluripotent stem cells (hiPSCs) derived from healthy and diseased individuals can give rise to many cell types, facilitating the study of mechanisms of development, human disease modeling, and early drug target validation. In this context, experimental model systems based on hiPSC-derived motor neurons (MNs) have been used to study MN diseases such as spinal muscular atrophy and amyotrophic lateral sclerosis. Modeling MN disease using hiPSC-based approaches requires culture conditions that can recapitulate in a dish the events underlying differentiation, maturation, aging, and death of MNs. Current hiPSC-derived MN-based applications are often hampered by limitations in our ability to monitor MN morphology, survival, and other functional properties over a prolonged timeframe, underscoring the need for improved long-term culture conditions. Here we describe a cytocompatible dendritic polyglycerol amine (dPGA) substrate-based method for prolonged culture of hiPSC-derived MNs. We provide evidence that MNs cultured on dPGA-coated dishes are more amenable to long-term study of cell viability, molecular identity, and spontaneous network electrophysiological activity. The present study has the potential to improve hiPSC-based studies of human MN biology and disease.
We describe the use of a new coating substrate providing improved conditions for long-term cultures of human iPSC-derived motor neurons, thus allowing evaluation of cell viability, molecular identity, spontaneous network electrophysiological activity, and single-cell RNA sequencing of mature motor neurons.
Keywords
Introduction
A major goal when studying human neurodegenerative diseases is to generate, culture, and study human neuronal cells displaying disease-relevant phenotypes. The use of human induced pluripotent stem cell (hiPSC)-derived neurons is providing the opportunity to model a variety of neurodegenerative diseases, thus facilitating the development of new therapeutics. For instance, loss of motor neuron (MN) functions is associated with several devastating neurodegenerative diseases including spinal muscular atrophy and amyotrophic lateral sclerosis (ALS) (Kanning et al., 2010; Nijssen et al., 2017). Several protocols exist to obtain MNs from hiPSCs (e.g., Li et al., 2005; Amoroso et al., 2013; Qu et al., 2014; Du et al., 2015), and these cells have been used to study the pathophysiology of MNs from ALS and spinal muscular atrophy patients (e.g., Ebert et al., 2009; Egawa et al., 2012; Chen et al., 2014; Kiskinis et al., 2014).
An important goal of MN disease modeling using hiPSCs is to implement culture conditions that can faithfully recapitulate in a dish the processes underlying MN degeneration. This objective requires the ability to monitor molecular and functional properties of MNs, such as gene/protein expression, morphology, and electrical activity, in long-term cultures of MNs “aged” in vitro. However, prolonged culture of hiPSC-derived MNs is challenging with most commonly used MN derivation protocols. Most notably, hiPSC-derived MNs typically coalesce into large cell aggregates, and eventually detach from their substrate after continued culture, thus making long-term studies technically challenging (Kuijlaars et al., 2016; Taga et al., 2019; Thiry et al., 2020). Studies relying on endpoint assays requiring immunocytochemical staining, electrophysiological recordings of networks with multi-electrode array (MEA) devices, or single cell RNA sequencing (scRNAseq) applications are hindered by the presence of these large MN clusters, highlighting a critical need to develop improved hiPSC-derived MN culture conditions that would address these limitations and facilitate long-term studies.
Here we describe the use of a dendritic polyglycerol amine reagent (Hellmund et al., 2014; 2015) as a new coating substrate providing improved cytocompatible conditions for long-term cultures of hiPSC-derived MNs, thus allowing continued evaluation of cell viability, molecular identity, and spontaneous network electrophysiological activity. These culture conditions also facilitate scRNAseq studies of mature MNs due to decreased proportions of damaged cells when compared to more conventional coating protocols. These results demonstrate the value of using a dendritic polyglycerol amine substrate for real-time, long-term qualitative and quantitative analysis of hiPSC-derived MN cultures.
Materials and Methods
dPGA Substrate Synthesis and Application
Dendritic polyglycerol amine (dPGA: ∼95 kDa, 50% amine content) (Hellmund et al., 2014; 2015) was synthetized, characterized, and applied to coverslips used for several different neuronal cultures as described in the accompanying manuscript by Clement and colleagues (Clement et al., submitted).
Generation of Motor Neurons from Human iPSCs
Human iPSC line NCRM-1 (male) was obtained from the National Institutes of Health Stem Cell Resource (Bethesda, MD, USA). Human iPSC line CS29iALS-C9n1.ISOxx, corresponding to a corrected, isogenic, line derived from a parental line generated from an ALS patient with C9ORF72 gene mutation, was obtained from Cedars-Sinai (Los Angeles, CA, USA). Human iPSCs were differentiated into neural progenitor cells (NPCs), and subsequently MN progenitor cells (MNPCs) and MNs, as described previously (Thiry et al., 2020). After 24 days of differentiation, MNPCs were dissociated and plated at 50,000 cells/well on coverslips coated only with Matrigel (Thermo-Fisher Scientific; Cat. No. 08-774-552) or with dPGA plus Matrigel. For the latter condition, coverslips were coated with 300 µL/coverslip of a solution of PBS containing dPGA at varying concentrations (10, 25 or 50 µg/mL), and returned to the incubator for at least one hour before rinsing with PBS and coating with Matrigel on top of dPGA. MNs were differentiated in a chemically defined neural medium including DMEM/F12 supplemented with GlutaMAX (1/1; Thermo-Fisher Scientific; Cat. No. 35050-061), Neurobasal medium (1/1; Thermo-Fisher Scientific; Cat. No. 21103-049), N2 (0.5x; Thermo-Fisher Scientific; Cat. No. 17504-044), B27 (0.5x; Thermo-Fisher Scientific; Cat. No. 17502-048), ascorbic acid (100 µM; Sigma-Aldrich; Cat. No. A5960), antibiotic–antimycotic (1x; Thermo-Fisher Scientific; Cat. No. 15240-062), 0.5 µM retinoic acid, 0.1 µM purmorphamine, 0.1 µM Compound E (Calbiochem; Cat. No. 565790), insulin-like growth factor 1 (10 ng/mL; R&D Systems; Minneapolis, MN; Cat. No. 291-G1-200), brain-derived neurotrophic factor (10 ng/mL; Thermo-Fisher Scientific; Cat. No. PHC7074) and ciliary neurotrophic factor (10 ng/mL; R&D Systems; Cat. No. 257-NT-050). The MN culture medium was replaced every other day for 6 days and the MN cultures were characterized by immunocytochemistry as described below. For MEA recording, MNs were dissociated, plated at ∼50,000 cells/well on Matrigel-coated or dPGA/Matrigel-coated wells of a CytoView MEA 24-well plate (Axion Biosystems; Atlanta, GA, USA; Cat. No. M384-tMEA-24 W), and cultured with the same MN medium described above. Culture medium was replaced every three days for one month, during which cells were recorded every week as described below.
Characterization of Human iPSC-Derived Cells by Immunocytochemistry
Immunocytochemical characterization of induced MNs was performed as described previously (Methot et al., 2018). The following primary antibodies were used: mouse anti-HOMEOBOX PROTEIN HB9 (HB9) (1/30; DSHB; Iowa City, IA, USA; Cat. No. 81.5C10-c); mouse anti-ISLET1 (ISL1) (1/30; DSHB; Cat. No. 39.4D5-c); rabbit anti-LIMB HOMEOBOX CONTAINING 3 (LHX3) (1/100; Abcam; Toronto, ON, Canada; Cat. No. ab14555); goat anti-FORKHEAD BOX PROTEIN 1 (FOXP1) (1/100; R&D Systems; Cat. No. AF4534); mouse anti-neurofilament protein (2H3) (1/35; DSHB; Cat. No. 2H3-c); and mouse anti-non-phosphorylated neurofilament H antibody SMI32 (1/100; BioLegend; San Diego, CA, USA; Cat. No. 801702). The indicated DSHB monoclonal antibodies were obtained from the Developmental Studies Hybridoma Bank, created by the NICHD of the NIH and maintained at The University of Iowa, Department of Biology, Iowa City, IA 52242. Secondary antibodies against primary reagents raised in various species were conjugated to Alexa Fluor 488, Alexa Fluor 555, or Alexa Fluor 647 (1/1,000, Invitrogen; Burlington, ON, Canada). For quantification, images were acquired using an Axio Observer Z1 microscope connected to an AxioCam camera and using ZEN software (Zeiss). For each culture and each condition, images of >500 cells in 3 random fields were taken with a 20X objective and analyzed with Image J.
Multi-Electrode Array Recording of Human iPSC-Derived Motor Neuron Cultures
After 24 days of differentiation, hiPSC-derived MNs were plated at ∼50,000 cells/well on Matrigel-coated or dPGA/Matrigel-coated wells of CytoView MEA 24-well plates (Axion Biosystems). Recordings from 16 electrodes per well were conducted using a Maestro (Axion BioSystems) MEA recording amplifier with a head stage maintaining a temperature of 32˚C at 5% CO2. MEA plates were allowed to equilibrate for ∼3 min prior to the 5 min recording of spontaneous activity. Data were sampled at 12.5 kHz, digitized, and analyzed using Axion Integrated Studio software (Axion BioSystems) with a band-pass filter (200–5000 Hz). The spontaneous electrophysiological activity of the hiPSC-derived cells was recorded every 7 days post-plating for 3 weeks: at day-32 (one-week post-plating), day-39 (two weeks post-plating), and day-46 (three weeks post-plating). The following electrophysiological parameters were analyzed: percentage of active electrodes, spike frequency, and firing rate.
Preparation of Single-Cells Suspensions
Suspensions of single cells were prepared from MNs cultured for 39 days (day 0 defined as start of NPC induction), approximately 2 weeks post-plating on either Matrigel only or dPGA (50 µg/mL)/Matrigel, as previously described (Thiry et al., 2020). Cells were counted to evaluate cell concentration and viability before scRNAseq.
Single Cell RNA Sequencing
Cells were sequenced on a single-cell level using the microdroplet-based protocol 10X Genomics Chromium Single Cell 3′ Solution, followed by Illumina sequencing at the McGill and Genome Quebec Innovation Center (https://cesgq.com/en-services). The 10X cDNA libraries were sequenced at a depth of 50,000 reads per cell on a HiSeq4000 system (Illumina).
In Silico Analysis
The raw scRNAseq data were first processed using the 10X pipeline (Cell Ranger) to demultiplex and align the sequences to the version GRCh38 of the human genome. The gene/cell matrices produced by Cell Ranger were used in the quality control processing and analysis. All the subsequent data analysis was conducted in R version 3.5.2 using Bioconductor Software Version 3.8. All packages are publicly available at the Bioconductor project (http://bioconductor.org). The workflow used for the analysis was adapted from a workflow previously developed to characterize hiPSC-derived MNs (Thiry et al., 2020) adapted from a published workflow designed to accommodate droplet-based systems such as 10X Chromium (Lun et al., 2016). Gene ontology analysis was performed on the 500 most up- or down-regulated genes between the two conditions using the web-based functional annotation tool DAVID (https://david.ncifcrf.gov/), as previously described (Huang et al., 2009).
Statistics
To detect differences in the proportion of total MNs, lateral motor column (LMC) MNs, or median motor column (MMC) MNs according to the substrate used for coating (i.e.; Matrigel alone or Matrigel in combination with either 10 µg/mL dPGA, 25 µg/mL dPGA, or 50 µg/mL dPGA), nonparametric Friedman test and Dunn's multiple comparisons post hoc test were used, because the variables did not fit a normal distribution (assessed by Kolmogorov-Smirnov test). Kruskal-Wallis and Dunn's multiple comparisons post hoc test was used to detect differences in the number of single-cells remaining attached to the coverslips after one-week post-plating, according to the substrate used for coating (same as above). Two-way ANOVA and Tukey's multiple comparison post hoc test were used to detect differences in the percentages of single-cells remaining attached to the coverslips at different time-points (one-, two- and three-weeks post-plating) according to the substrate used for the coating (i.e.; Matrigel alone or Matrigel and either 10 µg/mL dPGA, 25 µg/mL dPGA, or 50 µg/mL dPGA). The statistical significance was set at p <0.05.
Results
Differentiation of Human iPSC-Derived Motor Neurons Cultured on dPGA/Matrigel
Spinal MNPCs and MNs were generated from the hiPSC NCRM-1 line as described previously (Thiry et al., 2020). While no single marker is completely MN-specific, spinal MNs are commonly characterized by transient co-expression of the LIM homeodomain transcription factors HB9 and ISL1/2 (Arber et al., 1999; Thaler et al., 1999) and most MN derivation studies rely on a combination of HB9 and ISL1 immunostaining (referred to as ‘pan-MN’ marker) (Amoroso et al., 2013) to characterize the MN phenotype in vitro. After 25 days of differentiation using standard culture conditions, 64% ± 6% of induced cells were pan-MN marker-positive (eg, ISL1/HB9-positive) (Figure 1A and 1C; “Matrigel”). A large proportion of the hiPSC-derived ISL1+/HB9+ MNs (48% ± 9%) expressed FOXP1, a known marker of limb innervating MNs of the LMC (Dasen et al., 2003 and 2008; Amoroso et al., 2013) and did not express LHX3 (Thaler et al., 2002; Agalliu et al., 2009), thus exhibiting the typical LMC MN molecular profile (FOXP1+/LHX3−) (Figure 1B; top row; “Matrigel”; and Figure 1C; “LMC”). In contrast, only 14% ± 6% of the hiPSC-derived MNs were FOXP1− and expressed LHX3, corresponding to the molecular profile (LHX3+/FOXP1−) of MNs of the MMC (Figure 1B; top row; “Matrigel”; and Figure 1C; “MMC”).
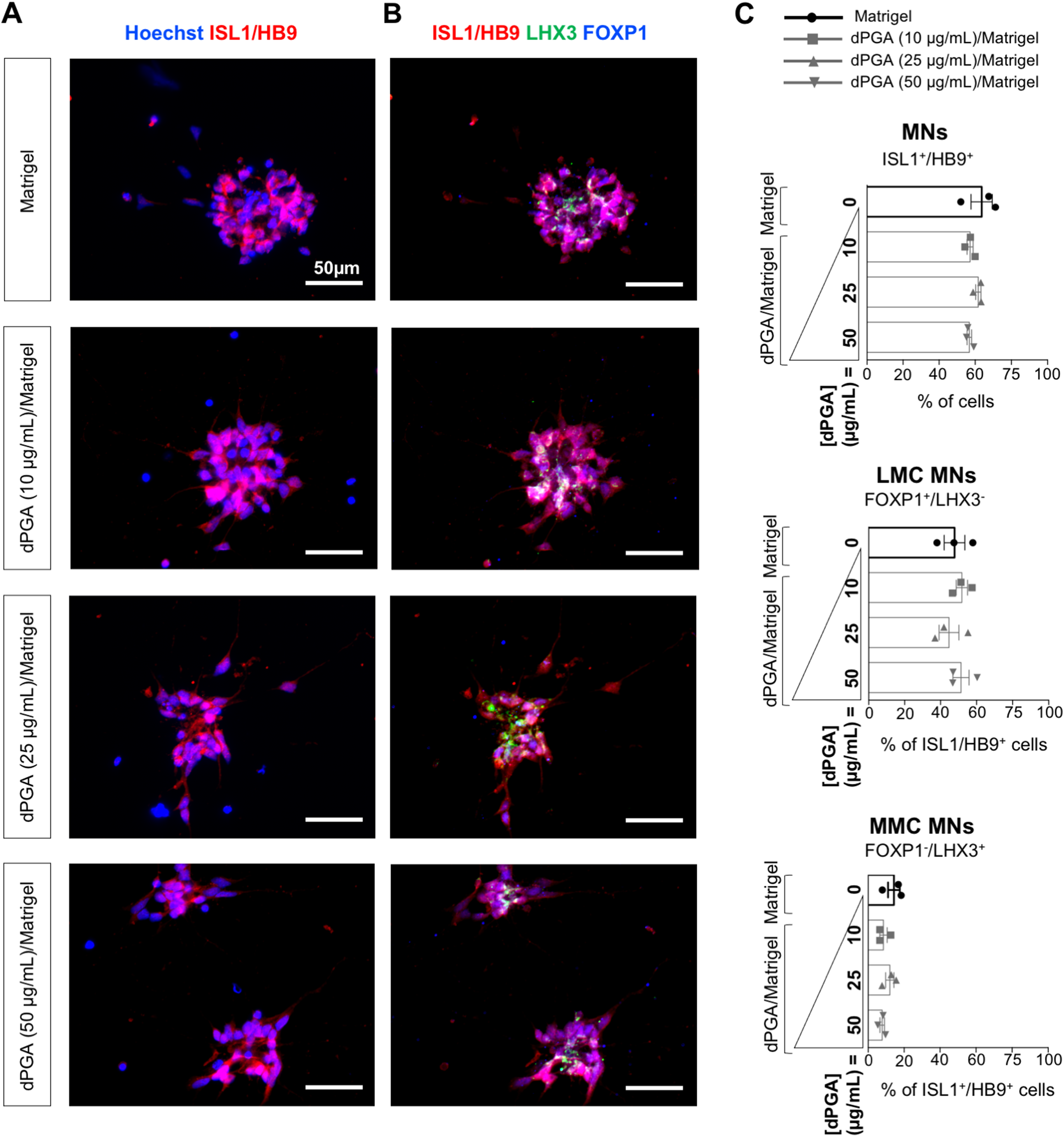
Efficient differentiation of hiPSCs into spinal motor neurons on dPGA/matrigel substrate. A-B) Representative images of MNs plated on Matrigel only vs dPGA (10, 25 or 50 µg/mL) and Matrigel combined, and co-stained with the known pan-MN marker ISL1/HB9 (red)
Our work and that of others has previously shown that hiPSC-derived MNs usually coalesce into large cell clusters, which gradually detach from the culture device (Kuijlaars et al., 2016; Taga et al., 2019; Thiry et al., 2020). This situation is most likely due to the degradation of the protein-based substrate (Matrigel or laminin) commonly used for coating, a process that can be mediated by enzymes secreted by the cells (reviewed in Schmidt et al., 2018; Clement et al., submitted). To address this shortcoming, we sought to test the effect of a non-peptide polymer substrate, dPGA, that was shown to improve primary neuron culture conditions, possibly due to its resistance to degradation by cell-secreted proteases (Clement et al., submitted).
We first asked whether the use of the dPGA compound as a coating substrate in combination with Matrigel would affect the differentiation of hiPSCs into MNs. Cells were plated on either Matrigel-coated or dPGA/Matrigel-coated coverslips, with varying concentrations of dPGA (10, 25 or 50 µg/mL), followed by immunocytochemistry after 25 days of culture (Figure 1). A similar proportion (∼60%) of the cells plated on Matrigel only or dPGA/Matrigel expressed the pan-MN marker ISL1/HB9 (Figure 1C). Furthermore, no statistically significant difference was found in the proportion of FOXP1+/LHX3− LMC MNs and FOXP1−/LHX3+ MMC MNs present in the cultures grown on either Matrigel only or dPGA/Matrigel, regardless of the concentration of dPGA used (Figure 1C).
Together, these observations show that the dPGA coating does not detectably interfere with the differentiation of hiPSCs into MNs, suggesting that it can be used for the culture of hiPSC-derived MNs.
Improved Long-Term Culture of Human iPSC-Derived Motor Neurons on dPGA/Matrigel
To test whether a dPGA coating would enhance long-term culture of hiPSC-derived MNs, we analyzed induced cultures up to three-weeks post-plating. As early as one-week post-plating on Matrigel only (i.e.; 32 days (D32) after the start of the differentiation protocol), hiPSC-derived MNs coalesced into large cell clusters, with their cell bodies crowded into clumps and their processes hyper-fasciculated into well-defined bundles (Figure 2A). In contrast, at the same time-point, hiPSC-derived MNs plated on dPGA/Matrigel remained more homogeneously distributed over the coverslip, with their processes creating widespread arborizations, regardless of the three tested dPGA concentrations (Figure 2B-D). The number of single cells remaining attached to the coverslip was systematically increased by the use of the dPGA substrate (Figure 2E). Interestingly, this effect appeared to be proportional to the concentration of dPGA used, with 10 µg/mL leading to a 4.7 ± 2.7-fold increase, 25 µg/mL leading to a 5.3 ± 1.8-fold increase and 50 µg/mL leading to a 7 ± 2-fold increase in the number of single-cells. Only the use of dPGA at 50 µg/mL led to a statistically significant effect, suggesting that this concentration is the most effective to improve hiPSC-derived MN culture conditions.
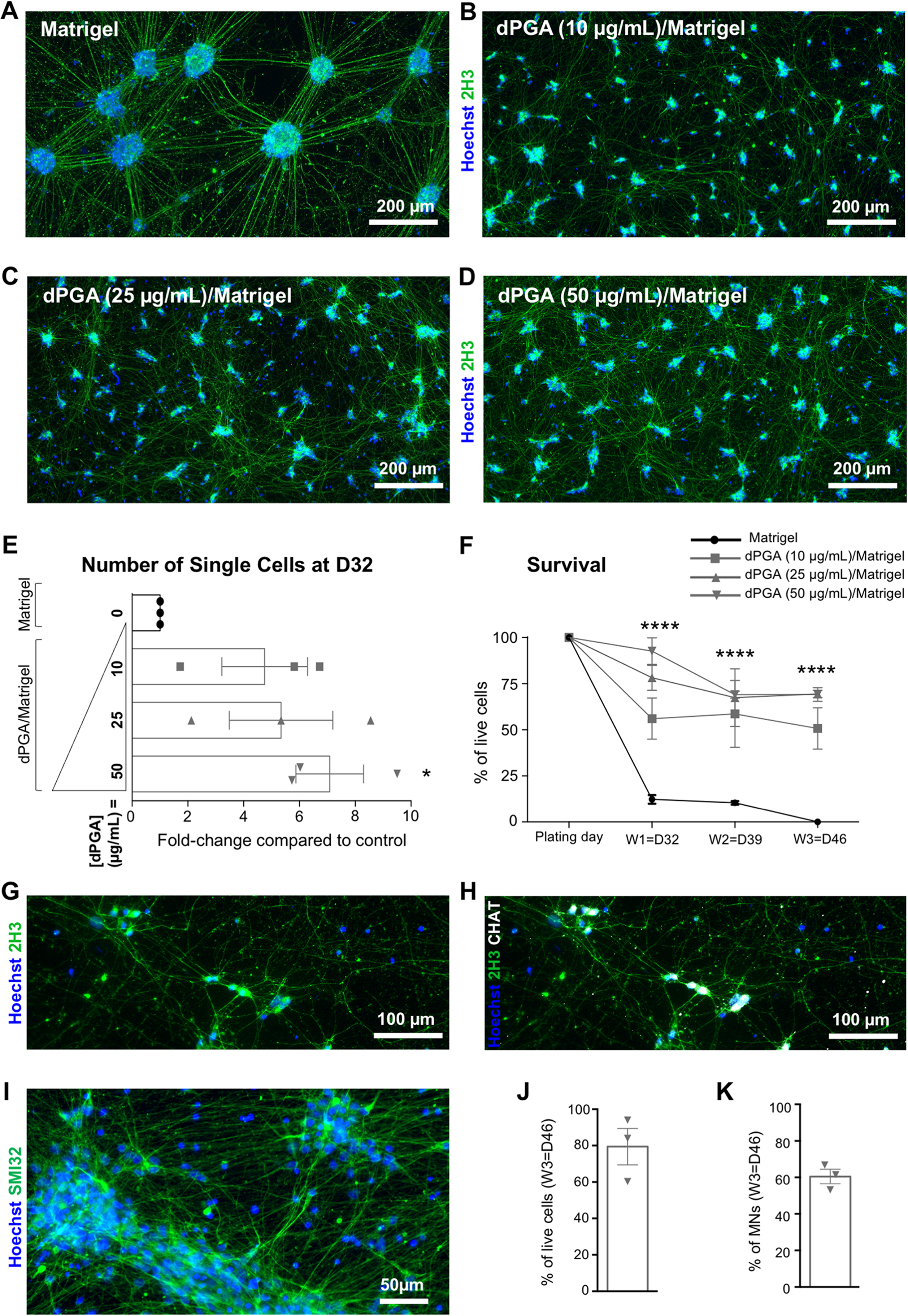
Enhanced long-term culture of human iPSC-derived motor neurons on dPGA/matrigel substrate. A-D) Representative images of NCRM1 hiPSC line-derived MNs one-week post-plating (i.e., 32 days of differentiation) on either Matrigel only
To evaluate whether this effect of the dPGA coating was stable over time, we next compared the percentages of cells remaining attached to Matrigel- or dPGA/Matrigel-coated coverslips after one week, two weeks and three weeks post-plating (Figure 2F). Both the type of coating substrate (e.g., Matrigel only vs. dPGA/Matrigel) and the culture time significantly affected cell adherence to the coverslips. As early as one-week post-plating, less than 10% of the cells plated on Matrigel-coated coverslips remained attached and this number decreased over time with all cells becoming detached at three weeks post-plating (Figure 2F, black line). In contrast, a minimum of 50% of the cells plated on dPGA/Matrigel-coated coverslips remained attached over-time, even three weeks after plating (Figure 2F, grey lines) (p < 0.0001; Two-way ANOVA). This effect was proportional to the concentration of dPGA used, although each of the three tested concentrations led to a statistically significant increased adherence when compared to Matrigel only. A dPGA concentration of 10 µg/mL was significantly less efficient than 25 or 50 µg/mL, and there was no statistical difference between the effect obtained with a dPGA concentration of 25 or 50 µg/mL. As shown in Figure 2G-H, MNs grown on dPGA (50 µg/mL)/Matrigel could be maintained in culture for at least 2-months post-plating without clumping, and they remained scattered over the coverslip and created widespread arborizations with their processes.
To confirm and extend these observations, MNs were derived from a separate human iPSC line (CS29iALS-C9n1.ISOxx), cultured for three weeks, and subjected to immunocytochemistry with a non-phosphorylated neurofilament protein (SMI-32) antibody, which is routinely used to mark MNs (Tsang et al., 2000). Human iPSC-derived MNs remained homogeneously distributed over the coverslips, with widespread arborizations of their processes, three weeks after plating on dPGA (50 µg/mL)/Matrigel (Figure 2I). An average of 79% ± 10% of the cells remained attached to the coverslip (Figure 2J), allowing quantitative analysis of day-46 cultures, which revealed that 60% ± 4% of the cells were SMI32+ mature MNs (Figure 2 K).
Together, these results provide evidence that using the dPGA substrate in conjunction with Matrigel improves the long-term culture conditions of MNs, allowing real-time, long-term qualitative and quantitative analysis of these cells.
Long-Term Multielectrode Array Recordings of Human iPSC-Derived Motor Neurons Cultured on dPGA/Matrigel
Electrophysiological characterization of hiPSC-derived MNs is crucial to assess their function. Multielectrode arrays are particularly suited for this purpose as they enable the recording of large neuronal populations through the detection of extracellular voltages. As reported by others (Kuijlaars et al., 2016; Taga et al., 2019), we observed that hiPSC-derived MNs plated on Matrigel-coated MEA wells formed large cell aggregates (Figure 3A). This made it difficult to record the electrophysiological activity of these cells, as reflected by a low number of electrodes with persistent activity during the three-week recording period (Figure 3C, black line). To test whether the use of dPGA/Matrigel coating could improve MEA analysis of hiPSC-derived MNs, we recorded hiPSC-derived MNs plated on either Matrigel- or dPGA/Matrigel-coated MEA plates, with varying concentrations of dPGA (10, 25 or 50 µg/mL), over a three-week period after plating (Figure 3). In contrast to the hiPSC-derived MNs plated on Matrigel alone, which coalesced into large cell clusters as early as one-week post-plating (Figure 3A), hiPSC-derived MNs plated on dPGA/Matrigel remained evenly distributed on the MEA wells (Figure 3B). As a result, each of the 16 electrodes in each of the dPGA/Matrigel-coated wells remained in the immediate vicinity of a number of cells for the three-week recording periods. This observation was reflected by a statistically significant increase in the number of electrodes with persistent activity over these recording periods in dPGA/Matrigel-coated wells (Figure 3C, grey lines) as compared to Matrigel-coated wells (Figure 3C, black line). This effect was consistent across the three tested concentrations of the dPGA substrate, without statistical difference between each concentration. Consequently, over the three-week recording periods, both the number of spikes (Figure 3D) and the mean firing rate (Figure 3E) were statistically significantly increased in dPGA/Matrigel-coated wells (grey lines) as compared to Matrigel-coated ones (black line). Together, these results suggest that using the dPGA substrate in combination with Matrigel enables stable long-term MEA recordings of hiPSC-derived MNs.
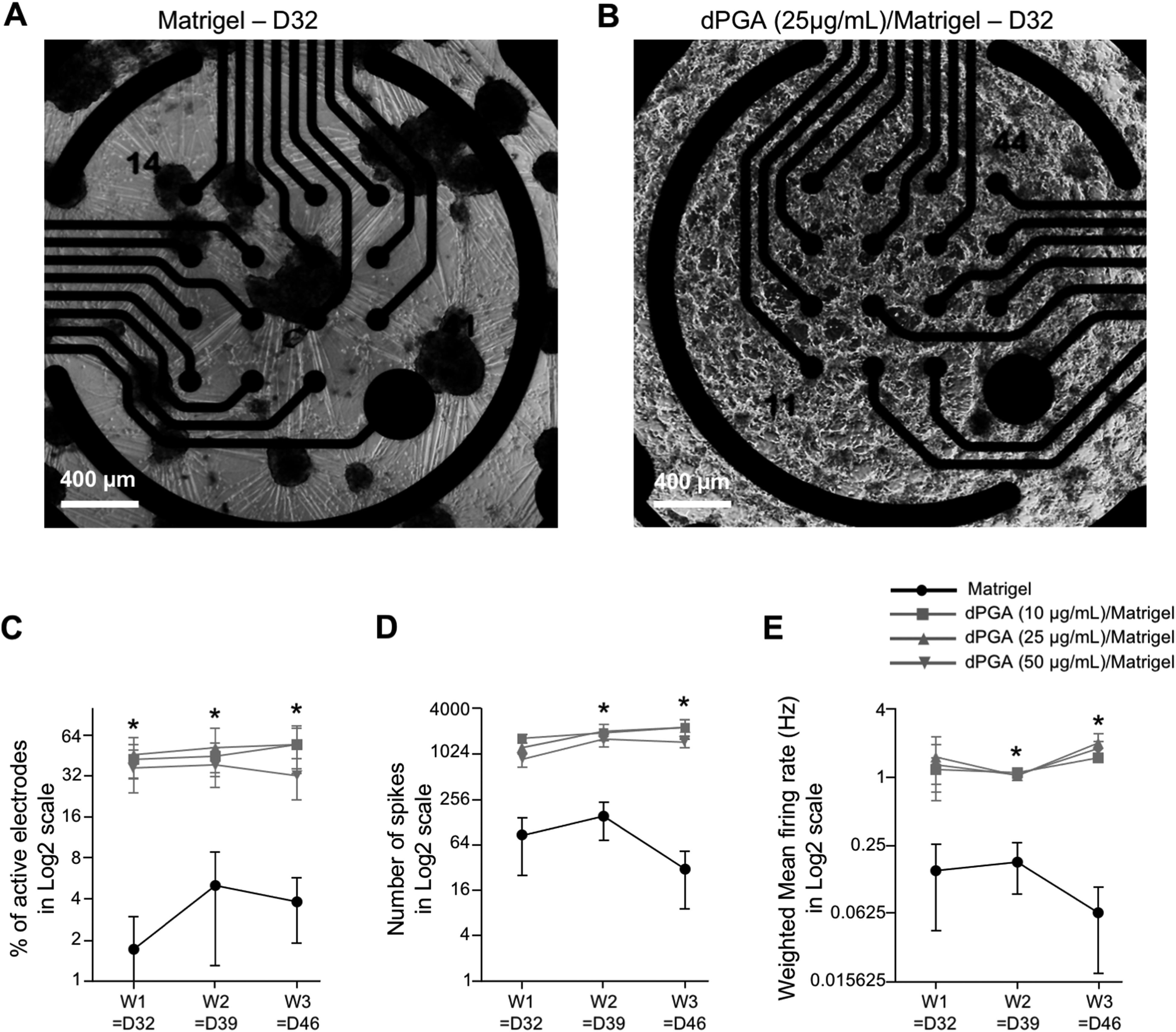
The dPGA/matrigel substrate enables long-term multielectrode array recordings of human iPSC-derived motor neurons. A-B) Representative images of hiPSC-derived MNs one-week post-plating (i.e., 32 days of differentiation) on MEA wells coated with either Matrigel only
Long Term Cultures of Human iPSC-Derived Motor Neurons on dPGA/Matrigel for Single-Cell RNA Sequencing
Microdroplet-based scRNAseq analysis provides a powerful tool to validate the quality and composition of hiPSC-derived MN cultures by enabling a precise characterization of the genome-wide expression profile of individual cells (Stegle et al., 2015; Thiry et al., 2020). As shown above, induced MNs already begin to coalesce into large cell clusters as early as one-week post-plating on Matrigel, making it challenging to perform scRNAseq studies due to the difficulty of isolating viable single cells. To address this situation, we evaluated whether the use of dPGA/Matrigel coating, which leads to reduced MN clustering, would facilitate the application of scRNAseq to the study of hiPSC-derived MNs. We submitted hiPSC-derived MNs plated on either Matrigel only (“Matrigel”) or 50 µg/mL dPGA/Matrigel (“dPGA/Matrigel”) to scRNAseq at 39 days of differentiation. After transcriptomic data (FASTQ files) were acquired from ∼6000 single cells for each sample, we used the Bioconductor software (Lun et al., 2016) to perform quality control (QC) steps to filter out damaged cells (Figure 4A to C). A total of 4375 cells of the “Matrigel” sample and 5857 cells of the “dPGA/Matrigel” sample passed QC. Interestingly, the median number of mapped reads per cell was lower in Matrigel as compared to dPGA/Matrigel conditions [Figure 4A; 6369 for Matrigel (top) and 9302 for dPGA/Matrigel (bottom)]. Similarly, the median number of genes expressed per cell was lower in Matrigel as compared to dPGA/Matrigel conditions [Figure 4B; 2685 for Matrigel (top) and 3168 for dPGA/Matrigel (bottom)]. Together, these QC metrics suggest a reduced proportion of damaged cells (i.e., the cells with lower numbers of detected genes and mapped reads) in the “dPGA/Matrigel” sample compared to “Matrigel”. In addition, the percentage of mitochondrial genes present in most cells was higher in Matrigel” compared to “dPGA/Matrigel” samples [Figure 4C; 10.3% for Matrigel (top) and 8.7% for dPGA/Matrigel (bottom)], suggesting a high proportion of damaged cells and/or debris in both samples, but in a reduced proportion in the “dPGA/Matrigel” sample, compared to “Matrigel”.
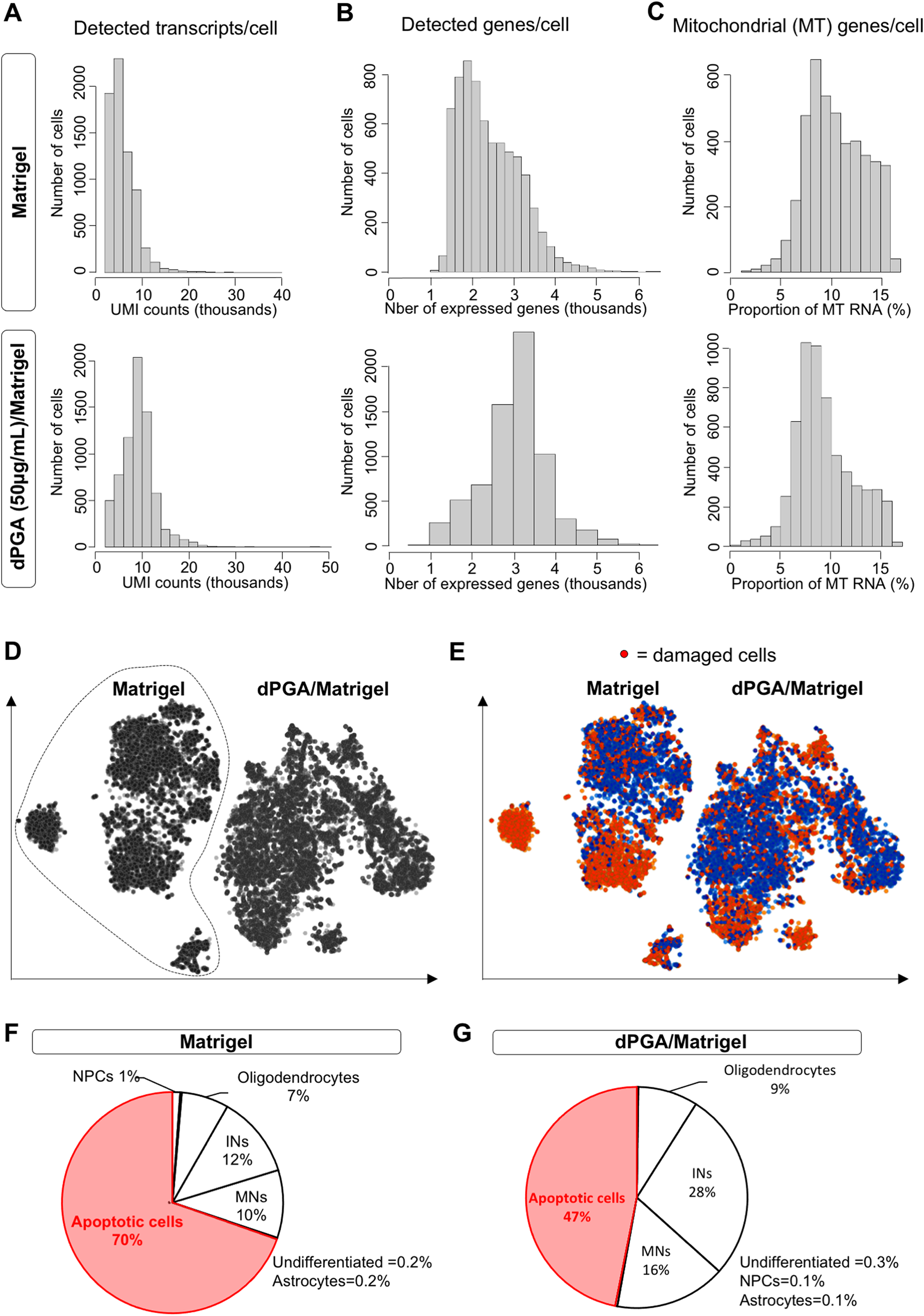
Reduced proportion of damaged cells after single cell RNA sequencing analysis of human iPSC-derived motor neuron cultures grown on dPGA/matrigel. A-C) Quality control metrics for hiPSC-derived MN cultures grown on either Matrigel only (top graphs) or dPGA (50 µg/mL)/Matrigel (bottom graphs) after 39 days of differentiation, with the number of detected transcripts per cell (
After QC, the remaining cells were submitted to principal-component analysis and t-distributed stochastic neighbor embedding (t-SNE) analysis (Figure 4D). In both samples, a large number of damaged/dying cells were identified, based on their low QC metrics and/or their high levels of expression of known markers of cell death such as BCL2, BAX, NGF, BOK, ATM, PSEN1, RB1, LIG4, CDK5, PARP1, BAD, MAP3K11, UBE2 M (Wei et al., 2001, D’Orsi et al., 2015; Frade & Barde, 1999) (Figure 4E). To further characterize this situation, we next performed integrative analysis to compare the cell proportions and gene expression differences between “Matrigel” (Figure 4F) and “dPGA/Matrigel” (Figure 4G) samples. Although a large proportion of cells were identified as “damaged cells” in both samples, this fraction was markedly reduced in the “dPGA/Matrigel” sample (47%; Figure 4G) as compared to “Matrigel” (70%; Figure 4F). Consequently, a larger proportion of cells were identified either as MNs and MNPCs (“MNs”; 16% in dPGA/Matrigel vs. 10% in Matrigel) based on their expression of the specific markers NEUROGENIN2 (NEUROG2), OLIG2, HB9, ISL1, ISL2 and CHOLINE ACETYL TRANSFERASE (CHAT), or as interneurons (“INs”; 28% in dPGA/Matrigel vs. 12% in Matrigel) based on their expression of the specific markers PAX2, PAX3, LBX1, EVX1, EN1, CHX10, GATA3, SOX14, SIM1, TLX3. Of note, other cell types were identified in relatively comparable proportions in both samples: oligodendrocytes accounted for approximately 7% of the cells in “Matrigel” and 9% in “dPGA/Matrigel”, while undifferentiated cells (iPSCs), NPCs and astrocytes were almost absent in both samples, as expected. Importantly, functional analysis of the most differentially expressed genes between the two culture conditions revealed a significant enrichment in Gene Ontology ‘Biological Processes’ terms related to apoptosis and the negative regulation of cell adhesion in the 500 most down-regulated genes in “dPGA/Matrigel”. In comparison, terms related to the regulation of transcription/translation were significantly enriched in 500 most up-regulated genes in the “dPGA/Matrigel” sample (Supplemental Figure 1).
Together, these scRNAseq results show that although it is technically challenging to isolate single MNs from ‘more mature’ hiPSC-derived cultures without causing cell damage in the process, the quality of single-cell suspensions for scRNAseq can be significantly improved by the use of dPGA/Matrigel coating during the maturation of hiPSC-derived MN cultures.
Discussion
Despite rapid advancements in iPSC-based technologies, current neuronal differentiation protocols tend to generate cells with properties of relatively immature neurons. This poses a problem when using iPSC-derived neurons to model age-related pathologies, including ALS (Arbab et al., 2014; Luisier et al., 2018; Mertens et al., 2015; Miller et al., 2013; Patani et al., 2012; Ho et al., 2016). Transcriptomic comparison of hiPSC-derived spinal MNs suggests that they are more similar to fetal than adult spinal cells (Ho et al., 2016; Luisier et al., 2018). Maturation- and age-related pathways are thought to play roles in MN disease pathology, highlighting the necessity to develop strategies to study ‘in vitro matured’ hiPSC-derived MNs to provide more disease-relevant iPSC-based models of ALS. However, hiPSC-derived MN maturation is made difficult in most current derivation protocols by the appearance of large MN aggregates, often as early as 5-7 days post-plating, which cause significant cell loss and hinder the analysis of single cells (Kuijlaars et al., 2016; Taga et al., 2019; Thiry et al., 2020). This shortcoming underscores a critical need to improve long-term culture of hiPSC-derived MNs to increase the fractions of viable single cells. Based on these observations, the present study aimed at developing enhanced culture conditions to facilitate the study of cell viability, molecular identity, and network electrophysiological activity in long-term hiPSC-derived MNs.
The extracellular matrix (ECM) performs a number of important functions, including mediation of cell–cell interactions, cell adhesion and migration, control of cell proliferation and differentiation, and apoptosis (Kharkar et al., 2013; Wang & Heilshorn. 2015). Over the past decades, the development of artificial ECM materials, for example via the use of ECM-mimicking substrates like Matrigel, has allowed the in vitro expansion and differentiation of iPSCs in a controllable, robust and scalable manner (reviewed in Schmidt et al., 2018). However, such protein-based substrates are susceptible to degradation by enzymes secreted by the cells, impacting on their use for long-term culture. To overcome this situation, non-peptide polymer substrates that are resistant to enzymatic degradation by the cells, like the cytocompatible dPGA, can be used in combination with peptide-based substrates to achieve a balance between resistance to protein degradation and lack of inherent biological activity of synthetic polymers. The accompanying manuscript by Clement and colleagues (Clement et al., submitted) provides further information on dPGA synthesis, application to coversplips used for cultures of a variety of neuronal types, as well as structure of dPGA-coated surfaces. In the present study, we have provided evidence suggesting that the use of this approach has the potential to specifically improve the study of hiPSC-derived MNs, by enhancing morphological, immunocytochemical, and survival studies, extracellular electrophysiological recordings, and single-cell RNA sequencing.
Our results have shown that dPGA coating in conjunction with Matrigel does not interfere with the differentiation of hiPSCs into MNs. More importantly, it results in a more homogeneous distribution of hiPSC-derived MNs, with widespread arborizations of their processes after the first week post-plating. These improved conditions result in a significant increase in MN survival over time, when compared to coating without dPGA. This advancement is expected to allow improved long-term qualitative and quantitative morphological and immunocytochemical analysis, as well as survival studies, of more mature hiPSC-derived MNs. This progress will benefit MN disease research, and ALS research in particular, based on hiPSC-derived MNs in a number of ways. For instance, it will facilitate the study of how different subtypes of spinal MNs, differing in gene expression, connectivity and function (William et al., 2003), are specifically vulnerable to degeneration in ALS (Chakkalakal et al., 2010). Being able to identify each subtype present in more mature iPSC-derived MN cultures, and to monitor changes in the morphology and survival rate of each MN subtype, will provide insight into the mechanisms responsible for driving differential susceptibility to degeneration. More generally, a growing number of studies of immature MNs derived from ALS patient iPSCs have shown their potential in recapitulating early morphological and disease-relevant molecular phenotypes, including RNA foci, protein inclusions, neurofilament aggregation, nucleoplasmic transport disruption, and cell death (Sareen et al., 2013; Chen et al., 2014; Kiskinis et al., 2014; Zhang et al., 2015). However, it remains to be determined how these phenotypes contribute to disease progression in more mature MNs. The availability of experimentally tractable ‘in vitro matured’ cultures will provide better model systems to answer these questions.
Beyond morphological, immunocytochemical and survival studies, the electrophysiological characterization of hiPSC-derived MNs is crucial to obtain accurate measures of their function. Multielectrode arrays are particularly suited for electrophysiological studies as they enable the extracellular recording of large populations of neurons, allowing for extended recordings that are important for a number of objectives, including drug testing. As shown in this study, and previously observed by others (Kuijlaars et al., 2016; Taga et al., 2019), hiPSC-derived MNs plated on a peptide-based substrate typically form a significant number of large cell aggregates that make it difficult to discern specific synaptic connections between the cells and to record their electrophysiological activity. We have shown that the plating of hiPSC-derived MNs on dPGA/Matrigel coated-MEA plates results in a more homogeneous distribution of the cells and a greater number of electrodes showing persistent activity overtime. Such improved experimental conditions will enable the monitoring of the electrophysiological activity of hiPSC-derived MNs over extended periods of time, an important goal of ALS research. For instance, analysis of MNs generated from C9ORF72 patient-derived iPSCs revealed that these cells display an initial period of hyperexcitability followed by a progressive loss in action potential output and synaptic activity at later timepoints (Devlin et al., 2015). These temporal changes could explain why several studies reported divergent findings regarding the inherent excitability of ALS iPSC-derived MNs (Sareen et al., 2013; Wainger et al., 2014; Naujock et al., 2016). While electrophysiological changes are a major phenotype in iPSC-based models of ALS, whether hyper- or hypo-excitability plays a crucial role in the disease remains unclear, highlighting the need to investigate the electrophysiological properties of ALS iPSC-derived MNs over extended periods of time in a stable and scalable manner.
The study of hiPSC-derived MNs using scRNAseq becomes increasingly challenging as cultures mature in vitro, most likely due to the harsher conditions needed to dissociate larger MN clusters. In previous studies, we have shown that scRNAseq data obtained from relatively immature hiPSC-derived MNs (28 days of differentiation) identified approximately 4% of dying cells (Thiry et al., 2020). In contrast, the present studies have revealed that a significantly larger proportion of more mature cells (39 days after differentiation) were identified as damaged cells using scRNAseq. However, the proportion of damaged cells in hiPSC-derived MNs cultures subjected to scRNAseq at later timepoints can be significantly reduced by the use of the dPGA/Matrigel combination as a coating substrate. This finding is expected to facilitate future scRNAseq studies of more mature hiPSC-derived MN cultures, thus enabling gene expression profiling of cells more likely to provide information on phenotypes that may be more relevant to MN disease pathophysiology.
In summary, the present study provides evidence that dPGA coating improves experimental conditions for the long-term culture of hiPSC-derived MNs, resulting in a more homogeneous distribution of the somas and a widespread arborizations of their processes, as well as a significant increase in MN survival over time. These improved experimental conditions will facilitate the study of more mature hiPSC-derived MNs, by enabling a better application of cell imaging techniques, electrophysiological recordings, as well as scRNAseq approaches, all of which are heavily reliant on the quality of long-term MN cultures. In turn, improved long-term culture of hiPSC-derived MNs will facilitate the study of phenotypes associated with neurodegeneration, thus providing a more informative experimental model system to develop cellular assays and facilitate early-stage drug discovery efforts in ALS and other age-related MN diseases.
Supplemental Material
sj-docx-1-asn-10.1177_17590914211073381 - Supplemental material for Optimization of Long-Term Human iPSC-Derived Spinal Motor Neuron Culture Using a Dendritic Polyglycerol Amine-Based Substrate
Supplemental material, sj-docx-1-asn-10.1177_17590914211073381 for Optimization of Long-Term Human iPSC-Derived Spinal Motor Neuron Culture Using a Dendritic Polyglycerol Amine-Based Substrate by Louise Thiry, Jean-Pierre Clément, Rainer Haag, Timothy E Kennedy and Stefano Stifani in ASN Neuro
Footnotes
List of abbreviations used
Acknowledgments
We thank Dr. Thomas Durcan for advice and for providing access to multielectrode array recording set-up, Dr. Ragoussis’ team at the McGill University and Genome Quebec Innovation Center for scRNA library preparation, as well as Rita Lo for invaluable assistance. We also acknowledge Cathleen Schlesener and Markus Hellmund for their help in the dPGA synthesis.
Author Contributions
LT designed and performed all experiments, data analysis, figures and wrote the first draft of the manuscript. SS, LT, J-PC and TEK conceived the study plan. RH provided and validated the dPGA. SS supervised data analysis and manuscript writing.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by grants to S.S. from the Canadian Institutes for Health Research and Fonds de la recherche en Sante-Quebec under the frame of E-Rare-3, the ERA-Net for Research on Rare Diseases, and from the Avrith Neuro-Cambridge Neuroscience Collaboration Initiative. SS is a James McGill Professor of McGill University.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
