Abstract
Ectonucleoside triphosphate diphosphohydrolase 2 (NTPDase2) hydrolyzes extracellular ATP to ADP, which is the ligand for P2Y1,12,13 receptors. The present study describes the distribution of NTPDase2 in adult rat brains in physiological conditions, and in hippocampal neurodegeneration induced by trimethyltin (TMT). The study also describes the regulation of NTPDase2 by inflammatory mediators in primary astrocytes and oligodendroglial cell line OLN93. In physiological conditions, NTPDase2 protein was most abundant in the hippocampus, where it was found in fibrous astrocytes and synaptic endings in the synaptic-rich hippocampal layers. In TMT-induced neurodegeneration, NTPDase2-mRNA acutely decreased at 2-dpi and then gradually recovered to the control level at 7-dpi and 21-dpi. As determined by immunohistochemistry and double immunofluorescence, the decrease was most pronounced in the dentate gyrus (DG), where NTPDase2 withdrew from the synaptic boutons in the polymorphic layer of DG, whereas the recovery of the expression was most profound in the subgranular layer. Concerning the regulation of NTPDase2 gene expression, proinflammatory cytokines IL-6, IL-1β, TNFα, and IFNγ negatively regulated the expression of NTPDase2 in OLN93 cells, while did not altering the expression in primary astrocytes. Different cell-intrinsic stressors, such as depletion of intracellular energy store, oxidative stress, endoplasmic reticulum stress, and activation of protein kinase C, also massively disturbed the expression of the NTPDase2 gene. Together, our results suggest that the expression and the activity of NTPDase2 transiently cease in neurodegeneration and brain injury, most likely as a part of the acute adaptive response designed to promote cell defense, survival, and recovery.
Keywords
Introduction
Extracellular purine and pyrimidine nucleotides, such as ATP and uridine triphosphate (UTP), regulate a wide variety of processes in the central nervous system (CNS), in health and disease (Ipata, 2011). In physiological conditions, nucleotides released from neurons (Fields, 2011; Pankratov et al., 2006; White, 1977), astrocytes (Koizumi, 2010), and microglia (George et al., 2015; Imura et al., 2013), participate in a multitude of processes, including cell proliferation, neurotransmission, and regulation of cell volume, blood-brain barrier permeability and cerebral blood flow (Carman et al., 2011; Narravula et al., 2000). In hypoxic/ischemic conditions, in the metabolic challenge (Juranyi et al., 1999; Melani et al., 2005) or after brain trauma (Davalos et al., 2005; Franke et al., 2006; Wang et al., 2004), ATP released from damaged or dying neurons, functions as an SOS signal for astrocytes and microglia (Bours, Dagnelie, et al., 2011; Bours, Swennen, et al., 2006; Rodrigues et al., 2015). The nucleotide initiates neuroinflammatory responses of microglia (Idzko et al., 2014; Koizumi et al., 2013) and induces the release of IL-1β, essential for morphological and functional activation of astrocytes (Cahill and Rogers, 2008; He et al., 2015). Moreover, the nucleotides produced by enzymatic degradation of ATP, i.e., ADP, AMP, and adenosine, also have a crucial role in the regulation of glial cell responses to an insult and inflammatory status of the tissue (Bours, Swennen, et al., 2006).
The actions of the extracellular purine nucleotides are mediated via P2 receptors, which comprise two families of ligand-gated receptor channels P2X1-7 and G-protein coupled receptors (GPRC) P2Y1−14 (Abbracchio et al., 2009; Khakh et al., 2001). ATP binds to and activates P2X1-P2X7 and a subset of ATP-sensitive P2Y purinoceptors, namely P2Y2 and P2Y11, the latter also being activated by UTP. The subtypes P2Y1, P2Y12, and P2Y13 mediate actions of ADP, while P2Y6 exhibits a preference towards UDP/ADP (Boeynaems et al., 2005). Typically, P2Y1, P2Y2, P2Y4, and P2Y6 are Gq/PLC-coupled membrane receptors, whereas P2Y12, P2Y13, and P2Y14 are the Gi/adenylyl cyclase-coupled receptors (Abbracchio et al., 2009). All purinoceptors, except P2Y4 and P2Y11 (Boehm, 2003) are abundantly expressed in the CNS, at neurons, astrocytes, oligodendrocytes, microglia, epithelial cells of the choroid plexus, and vascular endothelial cells (Burnstock and Knight, 2004), hence, the regulation of ligand availability for P1 and P2 receptor activation has a crucial impact on brain homeostasis and its response to injury (Illes et al., 2020). The second class of purinoceptors, P1, comprises four GPRC subtypes, namely A1, A2A, A2B, and A3, which mediate physiological actions of adenosine (Fredholm, 2007).
The availability of P1 and P2 receptor ligands is determined by the rate of ATP release and its hydrolysis by the ectonucleotidase enzymes. Major ectonucleoside triphosphate phosphohydrolases operating in the CNS are NTPDases1 and NTPDase2 (Robson et al., 2006; Zimmermann et al., 2012). Both ectonucleotidases hydrolyze purine and pyrimidine triphosphate and diphosphate nucleotides while differing in their substrate preference and cell localization (Kukulski and Komoszyński, 2003). Thus, NTPDase1 (CD39) (Kansas et al., 1991), mainly residing in microglia and vascular endothelial cells in the CNS (Langer et al., 2008), hydrolyzes ATP and ADP to AMP. The nucleotide monophosphate is further hydrolyzed to adenosine by ecto-5’-nucleotidase (eN/CD73) (Zimmermann, 1992). Accordingly, simultaneous induction of NTPDase1/eN, often seen in neuropathological conditions (Allard et al., 2017; Dragic et al., 2021b), eliminates P2 receptor ligands and potentiates P1-mediated anti-inflammatory actions of adenosine (Cunha, 2005).
Much less is known about the role of NTPDase2 in pathophysiological conditions (Kirley, 1997; Sévigny et al., 2002). The ectonucleotidase is mostly expressed in astrocytes
In the adult CNS, NTPDase2 exhibits wide distribution, but restricted cellular localization (Gampe, Hammer, et al., 2012; Gampe, Stefani, et al., 2015; Jakovljevic et al., 2017). Available immunohistochemical and
Our interest in NTPDase2 has been kindled by several recent
Materials and Methods
Animal Handling
A total of 44 two-month-old male rats of the Wistar strain were used for
Trimethyltin-Induced Neurodegeneration
On day zero (0-dpi), 36 animals received single
Preparation of Subcellular Fractions
Crude membrane fraction (P2) was prepared by following the protocol of Gray and Whittaker (1962). After isolation, brains were washed in the isolation buffer (0.32 M sucrose, 10 mM Tris-HCl, pH 7.4), dried, and the regions of interest were carefully dissected. The tissue was homogenized in 10 volumes of ice-cold isolation buffer (IB, 0.32 M sucrose in 5 mM Tris–HCl, pH 7.4) with a Teflon/glass homogenizer. Crude nuclear fraction and cell debris were removed by centrifugation at 1,000 × g for 10 min. The resulting supernatants were collected and centrifuged at 16,000 × g for 30 min to obtain crude membrane fraction (P2), which was resuspended in 5 mM Tris–HCl, pH 7.4. All steps were carried out at 4 °C. The samples were aliquoted and stored at −80 °C until use. The P2 fractions were separately isolated from four animals without pooling the tissue.
For purified hippocampal synaptosome (SYN) and gliosome (GLIO) preparations, the P2 fraction was placed on a discontinuous Percoll gradient (2, 6, 10, and 23% v/v Percoll in 0.32 M Tris-buffered sucrose and 1 mM EDTA, pH 7.4), and centrifuged at 35.000 × g for 5 min. The bands containing gliosomal and synaptosomal fractions were separated and recovered from 2- 6% Percoll interface and 10–23% Percoll interface, respectively. Both fractions were resuspended in the IB and washed from the Percoll reagent by double centrifugation at 12.000 × g for 15 min at 4 °C. The purity of the subcellular preparations was tested by immunoblotting, using specific molecular markers directed to GLIO (GFAP) and SYN (synaptophysin, SNAP25). GLIO was enriched in astroglial membrane fragments, while SYN was highly enriched in enclosed presynaptic and postsynaptic membrane fragments (Grkovic et al., 2019, Milanese et al., 2009). The protein content of the isolated subcellular fractions was determined by using bovine serum albumin (BSA) as a standard, as described previously. For functional assays, samples were used immediately after isolation, while for the immunoblotting procedures, samples were kept at −80 °C until use.
Immunohistochemistry/Immunofluorescence and Confocal Microscopy
Serial 25-μm sagittal paraformaldehyde-fixed cryosections were incubated in 0.1% H202 in methanol for 20 min at RT, washed 2 × 5 min in PBS, and blocked with 5% normal donkey serum (NDS) in PBS, for 1h at RT. After the incubation with rabbit anti-rat NTPDase2 antibodies (Ectonucleotidases-ab.com), at 4 °C overnight and 3 × 5 min washing in PBS, sections were incubated for 2h at RT in horseradish peroxidase (HRP)-conjugated secondary antibodies conjugated. The reaction was visualized with HRP chromogen, 3,3′-S-diaminobenzidine-tetrahydrochloride (DAB, Abcam, UK). The reaction was stopped with ddH2O, followed by dehydration in graded ethanol (70%, 95%, and 100%) and clearing in xylene. Sections were mounted with DPX medium (Sigma Aldrich, USA) and left to dry overnight. Sections were analyzed on LEITZ DM RB light microscope (Leica Mikroskopie & Systems GmbH, Wetzlar, Germany), under ×50 and ×200 magnifications, and digital images were captured by LEICA DFC320 CCD camera (Leica Microsystems Ltd., Heerbrugg, Switzerland) and LEICA DFC Twain software (Leica, Germany).
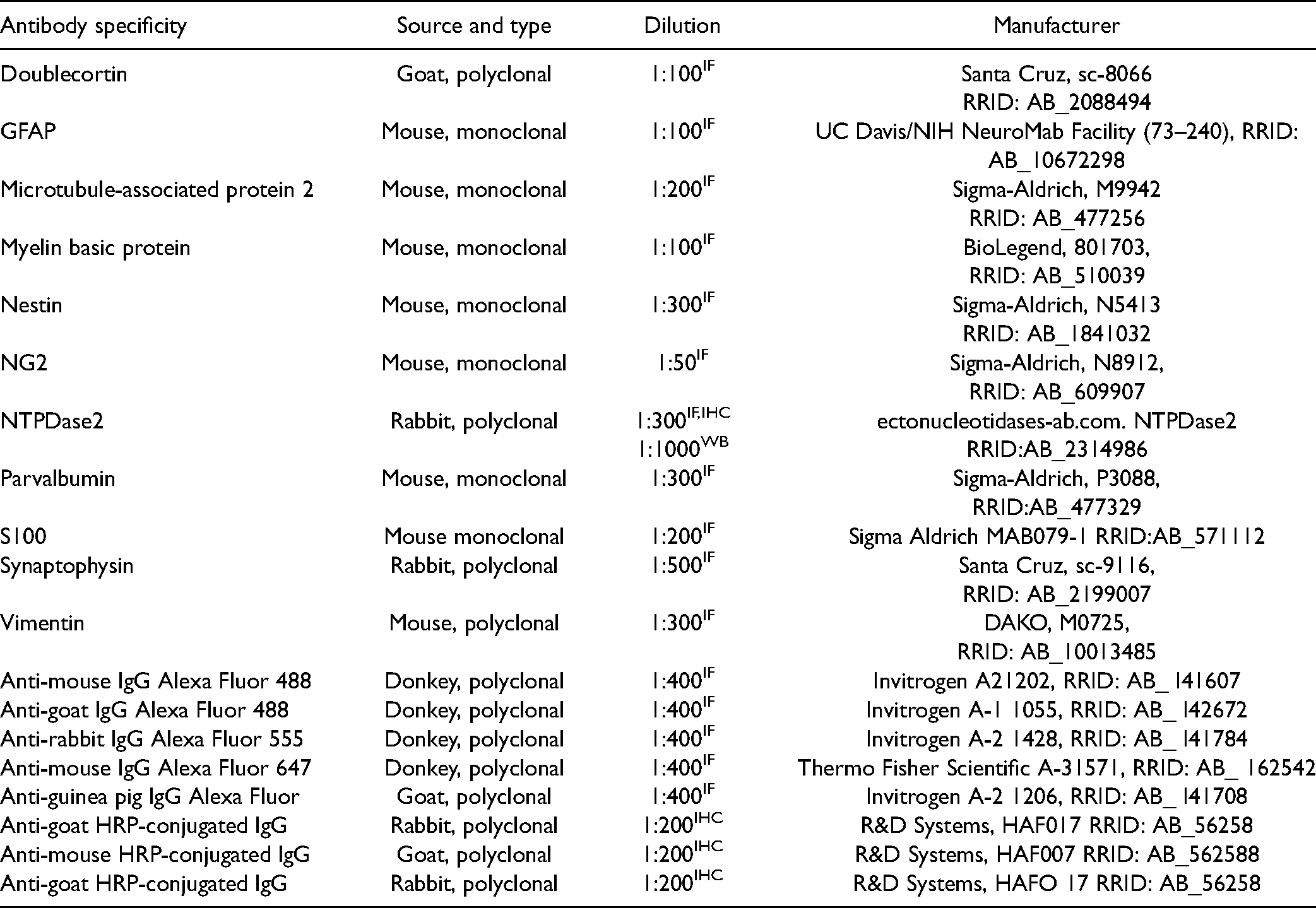
For confocal immunofluorescence, slides were air-dried 30 min, rehydrated in PBS (pH 7.4) for 10 min, and incubated in 5% NDS in PBS for 1h at RT. After incubation with primary antibodies (Table 1), overnight at 4 °C, and 3 × 5 min washes in PBS, sections were incubated with fluorophore-labeled secondary antibodies (Table 1), for 2h in a dark chamber at RT. After 3 × 5 min washing in PBS, sections were mounted with Mowiol (Sigma Aldrich, USA). Microscopic slides were stored at 4 °C until used for confocal laser scanning microscopy (LSM 510, Zeiss), with an argon laser (488 nm) and two helium-neon lasers (543 and 633 nm) at 400× magnification, equipped with a monochrome camera AxioCam ICm1camera (Carl Zeiss GmbH, Germany) and AxioVision software.
List of Primary and Secondary Antibodies.
Regions of interest were denoted according to The Rat Brain in Stereotaxic Coordinates by Paxinos and Watson (2004). High-resolution digital images (2,088 × 1,550 and 1,024 × 1,024 pixels) were saved as.tiff files. Basic image processing and linear adjustments were performed in
Cell Cultures and Treatments
Primary cortical astrocyte cultures were prepared from 1-day old neonatal rat puppies, as previously described (Adzic and Nedeljkovic, 2018), according to the protocols approved by the institutional Ethics Committee. Briefly, 1-day old rat cerebral cortices were dissociated by mincing and trituration and cells were seeded in a complete medium (Dulbecco's modified Eagle's medium, containing 10% heat-inactivated fetal bovine serum (FBS), 25 mmol/l glucose, 2 mmol/l L-glutamine, 1 mmol/l sodium pyruvate, 100 IU/ml penicillin and 100 μg/ml streptomycin) and cultured until a monolayer was formed (80–90% confluence). Primary microglia and oligodendrocytes were removed by vigorous shaking at 400 rpm for 16–20 h on a plate shaker (Perkin Elmer, Turku, Finland) and additional mechanical washing using the 1 ml pipette if needed. Adherent primary astrocytes were trypsinized (0.25% trypsin and 0.02% EDTA), and replated on 60-mm diameter dishes to multiply cells. Cells were treated after reaching 80–90% confluence with the following inflammatory factors: TNF-α, LPS (100 ng/ml), IL-1β, IL-6, IL-10, IFNγ (10 ng/ml), ATP, and adenosine (1 mM) for 8h (for rt-PCR) or 24h (for immunoblot).
OLN93 is a rat oligodendrocyte progenitor cell line, derived from spontaneously transformed cells in primary rat brain glial cultures (Richter-Landsberg and Heinrich, 1996). Cells were cultured and maintained in complete DMEM, at 37 °C and 5% CO2, as described elsewhere (Teske et al., 2018). The culture medium was changed every 2–3 days at 80–90% confluence. For the experiments, cells were seeded at 4 × 105 cells/well into Poly-D-lysine coated 6-well plates (Sigma-Aldrich, Munich, Germany). OLN93 were treated with TNF-α, IL-1β, IL-6, TGFβ (0.01 μg/ml), IFNγ (100 U/ml), and ATP (0.1 mM). To analyze whether blocking of mitochondrial respiratory chain activity alters the expression of NTPDase2, the following toxins were applied to OLN93 cells: Rotenone (complex-I inhibitor; 1 μM, diluted in dimethyl sulfoxide), antimycin (complex-III inhibitor; 10 μM, diluted in ethanol), sodium azide (complex-IV inhibitor; 10 mM, diluted in H2O), oligomycin (ATPase inhibitor; 10 μM, diluted in ethanol), tunicamycin (
For quantitative RT-PCR, cells were collected 8h after the treatment, in TRIzol® (astrocytes) or PeqGold® (OLN93). Total mRNA isolation, concentration determination, reverse transcription, and real-time PCR were conducted as described below. For Western blot analysis, cells were collected 24 h after the treatment, centrifuged at 500 × g for 5 min, and re-suspended in cold RIPA buffer, supplemented with a 0.5% w/v protease inhibitor cocktail. The suspension was centrifuged at 10,000 × g for 10 min, at 4 °C, the supernatant was separated from the pellet, and the protein concentration was determined using the protein assay kit (Pierce BCA protein assay kit).
Gene Expression Analyses
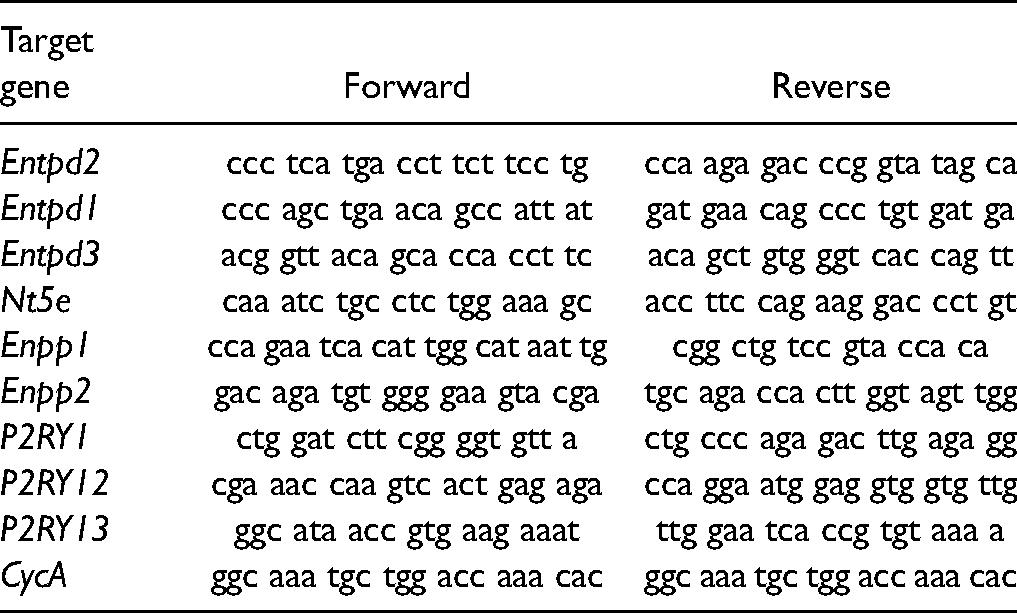
Total RNA was isolated from the hippocampal tissue (5/group) or collected cells, RNA concentrations were measured spectrophotometrically and the purity was determined by determining the A260/A280 and A260/A230 ratios. Volume equivalent to 1 µg of RNA was used for reverse transcription to generate cDNA (High Capacity cDNA Reverse Transcription Kit, Applied Biosystems, Foster City, CA, US), used for real-time PCR analysis (QuantStudioTM 3 Real-Time PCR System, Applied Biosystems, Foster City, CA, United States). The reaction mixture contained 2 μl cDNA (20 ng/μl), 5 μl QTM SYBR Green PCR Master Mix (Applied Biosystems, Foster City, CA, US), 0.5 μl primers (100 pmol/μl), and 2 μl RNase-free water (UltraPure, Invitrogen, Germany). The primer sequences were shown in Table 2. The amplification conditions were: 10 min of enzyme activation at 95 °C, 40 cycles of 15 s denaturation at 95 °C, 30 s annealing at 64 °C, 30 s amplification at 72 °C, and 5 s fluorescence measurements at 72 °C. To determine the specificity of the PCR reaction product we performed melting curve analysis and subsequent electrophoretic analysis. The results were expressed as the abundance of NTPDAse2 /CycA-mRNAs at each dpi relative to control (arbitrarily defined as 100%) ± SEM.
List of Primer Pairs for rt-PCR.

SDS-PAGE and Immunoblotting
Equivalent amounts of sample proteins (20 µg of P2 subcellular preparations or 10 μg of cell lysates) were resolved by denaturing SDS-PAGE and immunoblotted onto PVDF support membrane (0.45 mm, Millipore, Germany). After thorough washing in TBST (50 mM TRIS-HCl, pH 7.4, 150 mM NaCl, 0.05% Tween 20), the PVDF membrane was blocked (5% non-fat milk/TBST or 5% BSA/TBST), for 1 h at RT. The PVDF membranes were incubated with rabbit anti-rat NTPDase2 antibodies (Table 1), overnight at 4 °C. The membrane was washed 3 × 10 min in TBST and incubated with selected HRP-conjugated secondary antibodies (Table 1), for 1 h at RT. After several rounds of washing in TBST, the membrane was contacted with a peroxide reagent and luminal-containing solution (Bio-Rad Clarity Western ECL substrate), and the protein bands were visualized using chemiluminescence ChemiDoc-It®2 510 Imaging System. Protein bands obtained with the use of anti-β-actin-HRP conjugated antibodies at each membrane were used as a loading control. Densitometric analysis was performed using the
Statistical Analysis
In the immunoblot and rt-PCR experiments, the abundance of NTPDase2 was normalized to the level of β-actin and CyCA, respectively. The means ± SEM were calculated from
Results
Protein Abundance of NTPDase2 in Distinct Brain Areas
The specificity of rabbit anti-rat NTPDase2 antisera used for immunohistochemistry and immunofluorescence analyses was confirmed by immunoblotting. NTPDase2 was immunodetected in P2 fractions isolated from the olfactory bulb (Olf), cortex (Ctx), hippocampus (Hip), brain stem (BS), cerebellum (Crb), and caudate-putamen (CPu), with 1:500 dilution of the BZ3-4F/rN2-6L antisera. One protein band at about 55-kDa was detected in all tested samples (Figure 1A), while no band was detected in the presence of non-immune rabbit serum (data not shown), confirming the validity of the antisera for the use in NTPDase2-directed immunohistochemistry. The relative abundance of NTPDase2 protein was determined by calculating the OD ratio between the 55-kDa band and the β-actin band in each lane, expressed relative to the same value obtained for Ctx (arbitrarily defined as 1.0) (Figure 1B). The highest relative abundance of NTPDase2 protein was detected in Hip (1.44 ± 0.29,
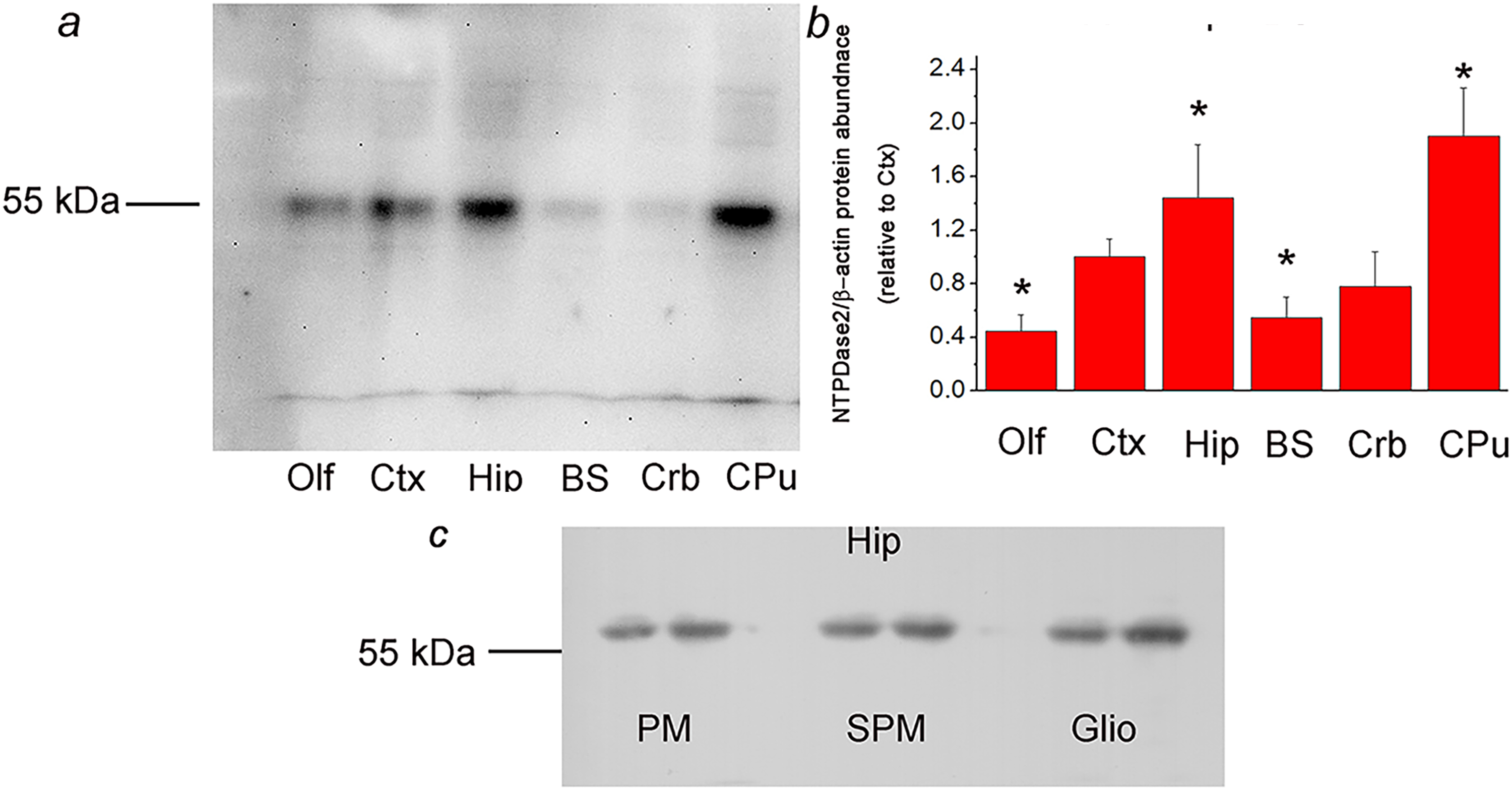
a: Crude membrane fractions isolated from the olfactory bulb (Olf), cortex (Ctx), hippocampus (Hip), brain stem (BS), cerebellum (Crb), and caudate-putamen (CPu), resolved by SDS-PAGE and probed with rabbit anti-rat NTPDase2 antisera (BZ3-4F/rN2-6L). Dash denotes the position of the specific 55 kDa band. b: Mean NTPDase2/β-actin protein abundance, expressed relative to Ctx (arbitrarily defined as 1.0) ± SEM (from
NTPDase2 protein was further immunoblotted in the plasma membrane (PM), synaptic membrane (SPM), and glial membrane (Glio) fractions, obtained by density gradient centrifugation of the hippocampal tissue. A single band migrating at about 55-kDa was detected in all samples, suggesting the presence of NTPDase2 protein in astrocyte membranes and synaptic membranes (Figure 1C).
NTPDase2 Immunohistochemistry and Immunofluorescence
The tissue expression of NTPDase2 in the forebrain is presented in Figure 2. Faint immunoreaction (
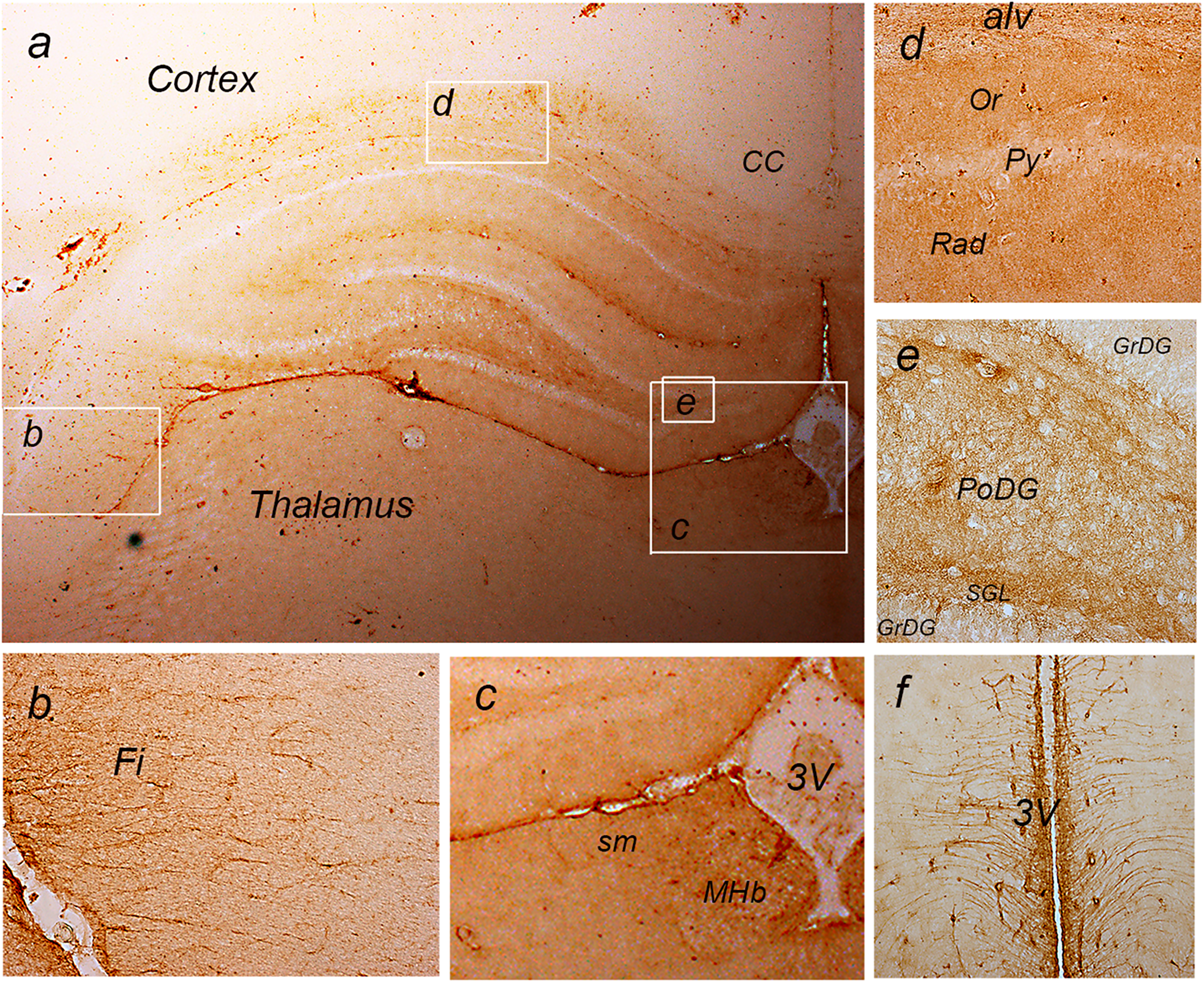
a: immunohistochemical labeling of NTPDase2 at coronal forebrain section. The white frames denote areas enlarged at b: The fimbria (
The association of NTPDase2 with glial cells and presynaptic elements was confirmed by double immunofluorescence directed to NTPDase2 in conjunction with astrocyte marker GFAP and presynaptic marker synaptophysin (Syn), respectively. Concerning astrocytic expression, we observed distinct patterns of immunoreactive signals (
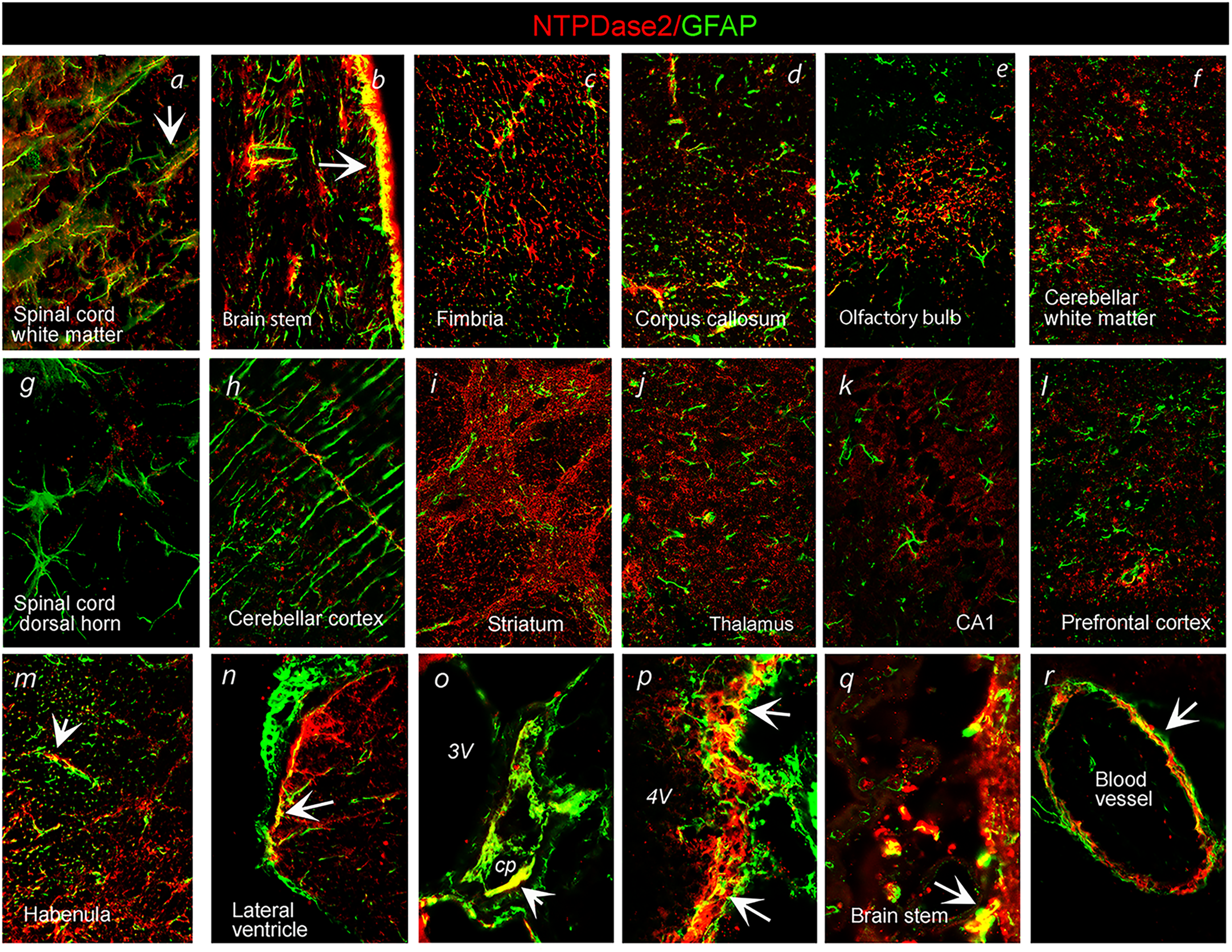
Double immunofluorescence labeling showing immunoreaction corresponding to NTPDase2 (
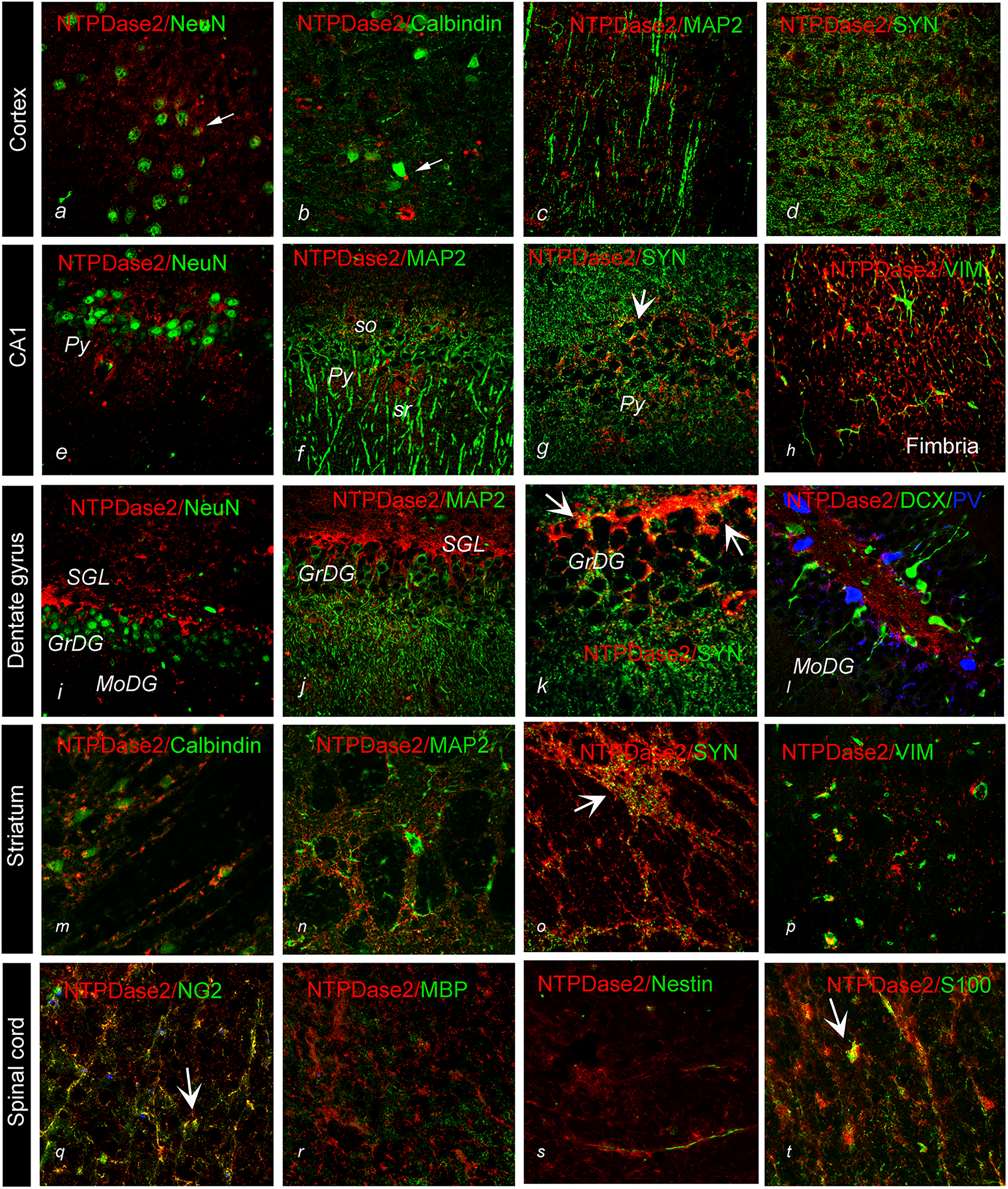
Double immunofluorescence labeling directed to NTPDase2 (
Concerning the association of NTPDase2 and Syn, significant overlap was found in the cortex, striatum, and hippocampus (Figure 4K). NTPDase2-
Expression of NTPDase2 After TMT-Induced Hippocampal Neurodegeneration
We next used the model of trimethyltin (TMT)-induced neurodegeneration to explore the expression of NTPDase2 in the hippocampal tissue. The neurotoxicant is known to induce progressive hippocampal cell death and astrocyte-driven neuroinflammation and microgliosis (Dragic, Zarić, et al., 2019; Dragic et al., 2021a). Two days after intoxication (2-dpi), NTPDase2-mRNA decreased to about 20% relative to control (
Hippocampal Expression of
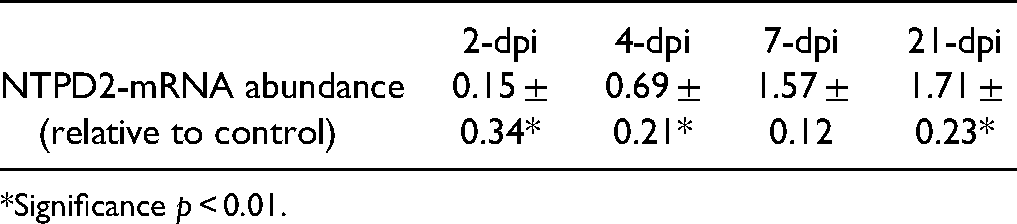

*Significance
Expression of NTPDase2 was more closely analyzed in the hippocampus at a histological level using NTPDase2-directed immunohistochemistry (Figure 5). The most prominent NTPDase2 labeling could be detected in the DG, particularly in the parenchyma of the polymorphic layer (
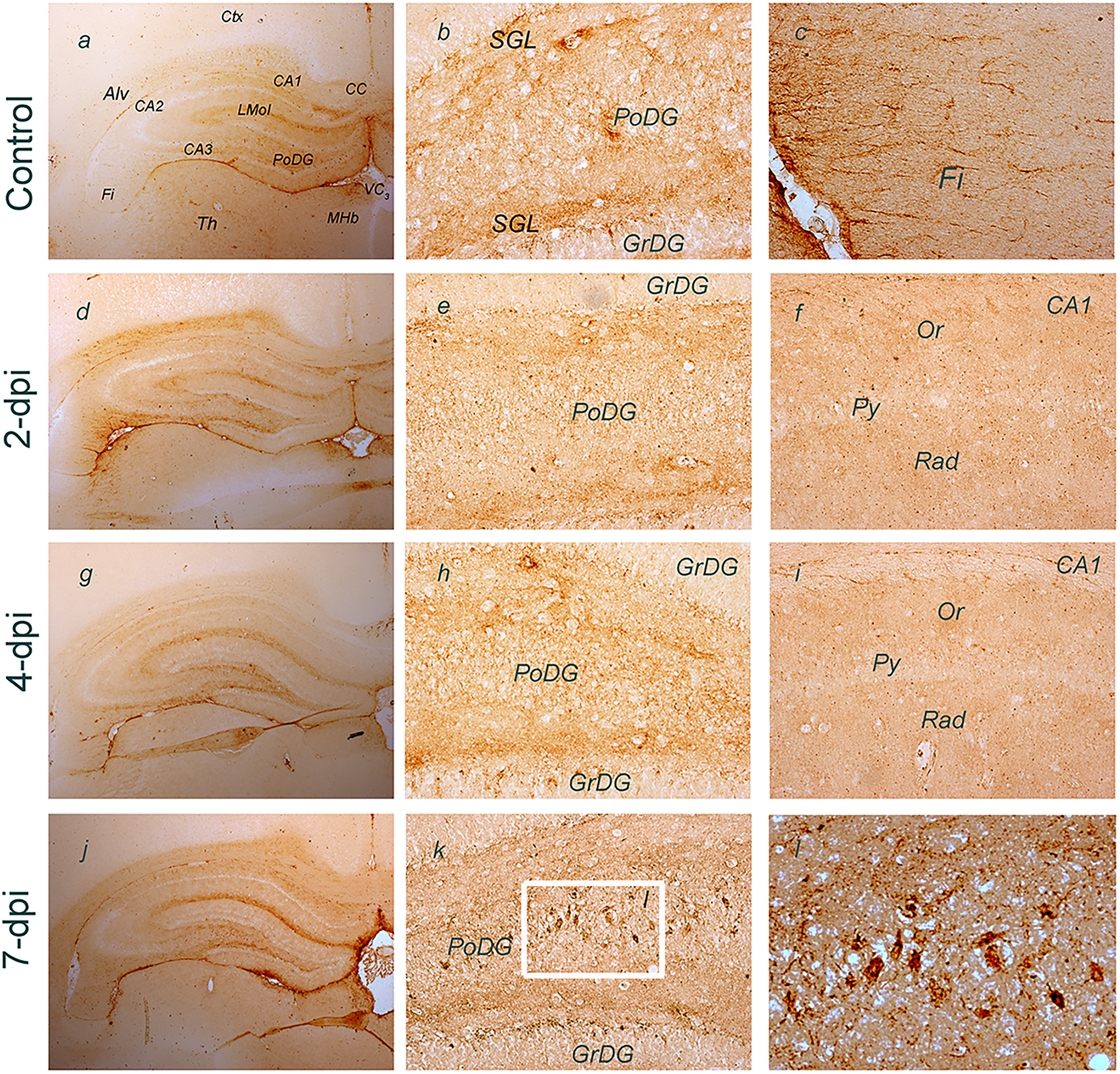
Immunohistochemical labeling of NTPDase2 at coronal hippocampal sections after TMT exposure. Abbreviations:
The cell types with altered NTPDase2 expression after TMT exposure were confirmed by double immunofluorescence (Figure 6). Double NTPDase2/GFAP-
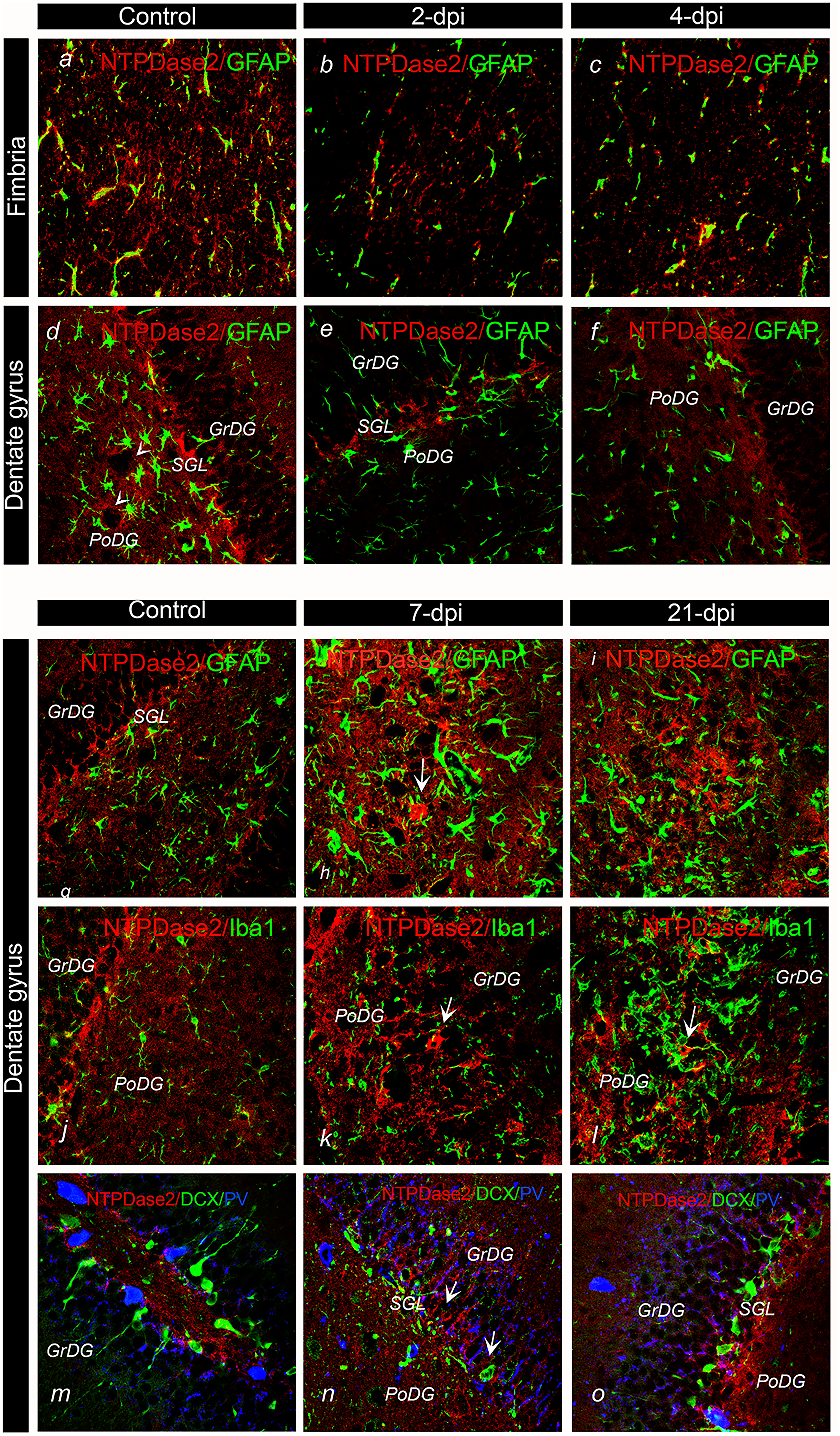
Immunofluorescence labeling directed to NTPDase2 (
Regulation of NTPDase2 by Inflammatory Mediators In Vitro
Factors that might regulate transcription of
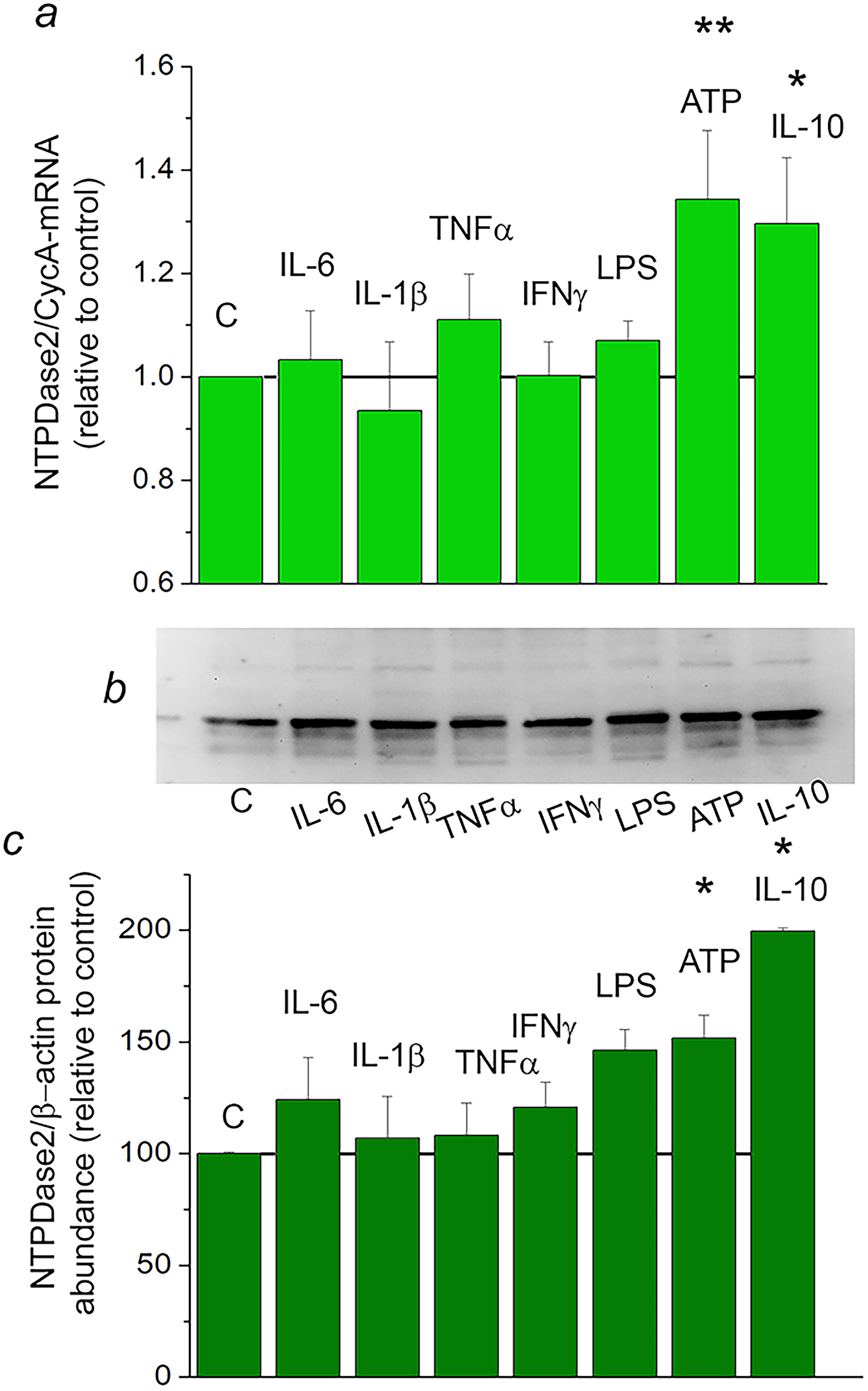
Regulation of NTPDase2 expression by inflammatory mediators in primary cortical astrocytes. a: NTPDase2/CycA-mRNA abundance in primary astrocytes treated with inflammatory factors for 8 h, expressed relative to non-treated control culture, ± SEM (from
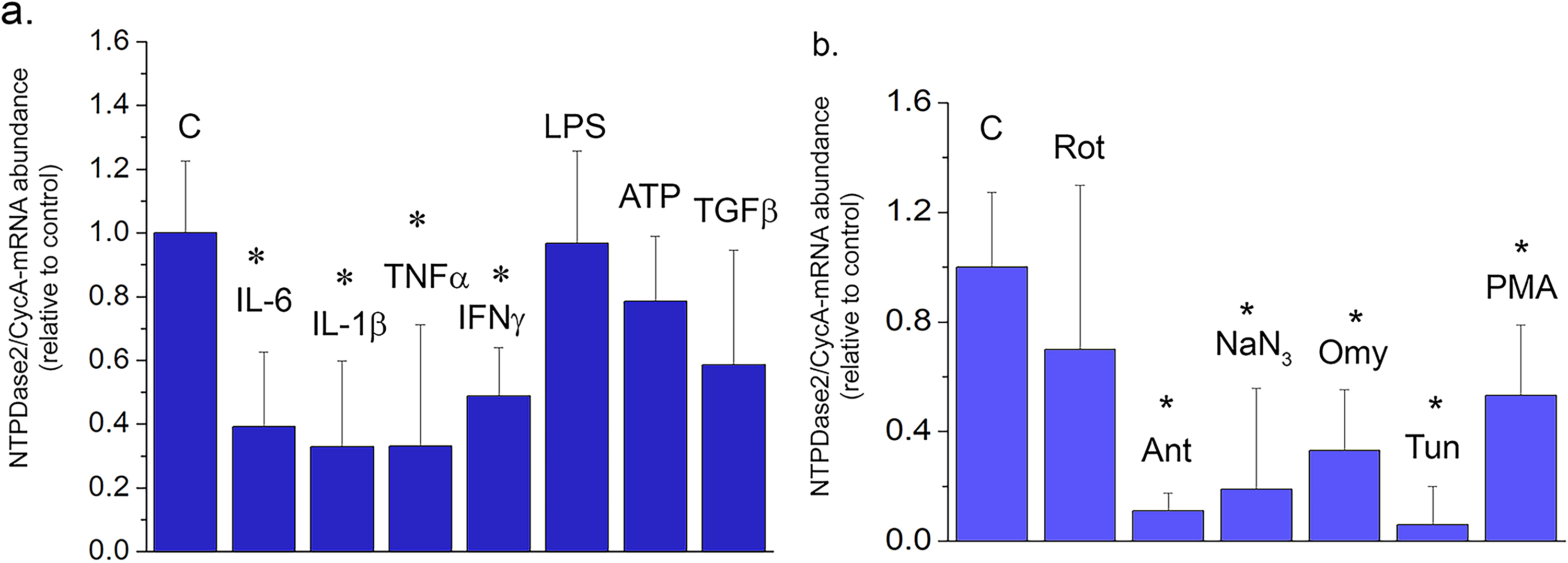
Regulation of NTPDase2 in OLN cell line. NTPDase2/CycA-mRNA abundance in OLN cells treated with (a) inflammatory mediators and (b) cell-intrinsic stressors, for 8 h. Results are expressed relative to non-treated control culture, ± SEM (from
Relative Transcript Abundance (2−ΔCt in Respect to CycA).

Significance level: *
Relative P2Y1R- and P2Y13R Transcript Abundance in Primary Astrocytes (Fold-Change Relative to Untreated Control).
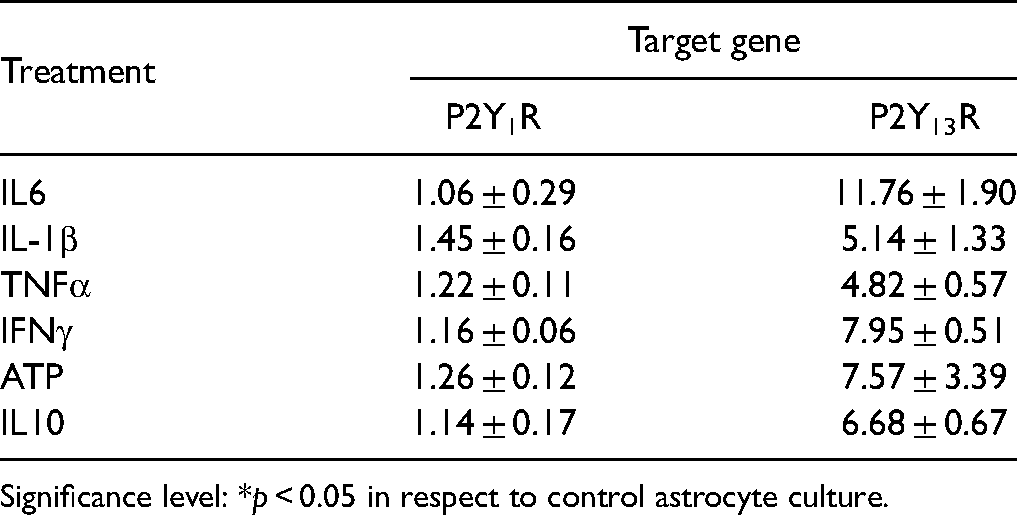

Significance level: *
Regulation of NTPDase2 was further assessed in the OLN93 cell line (Figure 8). The data showed that NTPDase2-mRNA and eN-mRNA were the only represented ectonucleotidase transcripts in the OLN93 cell line (Table 4). Exposure of OLN93 cells to pro-inflammatory cytokines IL-6, IL-1β, TNFα, and IFNγ decreased the abundance of NTPDase2-mRNA, whereas ATP and TGFβ did not exert a measurable effect, at least 8h after the exposure (Figure 8A). In light of the known sensitivity of OPC cells to oxidative stress, we also explored the impact of agents that affect cell metabolism and mitochondrial functions on
Discussion
In the present study, we sought to determine how NTPDase2 is affected and regulated under neuroinflammatory conditions
We next explored the NTPDase2 expression profile in the model of TMT-induced hippocampal neurodegeneration. Within the hippocampus as a whole, a significant drop in NTPDase2-mRNA abundance was observed at 2-dpi, followed by a gradual recovery to the control level from 4- to 7-dpi and an overcompensation at 21-dpi. A similar acute decrease in
To address the question of the pathophysiological significance of negative NTPDase2 regulation in neuroinflammatory conditions, we sought to identify cell types that downregulate NTPDase2 and to assess the overall neuroinflammatory context induced by TMT. Given that major changes in the expression of NTPDase2 occurred in DG, we focused the attention on this hippocampal subregion. Within DG, the expression of NTPDase2 decreased in presynaptic boutons innervating neurons in
Previous studies on the molecular mechanism of TMT action in astrocytes have shown that the neurotoxicant increases Ca2+ influx through L-type voltage-gated channels and inflicts oxidative stress and mitochondrial membrane depolarization and enhances the release of pro-inflammatory cytokines, including TNF-α, IFN-γ, IL-1β, and IL-6 (Dragic et al., 2021a; Geloso et al., 2011; Piacentini et al., 2008). Therefore, by using primary astrocytes and OLN93 oligodendroglial cells, in the present study we assessed the regulation of
In summary, the present study shows that expression of NTPDase2 is negatively regulated under neuroinflammatory conditions induced by TMT, mainly in fibrous astrocytes and presynaptic ending in
Footnotes
Acknowledgments
We would like to express gratitude to Prof. Jean Sevigny for providing rabbit anti-rat NTPDase2 antisera (BZ3-4F/rN2-6L) used in the study.
Author Contributions
All authors meet the International Committee of Medical Journal Editors (ICMJE) criteria for authorship for this article. N.N, I.G., and M.D. conceived and directed the project. M.D. and N.N. designed experiments, D.M and K.M. performed immunohistology, light, and confocal microscopy, M.Z.K. performed immunoblotting, N.M. performed rt-qPCR experiments. M.A., M.J., and M.K. performed the
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Ministry of Education, Science, and Technological Development of the Republic of Serbia, (grant number 451-03-1/2021-16/14, 451-03-68/2022-14/200178).
Data Availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
Ethics Approval
All animal procedures were performed in compliance with the European Communities Council Directive (2010/63/EU) and the National Laboratory Animal Science Association for the protection of animals used for experimental and scientific purposes and were approved by the Institutional Ethics Committee under authorization reference numbers 323-07-02057/2017-05 and EK-BF-2016/05.
