Abstract
No concerted investigation has been conducted to explore overlapping and distinct pathobiological mechanisms between repetitive mild traumatic brain injury (r-mTBI) and tau/amyloid proteinopathies considering the long history of association between TBI and Alzheimer’s disease. We address this problem by using unbiased proteomic approaches to generate detailed time-dependent brain molecular profiles of response to repetitive mTBI in C57BL/6 mice and in mouse models of amyloidosis (with amyloid precursor protein KM670/671NL (Swedish) and Presenilin 1 M146L mutations [PSAPP]) and tauopathy (hTau). Brain tissues from animals were collected at different timepoints after injuries (24 hr–12 months post-injury) and at different ages for tau or amyloid transgenic models (3, 9, and 15 months old), encompassing the pre-, peri-, and post-“onset” of cognitive and pathological phenotypes. We identified 30 hippocampal and 47 cortical proteins that were significantly modulated over time in the r-mTBI compared with sham mice. These proteins identified TBI-dependent modulation of phosphatidylinositol-3-kinase/AKT signaling, protein kinase A signaling, and PPARα/RXRα activation in the hippocampus and protein kinase A signaling, gonadotropin-releasing hormone signaling, and B cell receptor signaling in the cortex. Previously published neuropathological studies of our mTBI model showed a lack of amyloid and tau pathology. In PSAPP mice, we identified 19 proteins significantly changing in the cortex and only 7 proteins in hTau mice versus wild-type littermates. When we explored the overlap between our r-mTBI model and the PSAPP/hTau models, a fairly small coincidental change was observed involving only eight significantly regulated proteins. This work suggests a very distinct TBI neurodegeneration and also that other factors are needed to drive pathologies such as amyloidosis and tauopathy postinjury.
Mild traumatic brain injury (mTBI) is the most common (>80% cases) type of TBI within the military and civilian populations and is a major cause of disability and functional impairment. Epidemiological studies have reported that individuals exposed to a history of TBI are more susceptible to developing sporadic age-related neurodegenerative conditions in later life (Gedye et al., 1989; Mortimer et al., 1991; Fleminger et al., 2003; Li et al., 2017). The first clues linking TBI with pathological features of chronic neurodegenerative disease originated from studies reporting increased Aβ protein in the brains of patients who died acutely following TBI or in excised pericontusional injured tissue (Roberts et al., 1991, 1994; Ikonomovic et al., 2004; Dekosky et al., 2007). Neuroimaging studies with the amyloid sensitive Pittsburgh B compound have confirmed levels of amyloid beta in the gray matter and striata of TBI survivors comparable to mild cognitive impairment patients who go on to eventually develop dementia (Hong et al., 2014).
A neurodegenerative disorder, chronic traumatic encephalopathy (CTE), has also been reported, in a population of athletes exposed to repetitive mild TBI (r-mTBI; Omalu et al., 2011; McKee et al., 2013, 2016). CTE is marked by perivascular accumulation of phosphorylated tau (pTau) at the depths of the sulci. Although tau has been the major focus of autopsy cases of CTE, neuritic and diffuse Aβ plaques have also been reported at autopsy in 36% and 52% of CTE cases (respectively; Stein et al., 2015), and cerebral amyloid angiopathy has been linked with the pathological and clinical severity of CTE (Standring et al., 2019). Neuropathological reports have also shown similarities in other underlying histopathological features in TBI brains and age-related neurodegenerative diseases, such as white matter damage, vascular dysfunction, neuroinflammation, astrogliosis, and TDP43 pathology (Van Den Heuvel et al., 2007; Johnson et al., 2010; Perl, 2010; Omalu et al., 2011; Johnson et al., 2013; McKee et al., 2013; Smith et al., 2013; McKee et al., 2016), supporting the notion that TBI-induced neurodegeneration involves multiple pathobiological substrates. However, the precise nature of how TBI can lead to or precipitate this path toward tauopathy or amyloidosis remains elusive.
In-depth characterizations of biochemical cascades in TBI as it relates to tau and amyloid pathologies have been precluded by limitations inherent in human studies such as heterogeneity of TBI etiology, variances in demographics, lack of controlled prospective studies, and in vivo detection methods. We have therefore chosen a preclinical approach, in well-characterized mouse models of TBI and amyloidosis/tauopathy, for a time-dependent exploration of the overlap in pathogenic profiles and sequelae of events following TBI which may reveal how r-mTBI may lead to or precipitate these pathogenesis. We have extensively characterized a 5-hit mTBI model in both wild type and hTau mice, describing the behavioral and histopathological profiles from 24 hr to 24 months postinjury. Injured animals consistently exhibit persistent deterioration in reconsolidation of spatial memory and progressive white matter damage typified by thinning of the corpus callosum, gliosis, and axonal injury (Mouzon et al., 2012, 2014; Ojo et al., 2015; Ferguson et al., 2017; Mouzon et al.,2018a, 2018b, 2018c).
Several amyloid- or tau-driven transgenic models exist, such as the PSAPP and hTau models that represent amyloid and tau-based pathologies, respectively (Holcomb et al., 1998; Andorfer et al., 2003). The PSAPP model (PS1 [M146L], APP [K670N/M671L]) is a very well established and characterized mouse model of amyloidosis (Holcomb et al., 1998; Gordon et al., 2002; Sadowski et al., 2004; Trinchese et al., 2004) that demonstrates progressive age-dependent evolution of Alzheimer’s disease (AD) pathology accompanied by gliosis in an accelerated manner compared with single Tg2576 transgenic mice that carry the APP670/671 (“Swedish”) mutation alone. The hTau mice express all of the six isoforms of nonmutant human tau and lacks expression of murine tau (Andorfer et al., 2003). A few studies have characterized the age-related distribution and the pathology of hyperphosphorylation and tau aggregation, and glial activation in these mice, as well as the cognitive impairment (Duff et al., 2000; Andorfer et al., 2003; Polydoro et al., 2009).
At the outset of this study, we selected our wild-type C57BL6 mTBI model and the PSAPP and hTau models to study the complex molecular pathobiological response in TBI sequelae and tau/amyloid pathogenesis using an unbiased proteomics approach previously employed in our studies (Abdullah et al., 2011; Crawford et al., 2012; Zakirova et al., 2017; Ojo et al., 2018). During this longitudinal study, we observed and subsequently published a series of articles detailing the lack of amyloid and tau pathology from our neuropathological studies in our mTBI models up to 24 months postinjury (see Mouzon et al., 2012, 2014, 2018c). Herein, we use our proteomic platform to further pursue and explore the proteomic responses and any overlap or distinction between these different models.
For both TBI and tau/amyloid transgenic models, we have generated a comprehensive time line of omic responses by age or time postinjury, respectively. For the tau/amyloid transgenic models (hTau and PSAPP), we chose ages representative of pre-, peri-, and post-“onset” of the cognitive and pathological phenotypes (3, 9, and 15 months of age). For our TBI model (using C57BL/6 mice), we analyzed time intervals from 24 hr to 12 months postinjury, hypothesizing that 12 months would provide a sufficient postinjury timeframe to capture molecular signatures to enable comparison with the age-dependent neurodegenerative pathways observed in mice demonstrating tauopathy and amyloidosis. Owing to the lack of chronic amyloid and tau pathology in our 5-hit TBI model in wild-type mice (Mouzon et al., 2012, 2014, 2018c), our goal was not to show a direct TBI-dependent disease progression to AD in our injured mice. Thus, we have instead focused on exploring the distinctions and overlaps in the neurodegenerative pathways and sequelae in TBI and our chosen models of tauopathy and amyloidosis. We previously published plasma biomarkers in mice from this same TBI and tau/amyloid transgenic models (Ojo et al., 2018). We report our proteomic library of molecular changes from these studies in the brain following r-mTBI and in the pathological sequelae of amyloidosis and tauopathy and present in concert with our neuropathological findings largely distinct pathways between both models.
Methods
Animals
Wild-type C57BL/6 controls and hTau mice (Andorfer et al., 2003) were purchased from the Jackson Laboratories (Bar Harbor, ME). PSAPP mice expressing the double “Swedish” mutation in APP and the M146L mutation in PS1 were bred onto a C57BL/6 background in our vivarium. Their littermate controls were used as the wild-type controls. For the r-mTBI cohort, C57BL/6 mice (from Jackson Laboratories) were used. All the mice used in this study were male. All mice in the models of amyloidosis and tauopathy were aged until euthanasia at 3, 9, or 15 months, intended to represent ages pre-, peri-, and postonset of amyloidogenesis and tau pathology. For our r-mTBI study, mice were 3 months old at the time of their injury or sham injury and were euthanized at 24 hr, 3, 6, 9, or 12 months postinjury/sham. We have used exactly the same TBI and tau/amyloid transgenic mice and their chosen timepoints of evaluation from our previous article exploring blood proteomic biomarkers in TBI and models of amyloidosis and tauopathy (Figure 1). Please refer to our previous article for information on animal housing and conditions (Ojo et al., 2018). Proteomic analyses of r-mTBI mice were conducted in the hippocampus and cortex, while analyses of PSAPP/C57BL/6J and hTau/C57BL/6J mice were conducted in the cortex. Mice (N = 4/group) were assigned randomly to their groups, and all our analyses were designed blind to the experimenter. Studies performed herein were performed under a protocol approved by the Roskamp Institute Institutional Animal Care and Use Committee (IACUC - R054). All studies were in keeping with the Office of Laboratory Animal Welfare and National Institutes of Health guidelines.
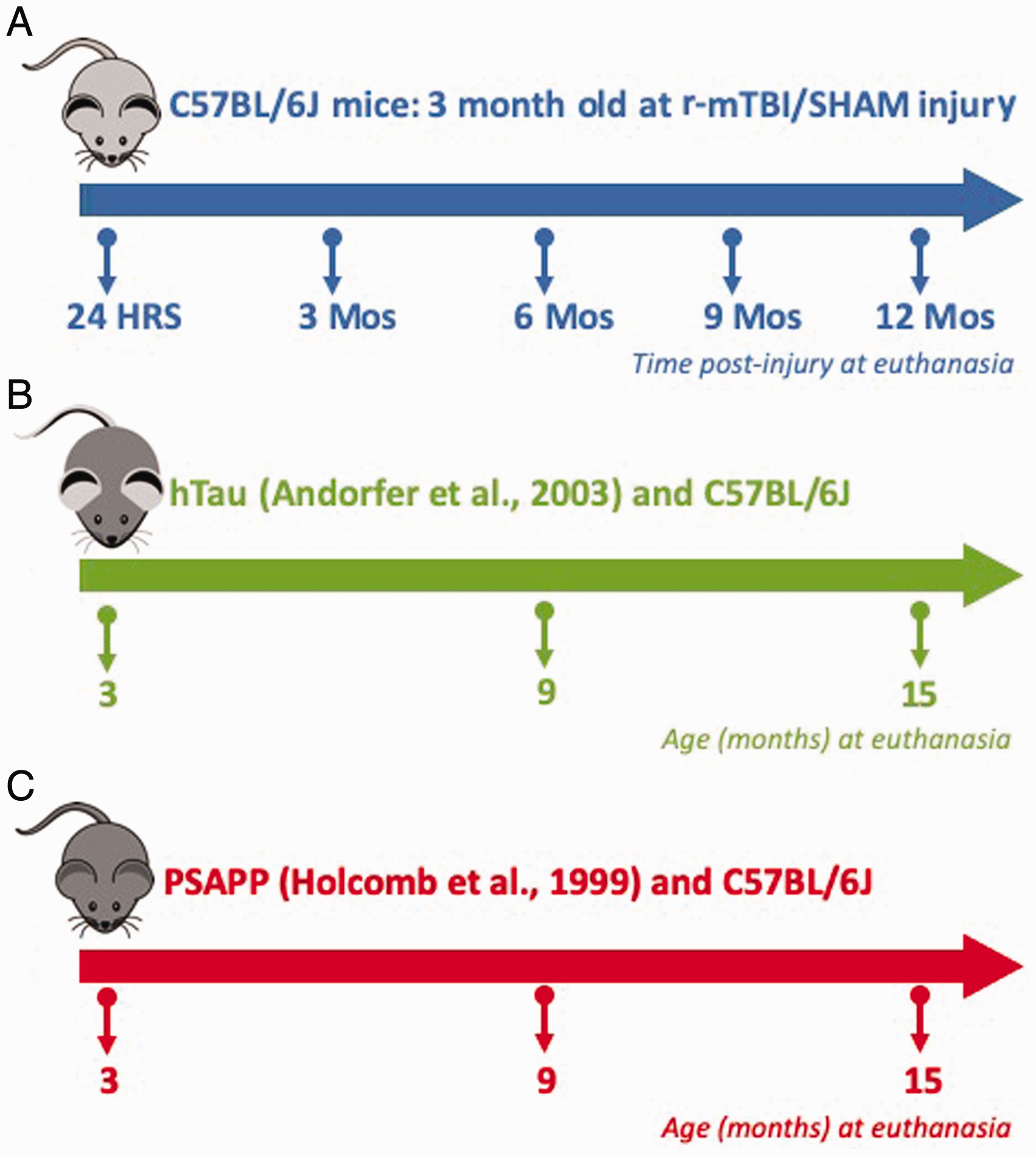
Proteomic Study Design and Time Lines of r-mTBI and Models of Amyloidosis and Tauopathy. This study used six groups of mice. The r-sham/r-mTBI group consisted of C57BL/6J mice at 3 months old, with euthanasia at 24 hr, 3, 6, 9, and 12 months post-TBI. Tauopathy group involved of hTau / C57BL/6J background strain mice with euthanasia at 3, 9, and 15 months old. Amyloidosis group involved PSAPP / C57BL/6J littermate mice with euthanasia at 3, 9, and 15 months old.
Experimental mTBI
Our experimental closed head injury paradigm followed procedures previously described (Mouzon et al., 2012). Procedures have been described in detail in Ojo et al., 2018. Briefly, we anesthetized sham/injury mice for 3 min with 1.5 L/min of oxygen and 3% isoflurane. We shaved the head of mice and placed them on a stereotaxic frame (Just For Mice Stereotaxic, Stoelting, Wood Dale, IL) that was mounted with an electromagnetic controlled impact device (Impact One Stereotaxic Motorized Impactor, Richmond, IL). Their body temperature was maintained at 37°C on a heating pad. A flat and blunt 5-mm metal impactor tip mounted onto the electromagnetic motorized device was zeroed on the shaved scalp, positioned above the midsagittal suture. After positioning, the metal tip was retracted and adjusted to a 1-mm depth strike. Scalp of mice were stretched to avoid delivering a trauma load that was inadequate and at an irregular angle. Parameters of injury were 5 m/s velocity, depth of 1.0 mm, 200 ms dwell time, and a projected 72 N strike force. This impact is sublethal; no skull fractures or subdural hemorrhages have been observed with our injury paradigm. r-mTBI mice were exposed to 5 hits over 9 days, with a 48-hr interinjury interval. r-sham mice were exposed to 3 min of anesthesia of the same frequency (×5). After injury, mice were placed on a 37°C heating pad before return to their normal cages.
Proteomics Experiments
Subcellular Protein Extraction From Brain Homogenates
Snap frozen brain tissues were weighed and homogenized in liquid chromatography and mass spectrometry (LC-MS) grade chilled water at a volume of 2.5× (cortices) and 4.5× (hippocampi) wet weight, respectively. Fifty microliters of homogenates were added to a 50-µl aqueous buffer solution containing 2× phosphate-buffered saline, 2× sodium chloride, and proteinase inhibitor (ThermoScientific, Waltham, MA). Samples in aqueous buffer were further homogenized with a probe sonicator and underwent centrifugation at 20,000g for 15 min at 4°C. Supernatant (nonmembrane fraction) was collected in a different Eppendorf tube to obtain cytosolic and nuclear protein fractions. Pelleted samples were resuspended in an aqueous solution containing 1× volume of ice-cold methanol (ThermoScientific, Waltham, MA) and 20,000g centrifugation for 15 min at 4°C. The precipitant was further resuspended in a 1× volume of ice-cold isopropanol and hexane (ThermoScientific, Waltham, MA) at a 2:1 dilution, followed by centrifugation at 20,000g for 15 min, and membrane protein pellet fractions were resuspended in 25 mM tetraethylammonium bromide (TEAB – Sigma-Aldrich, St Louis, MO) and 0.5% sodium deoxycholate (SDC – Sigma-Aldrich, St Louis, MO). Original supernatants (nonmembrane fractions) from the first centrifugation step were brought up in 0.5× volume of ice-cold methanol and centrifuged at 20,000g for 15 min, and the nuclear protein pellet fractions were resuspended in 25 mM TEAB and 0.5% SDC. Supernatant fractions from the last step was brought up in 7× volume of ice-cold methanol and centrifuged at 20,000g for 15 min, and cytosolic protein pellets were resuspended in 25 mM TEAB and 0.5% SDC. Membrane and cytosolic protein extracts were used in the entire study.
Trypsin Digestion
We added sequencing grade porcine trypsin (Promega, Fitchburg, WI) to aliquots from the hippocampus (50 µg) or cortex (100 µg), respectively (at a final protein concentration of 1 µg/µl), of depleted brain homogenate at an enzyme-to-substrate ratio of 1:100, followed by incubation at 37°C for 16 hr. Then, 10 µl of samples were taken to dryness in a vacuum centrifuge for labeling with tandem mass isobaric tags.
Tandem Mass Tag Labeling
To accommodate all timepoints/aged from all genotypes and sham/injury groups to be analyzed within the same batch, we used a multiplex tandem mass tag (TMT) kit. This enabled randomization of samples from all the different groups to be analyzed in the same batch with the inclusion of an internal reference per batch. Briefly, 6- and 10-plex TMT labeling kits (ThermoScientific, Waltham, MA) were used for analyses of samples from tau/amyloid transgenic (at 3, 9, 15 months of age) and sham/injury mice (24 hr, 3, 6, 9, 12 months postinjury), respectively. TMT labeling procedures have been previously described (see Ojo et al., 2018 for details). Sample IDs for each isobaric label tags were unknown to the experimenter. Labeled samples were taken to dryness in a vacuum centrifuge to remove acetonitrile before the SDC/TEAB cleanup procedure.
SDC–TEAB Cleanup
SDC/TEAB was removed from the dried TMT-labeled pooled samples using the steps described later. Dried samples were resuspended in 1% formic acid diluted in LC-MS grade water (at 100 µl) and centrifuged (20,000 × g RCF) for 1 min to remove precipitated SDC. Supernatant was collected, and 400 µl of ethyl acetate (ThermoScientific, Waltham, MA) was added. Samples were subsequently vortexed and centrifuged (at 20,000g) for 30 s to partition residual SDC into the upper (organic) layer; the resultant upper layers were discarded. This was repeated three more times, with the final lower phase taken to dryness and resuspended in 0.1% formic acid (Sigma-Aldrich, St Louis, MO) diluted in (100 µl) LC-MS grade water (ThermoScientific, Waltham, MA). Samples were then subsequently concentrated and desalted using C18 reversed phase ZipTips (MilliporeSigma, Burlington, MA) based on the manufacturer’s protocol. Eluates from the ZipTip procedure were resuspended in 0.1% formic acid (20 µl) and transferred into an autosampler vial, for analysis using a nano-UPLC MS on a Q-Exactive Orbitrap instrument (ThermoScientific, Waltham, MA).
Chromatography and Mass Spectrometry Methods (LC-MS/MS)
An LC-MS/MS (Q-Exactive) was used to analyze all samples in this experiment. Data-dependent acquisition settings for the experiments have been previously described in our earlier proteomic study (see Ojo et al., 2018 for more details). Our rationale for selecting these settings, especially our narrow isolation window combined with the ultra-long gradient, was to essentially minimize deleterious effects on quantitative accuracy that may result from coisolation of isobaric precursors without resorting to MS3-based methods.
Data Processing and Statistical Analysis
The animals were randomly assigned to multiplexes and the labels and analyzed blindly to prevent any unwanted experimental bias. We used the PMi Preview software to survey data files and add additional modifications to our search criteria. Preview results were also used to choose precursor/fragment ion mass tolerances (4 ppm, 0.02 Da, respectively) including dynamic modifications. Settings used to search the data using SEQUEST and MSAMANDA as the searches algorithms have been previously described (Ojo et al., 2018). We considered only unique peptides for quantification purposes. A false discovery rate (FDR) of 0.01 was set, with resulting datasets exported to JMP 8.0.2 (Cary, NY) for our statistical analysis. FDR of 0.01 is the default “stringent” setting in the proteome discoverer analytical software; it is common to use this stringent level of FDR for peptides, which considers that out of every 100 peptides identified 1 may not be real. We used only proteins found in at least 50% of the total number of plexes for our quantitative analysis. Data from raw ion counts were ln transformed, with each channel normalized using central tendency normalization method, whereby the medians were used. Ratios were generated by dividing datasets from each timepoint (3 month [3 m], 6 month [6 m], 9 month [9 m], or 12 month [12 m]) with 24 hr timepoint within the TBI and sham group separately per plex. Likewise for the study involving models of amyloidosis and tauopathy we divided data from each age-group (9 month [9 m], 15 month [15 m]) with 3 month [3 m] of age within the PSAPP and C57BL6 group separately per plex. Peptides that were not quantified in at least eight runs were omitted from the analysis. We used the mixed model analysis of variance (ANOVA) for analyses. The following linear model (1) was used per injury/genotype in animal models:
Analytical Approach
The dataset was analyzed using two different approaches. Our first approach was to divide values from each of the TBI animals with a randomly matched sham animal at a given timepoint to measure the response to TBI at each timepoint. However, because each TBI animal was being directly compared with a random sham animal instead of all TBI or sham animals being compared with a single reference animal, statistical noise was introduced to the data. Moreover, this type of ratio (TBI/sham), can only be analyzed using a method that tests if the ratio is significantly different from 0 (as the data is ln transformed). There were either no, or very small number of, proteins that were changing due to TBI using this statistical analysis approach. To reduce statistical noise and increase the power, the 24 hr timepoint was chosen as reference, and TBI and sham animals were analyzed separately using the mixed model ANOVA method that is more powerful than a one-sample t test. These identified proteins that were changing over time in the r-mTBI and the r-sham groups enabling upload of these lists to the ingenuity pathway analysis (IPA) software (Qiagen IPA, Redwood City, CA) where molecules and pathways responding over time could be identified along with differences between r-mTBI and r-sham. We confirmed the validity of this approach (using a reference sample) in other proteomic experiments where transgenic mouse models were used and significant protein lists included the transgene product. To be consistent with this type of analytical approach, we have also presented the PSAPP/C57BL6 and hTau/C57BL6 studies in a similar manner, as we also observed statistical noise in the direct comparisons of transgenic versus wild-type mice. Thus, we use the 3M of age mice as a reference, with the 9M and 15M datasets from PSAPP/C57BL6 littermates, hTau/C57BL6 animals expressed as a ratio of their 3M dataset. Each of the four groups was analyzed separately using the mixed model ANOVA method and significantly identified proteins uploaded into the IPA software to identify how age-dependent changes in molecules and pathways were influenced by the hTau or PSAPP transgene.
IPA Analysis
The three datasets of significantly modulated proteins (r-mTBI/sham, hTau/C57BL6, PSAPP/C57BL6 littermates) were uploaded to the IPA software (Ingenuity® Systems) to enable us to map proteins to established networks of protein interactions to determine the functional significance of hTau/amyloid/r-mTBI-dependent changes in protein expression from our studies. The core analysis settings for IPA analyses of uploaded significantly regulated proteins used the same procedures described in Ojo et al., 2018 to assign them onto established molecular pathways (“canonical pathways”) and biological functions in the knowledge base.
Enzyme-Linked Immunosorbent Assay
Validation of proteins was conducted by using enzyme-linked immunosorbent assay (ELISA). Details of procedures for each ELISA kit (given later) is described in the manufacturers guidelines: (a) myristoylated alanine-rich protein kinase C substrate (My BioSource; MBS2890340 – San Diego, CA); (b) SLC1A2/EAAT2/GLT-1—excitatory amino acid transporter 2 (LifeSpan BioSciences; LS-F6561 – Seattle, WA); (c) protein tyrosine phosphatase, nonreceptor type 23 (My BioSource; MBS9332331 – San Diego, CA).
Results
Time-Dependent Changes in Unique and Common Proteins in the Hippocampus of Sham and r-mTBI Mice Across Multiple Timepoints
A 10-plex TMT isobaric tag approach was used to study the protein profiles in the hippocampus of r-sham and injured mice at five separate timepoints postinjury (24 hr, 3, 6, 9, and 12 months; see Figure 2).
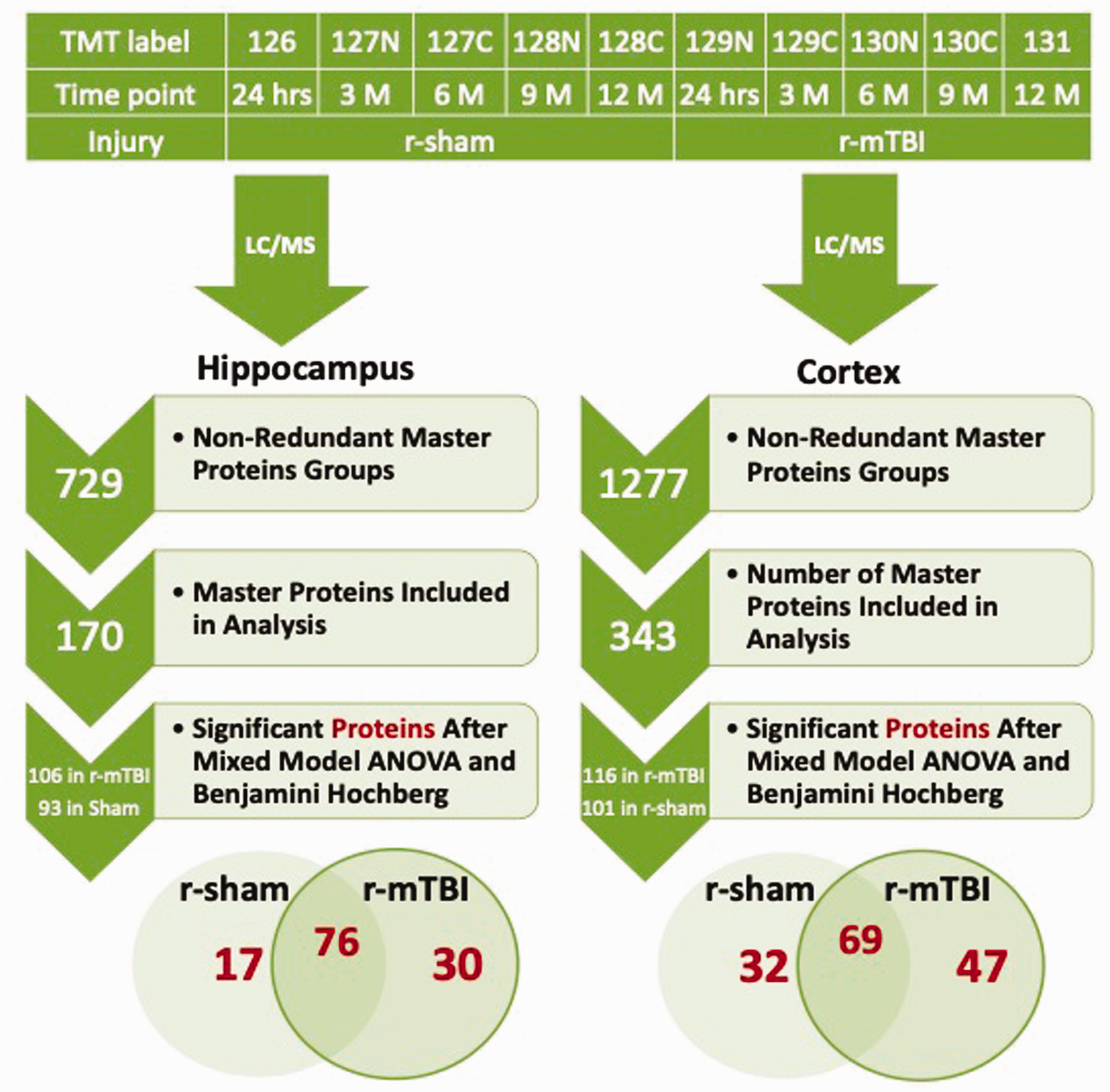
Tandem Mass Tag (TMT) Labeling, Liquid Chromatography-Mass Spectrometry, and Workflow for Proteomic Analyses From r-mTBI and r-sham Mice. Workflow shows randomization of both hippocampal and cortical samples for TMT multiplexing. Samples were pooled for multiplexing with 10-plex TMT isobaric mass tag labels. Workflow shows total number of nonredundant master protein groups, master proteins identified and analyzed, and Venn diagram depicting significantly proteins changing over the five multiple postinjury timepoints from sham/TBI mice. In the hippocampus, 17 unique proteins were significantly changing in shams, 30 unique proteins were changing in injured mice, and 76 proteins were common in sham and TBI mice. In the cortex, 32 unique proteins were significantly changing in sham mice, 47 unique proteins were changing in injured mice, and 69 were common in sham and TBI mice.
Our MS/MS-based proteomics analyses generated protein identities based on identified peptides, and to avoid redundancy, proteins within a group were ranked based on the number of peptide sequences, the number of peptide spectrum matches, their protein scores, and the sequence homology/coverage, with the top-ranking protein being identified as a master protein based on this criteria. A total of 729 nonredundant master protein were identified in the hippocampus in this study, 170 of these were identified within all plexes and groups examined.
We normalized samples from each timepoint to their 24 hr post-TBI or sham group, and analyses using mixed model ANOVA and after Benjamini-Hochberg correction identified 17 unique proteins significantly changing over time in sham injured mice, 30 unique proteins significantly changing in repetitive injured mice, and 76 common proteins significantly changing in both sham and injury groups over time (Figure 2; see also Supplementary Figure S1 for list of significant proteins in sham group and Supplementary Figure S2 for list of significant proteins in the injury group). A list of the unique significantly modulated proteins observed across multiple timepoints in the injury group is shown in Table 1.
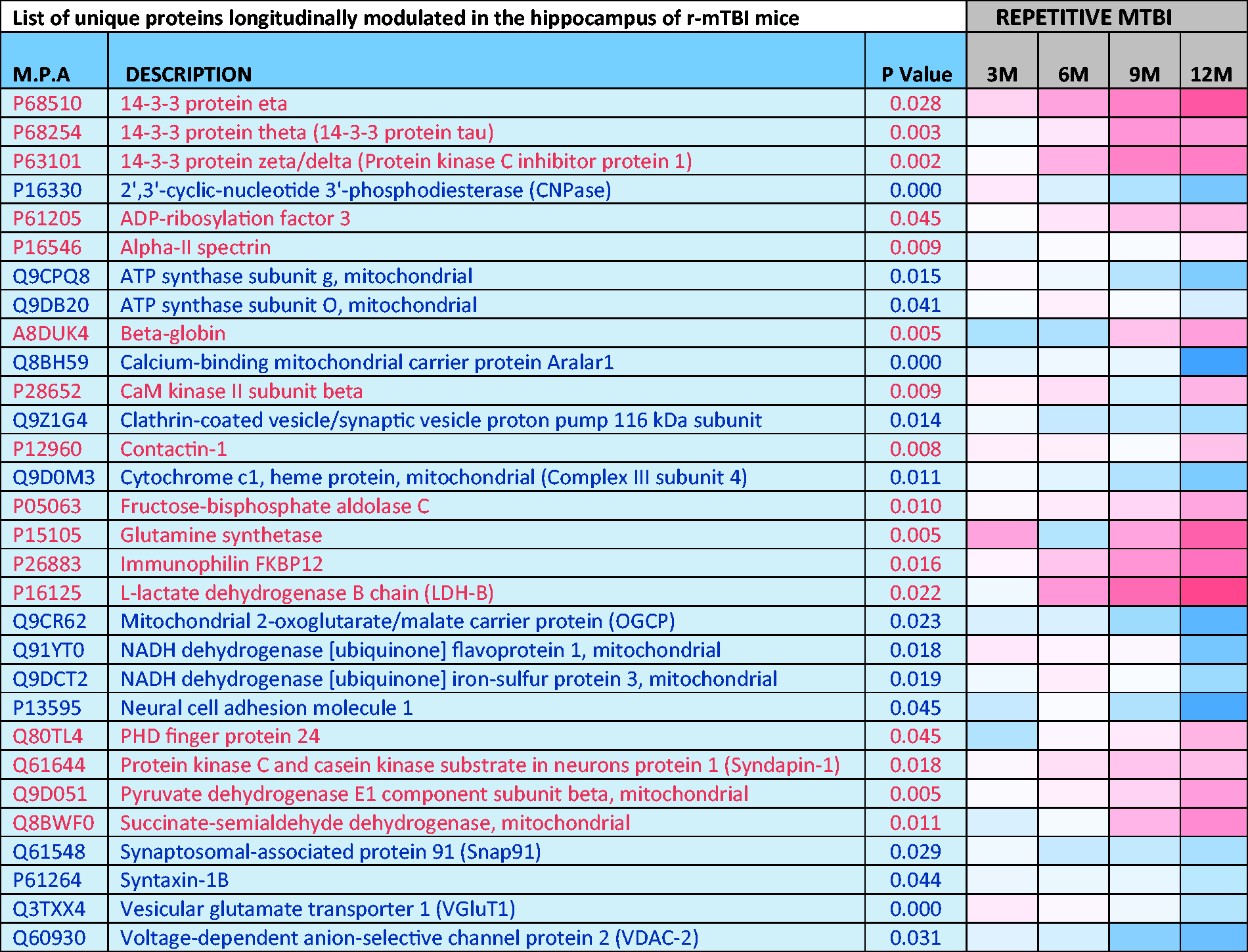
List of Unique Significantly Modulated (30) Proteins From the Hippocampus of Repetitive mTBI Mice, Compared With Their Sham Counterparts Across Multiple Timepoints.
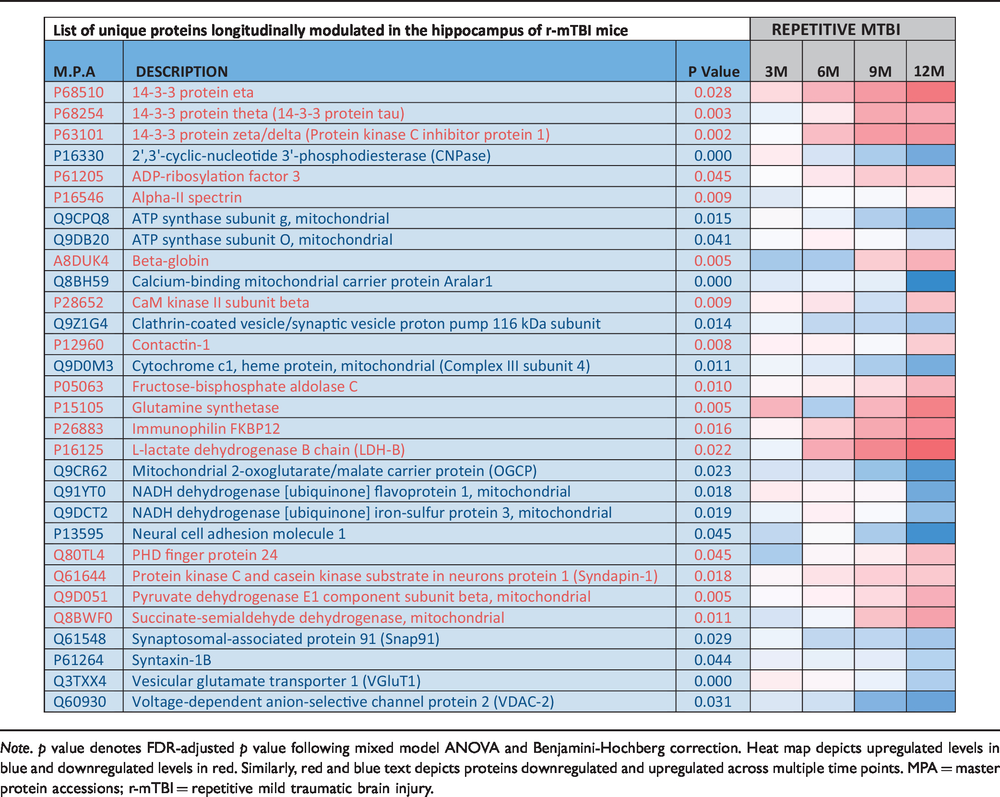
Note. p value denotes FDR-adjusted p value following mixed model ANOVA and Benjamini-Hochberg correction. Heat map depicts upregulated levels in blue and downregulated levels in red. Similarly, red and blue text depicts proteins downregulated and upregulated across multiple time points. MPA = master protein accessions; r-mTBI = repetitive mild traumatic brain injury.
Biofunctions, Canonical Pathways, and Pathobiological Processes Modulated in the Hippocampus Samples From r-mTBI and r-Sham Mice Across Multiple Timepoints
IPA generated a list of 16 biofunctions and pathobiological processes modulated in the hippocampus of r-mTBI versus sham injury groups across multiple timepoints, and this is shown in Figure 3. Some of the diseases/biofunctions include exocytosis, engulfment of cells, neurodegeneration, synaptic depression, and formation of vesicles. IPA identified the top three significantly modulated canonical pathways generated from our list of significantly regulated proteins in the hippocampus of injured versus sham groups. These pathways include phosphatidylinositol-3-kinase (PI3K)/AKT signaling, protein kinase A signaling, and Peroxisome proliferator-activated receptor alpha and Retinoid X receptor alpha (PPARα/RXRα) activation (see Figure 4).
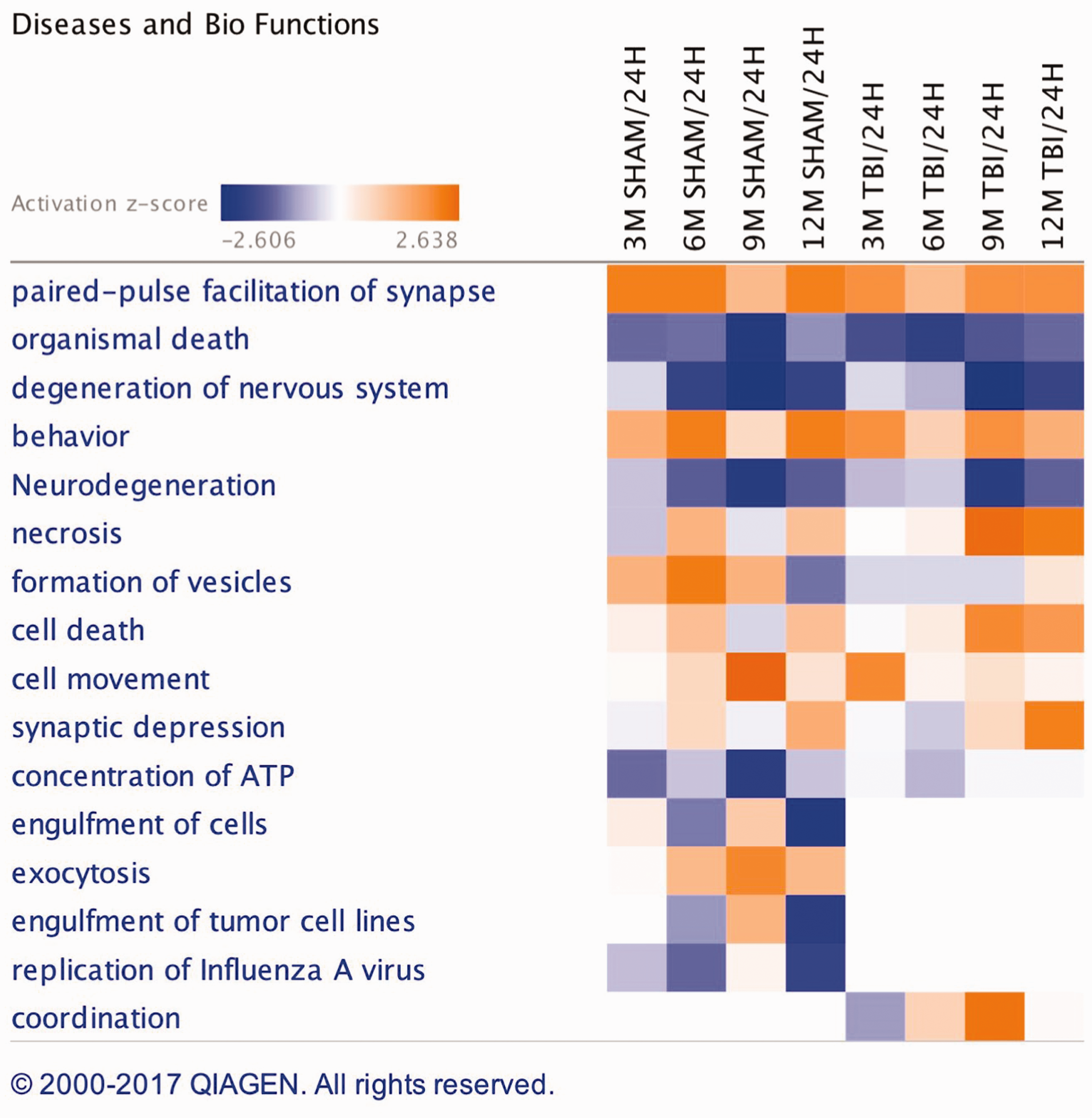
Biofunctions and Disease Altered in the Hippocampus of Sham/Injury Mice. Biofunctions and diseases identified from the list of generated significant proteins in sham/injury groups over multiple timepoints using ingenuity pathway analyses (IPA).
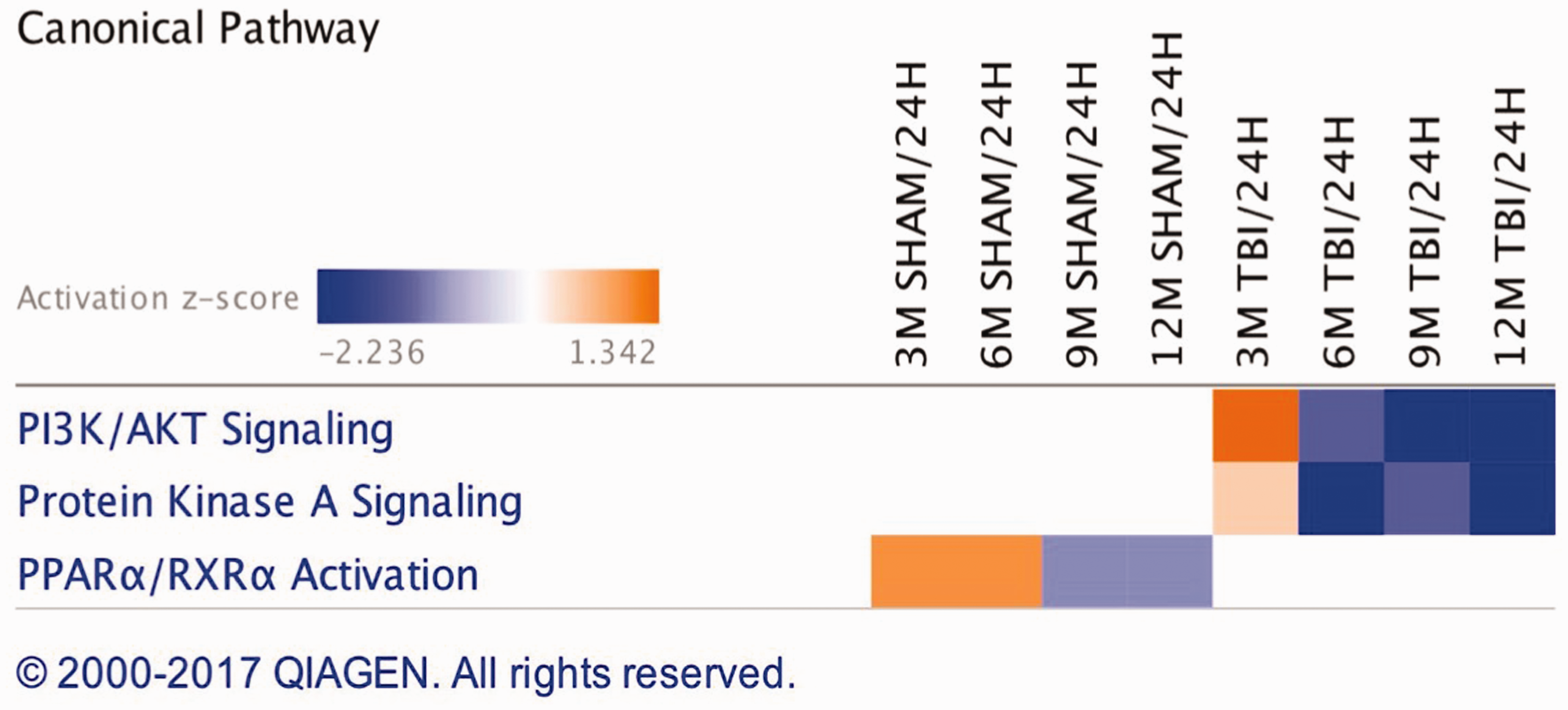
Canonical Pathways Altered in the Hippocampus of Sham/Injury Mice. Canonical pathways generated from the list of generated significant proteins in sham/injury groups over multiple timepoints using IPA. Top three pathways include PI3K/AKT signaling, protein kinase A (PKA) signaling, and PPARα/RXRα activation.
Time-Dependent Changes in Unique and Common Proteins in the Cortices of r-mTBI and r-Sham Mice Across Multiple Timepoints
A 10-plex TMT isobaric tag approach was used to study the profiles in the cortices of r-TBI and sham mice at five timepoints postinjury (24 hr, 3, 6, 9, and 12 months; see Figure 2). In this study, a total of 1,277 nonredundant master protein groups were identified in the cortex, and 343 of these master protein groups were identified within all plexes and groups. As mentioned earlier, a mixed model ANOVA approach was used to analyze the master proteins present in all plexes to identify prospective changes in unique and common proteins between sham and TBI groups over multiple timepoints.
Samples were once again normalized with their 24 hr postinjury/sham group, and analyses of age-related changes within injured and sham groups using mixed model ANOVA after Benjamini-Hochberg correction identified 32 unique proteins changing in the sham group, 47 unique proteins changing in the repetitive injured group, and 69 common proteins changing in both sham and injury groups over time (Figure 2; see also Supplementary Figure S3 and S4 for list of proteins). A list of the significantly regulated proteins uniquely changing in the injured group is shown in Table 2.
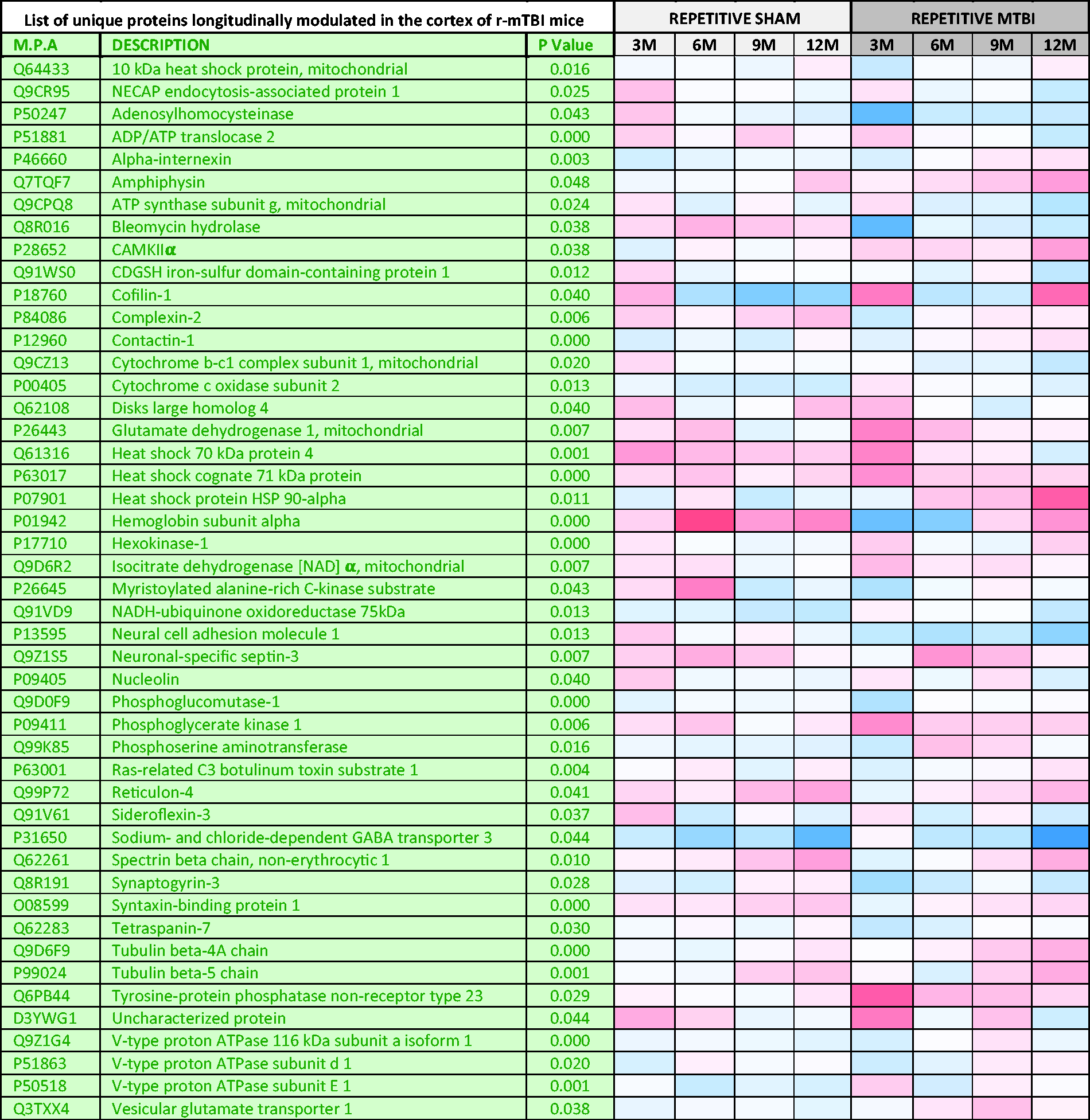
List of Unique Significantly Modulated (47) Proteins From the Cortex of Repetitive mTBI Mice, Compared With Their Sham Counterparts Across Multiple Timepoints.
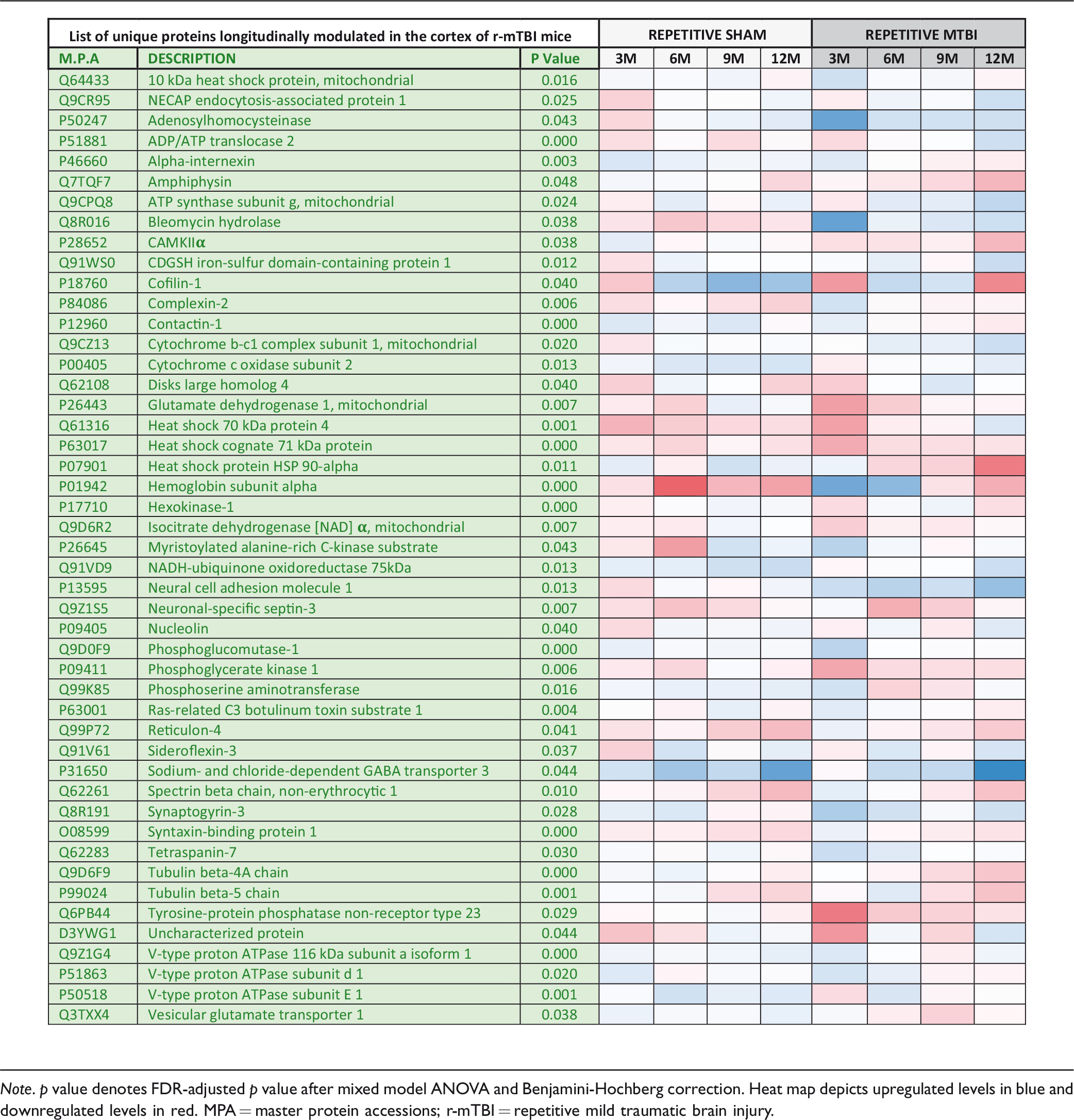
Note. p value denotes FDR-adjusted p value after mixed model ANOVA and Benjamini-Hochberg correction. Heat map depicts upregulated levels in blue and downregulated levels in red. MPA = master protein accessions; r-mTBI = repetitive mild traumatic brain injury.
To validate our proteomics LC-MS/MS results, we used an antibody-based approach to examine the levels of three proteins in cortical tissue (myristoylated alanine-rich protein kinase C substrate; protein tyrosine phosphatase, nonreceptor type 23, and excitatory amino acid transporter 2). These proteins were chosen on the basis that they were highly abundant in our samples, significantly regulated in sham and/or TBI, and also easily detectable by a reputable and well-established antibody-based (ELISA) method. These three proteins showed trends consistent with our TMT data at four separate timepoints investigated in the groups specifically analyzed (Supplementary Figure 5). We also measured levels of heat shock protein 90 (HSP 90) by ELISA method (data not shown). Our results showed statistically similar trends between TMT and ELISA methods; however, the direction of change was observed in opposite directions, with a reduction observed by TMT analyses and an increase by ELISA over time.
Biofunctions, Canonical Pathways, and Pathobiological Processes Modulated in the Cortical Samples From r-mTBI and r-Sham Mice Across Multiple Timepoints
A list of 37 biofunctions and pathobiological processes modulated in the cortices of r-mTBI versus sham groups across multiple timepoints is shown in Figure 5. Some of these include endocytosis, myelination, neurodegeneration of axons, engulfment of cells, migration of neuroglia, clathrin-mediated endocytosis, axonogenesis, and receptor-mediated endocytosis. Ingenuity pathway analyses (IPA) identified the top three canonical pathways significantly regulated within the cortex of TBI versus sham groups after analyses of significantly modulated proteins. These pathways include protein kinase A signaling, protein gonadotropin-releasing hormone (GnRH) signaling, and B cell receptor activation (see Figure 6).
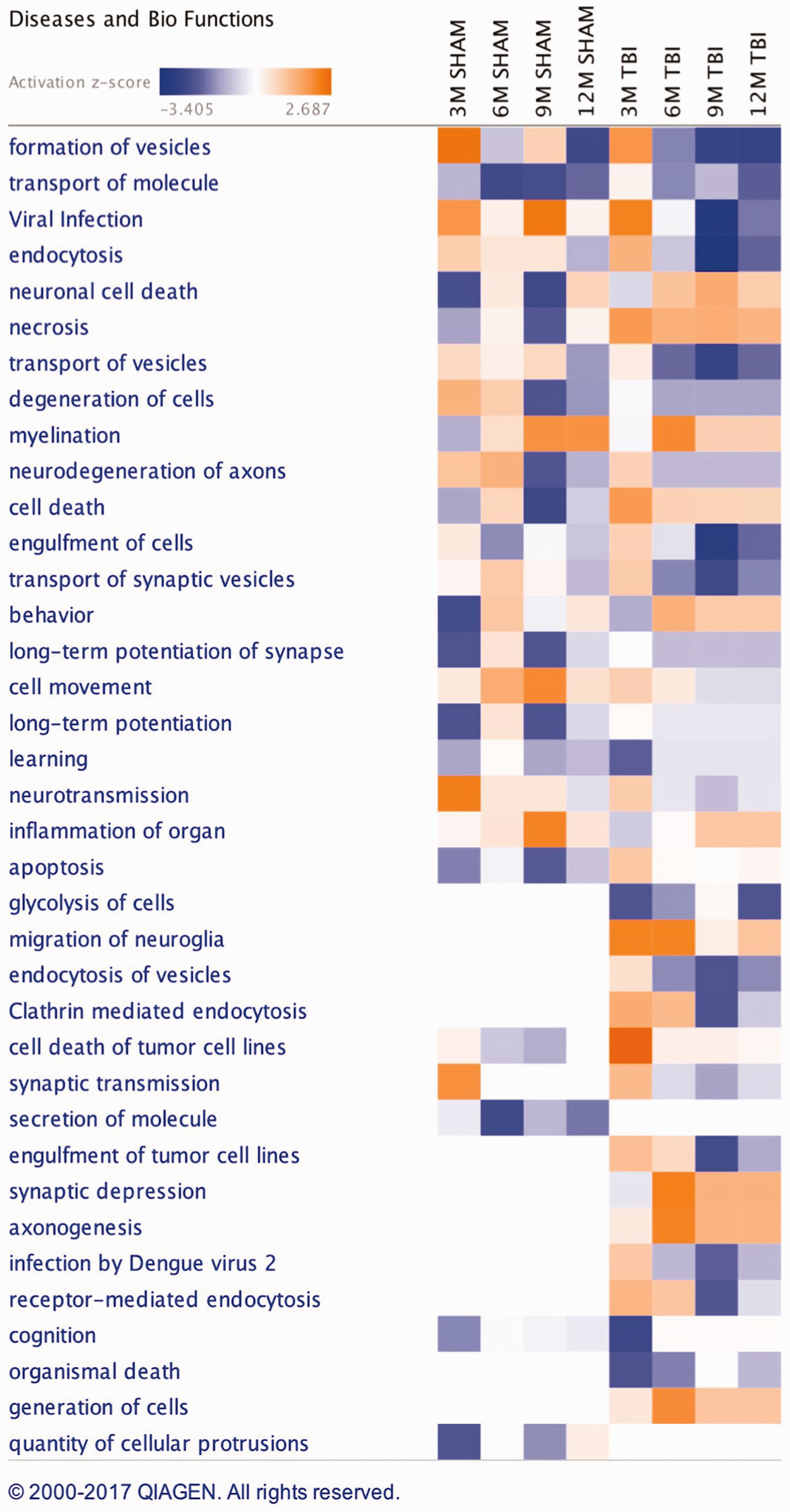
Biofunctions and Disease Altered in the Cortex of Sham/Injury Mice. Biofunctions and diseases identified from the list of generated significant proteins in sham/injury groups over multiple timepoints using ingenuity pathway analyses (IPA).
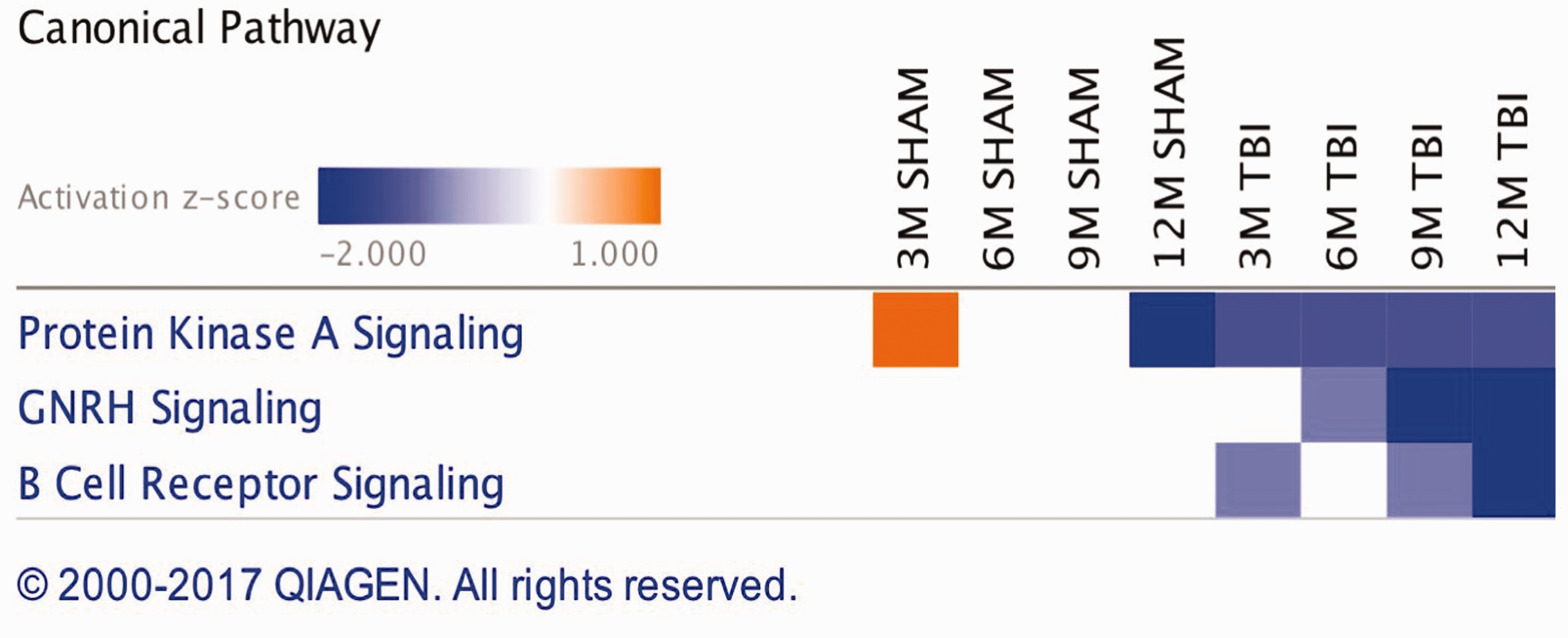
Canonical Pathways Altered in the Cortex of Sham/Injury Mice. Canonical pathways generated from the list of generated significant proteins in sham/injury groups over multiple timepoints using IPA. Top three pathways include protein kinase A (PKA) signaling, GnRH signaling, and B cell receptor signaling.
Time-Dependent Changes in Unique and Common Proteins in the Cortex of hTau and PSAPP Mice at Different Ages
We used a 6-plex TMT isobaric tag multiplex approach to study the profiles between hTau mice versus C57BL6 wild-type mice and PSAPP versus C57BL6 wild-type littermate mice at 3 different ages (3, 9, and 15 months; see Figure 7).
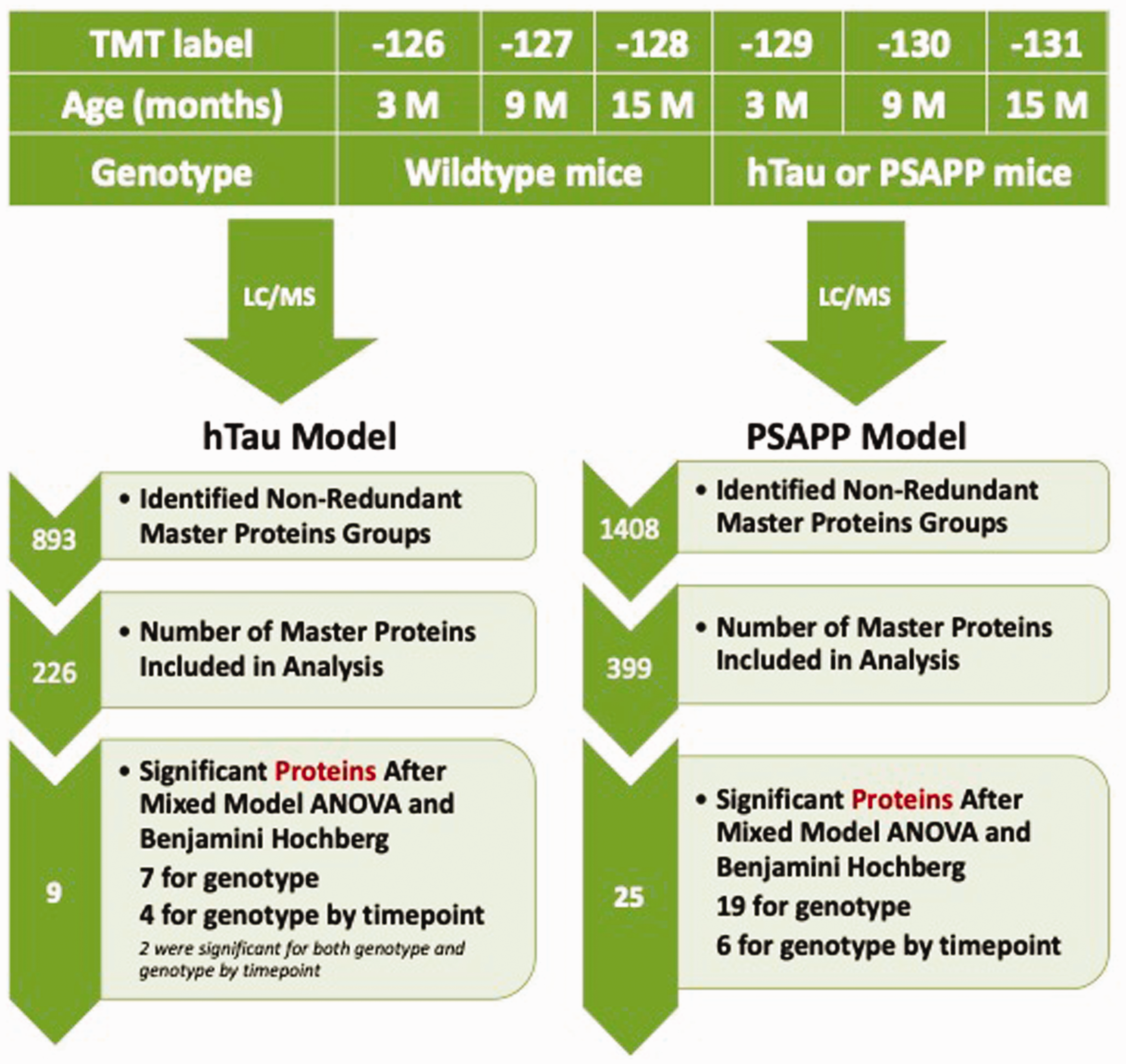
Tandem Mass Tag (TMT) Labeling, Liquid Chromatography-Mass Spectrometry, and Workflow for Proteomic Analyses in hTau and PSAPP Mice. Workflow shows randomization of cortical samples for TMT multiplexing. Samples were pooled for multiplexing with 6-plex TMT isobaric mass tag labels. Workflow shows total number of nonredundant master protein groups, master proteins identified and analyzed, and Venn diagram depicting significantly proteins changing over the 3 age groups in hTau and PSAPP mice. In PSAPP mice, 19 proteins were significantly changing with genotype (PSAPP vs. wild type), and 6 proteins showed significant interaction with aging and genotype. In the hTau group, seven proteins were significantly regulated with genotype (hTau vs. wild type), and four proteins showed significant interaction with aging and genotype.
A total of 1,408 and 893 nonredundant master protein groups were identified from the PSAPP and hTau study, respectively; 399 identified master protein groups were present in all plexes and groups of the PSAPP study and 226 in the hTau study (Figure 7).
Dataset were analyzed separately using the mixed model ANOVA after Benjamini-Hochberg correction to identify proteins changing with age in the two transgenic and respective wild-type groups.
In the PSAPP group, 19 proteins were significantly changing with genotype alone, and 6 proteins were identified that showed a significant interaction between genotype and aging (Figure 7). In the hTau group, seven proteins were significantly changing with genotype alone, and four proteins were identified that showed significant interaction between genotype and aging (Figure 7). A list of genotype-specific age-related significantly regulated proteins in the cortices of PSAPP versus wild-type groups and hTau versus wild-type groups is shown in Tables 3 and 4, respectively. Unfortunately, analyses of the hippocampal samples from these mice were not possible because they were processed differently from TBI samples and also used for lipidomic analyses.
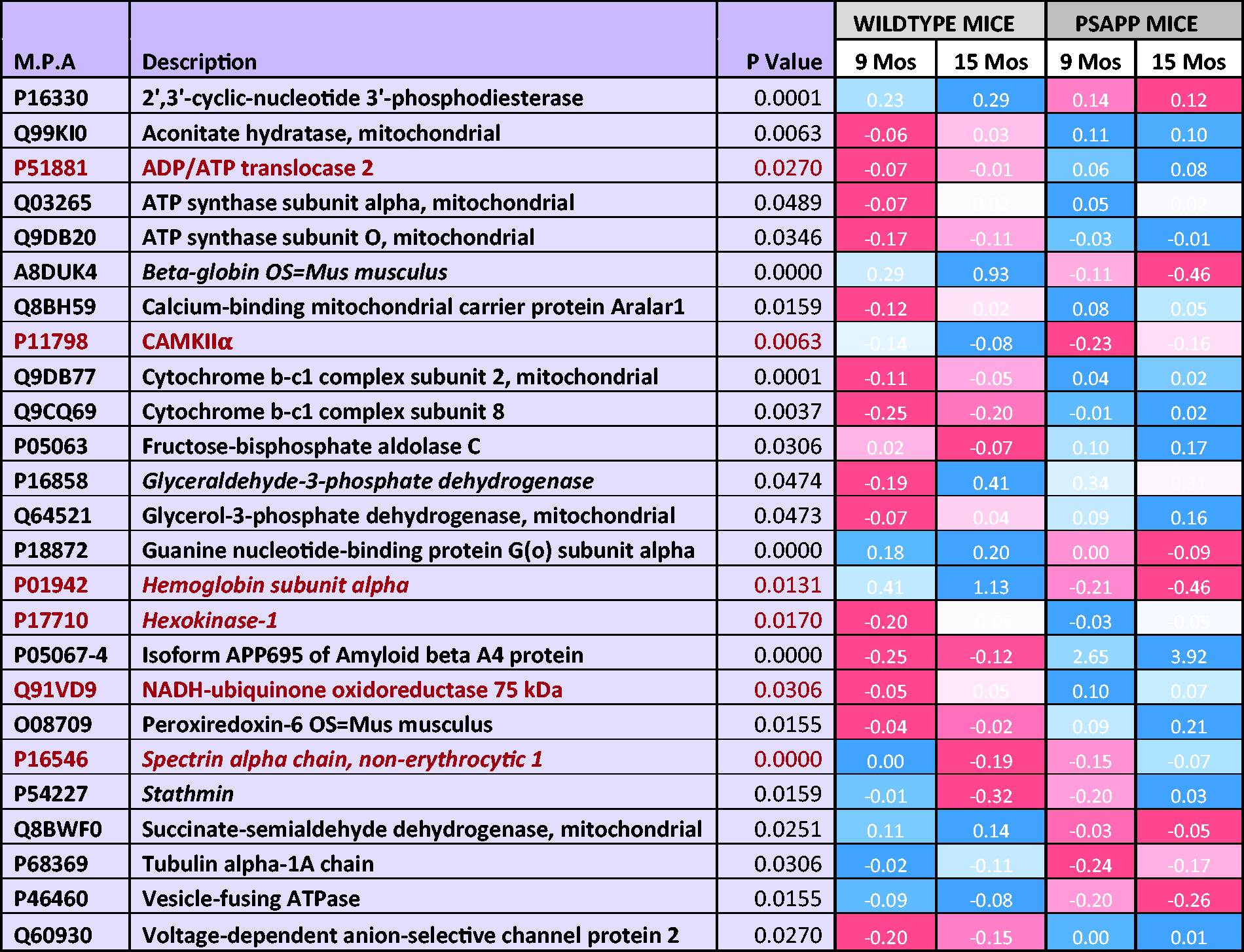
List of Significantly Modulated Proteins From the Cortex of PSAPP Mice Compared With Wild-Type Controls.
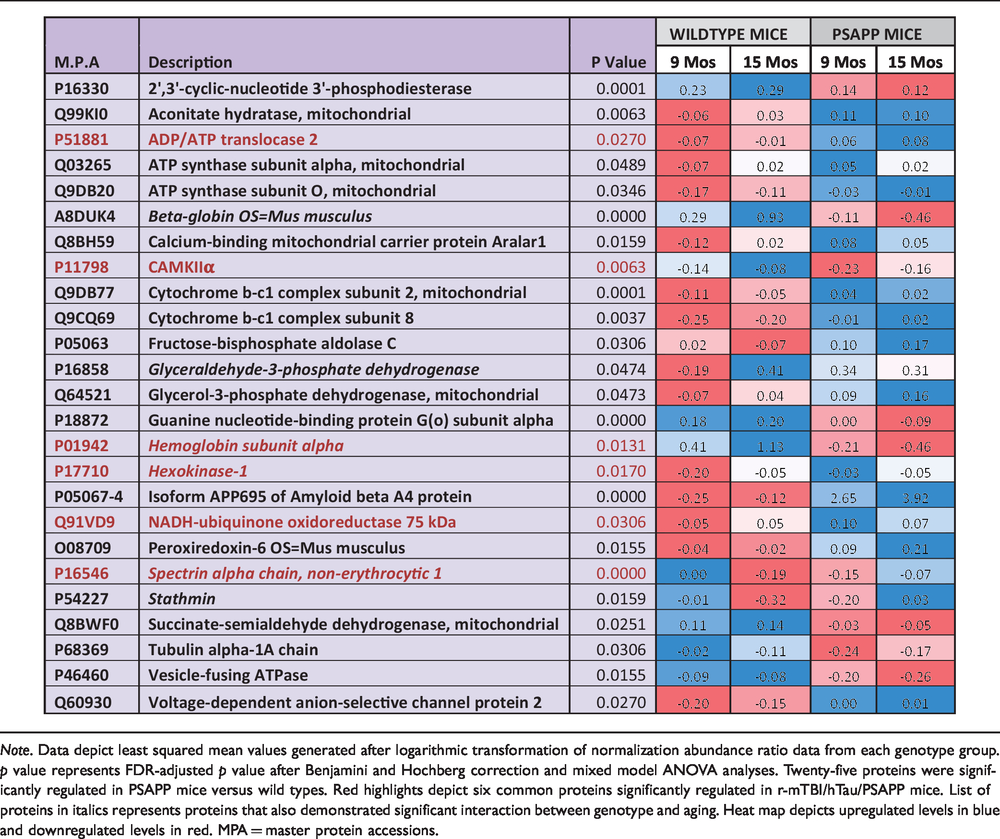
Note. Data depict least squared mean values generated after logarithmic transformation of normalization abundance ratio data from each genotype group. p value represents FDR-adjusted p value after Benjamini and Hochberg correction and mixed model ANOVA analyses. Twenty-five proteins were significantly regulated in PSAPP mice versus wild types. Red highlights depict six common proteins significantly regulated in r-mTBI/hTau/PSAPP mice. List of proteins in italics represents proteins that also demonstrated significant interaction between genotype and aging. Heat map depicts upregulated levels in blue and downregulated levels in red. MPA = master protein accessions.
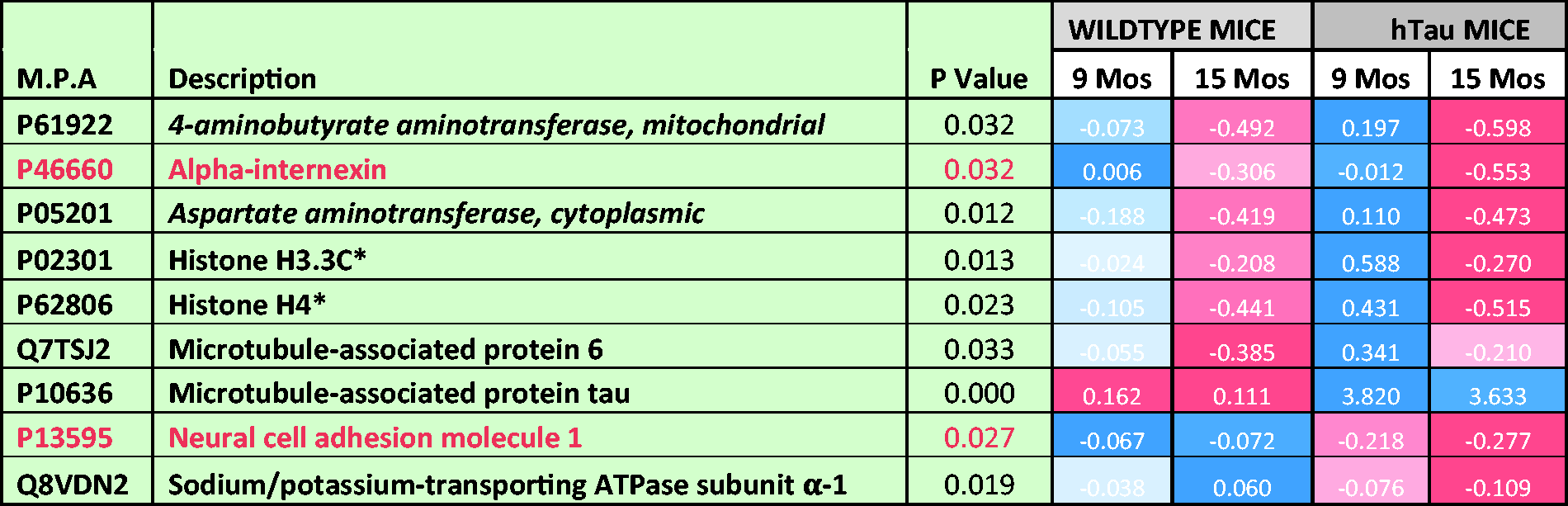
List of Significantly Modulated Proteins From the Cortex of hTau Mice Compared With Wild-Type Littermate Controls.
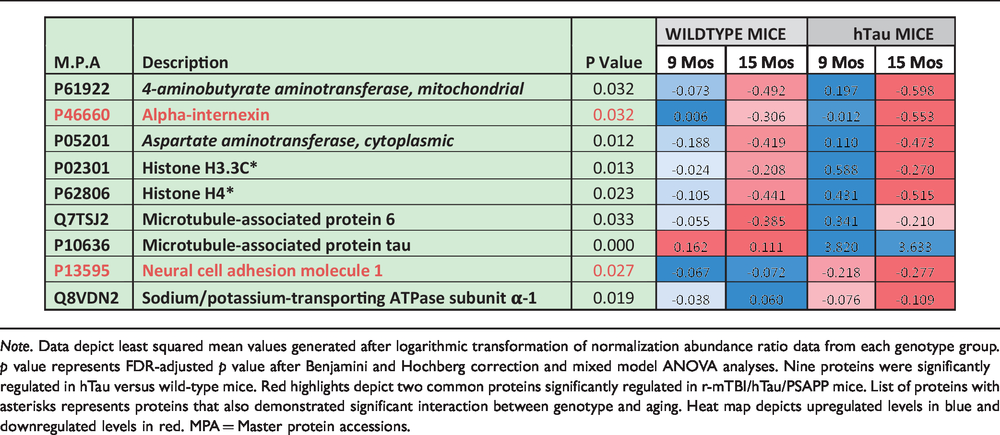
Note. Data depict least squared mean values generated after logarithmic transformation of normalization abundance ratio data from each genotype group. p value represents FDR-adjusted p value after Benjamini and Hochberg correction and mixed model ANOVA analyses. Nine proteins were significantly regulated in hTau versus wild-type mice. Red highlights depict two common proteins significantly regulated in r-mTBI/hTau/PSAPP mice. List of proteins with asterisks represents proteins that also demonstrated significant interaction between genotype and aging. Heat map depicts upregulated levels in blue and downregulated levels in red. MPA = Master protein accessions.
Commonalities Between r-mTBI Model and hTau/PSAPP Models
To investigate any potential molecular pathobiological relationship underlying the sequelae of events in TBI and models of amyloidosis and tauopathy, we compared using mixed model ANOVA the significant cortical proteomic profiles that were uniquely r-mTBI dependent or either PSAPP or hTau dependent (Figure 8). Only six proteins were common to the r-mTBI mouse model and the PSAPP mouse model (ADP/ATP translocase 2, CAMKII, hemoglobin subunit a, hexokinase-1, NADH-ubiquinone oxidoreductase, and spectrin alpha chain, nonerythrocytic 1; see Table 3 and Figure 8), while only two were common to the r-mTBI model and the hTau mouse model (alpha-internexin and neural cell adhesion molecule 1; see Table 4 and Figure 8). A summary of the proteomic and altered biofunctions observed in all three models is detailed in Figure 9.
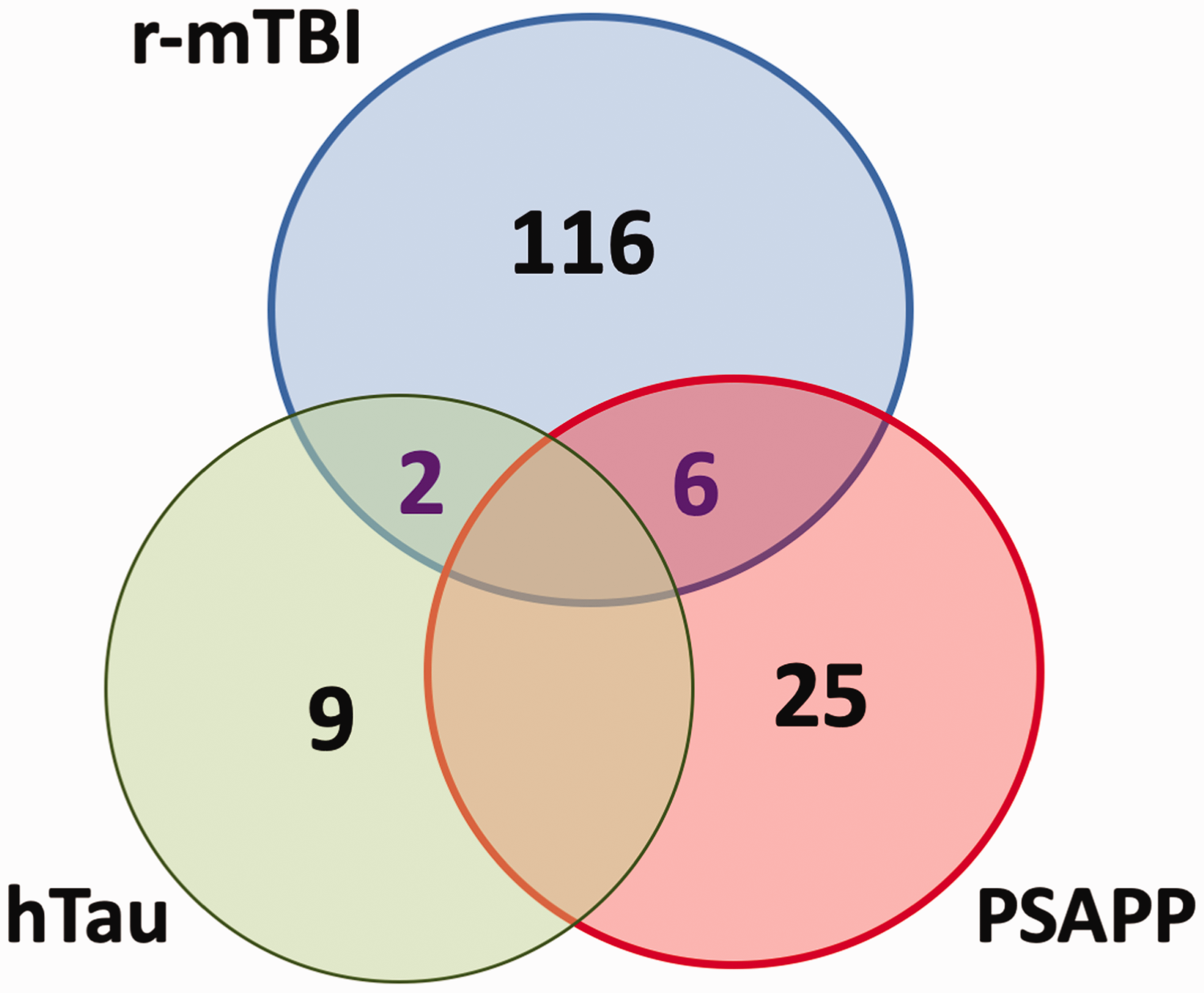
Venn Diagram Depicting Overlap Between Proteins Changing Over Time in r-mTBI (C57BL/6J Mice), Tau (hTau Mice), and Amyloid (PSAPP Mice) Models. Totally, 116 proteins were significantly changing in the injury group, 25 proteins in the amyloid model, and 9 in the tau model over time. Only six proteins overlapped between injury and amyloid models, while two overlapped between injury and tau models.
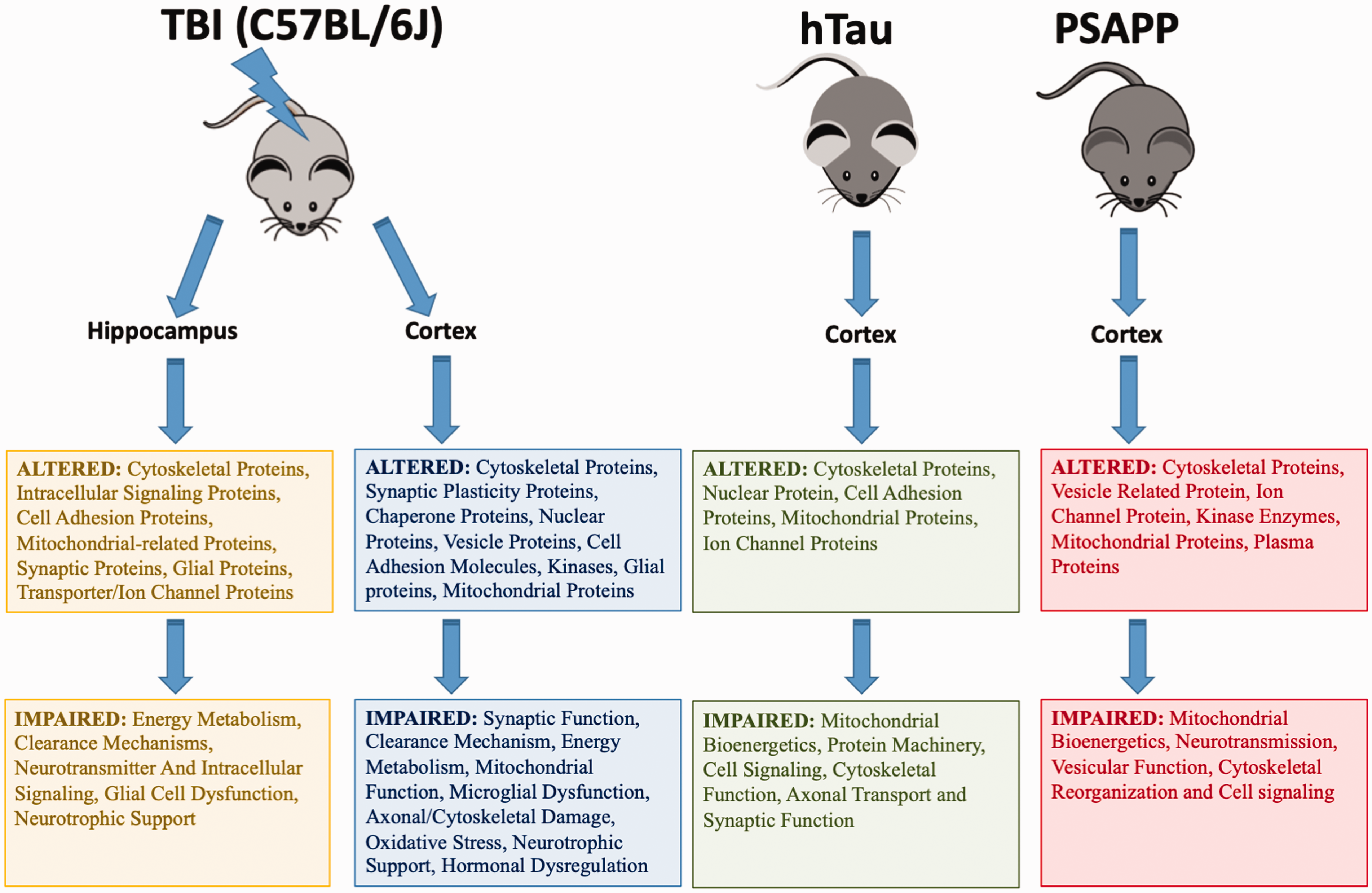
Graphic Summary of the Class of Proteins and Impaired Functions Observed in the r-mTBI, Tau, and Amyloid Models.
Discussion
Our previous neuropathological assessment of our 5-hit r-mTBI model revealed a lack of tau or amyloid pathology up to 24 months postinjury. Thus, in this work, we aimed to use our proteomic platform to further explore the underlying molecular responses to r-mTBI in our mouse model (Mouzon et al., 2012; Ojo et al., 2013; Mouzon et al., 2014; Ojo et al., 2015), with two unique preclinical models of amyloidosis (PSAPP) and tauopathy (hTau). We employed an unbiased LC-MS/MS-based approach and have generated a detailed time line of molecular responses in the cortex and hippocampus from 24 hr to 12 months postinjury in mouse models of r-mTBI and in transgenic mouse models of amyloidosis and tauopathy encompassing pre-, peri-, and post-“onset” of pathological phenotypes (3 to 15 months of age). Our data reveal largely distinct r-mTBI-dependent and tau/amyloid-driven and dependent molecular-level changes which are of interest to the chronic consequences of r-mTBI in humans.
Proteomic Changes in the Hippocampus and Cortex Following r-mTBI
Our prior work with this wild-type 5-hit r-mTBI model have been previously published and demonstrates deficiency in hippocampal-dependent spatial learning and memory processing (Barnes Maze) at 24 hr postinjury, which persists until 6 months, and progressively worsens at 12, 18, and 24 months postinjury timepoints (Mouzon et al., 2012, 2014, 2018c). We have also extensively characterized the histopathological features of this model at 24 hr, 3, 6, 12, and 24 months postinjury; injured animals consistently demonstrate persistent white matter damage involving thinning of the corpus callosum, gliosis, and axonal injury (Mouzon et al., 2012, 2014; Ojo et al., 2015; Ferguson et al., 2017; Mouzon et al., 2018a, 2018b, 2018c). However, gross neuropathological assessment of the gray matter (hippocampus and also cortex) revealed no overt signs of proteinaceous (tau, amyloid, etc.) lesions resulting from the chronic effects of injury. Injury-dependent astroglial and microglial activation, transient increase in phospho-tau levels, ultrastructural abnormalities to dendrites, and electron dense dark neurons, although observed at earlier timepoints (either 24 hr or 3 months) postinjury, were not observed beyond >6 to 24 months after injury in the gray matter (Mouzon et al., 2012, 2014; Ojo et al., 2015; Mouzon et al., 2018c). This indicates that the anatomical correlates responsible for the early neurobehavioral changes in our model may be attributed to subtleties in molecular responses that cannot be otherwise detected by traditional gross histopathological analyses.
In this study, we focused our omic analyses within each specific group (i.e., TBI or sham animals) by creating a dataset of proteomic responses to injury or normal aging over multiple extended timepoints postexposure. This enabled us to identify common and unique proteins significantly modulated in shams and/or injured groups. Within the hippocampus, we identified 30 proteins unique to the TBI group only, and these were changing over multiple timepoints postinjury. These class of proteins included cytoskeletal proteins, downstream signaling proteins, cell adhesion proteins, mitochondrial-related proteins, synaptic proteins, glial-associated proteins, and transporter/ion channel proteins. In addition, injured mice also showed a lack of age-related change in 17 proteins that were unique to and significantly altered in sham mice, indicating a suppression or dysregulation in normal age-related processes which may drive pathological aging. These classes of proteins included calcium-associated and cytoskeletal proteins, SNARE proteins, kinase enzymes, proteins involved in bioenergetics, and ubiquitin proteasome-related proteins. The top scoring diseases and biofunctions (i.e., those most significantly altered between sham and injured mice) included exocytosis, paired-pulse facilitation of synapse, engulfment of cells, concentration of ATP, neurodegeneration, synaptic depression, and formation of vesicles. The top canonical pathways included PI3K/AKT signaling, protein kinase A signaling, and PPARα/RXRα activation. These findings implicate pathophysiological responses involving impaired energy metabolism, clearance mechanisms, neurotransmitter and intracellular signaling, glial cell dysfunction, and diminished neurotrophic support after injury within the hippocampus. Our interrogation of the cortex revealed 47 proteins that were changing in the r-mTBI group compared with the sham controls over time. They can be categorized mainly as filament and cytoskeletal proteins, synaptic plasticity proteins, chaperone proteins, nuclear proteins, synaptic vesicle proteins, cell adhesion molecules, kinase enzymes, and mitochondrial proteins involved in energy metabolism. We also observed that 32 significant proteins uniquely changing with age in sham mice did not show any significant changes in injured mice with aging. These classes of proteins include microglia/macrophage inhibitory protein factors, plasma membrane and cytoskeletal proteins, antioxidant proteins, synaptic proteins, growth factors, and proteins involved in energy metabolism. We employed an antibody-based approach to validate our findings for three different proteins that were highly abundant in the tissue and detectable by ELISA method, which confirmed similar trends in some of the groups assessed. It is worthy of note that there was a discrepancy in our validation attempt for HSP 90 by ELISA method, which showed changes in opposite directions by TMT versus ELISA, although changes were both significant over the timepoints of analyses by ANOVA. The reason behind these differences is unknown. We attribute these to issues surrounding the lack of sensitivity of the ELISA detection method, age of samples at validation with ELISA and stability of HSP 90 proteins, or possibility of sample degradation due to multiple freeze thaw cycles (up to 2 cycles). It is also worthy of note that we confirmed, in our TMT experiments, the increase in levels of our transgene products (e.g., hTau and hAPP) from the cortices/hippocampi of our models of tauopathy and amyloidosis at different ages and also noted TBI-related markers (e.g., glial fibrillary acidic protein [GFAP]) previously shown in our histopathological studies (Mouzon et al., 2012, 2014, 2018c) was also confirmed in our TMT experiments.
IPA generated a list of diseases/biofunctions significantly altered between injured and sham mice, some of which included endocytosis, myelination, engulfment of cells, neurodegeneration of axons, migration of neuroglia, clathrin-mediated endocytosis, axonogenesis, and receptor-mediated endocytosis. The top three canonical pathways include protein kinase A signaling, protein GnRH signaling, and B cell receptor activation. Our proteomic profiles of the cortical tissue implicate a complex and wide array of pathophysiological responses after injury, possibly involving alterations in synaptic function, clearance mechanism, energy metabolism and mitochondrial function, microglial/immune dysfunction, axonal/cytoskeletal damage, oxidative stress, and dysfunction in neurotrophic, hormonal, and downstream intracellular signaling processes. Because we have used whole cortical regions for these proteomic experiments, it is possible that our TBI-dependent profiles in injured mice may be diluted, due to our injury being administered at the midline, beneath the parietal cortex. In parallel plasma biomarker studies from the same mice used in this model, we identified 31 proteins significantly changing in the r-mTBI animals over time, and, when compared with changes observed over time in sham animals, 13 proteins of these were unique to the TBI mice (Ojo et al., 2018). The top pathways significantly modulated by these unique TBI-dependent changes were liver X receptor and retinoid X receptor (LXR/RXR) activation, production of nitric oxide (NO) and reactive oxygen species (ROS) and complement systems. Due to the chronic timepoints selected for these studies and the mild nature of our injuries, there was no direct overlap in proteins in the blood and brain, as utility of brain-derived proteins in the blood is more prominent at acute timepoints. Nonetheless, due to the alterations in RXR activation in the hippocampus of our TBI mice and blood samples, including immune-related changes typified by activation of complement systems in the blood and alterations in microglia/macrophage inhibitory factors and B cell receptor activation in the cortex, we can surmise that there is a consistent overlap in these pathogenic processes in the blood and brain that may be interrogated as a putative biomarker of r-mTBI-mediated neurodegeneration. However, further validations studies will be needed to confirm this.
There have been few previous studies employing proteomic approaches in preclinical models of mTBI, particularly those that have explored chronic timepoints postinjury. A recent study using isobaric tags for relative and absolute quantitation (iTRAQ)-based proteomic approach in r-mTBI (3 hits) rats using a metallic pendulum-striker at 1, 7, and 180 days post-TBI demonstrated 237 proteins significantly changing between control and r-mTBI mice, with a prominent role of upregulation in cAMP signaling pathway (Song et al., 2018). Other biological activities, such as myelination of axons, cell adhesion signaling, autophagic clearance, microtubule depolymerization, and neurodevelopment of the brain, were also observed. Another study used a microwave and magnetic (M2) proteomics approach in 2-month-old mice exposed to a single closed skull impact mouse model using a closed cortical injury device (2 mm depth and 4.5 m/s) that closely resembles our experimental model used herein (Evans et al., 2014). The authors collected brain tissue at 1, 7, 30 and 120 days postinjury and reported significant changes in myelin cytoskeletal and neurofilament proteins, antioxidant enzymes, proteins involved in bioenergetics, and neurotrophic factors in the ipsilateral brains of injured mice up to 30 days postinjury, and this correlated with motor abnormalities. It is noteworthy that this model demonstrated evidence of neuron loss and edema, indicating that it is a more severe model than our mild(TBI). Nonetheless, proteomic profiles in injured mice from this study and ours appeared to show a large degree of overlap. In another proteomics study, using rats exposed to closed cortical injury, the authors revealed mitochondrial fraction proteins oxidatively modified in the cortex and hippocampus after injury (Opii et al., 2007). Although they also used a more severe model compared with ours, they likewise confirmed early significant changes in several proteins involved in energy metabolism in both brain regions analyzed after 3 hr postinjury. Even though no chronic timepoints were examined in this study, their work nevertheless supports the early involvement of dysfunctional mitochondrial bioenergetics in TBI which is known to precipitate a host of different responses such as oxidative stress, calcium dis-homeostasis, excitotoxicity, and ATP depletion (Opii et al., 2007). Another model that explored effects of fluid percussion (moderate) injury in rat cortical tissue 48 hr after injury using 2D gel electrophoresis and MS/MS analysis also reported significant changes in mitochondrial proteins associated with energy metabolism, cytoskeletal proteins, hormonal signaling receptors, and synaptic proteins (Ding et al., 2015).
Quantitative TMT proteomic studies have also been conducted in single low-intensity blast (mTBI) injury studies at 2 and 24 hr, and 7 and 30 days postinjury (Song et al., 2019). Animals revealed mitochondrial and axonal ultrastructural changes. Remarkable dynamic changes in 2,216 proteins and 459 phosphorylated proteins at variety of timepoints postinjury were observed. The key canonical pathways disrupted included mitochondrial dysfunction, oxidative stress, and axonal/cytoskeletal/synaptic dysregulation. Particularly with respect to proteomic changes in mitochondria dysfunction, altered fission/fusion dynamics, reduced mitophagy, diminished oxidative phosphorylation, and compensated respiration-relevant enzymatic activities were thought to underlie these dysfunctions, and these changes were very prominent at 7 and 30 days postinjury. In another LC-MS/MS blast-induced mTBI study examined acutely after injury, using isobaric mass tags for relative protein quantification, alterations were observed in axonal and synaptic proteins associated with cortical cytoskeleton organization, and synaptic vesicle exocytosis (Chen et al., 2018). Microtubule-associated protein 1B, neurofilaments, stathmin, synaptotagmin I, myelin basic protein, calcium-calmodulin-dependent protein kinase, and actin binding proteins were the most significantly regulated proteins notably observed in blast versus sham mice. Although these findings were conducted in different mTBI models at more acute timepoints compared with ours, they all appear to demonstrate some similar threads of pathobiological changes that may occur in continuum over time, albeit to a different degree in different models and TBI paradigms.
The translational significance of these preclinical findings cannot yet be confirmed, as there have been no proteomic approaches in postmortem brain tissue from human mTBI. Only severe injuries such as those investigating pericontusional (penumbra) regions following TBI have been conducted, and the authors reported evidence of altered cytoskeletal architecture, oxidative stress/damage, depletion of antioxidant-glutathione, dysfunction of mitochondria, and synaptic loss (Harish et al., 2015). Although this human study was conducted at an acute timepoint following a more severe injury paradigm, the overlap with some of the molecular responses identified in our model suggests that these processes may be common responses even across different types of injury and may thus offer novel insights into TBI pathobiology in general.
Cortical Proteomic Profiles in Models of Tau and Amyloid Pathology
At the outset of this study, we hypothesized that comparisons of the detailed time lines of proteomic responses following r-mTBI and in models of amyloidosis and tauopathy would identify underlying pathobiological mechanisms common to both pathobiological events. However, during the length of this study, we observed from our neuropathological studies a lack of amyloid and tau pathology in our 5-hit r-mTBI model following investigations up to 24 months postinjury (see Mouzon et al., 2012, 2014, 2018c). We thus continued our investigations by using our proteomic platform to further pursue and explore the proteomic responses and any indicators of overlap or distinction between these different models.
First, the PSAPP mice are reported to show normal acquisition in the performance of cognitive tasks between 6 to 9 months when neuropathology is minimal. However, by 15 to 17 months when the amyloid pathology is widespread, there is a progressive age-related decline in water maze learning and corresponding working memory when compared with young 5- to 7-month-old mice (Holcomb et al., 1999; Arendash et al., 2001). The hTau mice develop tau pathology consistent with human disease (Andorfer et al., 2003). The distribution and pathology of hyperphosphorylation, aggregation of paired helical filaments, and neurofibrillary tangles in these mice develops in an age-dependent manner and is evident 9 to 15 months of age (Andorfer et al., 2003; Andorfer, 2005; Polydoro et al., 2009). Decline in reconsolidation of object-recognition memory and spatial learning and memory has been reported from 12 month old (Polydoro et al., 2009).
Our analyses at 3, 9, and 15 months of age were designed to capture pre, peri, postonset of neurological damage in both models. The 25 proteins significantly changing between PSAPP and wild-type littermates (19 by genotype, 6 with age and genotype) suggest amyloid-dependent alterations in mitochondrial bioenergetics, neurotransmission, cell signaling, vesicular function, and aberrant cytoskeletal reorganization. In the hTau model, the seven proteins that significantly altered compared with wild-type mice and four with age and genotype interaction indicate pathogenic tau-dependent changes in mitochondrial bioenergetics, protein machinery, cell signaling, cytoskeletal reorganization, and axonal transport. In parallel studies from the plasma, we identified 18 plasma proteins significantly changing in our PSAPP mice and 19 proteins changing in our hTau mice compared with counterpart wild-type littermates/controls with aging (Ojo et al., 2018). The top regulated canonical pathways identified by IPA in PSAPP plasma samples with aging compared with their wild-type littermates included production of NO and ROS, acute phase response signaling, and LXR/RXR activation. In hTau plasma, we identified coagulation system, LXR/RXR activation, and production of NO and ROS significantly modulated with aging compared with wild-type controls. No direct overlap was observed in proteins changing in the blood and brain. As both models are driven by the insertion of a transgene and or mutation, the unique pathogenic changes observed in the plasma with aging may thus indicate the consequence of tau and amyloid pathologies, which may be used as a correlative biomarker of these processes in the blood. However, validation of these processes in human studies is needed to corroborate our findings.
Previous proteomic approaches in animal models and using human brains exhibiting tau and amyloid pathologies confirm some of our findings. Preceding studies using the PSAPP mouse model compared with age-matched controls demonstrated increased levels of protein oxidation of metabolic enzymes and cytoskeletal proteins (Abdul et al., 2008; Sultana et al., 2011), coupled with significant reductions in proteins involved in energy metabolism such as glyceraldehyde-3-phosphate dehydrogenase and enolase (Abdul et al., 2008; Robinson et al., 2011). In addition, redox proteomics studies in this model also revealed an age-related significant alteration in proteins involved in mitochondrial bioenergetics (Swomley and Butterfield, 2015). In a recent study involving 8-month-old 5xFAD mice (where amyloid pathology is observed as early as 2 to 3 months), IPA of proteins identified to be significantly modulated in the hippocampus of Tg mice versus controls revealed mitochondrial dysfunction and Gi-mediated intracellular signaling which is involved with the inhibition of cAMP production from ATP and reduction in neuronal excitability and neurotransmitter release (Neuner et al., 2017). In 3xTgAD mice, significant alterations were reported (postonset of amyloidosis) in proteins associated with synaptic plasticity, neurite outgrowth, and microtubule dynamics (Martin et al., 2008). Intracerebral injections of Aβ (1–40) have also been shown to decrease levels of proteins linked with the production of ATP and cytoskeletal integrity (Shi et al., 2011). Aβ (1-42) injections into the nucleus basalis magnocellularis of rodents also showed significant changes to proteins such as glutamine synthetase, beta-synuclein, 14-3-3 zeta, phosphoglycerate mutase 1, pyruvate dehydrogenase, and glyceraldehyde-3-phosphate dehydrogenase in the cortex, nucleus basalis magnocellularis, and hippocampus (Boyd-Kimball et al., 2005). In vitro studies exploring the effects of different Aβ species exposed to cell cultures have also corroborated some of these molecular findings using targeted and unbiased proteomic approaches. For example, exposure of Aβ (25–35), Aβ (1–40) or oligomeric Aβ (1–42) to neuronal cell cultures results in the production of free radicals/ROS/protein carbonyls (Hensley et al., 1994; Harris et al., 1995; Yatin et al., 1999, 2000; Butterfield and Boyd-Kimball, 2004), oxidation of proteins associated with energy metabolism and cytoskeletal function (Boyd-Kimball et al., 2005; Sultana et al., 2006a, 2006b, Sultana and Butterfield, 2009), decreased levels of metabolic enzymes, and increased chaperone proteins (Földi et al., 2011). Immortalized microglial cells exposed to Aβ (25–35) also demonstrated downregulation of metabolic enzymes, redox proteins, and chaperone proteins (Di Francesco et al., 2012). These together implicate a pivotal role in mitochondrial bioenergetics, synaptic plasticity, and protein clearance mechanisms in association with amyloidogenesis.
Very few proteomic studies have been conducted in tau transgenic mice. One study used a NSE-htau23 model that displays NFT deposition (Chang et al., 2013). Proteomic analyses at 6 and 12 months of age identified downregulation of glutathione S-transferase P1 and carbonic anhydrase II with the progression of NFT pathology, upregulation in secerin-1 and V-type proton ATPase subunit E 1 (ATP6VE1) in early stages, and downregulation in the chronic stages of tauopathy. As with our analyses, these changes suggest a unique and important role in dysfunctional energy metabolism in association with age-related tauopathy.
A plethora of proteomics studies conducted on human tissue exhibiting tauopathy and amyloidosis also validate our preclinical studies. Cytoskeletal protein phosphorylation enzyme Pin-1 and microtubule-associated protein, β-tubulin, were significantly reduced, while glycolytic enzymes and molecular chaperone HSP 70 were significantly increased in autopsy brains (Sultana et al., 2007). Similar changes have been reported in detergent insoluble fractions of AD brains with tau/amyloid pathology showing alteration in proteins involved in energy metabolism (Woltjer et al., 2005). Positron emission tomography scanning supports these finding implicating deficiency in energy metabolism, as deficits in cerebral glucose uptake is a typical feature observed in the brains of patients with tauopathy and amyloidosis (Watson and Craft, 2004). A redox proteomic approach of human brain tissue with pathologically confirmed tau and amyloid pathology also identified oxidation of a variety of proteins and oxidative stress as a driving force in the neuropathological sequelae to tauopathy and amyloidosis (Castegna et al., 2002a, 2002b; Butterfield and Castegna, 2003; Castegna et al., 2002c; Castegna et al., 2003; Sultana et al., 2006b). In another study, Begcevic et al. (2013) used a semiquantitative proteomic analyses to interrogate human brain tissue with confirmed tau and amyloid pathology and age-matched controls. The authors reported alterations in metabolic processing of amyloid precursor protein, diminished oxidative phosphorylation, dysfunction in calcium homeostasis and mitochondrial function, and aggregation of Aβ. Among the biological activities altered, cellular and primary metabolic processes, vesicle-mediated transport, oxidative phosphorylation, and positive regulation of ubiquitin activity were highly significant.
Thus, it appears that from our findings herein and our selection of reports from in vitro animal and human studies, a common theme emerges regarding the main pathophysiological changes in the brains exhibiting tau- and amyloid-driven pathological lesions, which can be categorized by significant changes in mitochondrial function, energy metabolism, signal transduction, synaptic function, structural cytoskeletal organization, proteasome function, and protein clearance mechanisms (Swomley and Butterfield, 2015).
Overlap in Brain Proteomic Profiles in Models of r-mTBI, Tauopathy, and Amyloidosis
When we investigated the molecular comparisons between r-mTBI and two unique models of amyloidosis and tauopathy in the cortex, we found only eight total proteins that were highly statistically significant and common, namely ADP/ATP translocase 2, CAMKII, hemoglobin subunit a, hexokinase-1, NADH-ubiuinone oxidoreductase, and spectrin alpha chain, nonerythrocytic 1, alpha-internexin, and neural cell adhesion molecule 1. These proteins were involved in mitochondrial bioenergetics, aberrant cytoskeletal reorganization, and alterations in intracellular signaling cascades and transduction. Our findings ostensibly support the notion that the sequelae of events following TBI-mediated neurodegeneration and in transgenic models of disease-undefined “amyloidosis” (PSAPP mouse) and “tauopathy” (hTau mouse) only show very subtle similarities in molecular responses, suggesting a more complex picture of the proteomic response, with likely distinct and separate neurodegenerative processes occurring following TBI-neurodegeneration and tau/amyloid pathogenesis.
Limitations
As a common caveat inherent to all proteomic studies, we acknowledge the possibilities of false negative results, inaccuracies in protein identification criteria, and possible issues surrounding the sensitivity of our approach for identifying low abundant proteins. This is primarily attributed to biological and technical limitations that are inherent to current approaches that are unable to yield whole coverage of the full proteome (de Godoy et al., 2006; Cox and Mann, 2011). As such, caution should be used when interpreting any proteomic dataset. Several reasons may explain these caveats. For example, peptides with a low abundant protein may elute at the same time with those from a highly abundant protein and, thus, will be omitted for sequencing by our MS procedure. Also, unique peptides generated by trypsin digestion may possess certain qualities that preclude their adequate ionization, making them incompatible for detection by MS. On the other hand, peptides that ionize well may not be unique and, thus, will not be reliably attributed to a particular protein (de Godoy et al., 2006). It is worth mentioning that we have tried to avoid or minimize any issues surrounding false positive protein identifications by using a very strict FDR cutoff point of 0.01 in this study. In addition, the use of homogenates from entire brain regions may result in masking of more subtle molecular responses that might be cell specific or primarily subcellular in nature.
The use of a sample size of four/group in this study warrants discussion. We selected this sample size based on our literature search of previous proteomic experiments (mentioned earlier) reporting a similar group size. It is worthy of note that our model has been well characterized and demonstrates consistent pathological and behavioral phenotypes (Mouzon et al., 2012, 2014, 2018c; Ojo et al., 2015), which is further supported by the large number of unique and statistically significant proteins identified in the mTBI group. While a larger sample size or higher powered study of larger cohorts merit additional investigation, this work nonetheless supports our proof-of-concept that an unbiased proteomics approach involving a detailed time line of molecular events in preclinical models of disease can serve as an invaluable tool to explore putative changes to proteins and molecular pathways significantly altered during the pathogenesis of disease.
We also acknowledge from hindsight of this study (owing to the lack of tau pathology and amyloid plaques in our TBI model—see Mouzon et al., 2012, 2014, 2018c) that the wild-type mouse model we have chosen may not represent the best platform for exploring how r-mTBI contributes to tau or amyloid pathogenesis. Also, Aβ42 protein is not detectable in our wild-type TBI mice using commercially available ELISA’s, and murine Aβ40 (which can be detected) does not show any change after injury (Mouzon et al., 2014, 2018c). Thus, our reports can focus only on the sequelae of events after TBI (i.e., time-dependent molecular changes) as they compare/relate to molecular changes in the models of tauopathy/amyloidosis.
Our TBI work in hTau mice have observed augmentation of tauopathy in aged rather than young animals (Ojo et al., 2013; Mouzon et al., 2018a, 2018b) and also with a more chronic mTBI paradigm (Ojo et al., 2016a), suggesting that the ability to demonstrate TBI-dependent proteinopathy might be dependent on several factors such as age at injury, frequency of hits, and use of relevant humanized (nonmutant) models (Ojo et al., 2016b). Despite the lack of tau and amyloid pathology in our wild-type TBI model, we have previously reported pathological changes such as axonal injury, gliosis, and white matter degeneration, along with spatial memory deficits from 24 hr to 24 months postinjury (Mouzon et al., 2012, 2014, 2018c), which are all relevant to the complex pathobiological mechanisms involved in TBI-mediated neurodegeneration. We thus consider that our findings are still vital to present to the TBI community, and we are confident that with the development of better models of TBI exhibiting relevant proteinopathies, our future work will be able to closely address this elusive interrelationship at the molecular level.
Another limitation of our study design was the lack of statistical comparison between our TBI and amyloidosis/tauopathy models and also the timepoints/ages in our TBI versus tau/amyloid transgenic groups not included that precluded direct comparisons at 6 and 12 months of age. This was due to our initial study design for our TMT workflow, where we ran samples in batches consisting of (nonnormalized) TBI and sham groups alone in a single 10-plex experiment, or separately Tau or PSAPP versus their wild-type counterparts, in a single 6-plex study. Given this limitation, we were only able to conduct our comparative statistical analyses focusing on sham- and TBI-“specific” changes over the five multiple timepoints investigated and also naïve wild-type control, tauopathy, and amyloidosis-dependent changes over the three chosen ages, looking for unique overlaps in the trajectory of temporal profiles. A better approach from hindsight would have been to include a reference sample in all plexes for normalization of each sample, to enable generation of normalized ratio’s and direct comparison of group differences between different experiments.
Although we have not explored the impact of exposure to five separate anesthesias on molecular changes in the brain, we do not anticipate major differences between r-sham and naïve unexposed mice. Nonetheless, we suggest that inclusion of a naïve group in future studies would have been beneficial to address this issue.
Conclusion
This work was designed to investigate the detailed time-dependent molecular events after r-mTBI, and their overlap with the pathogenic events in animal models of amyloidosis and tauopathy. Herein, we detail a detailed time line of changes following r-mTBI and in tau/amyloid-driven transgenic animal models. We chose to investigate timepoints encompassing pre-, peri-, and post-“onset” of the cognitive and pathological phenotypes. At the outset, we observed a lack of amyloid and tau pathology in our injury model.
In our studies, we report our molecular library of changes from these studies and demonstrate mainly distinct response to TBI in our injury model compared with models of amyloidosis and tauopathy, signifying a distinct TBI-mediated neurodegeneration, and a very complex proteomic response in both cases. This work also suggests that other factors are needed to drive the tau and amyloid pathologies observed in the small percentage of patients exposed to repetitive concussive injuries. Nonetheless, the significantly regulated pathways we have identified in our TBI model (e.g., PI3K/AKT and protein kinase-A signaling, PPARα/RXRα activation, GnRH signaling and B cell receptor signaling) may serve as unique targets for therapeutic intervention against the consequences of r-mTBI.
Given the lack of amyloid and tau pathology in our wild-type injury model possibly owing to the intrinsic properties of murine Aβ, we were unable to explore the impact of r-mTBI on risk for AD. We therefore recognize that more studies need to be conducted in relevant models, such as the humanized Tau and humanized APP knock in mice to investigate the contribution of these pathogenic mechanisms as relevant factors in the population of TBI patients at risk of developing AD in later life. Although beyond the scope of this current study, we acknowledge that further work will be needed to confirm changes in proteins implicated in significantly modulated pathways, their messenger RNA signals, and cellular origin, including validation of our findings in autopsy brain tissue to confirm human relevance. We are pursuing these studies in our current work.
Summary
Thirty hippocampal and 47 cortical proteins were significantly modulated over time in the r-mTBI compared with sham mice, implicating PI3K/AKT, protein kinase A, PPARα/RXRα activation, GnRH, and B cell receptor signaling.
In PSAPP mice, 19 proteins significantly changed in the cortex, but only 7 proteins in hTau mice compared with wild-type littermates.
Overlap between r-mTBI model and the PSAPP/hTau models showed a fairly small coincidental change involving eight significantly regulated proteins, suggesting a very distinct TBI neurodegeneration compared with tau or amyloid pathogenesis.
Supplemental Material
ASN914768 Supplemental Material - Supplemental material for Unbiased Proteomic Approach Identifies Pathobiological Profiles in the Brains of Preclinical Models of Repetitive Mild Traumatic Brain Injury, Tauopathy, and Amyloidosis
Supplemental material, ASN914768 Supplemental Material for Unbiased Proteomic Approach Identifies Pathobiological Profiles in the Brains of Preclinical Models of Repetitive Mild Traumatic Brain Injury, Tauopathy, and Amyloidosis by Joseph O. Ojo, Gogce Crynen, Moustafa Algamal, Prashanti Vallabhaneni, Paige Leary, Benoit Mouzon, Jon M. Reed, Michael Mullan and Fiona Crawford in ASN Neuro
Footnotes
Declaration of Conflicting Interests
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: Jon M. Reed was employed by Boehringer Ingelheim Pharmaceuticals. All other authors declare no competing interests.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This project was supported by the Congressionally Directed Medical Research Program (CDMRP) award to Fiona Crawford (DOD/W81XWH-13-1-0253) and by the Roskamp Foundation. The content presented does not express the views of the Department of Defense or Veteran Affairs or the United States Government.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.