Abstract
Background:
Circulating tumor DNA (ctDNA) is predictive of recurrence in resected stage III melanoma, yet its role in the neoadjuvant setting for clinical stage III (cSIII) is unclear.
Objective:
Assess the association between ctDNA and outcomes following neoadjuvant immunotherapy (IO) + targeted therapy (TT) in cSIII melanoma.
Design:
Patients in the NeoACTIVATE study were treated with neoadjuvant IO + TT, underwent lymphadenectomy, and received adjuvant immunotherapy.
Methods:
Patients with ctDNA testing performed at baseline, pre- and post-operation were analyzed. Baseline positron emission tomography–computed tomography volumetrics and surgical, major pathological responses (MPR) were assessed.
Results:
Thirteen patients had serial ctDNA, 10 (77%) were detectable at baseline, and 9/10 (90%) had ctDNA clearance. Seven (54%) achieved MPR, all with undetectable preoperative ctDNA, yet 3/7 (43%) had disease recurrence.
Conclusion:
ctDNA and MPR are poor predictors of recurrence following neoadjuvant IO + TT for cSIII melanoma patients. Further studies are warranted to define the role of ctDNA in this setting.
Keywords
Introduction
Circulating tumor DNA (ctDNA) has become increasingly utilized in the care of melanoma patients; however, additional data are needed to inform treatment decision-making based on ctDNA results. ctDNA has been studied for both prognostication and on-treatment surveillance in the metastatic setting1,2 and as a biomarker to predict recurrence in resected stage III disease.3,4 The use of ctDNA to predict recurrence in the neoadjuvant setting in clinical stage III (cSIII) melanoma is of special interest, as these patients present with a clinically overt lymph node involvement 5 and are at higher risk of recurrence when compared to patients with an occult disease. 6 In recent years, immunotherapy-based trials examining different neoadjuvant regimens have shown that achieving a major pathological response (MPR) is the best predictor of recurrence-free survival (RFS),7–9 yet clinically available baseline and post-neoadjuvant predictive biomarkers are missing. The use of ctDNA in this setting has been previously reported in a pooled analysis of 40 patients from 2 immunotherapy-based studies—PRADO and OpACIN-NEO, 10 where no patient who converted from detectable baseline ctDNA to undetectable post-surgery ctDNA recurred. In contrast, after a 54-month median follow-up, all patients who had detectable post-surgery ctDNA recurred. For patients who had consistent undetectable ctDNA levels, the test did not perform as well, where 26% did recur despite having consistent undetectable results. Similar results have been observed with neoadjuvant treatment for resectable lung cancer, where ctDNA clearance was associated with achieving an MPR and improved survival. 11
We have previously reported on the outcomes of cSIII melanoma patients treated in the NeoACTIVATE study with a neoadjuvant combination of immunotherapy (IO) and targeted therapy (TT): Arm A (BRAF V600E/K mutated (n = 15): atezolizumab + vemurafenib + cobimetinib) and Arm B (BRAF wild type (n = 15): atezolizumab + cobimetinib). 12 While patients in Arm A had a high rate (87%) of MPR, this did not correlate with improved RFS, and 50% of these patients with MPR eventually recurred. 13 This finding is in contrast to previous reports associating MPR with improved RFS for those treated with purely immunotherapy-based regimens.8,14 Given the limited predictive value of MPR in our study, we aimed to investigate additional biomarkers of response. Here, we report data from the correlative ctDNA analyses and their association with pre-treatment imaging parameters, the pathological response, and the 2-year risk of recurrence. We hypothesized that having baseline detectable ctDNA would correlate with higher disease volume on baseline imaging, and that both metrics would be associated with worse outcomes in terms of achieving an MPR and risk of recurrence.
Methods
Patients
Patients will cSIII melanoma were assigned to one of two arms based on the BRAF status: Arm A—BRAF V600E/K mutated, Arm B—BRAF wild type. All patients were treated with neoadjuvant IO + TT combination, underwent therapeutic lymph node dissection followed by 6 months of adjuvant immunotherapy. The clinical endpoint reported herein was any recurrence within 2 years following surgery, with a median follow-up time of 996 days post-surgery.
ctDNA testing
Only patients who had tumor-informed ctDNA test results available at three predefined time points (baseline/pre-treatment, post-neoadjuvant/preoperative, and postoperative) were included in the analysis. Testing was performed on plasma samples stored in −80°C using the Signatera assay, which consists of a multiplex polymerase chain reaction next-generation sequencing method as previously described.1,15 ctDNA was quantified as the mean tumor molecules (MTM)/ml of plasma. Any result >0 MTM/ml was considered positive.
Pathological data
All patients who were operated on had their surgical specimen assessed for pathological response per the International Melanoma Neoadjuvant Consortium guidelines, 16 with MPR being the dichotomous criterion of interest for the analyses.
Imaging data
Pre-treatment positron emission tomography–computed tomography (PET-CT) data were analyzed by a radiologist with training in nuclear medicine. The radiologist conducted semi-automatic data extraction using a MIM workflow (version 7.3.4; MIM Software Inc., Beachwood, OH, USA). Data collected on each scan consisted of the number of positive nodes, mean and maximum standardized uptake values (SUV), metabolic tumor volume (MTV), and total lesion glycolysis (TLG), and these were exported from MIM RTstruct objects for each lymph node identified. MTV was calculated according to Positron Emission Tomography Response Criteria in Solid Tumors (PERCIST 1.0) criteria. 17 TLG was calculated by multiplying the MTV for each lymph node by its corresponding SUVmean. The total MTV (in ml) and total TLG (in ml × SUV) of each patient were used in the analyses and were defined as the sum of their respective values for all lymph nodes that were identified. 18
Statistical analysis
Due to the small sample size and skewed distribution of the continuous parameters, the non-parametric Wilcoxon rank sum test was used to compare group medians, with most data reported descriptively. Simple linear regression was used to assess the correlation between baseline PET-CT parameters and baseline ctDNA levels. Due to the skewed distribution of the ctDNA results, a natural log transformation with adjustment for zero values was used. For all analyses, a p-value = 0.05 was considered significant. Results were reported according to the REMARK guidelines. 19
Results
At baseline, 14 patients with cSIII melanoma had ctDNA testing performed, with 11 having a detectable result, leading to a sensitivity of 79%. One patient had dropped out of the study prior to treatment initiation, leading to a total of 13 patients (6-arm A (all BRAF V600E mutated), 7-arm B) who had ctDNA testing performed at all three predefined time points and were analyzed. Eight were females (61%), and the median age was 63 (range: 44–76).
At baseline, 10/13 (77%) patients had detectable ctDNA with a median MTM/ml of 0.76 (range: 0–876.19). Of these 10 patients, 5 (50%) had ctDNA clearance to undetectable levels at the post-neoadjuvant/preoperative time point (median 0 MTM/ml, range: 0–81.5) and all but one Arm B patient (90%) had complete ctDNA clearance postoperatively (the single patient had 0.17 MTM/ml).
Of the 13, 5 (38%) had recurrence within 2 years post-surgery, with 2 recurrences in patients who had consistently undetectable ctDNA (one in each arm), 2 in patients who had ctDNA clearance preoperatively (one in each arm), and the 1 Arm B patient who had consistently detectable levels. Two patients (40%) had distant recurrence as their first recurrence; both were Arm B patients who had undetectable ctDNA pre- and post-surgery.
When assessing the correlation between ctDNA and the pathological response, seven patients (54%) had an MPR, five in Arm A, and two in Arm B. All patients who achieved an MPR had undetectable levels at the preoperative time point, with four having ctDNA clearance (57%) and the other three (43%) being consistently undetectable. All seven had undetectable levels postoperatively as well. Nevertheless, of these seven patients, three (43%) had disease recurrence, two had consistently undetectable levels (one in each arm), and one Arm A patient had ctDNA clearance preoperatively. Individual patient-level natural log transformation of the ctDNA levels across the time points with indicators for arm, MPR, and recurrence is presented in Figure 1(a), and based on MPR status, Figure 1(b; baseline), and Figure 1(c; preoperative).
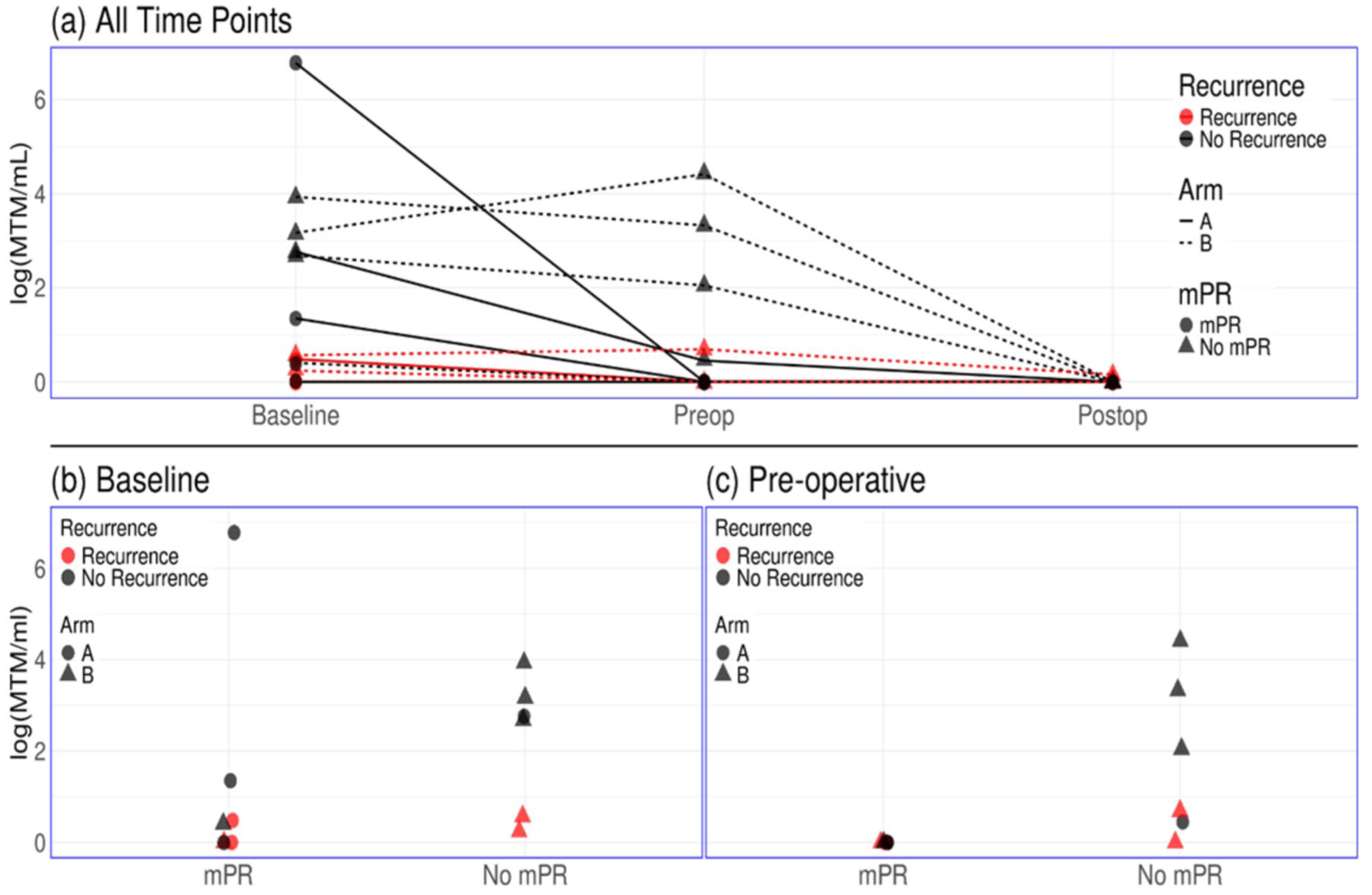
ctDNA at various time points. (a) ctDNA of all patients at the three timepoints with indicators for arm, MPR and recurrence status. (b) ctDNA at the baseline timepoint with indicators for arm and recurrence status. (c) ctDNA at the pre-operative timepoint with indicators for arm and recurrence status.
PET-CT data were available for 11 of 13 patients (85%) at baseline/pre-treatment, of which 4 (36%) had achieved an MPR. At baseline patients who achieved MPR had numerically lower median MTV (31 ml; range: 0.3–105.3) and median TLG (200.8 ml × SUV; range: 0.6–791.4) when compared to those who failed to achieve MPR (MTV: 72.4 ml; range: 1.5–219.3; TLG: 478.9 ml × SUV; range: 4.9–1227), yet these differences were not significant (p = 0.46 for both). On the other hand, patients who experienced disease recurrence within 2 years post-surgery (n = 4) had surprisingly lower values when compared to those who did not recur (n = 7). The median MTV was 3.1 ml (range: 0.3–21.7) for those who experienced recurrence and 105.3 ml (range: 23.8–219.3) for those who did not recur. The same was true of median TLG, which was 10.9 ml × SUV (range: 0.6–117.6) for those who recurred and 791.4 ml × SUV (range: 162.3–1227) for those who did not recur. Both values were statistically significant with p-values of 0.006.
When assessing for associations between PET-CT parameters and the natural log transformed ctDNA results—we found that baseline ctDNA was positively correlated with both MTV (p-value = 0.03, Pearson coefficient = 0.64, 95% CI: 0.06–0.90) and TLG (p-value = 0.02, Pearson coefficient = 0.68, 95% CI: 0.14–0.91). These findings are in line with previous reports about ctDNA shedding correlating with the extent of lymph node involvement, 20 yet our findings were inherently limited by the small sample size. A scatterplot of the baseline MTV and log-transformed ctDNA levels (adjusted for zero values) with indicators for MPR and recurrence is presented in Figure 2.

Scatterplot of baseline metabolic tumor volume versus baseline ctDNA.
Discussion
ctDNA is emerging as a useful biomarker in advanced melanoma, yet the sensitivity of the test in cSIII is limited, with previous reports showing sensitivity of about 33%–48%,4,11 while our study had a higher sensitivity of 79%, albeit with a smaller sample size. When examining ctDNA as a predictor of recurrence, previous reports on ctDNA dynamics following neoadjuvant treatment with purely immunotherapy-based regimens have shown that detectable postoperative ctDNA is highly predictive of recurrence, whereas postoperative undetectable ctDNA is only predictive of a reduced risk of recurrence if there was a clearance of a previously positive test. 10 This leaves achieving an MPR as the most reliable predictor of RFS in those treated with neoadjuvant immunotherapy. 14
In our study, we reported on patients treated with a combination of IO and TT and the association between ctDNA dynamics, baseline PET-CT volumetrics, achieving an MPR, and the 2 years post-surgery risk of recurrence. Our main findings were that baseline lymph node MTV and TLG both correlate with baseline ctDNA levels, yet in our sample, we observed generally low baseline ctDNA levels for those who recurred compared to those who did not (Figure 1(b) red). However, all patients with undetectable baseline ctDNA achieved MPR (Figure 1(b)), and all patients who achieved MPR had undetectable preoperative ctDNA levels (Figure 1(c)).
While the correlation between baseline ctDNA and PET-CT volumetrics was as we hypothesized, we did find that, in contrast to previous reports, ctDNA clearance postoperatively did not predict a reduced risk of recurrence. While for patients who had consistent undetectable levels, previous reports hypothesized that the lack of ctDNA shedding might be due to host factors such as age, weight, and comorbidities, 10 for those who had ctDNA clearance post-surgery, we hypothesized that this might be related to the failure of TT to completely eradicate resistant tumor clones, similarly to the phenomenon seen in the metastatic setting. In addition, we did find a surprising inverse association between the extent of the baseline tumor volume and the risk of recurrence, suggesting that baseline tumor volume is not a valid predictor of recurrence in this setting, or hinting at the possibility that a higher baseline disease volume could induce a stronger and more durable anti-tumor immune response. Alternatively, this finding could also be attributed to chance, given the small sample size, or because our study bundled together two different Arms that had distinct biological properties.
Our findings were limited by the small sample size and, as such, all data should be interpreted with caution. Due to the limited availability of specimens, we were only able to perform the serial ctDNA assay on a portion of the patients who participated in the NeoACTIVATE study, yet these were not selected based on any features other than sample availability. To the best of our knowledge, this is the first report of the utility of ctDNA as a predictive biomarker in patients with cSIII melanoma receiving neoadjuvant treatment with a combination of IO + TT.
Conclusion
For patients with cSIII melanoma treated with neoadjuvant IO combined with TT, both the degree of the pathological response and post-neoadjuvant/preoperatively ctDNA clearance are poor predictors of the risk of recurrence. The role of this combination therapy in the neoadjuvant setting is unclear and warrants further investigations into possible predictive biomarkers that will help tailor treatment intensity and duration based on personalized baseline features. Specifically, more research is needed to better define the role of ctDNA in neoadjuvant-treated cSIII melanoma patients, including investigation of the prognostic abilities of different ctDNA assays with possible integration of other predictive markers such as RNA signatures or radiometabolic parameters.
Supplemental Material
sj-docx-1-tam-10.1177_17588359251412729 – Supplemental material for Biomarker analysis of circulating tumor DNA in clinical stage III melanoma patients treated with neoadjuvant immunotherapy combined with targeted therapy
Supplemental material, sj-docx-1-tam-10.1177_17588359251412729 for Biomarker analysis of circulating tumor DNA in clinical stage III melanoma patients treated with neoadjuvant immunotherapy combined with targeted therapy by Ronen Stoff, David M. Routman, Mariana Yalon, Aaron W. Bogan, Matthew S. Block and Tina J. Hieken in Therapeutic Advances in Medical Oncology
Footnotes
Acknowledgements
We want to thank the staff at the Mayo Clinic Comprehensive Cancer Center who took part in the conduct of this study, and thank all participating patients and their caretakers.
Declarations
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
