Abstract
Background:
Platinum-based compounds have been instrumental in clinical oncology, underscoring the critical need to identify predictive biomarkers for these agents to advance personalized medicine.
Objectives:
This study aimed to identify predictive biomarkers for platinum-based chemotherapies in gastrointestinal cancers.
Design:
This study was designed to explore candidate single-nucleotide polymorphisms (SNPs) through a genome-wide association study (GWAS), followed by re-evaluation using external nationwide cohorts. Subsequently, the biological functions of the selected SNPs were analyzed utilizing the online multi-omics databases.
Methods:
First, we conducted a GWAS in a discovery cohort of esophageal cancer (EC) patients undergoing platinum-based chemoradiation therapy, followed by Cox proportional-hazards analyses for overall survival and progression-free survival. Potential candidate polymorphisms identified in the discovery phase were validated in an external cohort derived from the nationwide BioBank Japan (BBJ).
Results:
Validation analysis revealed a significant association between the T allele of rs12876842 and tumor control rates in a gastric cancer cohort (p = 0.006). In other BBJ cohorts, including the chemoradiotherapy-treated BBJ-EC cohort, the odds ratio trends were consistent with the hazard ratios observed in the discovery cohort, indicating concordance of the risk allele across cancer types, although statistical significance was not reached. Analysis of online databases suggests that the C allele, identified as a risk allele, exhibits higher binding affinity to KLF4, while POLR1D, a downstream target of KLF4, may contribute to adverse prognostic signatures across multiple cancers. Besides, a significant association between genotype of rs12876842 and POLR1D expression in the gastroesophageal junction is confirmed in Genome Tissue Expression (p = 0.0000113), and prior studies have linked POLR1D expression with disease progression, neoplastic transformation, chemotherapy resistance, and poor clinical outcomes in various malignancies.
Conclusion:
Given the centrality of platinum-based drugs in cancer therapy, our findings align with biochemical evidence and prior literature, highlighting rs12876842 as a promising prognostic biomarker.
Keywords
Introduction
Platinum-based derivatives, such as cisplatin and oxaliplatin, have played pivotal roles in clinical oncology since their introduction into the field.1–3 These compounds remain crucial treatment options not only for adjuvant or neoadjuvant chemotherapy (ChT) in perioperative settings but also for patients with distant metastases.4–6 Consequently, identifying predictive biomarkers for platinum derivatives is imperative to establish personalized medicine within oncology.
Recent advances in genomic science and biomedical technology have made genome sequencing increasingly accessible at the bedside.7,8 The cumulative data from clinical sequencing analyses, primarily through candidate gene or single-nucleotide polymorphism (SNP) approaches, have led to the understanding that specific SNPs may influence the therapeutic efficacy of platinum derivatives. Indeed, previous studies have reported that the therapeutic response to platinum-based therapies can be predicted by the genetic status of BRCA1/2.9–12 However, BRCA1/2 variants are relatively frequent in breast, ovarian, prostate, or pancreatic cancers but are rare in gastrointestinal (GI) cancers such as esophageal or gastric cancers. 13 Therefore, there remains a need to identify additional predictive markers for platinum derivatives, particularly in GI cancers. The candidate gene approach mentioned earlier has proven inadequate for this purpose, prompting us to explore relevant SNPs through a genome-wide association study (GWAS) using our newly developed population-optimized SNP array, the “Japonica SNP-array.” 14 This array enabled us to catalog a more comprehensive set of genetic polymorphisms relevant to predicting the efficacy of platinum-based therapies, especially within the Japanese population.
In this study, we employed a whole-genome approach in a discovery cohort, comparing patients with esophageal cancer (EC) who achieved progression-free survival (PFS) for more than five years with those who experienced disease recurrence within five years after initial platinum-based definitive chemoradiation therapy (CRT). Survival outcomes were subsequently evaluated using multivariate hazard analysis, incorporating the genotype of each candidate SNP, age at the initial therapy, and clinical stage. A candidate polymorphism was then re-evaluated in an external validation cohort obtained from the nationwide clinicogenomic database, BioBank Japan (BBJ).15,16 BBJ comprises clinicogenomic data from numerous hospitals across Japan, where cancer patients were treated with various platinum-based therapeutic regimens, reflecting on “real-world” clinical practice. As a result, we identified a candidate SNP potentially regulating POLR1D, a gene implicated in ribosome biogenesis, which we demonstrate to be associated with survival outcomes across various cancer types through reference to external clinicogenomic databases in this study.
Materials and methods
Reporting guideline
We have followed the reporting guideline, REMARKS, suggested by the EQUATOR website (https://www.equator-network.org). 17 The reporting of this study conforms to the REMARKS statement (Supplemental Table).
Ethical approval and consent to participate
The details of ethical approval and the characteristics of patients in cohort A were delineated in our preceding article. 18 In brief, the study received approval from the Institutional Review Board (IRB) of the Tohoku University Graduate School of Medicine (permission numbers: 2003-003, 2005-124, 2016-1-331), the IRB of the National Hospital Organization Mito Medical Center (permission 20060529), the National Hospital Organization Sagamihara National Hospital (permission 2018-051), and the National Center for Global Health and Medicine (permission NCGM-A-003267). A total of 118 EC patients, who were treated with definitive concurrent platinum-based CRT as the first-line treatment and provided written informed consent, were enrolled as Cohort A in this study.
All authors have confirmed that this study was conducted following the Declaration of Helsinki.
Two cohorts of EC treated by CRT as first-line
Two cohorts of EC patients treated with definitive CRT as first-line therapy were analyzed in this study. Cohort A comprised all 118 EC patients treated with platinum-based definitive CRT as the first-line treatment, as reported in our previous publication. 18 From this group, 95 patients were selected for a GWAS and designated as Cohort B. This GWAS in Cohort B compared 59 patients who had remained progression-free for more than five years with 36 patients who experienced disease progression within five years after the initial platinum-based CRT. Twenty-three patients who had remained progression-free for less than five years were excluded from Cohort A (Figure 1(a), Table 1).

CONSORT flow diagram of this clinicogenomic study. (a) Investigation of potential predictive biomarkers for platinum-based therapies in ECs. (b) Compilation of diverse cancer cohorts from the nationwide clinicogenomic database, BBJ. An asterisk indicates that the compilation of the BBJ-EC cohort was described previously. 17
Patients’ characteristics of cohorts A and B and BBJ cohort of EC patients treated by CRT.

The details of cohort A were delineated in our preceding article. 18
p-Values were calculated to compare cohort A with BBJ-EC cohort.
BBJ, BioBank Japan; CRT, chemoradiation therapy; EC, esophageal cancer.
Cohorts of diverse cancers from nationwide “BBJ”
The BBJ-EC cohort was compiled as previously described. 18 The assembly of cohorts for various cancers other than EC is depicted in Figure 1(b). In summary, among the 28,966 registered cancer patients in BBJ, excluding individuals deemed inadequate—such as those lacking CT-based Response Evaluation Criteria in Solid Tumors (RECIST) evaluations, those treated with preoperative neoadjuvant or postoperative adjuvant ChT, those treated with non-platinum derivatives, or those receiving second- or later-line ChT (Figure 1(b), Table 1).
SNP genotyping and imputation
The details of SNP genotyping and imputation in cohort A were described previously 18 : all samples were genotyped using the Japonica Array v1 (Toshiba Corp., Tokyo, Japan). We conducted genotype calling using Affymetrix Power Tools (version 1.18.2; Thermo Fisher Scientific Inc., Waltham, MA, USA). All samples met the manufacturer’s quality control criteria (dish QC ⩾0.82 and sample call rate ⩾97%). SNPs that were categorized as “Recommended” by the SNPolisher package (version 1.5.2; Thermo Fisher Scientific Inc.) were used for subsequent analyses. No cryptic relatives were found among samples using the Maximum Unrelated Set Identification (IMUS) method implemented in PRIMUS v1.8.0 24 with default settings. Haplotype phasing was conducted using SHAPEIT (v2.r644) as a pre-phasing for the genotype imputation after filtering SNPs with a call rate <97.0%, and a Hardy–Weinberg Equilibrium test result of p < 10−6, or minor allele frequency <0.5%. Genotype imputation was performed using IMPUTE2 (ver. 2.3.1) using a phased reference panel of 1070 Japanese individuals.
Genome-wide association study
A genome-wide association test was conducted for the initial screening of potential candidate SNP markers associated with prognosis outcomes, specifically CRT-refractory disease within five years (n = 36) or CRT-response, which includes patients with PFS for more than five years (

Genome-wide association study, comparison between 59 esophageal cancer patients who remained progression-free for over five years and 36 patients who experienced disease progression within five years following definitive chemoradiation therapy (cohort B), followed by eQTL study, Cox hazard analyses by cohort A, and validation study by various BBJ cohorts.
Statistical analyses
Following the GWAS, we assessed the hazard ratios (HRs) for PFS and overall survival (OS) using Cox proportional-hazards multivariate analyses, incorporating the genotype of each candidate SNP alongside stratification factors such as age at CRT, and clinical stage at CRT, as well as Kaplan-Meyer analyses, in Cohort A. The evaluations of HRs using a Cox proportional-hazards model and of survival analyses by the Kaplan-Meyer method were conducted with SSCP (version 27; IBM, Armonk, NY, USA).
The validation cohorts for EC or diverse cancers from BBJ
We selected all EC patients treated with platinum-based CRT as first-line therapy and evaluated by the RECIST version 1.1 of post-treatment imaging from a total of 1338 EC patients in the BBJ project, as previously described. 18 In this study, we compiled the EC cohort from patients treated with concurrent platinum-based CRT, excluding those who received ChT without concurrent radiation.
We compiled the BBJ cohorts for various cancers from all registered cancer patients other than EC (n = 28,966), as illustrated in Figure 1(b). We excluded patients who were not evaluated by CT-based RECIST version 1.1 criteria by radiologists or physicians at each hospital, those treated with adjuvant ChT after surgery or neoadjuvant ChT before surgery, those not treated with platinum derivatives, those who received second-line or later-line ChT, or those lacking detailed information on ChT, as depicted in Figure 1(b).
Tumor response, as assessed by RECIST version 1.1 by radiologists or physicians at each hospital where the registered patients were treated, was adopted as the endpoint of clinical outcome. This approach was taken because long-term prognosis and follow-up data were incomplete. In contrast, therapeutic efficacy was precisely evaluated by post-treatment imaging according to RECIST in these BBJ cohorts.
The Transfac analysis
To predict transcription factor (TF) binding, we used TRANSFAC Professional (QIAGEN, Valencia, CA, USA; http://www.gene-regulation.com/pub/databases.html). 20
The online multi-omics datasets
We evaluated the potential effects induced by each genotype of rs12876842 using online multi-omics datasets, such as the Harmonizome and Genome Tissue Expression (GTEx). In the Harmonizome (http://amp.pharm.mssm.edu/Harmonizome), target genes of the TF KLF4, as suggested by Transfac analysis, are inferred based on CHEA TF targets.21,22 Subsequently, we examined the expression of POLR1D in various human organs using data from the Adult GTEx Project (https://gtexportal.org/home/aboutAdultGtex), which identified POLR1D as one of the target genes of KLF4 according to CHEA TF targets (https://maayanlab.cloud/Harmonizome/gene_set/KLF4/CHEA+Transcription+Factor+Targets). The data used for the analyses described in this manuscript were obtained from the GTEx Portal v.8 (https://www.gtexportal.org/home/eqtlDashboardPage).
The Human Protein Atlas pathology section provides information on the association between genome-wide RNA expression levels and the survival of nearly 8000 cancer patients representing 17 major types of cancer. Detailed information is available at https://www.proteinatlas.org/humanproteome/pathology/method; in summary, based on the FPKM value of each gene, quantified from raw sequencing data obtained from https://gdc-portal.nci.nih.gov/, patients were classified into two groups, and their prognoses were analyzed. 23 We examined the association between POLR1D expression and patient prognoses across several cancer types (https://www.proteinatlas.org/ENSG00000186184-POLR1D/pathology/).
GSE45670 and GSE104958, available on the Gene Expression Omnibus (GEO) portal (https://www.ncbi.nlm.nih.gov/geo/), were employed to obtain expression data from ECs and normal esophageal mucosae prior to neoadjuvant CRT (GSE45670) or neoadjuvant ChT (GSE104958), utilizing microarray technology, and for the pathological evaluation of therapeutic effects on EC tumors post-esophagectomy, respectively.
Results
SNPs with expression quantitative trait locus selected by GWAS comparing those with progression-free more than five years and those with disease progression within five years after definitive CRT
Twenty-nine SNPs were identified through GWAS with p-values less than E-4, in which 59 EC patients who remained progression-free for more than five years and 36 who experienced disease progression within five years, each following first-line CRT, were examined (Table 2, Figure 2). Of these, 15 SNPs were significantly associated with gene expressions, as suggested by expression quantitative trait locus (eQTL) analyses in the online multi-omics database, GTEx. The data used in this manuscript were obtained from the GTEx Portal via dbGaP accession number phs000424.v8.p2 on July 27, 2024.
Twenty-nine SNPs suggested by GWAS and eQTLs in GTEx.

eQTL, expression quantitative trait locus; GTEx, Genome Tissue Expression; GWAS, genome-wide association study; SNP, single-nucleotide polymorphism.
Four SNPs confirmed by Cox hazard-proportional multivariate analyses for OS and PFS in cohort A
The 15 SNPs selected above were subsequently subjected to Cox proportional-hazard multivariate analyses for OS and PFS in 118 EC patients (cohort A). Only rs12876842 and the other three SNPs, namely rs4978306, rs11992426, and rs34503516, were identified with p-values less than 0.00166 after Bonferroni correction, in OS and PFS, respectively (Table 3). Notably, only rs12876842 among these four SNPs is significant in both PFS and OS, which is also confirmed by Kaplan-Meyer analyses as shown in Figure 3, while the other three SNPs are closely located, with chromosomal positions ranging from 94,310,204 to 94,319,870 in chromosome 8.
Cox hazard-proportional multivariate analysis for overall survival (a) and progression-free survival (b).

CI, confidence interval; HR, hazard ratio; SNP, single-nucleotide polymorphism.

Kaplan–Meier analyses stratified by rs12876842 genotypes in Cohort A, comprising 118 esophageal cancer patients primarily treated with CRT and followed for the maximum feasible duration, revealed median survival times of 108.1 and 30.5 months for OS, and 106.3 and 7.0 months for PFS, in the TC and CC genotypes, respectively. Median survival for the TT genotype was not reached for either OS or PFS.
Significant association of rs12876842 with platinum-based therapeutic effects in BBJ-gastric cancer cohort
We re-evaluated the SNPs rs12876842 and rs4978306 that represent the other two SNPs rs11992426 and rs34503516 based on chromosomal location, in independent external nationwide BBJ-EC and diverse cancer cohorts. This comprehensive BBJ clinicogenomic database incorporates cancer diagnoses as well as therapeutic information, including chemotherapeutic regimens administered and CT-based assessments of therapeutic effects conducted by radiologists or physicians at each hospital. The odds ratio for tumor response rate and tumor control rate was calculated from independent reports provided by each hospital, without any influence from the authors. The results are summarized in Table 4. The odds ratio for the tumor control rate is statistically significant, with a p-value of 0.006 in the gastric cancer cohort, where patients were treated with platinum-based chemotherapies and assessed using CT-based RECIST criteria. Moreover, the directions of the odds ratios are consistent with the HRs from the Cox proportional-hazards multivariate analyses, indicating that the risk allele is comparable between the EC CRT cohort and the BBJ diverse cancer cohort, despite the lack of statistical significance in BBJ-EC and colorectal cohorts (Table 4).
CT-based RECIST evaluations for GI-tract cancers in BBJ.
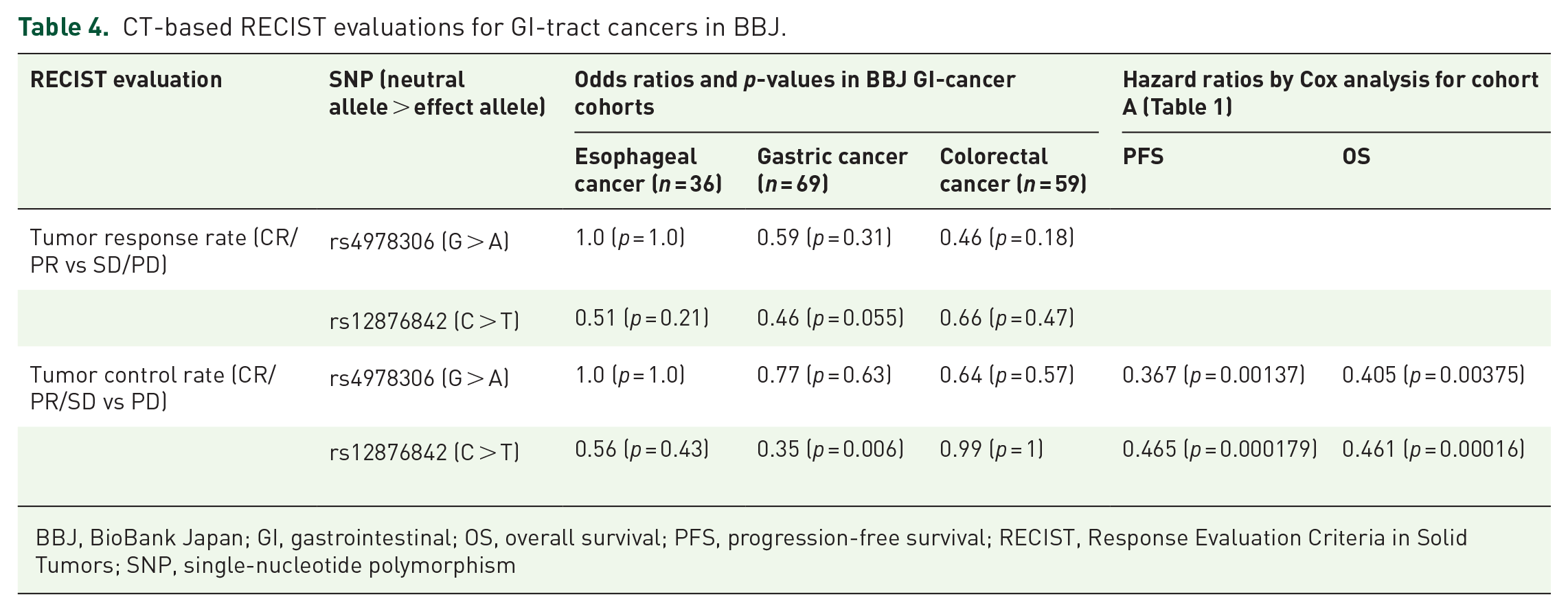
BBJ, BioBank Japan; GI, gastrointestinal; OS, overall survival; PFS, progression-free survival; RECIST, Response Evaluation Criteria in Solid Tumors; SNP, single-nucleotide polymorphism
The Transfac analysis suggested that the genotype at rs12876842 might influence the affinity for TF KLF4
TF binding sites near rs12876842, as predicted by Transfac (https://genexplain.com/transfac/), suggest that the “C” genotype at rs12876842 is expected to exhibit a higher affinity for TFs such as KLF4 compared to the “T” genotype, as illustrated in Figure 4. POLR1D has been identified as one of the downstream genes regulated by KLF4, as described above (https://maayanlab.cloud/Harmonizome/gene_set/KLF4/CHEA+Transcription+Factor+Targets).
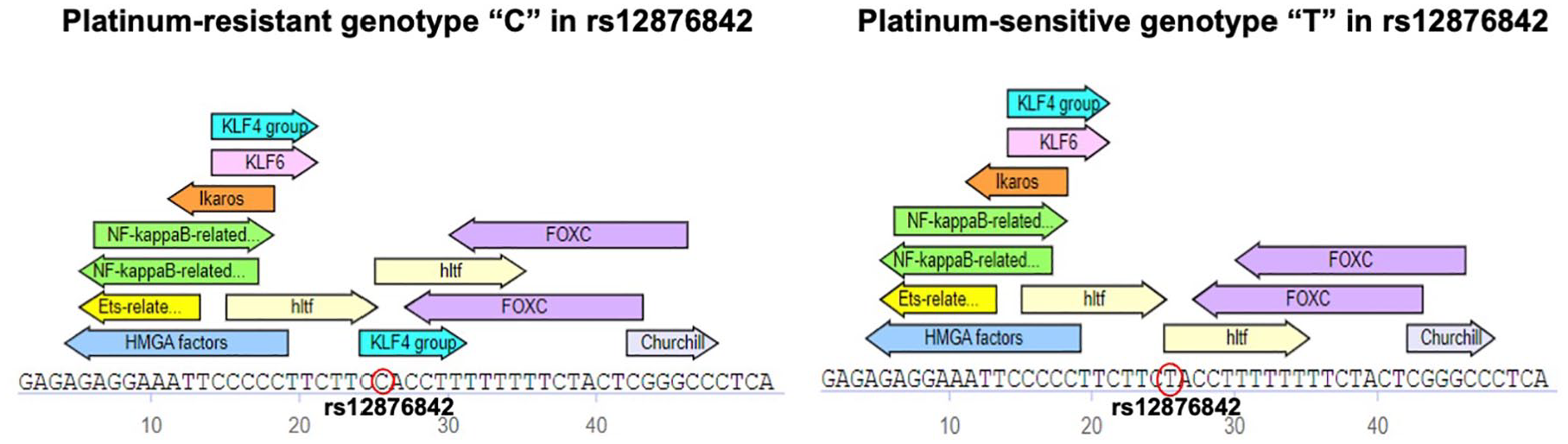
Transcription factor binding sites in the vicinity of rs12876842, as predicted by Transfac (https://genexplain.com/transfac/), suggest that the genotype “C” at rs12876842 is anticipated to demonstrate a higher affinity for transcription factors such as KLF4 in comparison to genotype “T.” POLR1D has been identified as one of the downstream genes modulated by KLF4.
Expression level of POLR1D is significantly associated with rs12876842 in the gastroesophageal junction mucosae
POLR1D is identified as a target of the KLF4 TF in the CHEA TF Targets dataset (https://maayanlab.cloud/Harmonizome/gene_set/KLF4/CHEA+Transcription+Factor+Targets). 21 Furthermore, a significant association between the genotype of rs12876842 and the expression of POLR1D in the gastroesophageal junction (p = 0.0000113) has been demonstrated in the eQTL analysis by GTEx v.8 (https://www.gtexportal.org/home/eqtlDashboardPage).
Reduced expression of POLR1D was significantly associated with improved prognosis in patients with several types of cancers
Reduced expression of POLR1D is associated with improved prognosis in various human malignancies, including head and neck cancer (upper left), gastric cancer (upper right), colorectal cancer (lower left), and cervical cancer (lower right), with respective p-values of 0.00021, 0.33, 0.081, and 0.019, as reported by the Human Protein Atlas version 24.0 (https://www.proteinatlas.org/ENSG00000186184-POLR1D/pathology/; Figure 5). Images are available from Protein Atlas version 24.0 as described in the method. Platinum-based derivatives such as carboplatin, oxaliplatin, and cisplatin are integral to the chemotherapeutic regimens for these cancers.

Reduced expression of POLR1D is associated with improved prognosis in various human malignancies, including head and neck cancer (upper left), gastric cancer (upper right), colorectal cancer (lower left), and cervical cancer (lower right), with respective p-values of 0.00021, 0.33, 0.081, and 0.019, as reported by the Human Protein Atlas version 24.0 (head and neck cancer: https://www.proteinatlas.org/ENSG00000186184-POLR1D/cancer/; gastric cancer: https://www.proteinatlas.org/ENSG00000186184-POLR1D/cancer/stomach+cancer; colorectal cancer: https://www.proteinatlas.org/ENSG00000186184-POLR1D/cancer/colorectal+cancer; cervical cancer: https://www.proteinatlas.org/ENSG00000186184-POLR1D/cancer/cervical+cancer). Platinum-based chemotherapeutic agents, including carboplatin, oxaliplatin, and cisplatin, play pivotal roles in treating these cancers. The blue and red lines indicate survival probabilities for each cancer with low and high POLR1D expression, respectively. The cutoff values defining high and low gene expression were based on the default settings provided by the Human Protein Atlas version 24.0 (https://www.proteinatlas.org/humanproteome/cancer/method).
As for ECs, we retrieved the data from the GEO portal site; the total number of ECs analyzed by microarray was 68 (28 in GSE45670 and 40 in GEO104958), along with 15 normal mucosa samples (10 in GSE45670 and 5 in GSE104958). The baseline expression level of POLR1D was established using the average and standard deviation (SD) calculated from the normal mucosae in each GSE study. The absence of POLR1D expression was evaluated and compared with pathological diagnoses following neoadjuvant ChT or CRT (Table 5). The results indicated that the cases with absent POLR1D expression showed a trend toward a response to neoadjuvant ChT or pathological complete response with odds ratios of 4.950 and 2.708, respectively. However, statistical significance was not reached with p-values of 0.125 and 0.212 in GSE104958 and GSE45670, respectively.
Absent POLR1D in EC tissues before CRT/ChT is associated with pCR or therapeutic effects in online omics databases.

ChT, chemotherapy; CI, confidence interval; CRT, chemoradiotherapy; EC, esophageal cancer; pCR, pathological complete response; SD, standard deviation.
GWAS data availability
The GWAS data utilized in this study have been deposited in the Medical Genomics Japan Variant Database (https://mgend.ncgm.go.jp) under the accession number MGS000084.
Discussion
In this study, we conducted a GWAS to examine five-year disease-free survival in patients with EC who received definitive CRT as first-line treatment (Figures 1 and 2). This analysis was complemented by hazard assessments for OS and PFS using multivariate analyses incorporating genotypes of candidate SNPs, EC clinical stages, and patient ages at initial therapy (Table 3). The GWAS and hazard evaluation cohorts overlapped, necessitating an independent validation cohort. Accordingly, we established an external validation cohort from BBJ, one of the world’s largest disease biobanks, which includes 1338 EC patients and 28,966 patients with various cancers other than EC. This validation cohort enabled us to explore associations between platinum derivatives and objective therapeutic evaluations, such as RECIST criteria (Figure 1(b)). The rationale for using RECIST-based post-treatment imaging evaluations as a prognostic endpoint has been detailed previously. 18 In summary, BBJ cancer patients lacked extensive long-term follow-up for survival analyses, making an objective, widely accepted therapeutic evaluation, such as RECIST, essential for a nationwide cohort spanning multiple hospitals across Japan.
For this study, we assembled three cohorts. Through GWAS, followed by Cox proportional-hazards multivariate analyses for OS and PFS, and utilizing the BBJ cohort, we identified and validated one SNP, rs12876842 (Table 3 and Figure 3). The BBJ validation revealed a statistically significant association of the T allele of rs12876842 with tumor control rate (complete response, partial response, or stable disease) in the gastric cancer cohort (p = 0.006). This finding suggests that the T allele of rs12876842 is correlated with improved prognoses in platinum-treated gastric cancer patients (Table 4). In other BBJ cohorts, such as those with esophageal and colorectal cancers, the odds ratio directions aligned with HRs from Cox multivariate analyses (Table 4), indicating consistency of the risk allele between the EC CRT cohort (cohorts A and B in this study) and the broader BBJ cancer cohort, though statistical significance was lacking. The lack of statistical significance between the alternative allele (T) and tumor regulation in the BBJ esophageal and colorectal cancer cohorts may be due to the limited number of BBJ cancer patients treated with platinum derivatives. Although BBJ is among the most prominent global biobanks, covering 47 diseases, including various cancers, cerebral infarction, cerebral aneurysm, etc.,15,16 the patient count per disease cohort remains relatively small (Figure 1). Moreover, the BBJ cohorts likely reflected real-world clinical practices, with treatment strategies exhibiting considerable heterogeneity. These limitations necessitate further validation of our findings in larger and prospective cohorts in the future.
Subsequently, we assessed the biological, biochemical, and oncological functions associated with rs12876842 genotypes using online databases such as Transfac, GTEx, CHEA transcription target dataset, and the Human Protein Atlas. The findings were as follows: first, rs12876842 genotypes influence affinity for the TF KLF4. The C allele, identified as a risk allele in this study, demonstrates higher KLF4 affinity per Transfac analysis (Figure 4). Second, POLR1D, identified as a downstream target gene of KLF4 based on the CHEA transcription target dataset (http://amp.pharm.mssm.edu/Harmonizome), was validated by GTEx expression data, revealing that the C allele of rs12876842 is significantly associated with elevated POLR1D expression in esophageal and gastric tissues, particularly at the esophagogastric junction (p = 0.0000113), as evidenced by GTEx eQTL analysis (https://www.gtexportal.org/home/eqtlDashboardPage). Elevated POLR1D expression correlates with poorer prognoses across various cancers, such as cervical, gastric, colorectal, and head and neck, consistent with our findings (Figure 5). In external EC cohorts, comprehensive microarray-based gene expression data prior to neoadjuvant CRT/ChT and post-treatment pathological diagnoses are available online as GSE104958 and GSE45670, corresponding to platinum-based CRT and a cisplatin, docetaxel, and 5-fluorouracil regimen, respectively. The findings, consistent with ours, suggest that POLR1D overexpression is associated with poor prognosis, although statistical significance was not reached (Table 5). Thus, the candidate SNP identified in this study, likely associated with POLR1D expression, may contribute to the adverse prognostic signatures observed across cancer types. Previous studies have proposed that POLR1D expression levels may correlate with disease progression, neoplastic transformation, chemotherapy resistance—especially against platinum-based agents—and poorer outcomes in multiple cancers.24–33 Consequently, POLR1D is emerging as a potential target for novel anticancer therapies.34–37
Conclusion
As platinum-based derivatives are central to chemotherapy across various malignancies, the prognostic analyses from our EC cohort and the diverse BBJ cancer cohorts are consistent with the biochemical findings and previously reported studies discussed above, highlighting rs12876842 as a potential prognostic biomarker. Nonetheless, further investigation is warranted to explore the clinical applicability of SNP rs12876842 in personalizing platinum-based chemotherapies, as statistical significance was not achieved in several analyses, likely due to the limited number of enrolled patients and treatment heterogeneity reflecting real-world clinical practices in nationwide BBJ cohorts, which represent a limitation of this study.
Supplemental Material
sj-docx-1-tam-10.1177_17588359251355079 – Supplemental material for A POLR1D-regulating single-nucleotide polymorphism as a predictive marker candidate for platinum-based chemotherapy in gastrointestinal cancers
Supplemental material, sj-docx-1-tam-10.1177_17588359251355079 for A POLR1D-regulating single-nucleotide polymorphism as a predictive marker candidate for platinum-based chemotherapy in gastrointestinal cancers by Takahiro Mori, Kazuko Ueno, Masao Nagasaki and Koichi Matsuda in Therapeutic Advances in Medical Oncology
Footnotes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
