Abstract
Background:
The widespread use of next-generation sequencing (NGS) panels to guide targeted therapies in advanced malignancies raises questions regarding their real-world impact on clinical outcomes and affordability.
Objectives:
The study aims to find trends in targetable genetic alterations among late-stage cancer patients in the Jordanian population and to assess the clinical effectiveness of NGS testing in Jordan.
Design:
This article is a retrospective observational analysis. Data extracted from patient files were used to determine genetic alteration trends and for a comparative assessment of progression-free survival (PFS) between patients receiving NGS-guided personalized treatment compared to those receiving conventional treatment.
Methods:
Two hundred thirteen nonselected late-stage cancer patients from three Jordanian hospitals (2019–2024) were included in this study. Key data points included demographics, genetic variations, and treatment details. Statistical analysis involved the use of the Cox proportional hazards model to evaluate PFS between the two groups of patients.
Results:
Clinically significant genetic alterations were found in 89 patients (42%), with the most common genes being KRAS (20%), PIK3CA (9%), ERBB2 (7%), EGFR (6%), and BRCA1 and 2 (1.5% each). Microsatellite instability-high status was seen in 8 patients (3.7%). Out of the 89 patients with targetable genetic variations, 80.9% received their recommended NGS-guided treatments, while 19.1% did not due to cost or drug unavailability. PFS was significantly improved for patients receiving NGS-guided targeted treatment compared to conventional treatment in the general population with a hazard ratio (HR) of 0.216, the colorectal cancer population (HR of 0.185), the breast cancer population (HR of 0.239), and the lung cancer population (HR of 0.089).
Conclusion:
NGS testing is a very beneficial tool for patients with advanced malignancies to select targeted therapies and potentially improve survival. However, the incorporation of NGS testing in Jordan faces challenges such as high testing and treatment costs and drug unavailability.
Keywords
Introduction
Next-generation sequencing (NGS) enables the identification of specific genetic alterations, allowing for the development of targeted therapies for cancer patients. It has become an important tool in personalized medicine. 1 This treatment approach is different from conventional cancer treatments, which may not account for the disease heterogeneity between patients, leading to ineffective, expensive treatments and potentially avoidable side effects. 2 For late-stage cancer patients, NGS testing allows for rapid decision-making by offering a complete genomic profile that identifies alterations driving tumor growth and potential therapeutic targets, as well as recommending various therapies. 3
Despite its potential, the adoption of NGS in clinical practice faces several challenges, including cost, drug availability, and unnecessary testing. In addition, not all identified alterations are actionable, and for some patients, NGS testing may not lead to a viable therapeutic option.
The overall aim of this study is to evaluate the clinical effectiveness of NGS tests in cancer treatment in Jordan. Conducting this study in Jordan is particularly important, as, to our best knowledge, we are not aware of any research conducted on this specific demographic that covers NGS testing. Jordan has been listed as a low-middle income country, 4 where NGS-driven personalized treatment might not always be accessible due to cost and availability; therefore, it is crucial that we verify the clinical impact of NGS testing. Prospective studies and cost analysis are required before including it in public health policies in Jordan.
Methods
Design
This study is a retrospective observational analysis. It aims to evaluate the practical effectiveness of NGS testing in real-world clinics in guiding personalized cancer treatment compared to conventional treatment in Jordan. Data was gathered from medical records for patients diagnosed with stage 4 cancer. Patients were categorized into two groups based on the type of treatment they received: NGS-guided personalized treatment and conventional treatment. This allowed for a comparative assessment of progression-free survival (PFS) outcomes across both groups.
PFS is defined as the duration from the start of treatment until disease progression or death. 5
This study was approved by the Institutional Review Board (IRB) at Al-Khalidi Hospital.
The reporting of this study conforms to the Strengthening the Reporting of Observational Studies in Epidemiology (STROBE) statement 6 (Supplemental File).
Population characteristics
The study included 213 patients with various types of stage 4 cancer from three private hospitals in Amman, Jordan, conducted from 2019 to 2024.
NGS testing was performed using Guardant360 tests (Guardant360 CDx or Guardant360 TissueNext), developed by the Guardant Health Clinical Laboratory in Redwood City, CA, USA. 7
For patients who received NGS-guided personalized treatment, the choice of therapy given was based on the attending physician’s choice; some received treatment with strong clinical evidence specific to their tumor type and genetic variation (group A), while others received treatment with clinical evidence for other tumor types with the patients’ genetic variation (group B).
Key data points, including gender, age, diagnosis, clinically significant genetic variations, tumor mutational burden (TMB), microsatellite stability status, and suggested therapies, were extracted from patient files. In addition, data on the administration of recommended targeted therapies were collected from the patient’s primary physicians.
Statistical analysis
To make sure age and gender were not significantly different between the two groups, independent sample t-tests were carried out. The patients were grouped based on treatment type, and the mean differences in age and gender between the two groups were tested for statistical significance.
A Cox proportional hazards model was fitted to all data to evaluate the clinical effectiveness of personalized treatment compared to conventional treatment. The covariates used in the model were as follows: age, gender, treatment type (NGS-guided personalized treatment vs conventional treatment), and PFS status.
Regarding PFS, a maximum follow-up date of October 1, 2024, was used to determine if there was disease progression or death.
This analysis was done on Python (version 3.12), 8 using the SciPy library 9 for the statistical analysis and the lifelines library for survival analysis. 10
As well as an analysis of all the patients combined, separate Cox proportional hazards analyses were conducted for each of the most prominent cancer types in our study population: colorectal, breast, lung, and pancreatic cancer in that respective order.
We calculated hazard ratios (HR) with 95% confidence intervals (CI) to quantify the effect of the covariates on the risk of disease progression. A significance level of p < 0.05 was considered statistically significant.
Results
Demographics
Our study included 213 patients with a median age of 58 years, with our sample being split into 52.6% (n = 112) males and 47.4% (n = 101) females.
Details about the distribution of cancer types in our study population are shown in Table 1.
Cancer types in the study population.
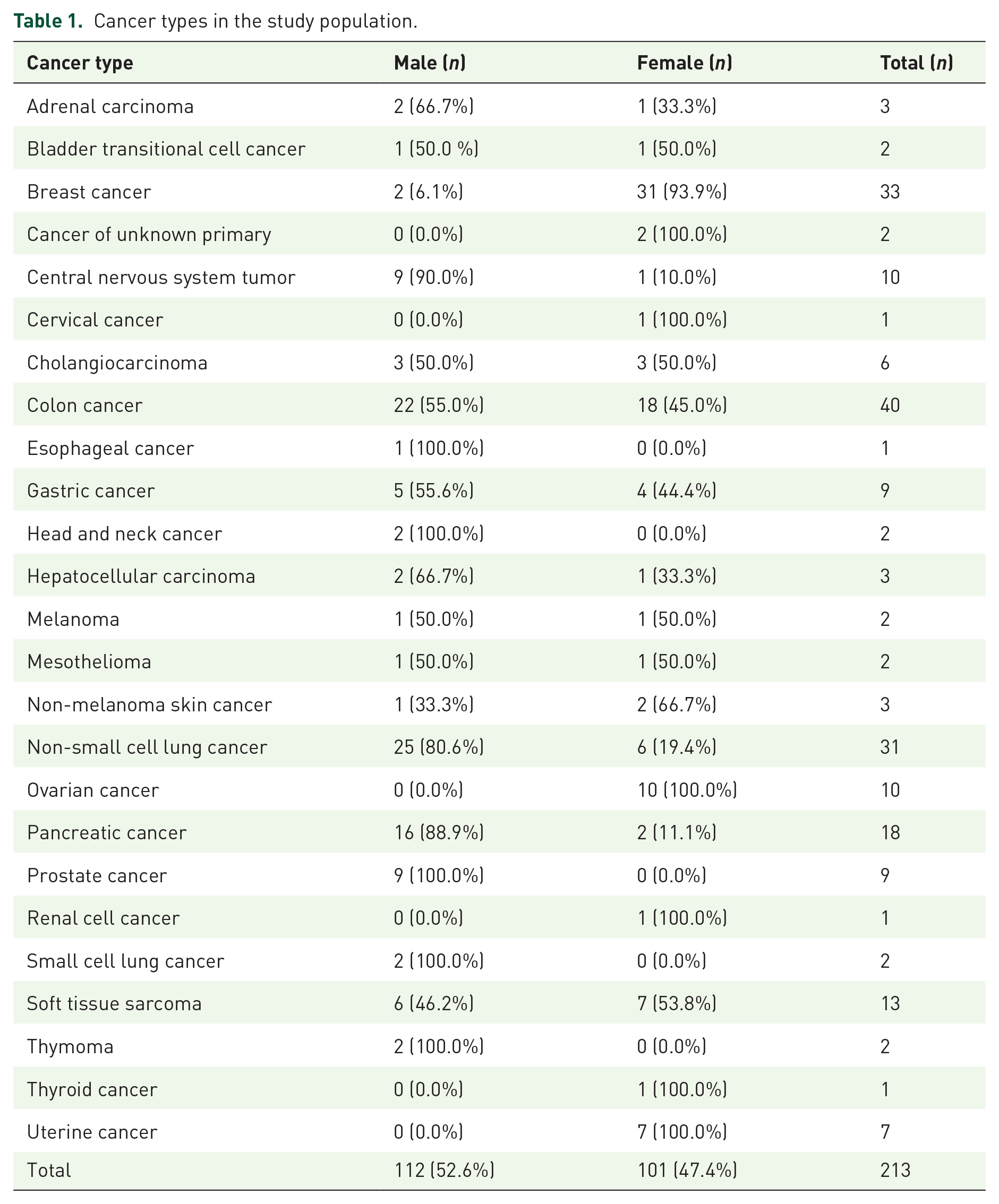
Of the 213 patients, 89 patients (41.8%) exhibited clinically significant variations (group A and group B); of those, 72 (80.9%) received their recommended treatment: 55 were classified as group A and 17 as group B. Seventeen patients (19.1%) could not access the recommended drugs (8 due to high treatment cost and 9 due to drug unavailability in Jordan), causing them to undergo conventional treatment with the 124 patients that had no targetable genetic variations, resulting in 141 (66.2%) patients overall.
The independent sample t-test for age and gender distribution between the patients receiving personalized treatment and patients receiving conventional treatment showed no significant differences in age (T = 0.43, p = 0.665) nor gender distribution (T = 0.83, p = 0.409). This indicates the two groups are well balanced regarding age and gender distribution.
Clinically significant genetic alterations
The most significant genetic alterations among the study population are shown in Table 2.
The clinically significant genetic alterations detected through NGS testing in our study population.

ALK, anaplastic lymphoma kinase; BRAF, V-Raf Murine Sarcoma Viral Oncogene Homolog B; BRCA1, breast cancer gene 1; BRCA2, breast cancer gene 2; EGFR, epidermal growth factor receptor; EML4, echinoderm microtubule-associated protein-like 4; ERBB2, v-erb-b2 avian erythroblastic leukemia viral oncogene homolog 2; FGFR3, fibroblast growth factor receptor 3; HCC, hepatocellular carcinoma; KRAS, Kristen Rat Sarcoma Viral oncogene homolog; MET, mesenchymal-epithelial transition; NGS, next-generation sequencing; NRAS, neuroblastoma ras viral oncogene homolog; NSCLC, non-small cell lung cancer; NTRK1, neurotrophic receptor tyrosine kinase 1; NTRK2, neurotrophic receptor tyrosine kinase 2; PIK3CA, phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit alpha; TCC, transitional cell carcinoma.
High TMB (defined as higher than 10 Muts/Mb) 11 was seen in five non-small cell lung cancer (NSCLC) patients (four non-squamous and one squamous), one melanoma patient, one colon cancer patient, one bladder transitional cell carcinoma (TCC) patient, two soft tissue sarcoma patients, and two pancreatic adenocarcinoma patients.
Microsatellite instability-high status tumors were seen in four patients with colon cancer, one with gastric adenocarcinoma, one with breast cancer, one with bladder TCC, and one with small cell lung cancer.
Homologous recombination deficiency (HRD)–positive tumors were seen in three breast cancer patients, three pancreatic adenocarcinoma patients, three prostate adenocarcinoma patients, one hepatocellular carcinoma (HCC) patient, one NSCLC patient, one endometrial adenocarcinoma patient, and one ovarian cancer patient.
Survival analysis
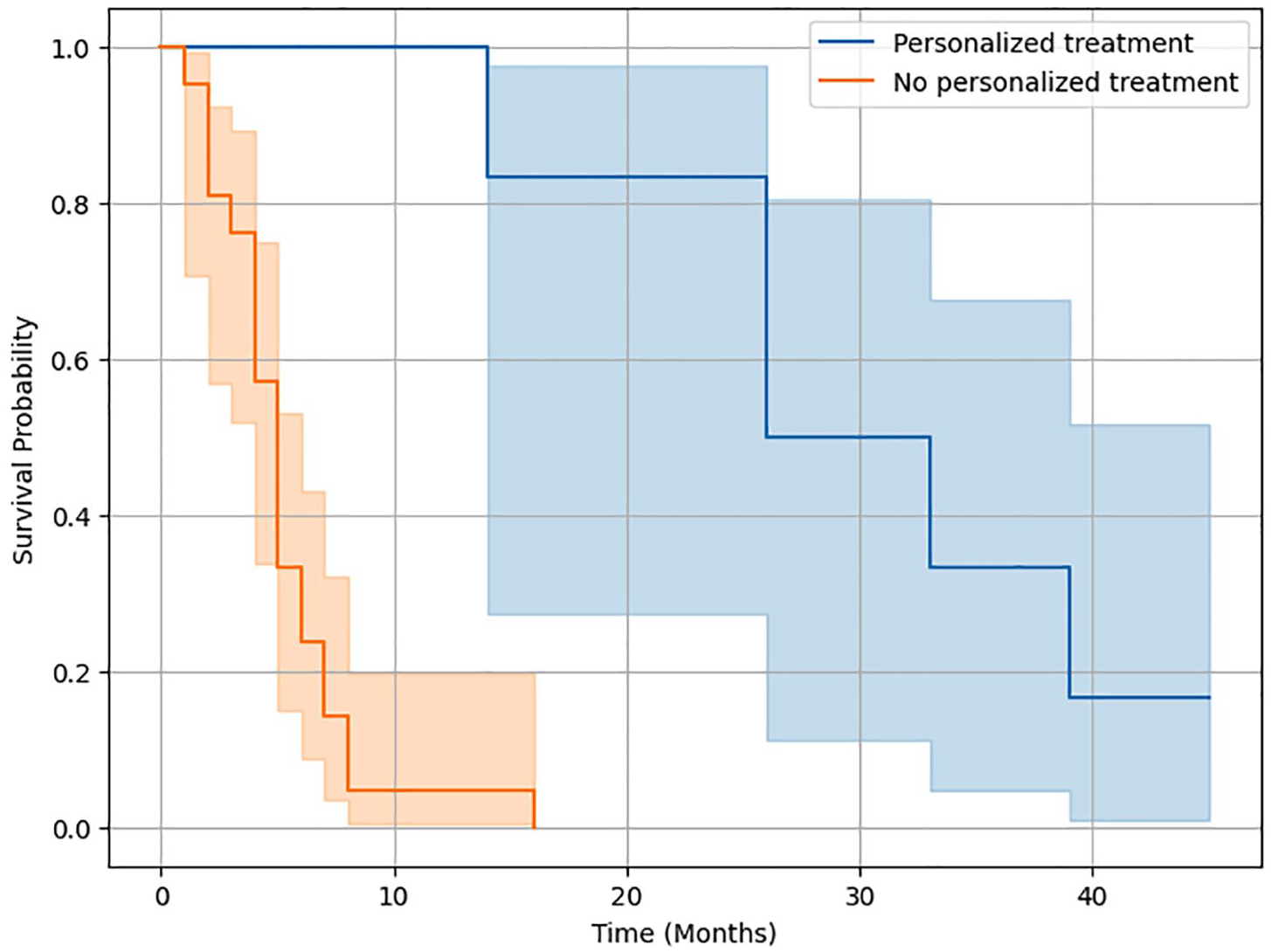
We investigated the effect of gender, age, and type of treatment on PFS in the whole population (n = 213) using Cox proportional hazards analyses. NGS-guided personalized treatment showed a significant association with improved PFS, with an HR of 0.216 (95% CI: 0.157–0.299, p < 0.001), suggesting that patients receiving personalized treatment had a significantly lower risk of progression compared to patients receiving conventional treatment. The difference in survival probabilities in favor of the personalized treatment group is further shown by the Kaplan-Meier survival curves by treatment type in Figure 1.

Kaplan-Meier survival curves by treatment group for the whole population.
Regarding the association of gender with PFS, the HR was 0.853 (95% CI: 0.650–1.121), with a p-value higher than 0.05 (p = 0.255), showing a statistically insignificant lower risk for one gender over the other. Age also indicated a minimal effect on PFS, with an HR of 0.993 (95% CI: 0.984–1.002) and no statistically significant association (p = 0.125).
For the Cox proportional hazards analysis for patients with colorectal cancer (n = 40), personalized treatment showed a statistically significant association with improving PFS values, with an HR of 0.185 (95% CI: 0.067–0.512, p < 0.001). However, age did not show any significant improvement at an HR of 1.013 (95% CI: 0.985–1.042, p = 0.900). Gender was also insignificant at an HR of 0.979 (95% CI: 0.489–1.959, p = 0.951) (Figure 2).

Kaplan-Meier survival curves by treatment group for colorectal cancer patients.
The Cox proportional hazards analysis for patients with breast cancer (n = 33) showed that personalized treatment showed a statistically significant association with improving PFS values, with an HR of 0.239 (95% CI: 0.099–0.579, p = 0.002). However, age did not show any significant improvement at an HR of 1.010 (95% CI: 0.983–1.039, p = 0.463). Gender was also insignificant at an HR of 1.273 (95% CI: 0.304–5.325, p = 0.741) (Figure 3).

Kaplan-Meier survival curves by treatment group for breast cancer patients.
The Cox proportional hazards analysis for patients with NSCLC (n = 31) suggested that personalized treatment showed a statistically significant association with improving PFS values with an HR of 0.089 (95% CI: 0.028–0.284, p < 0.001). However, age did not show any significant improvement at an HR of 1.026 (95% CI: 0.982–1.072, p = 0.253). Gender was also insignificant at an HR of 0.707 (95% CI: 0.254–1.969, p = 0.507) (Figure 4).
Kaplan-Meier survival curves by treatment group for NSCLC patients.
Finally, for the Cox proportional hazards analysis for patients with pancreatic cancer (n = 18), personalized treatment showed no statistically significant association with improving PFS values with an HR of 0.240 (95% CI: 0.046–1.247, p = 0.090) (Figure 5). Age did not show any significant improvement at an HR of 0.968 (95% CI: 0.046–1.247, p = 0.205). Gender was also insignificant at an HR of 1.376 (95% CI: 0.360–5.261, p = 0.641).

Kaplan-Meier survival curves by treatment group for pancreatic cancer patients.
A summary of the Cox proportional hazards models for all groups of the study are shown in Table 3.
Cox proportional hazards model analyses results.
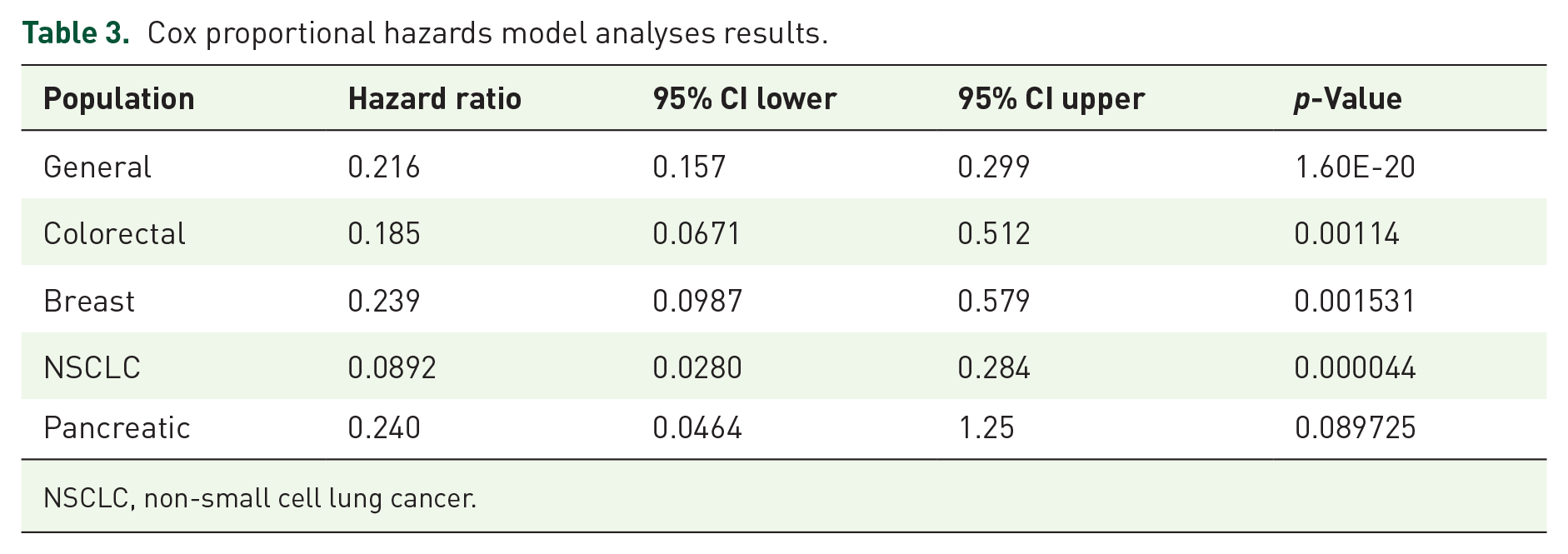
NSCLC, non-small cell lung cancer.
Discussion
Our study showed interesting findings regarding clinically significant genetic alterations. High TMB and microsatellite instability-high cancers were found in 12 and 8 patients, respectively. These are very important targets for Food and Drug Administration (FDA)-approved immunotherapies and have strong clinical evidence for survival benefit.12,13 These results are usually difficult to detect using conventional testing methods available in Jordan.
HRD-positive cancer is considered a candidate for treatment with Poly (ADP-ribose) polymerase (PARP) inhibitors. 14 Our study identified 11 patients with significant cancers who are HRD-positive (breast, pancreas, prostate, and ovarian cancers). HRD score is not available in Jordan, and unless we use NGS testing, these patients will miss the opportunity for significant targeted therapy that would not be achieved using conventional labs available locally.
Thirty-five patients (16%) were found to have actionable genetic alterations with FDA-approved targeted therapies that would not have been identified using conventional methods in Jordan; these include five NSCLC patients with KRAS (G12C) mutations, five patients with BRAF V600E mutations, eight breast cancer patients with PIK3CA mutations, six patients with ERBB2 alterations (gastric, esophageal, and NSCLC), six breast cancer patients with BRCA1 or BRCA2 mutations, two bladder TCC patients with FGFR3 mutations, one NSCLC patient with MET mutation, and two patients with NTRK mutations.
Our findings on common alteration types in various cancer types are consistent with existing literature. Regarding KRAS mutations, our study showed a high occurrence in colon cancer patients (54.5%), pancreatic cancer patients (22.7%), and NSCLC patients (15.9%). Yang et al. 15 found similar results, stating that 81.72% of patients with KRAS mutations are pancreatic cancer patients, followed by colon cancer patients at 37.97% and 21.20% of patients with NSCLC. The G12D variation of the KRAS mutation was the most common in both studies (43.2% in our study compared to approximately 29.2% in Yang et al.’s study). Both studies showed a high number of G12D mutations in both colon and pancreatic cancers, with a lower number of patients with NSCLC. The G12C variation was seen to be the most prominent in NSCLC patients in both studies (71.4% compared to 45.0%). Our study reported a high number of G12V variant mutations in colon cancer patients (50.0%) and pancreatic cancer patients (37.5%), with NSCLC patients representing 12.5% of patients. Yang et al. reported G12V being common in pancreatic cancer patients (31.96%) and colon cancer patients (18.5%). Their study also showed that the G12V variation was the second most common variation in NSCLC patients, which is supported by our findings.
As can be seen from the results, this study shows the significant clinical benefit of NGS-guided personalized treatment in improving PFS for late-stage cancer patients in Jordan. Throughout the entire study population, patients receiving personalized treatment had a statistically significant lower disease progression risk compared to patients receiving conventional therapy, demonstrated by an HR of 0.216 (95% CI: 0.157–0.299, p < 0.001). This indicates that matching therapies based on the genetic profiling of patients produced by NGS testing substantially increases survival among patients.
The analysis for specific cancer-type populations further solidifies our initial findings. For colorectal, breast, and lung cancer patients, personalized treatment was consistent with improved PFS values, with HRs between 0.089 and 0.239, all statistically significant with p-values under 0.05.
These findings are aligned with existing literature that shows the improved clinical capabilities associated with using NGS testing. Gibbs et al. 16 carried out a literature review of publications that covered the PFS and overall survival (OS) of patients receiving NGS testing. They found that 11 out of 16 publications showed significantly increased PFS and OS for patients who were matched with NGS-guided targeted treatment. This matches our observations that showed significantly improved PFS outcomes with NGS-guided therapies compared to conventional therapies.
Conversely, Kang et al. 17 present a nuanced view, indicating that while NGS improves the detection of specific genetic mutations, it does not improve survival across all patient populations. In their study conducted in South Korea, it was found that NGS did not improve the OS of a general population with late-stage cancer, except in specific subgroups that did not have viable existing targeted therapies. This is an important distinction in our study in Jordan, as it highlights the need for NGS in patient groups that are most likely to benefit, rather than a generalized application across all late-stage cancer patients. This is shown by the proportion of patients in our study who did the NGS test but did not gain any benefit from it due to the absence of any significant targetable genetic alterations; they represented around 58.2% of our study population.
Recent advances in genomic sequences have greatly increased our understanding of how targetable mutations significantly enhance treatment outcomes. For example, KRAS mutations, which were long considered “undruggable,” have become frontrunners in targeted genetic therapies with the development of KRAS inhibitors such as Sotorasib and Adagrasib, offering promising results in cancers such as NSCLC and colorectal cancer. 18 Incorporating these findings into a clinical effectiveness study showcases the technical and real-world problems that might hinder the incorporation of these advances into patient care.
A substantial portion of patients were unable to access their recommended therapies. Out of 89 patients who had clinically significant genetic alterations and received recommended therapies based on NGS results, 17 patients (19.1%) could not continue with the personalized treatment. This was mainly due to systemic barriers, as eight patients could not cover the high cost of targeted therapies, and nine patients could not access the drugs due to unavailability in Jordan.
Furthermore, the study shows that there is no significant influence of age and gender on PFS values in our sample population, suggesting that NGS-guided treatment benefits are broadly applicable across demographic groups.
These findings show the challenges with implementing NGS-guided personalized treatment in low-middle-income countries. NGS testing alone costs 2819.72 USD (2000 Jordanian Dinars) per test in Jordan, which is more than half of the GDP per capita (4482 USD in 2023), 19 and that combined with the high cost of therapies often not covered by insurance poses a very high price for patients. Likewise, the unavailability of certain medications in Jordan suggests a gap in the distribution of essential medications for personalized cancer treatment in the region.
These obstacles should be addressed at a national level in Jordan using targeted policy interventions. Pharmaceutical companies could offer low-cost or no-cost testing for patients who may become eligible for their targeted therapies due to NGS findings. While that could solve one aspect of the problem, it is also crucial to support the medical institutions carrying out these tests. This could be done by allocating training grants and funding to hospitals and diagnostic laboratories to help them in acquiring and operating NGS equipment locally, which would ultimately lower testing costs. Lastly, insurance companies are encouraged to cover NGS panels for indicated patients.
One of the primary limitations of this study is its retrospective nature, which may introduce selection bias and limit the generalizability of the findings. Given this, confounding factors that may affect results, such as median household income and comorbidities, were not available during data collection. In addition, potential bias is introduced in the selection of patients who can afford the high cost of NGS testing and personalized treatment, which is not the case for most of the Jordanian population. Also, the study population included patients with late-stage disease, but at different lines of therapies; in fact, most patients agreed to do the NGS test only when they had exhausted their standard-of-care therapies.
Conclusion
There is a substantial clinical benefit to using NGS in personalized treatment for late-stage cancer patients. The findings show improved PFS for patients receiving NGS-guided therapies compared to conventional treatments.
While the potential of NGS testing in personalized medicine is evident, there are various challenges that face the incorporation of such technology into clinical practice in low-middle-income countries such as Jordan, such as high testing and treatment costs and drug unavailability.
The study emphasizes the urgent need for policy interventions to improve access to NGS testing and associated recommended therapies.
Addressing cost-related issues and improving the availability of targeted treatments could significantly enhance patient care. Finally, future research with larger populations is essential to confirm these findings and extend the benefits of NGS-guided treatment to broader populations.
Supplemental Material
sj-docx-1-tam-10.1177_17588359251337271 – Supplemental material for Next-generation sequencing in the management of cancer: an evaluation of the clinical effectiveness in Jordan
Supplemental material, sj-docx-1-tam-10.1177_17588359251337271 for Next-generation sequencing in the management of cancer: an evaluation of the clinical effectiveness in Jordan by Yousef Abbasi, Omar Altamimi, Majdi Al-Shehab, Hamza Nabulsi and Salah Abbasi in Therapeutic Advances in Medical Oncology
Footnotes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
