Abstract
Background:
Conventional treatments for follicular thyroid cancer (FTC) can be ineffective, leading to poor prognosis. The aim of this study was to identify mutations associated with FTC that would serve as novel molecular markers of the disease and its outcome and could potentially identify new therapeutic targets.
Methods:
FLT3 mutations were first detected in a 29-year-old White female diagnosed with metastasized, treatment-refractory FTC. Analyses of FLT3 mutational status through next-generation sequencing of formalin-fixed, paraffin-embedded FTC specimens were subsequently performed in 35 randomly selected patients diagnosed with FTC.
Results:
FLT3 mutations were found in 69% of patients. FLT3 mutation-positive patients were significantly older than those that were FLT3 mutation-negative [median age at diagnosis 54 (36–82) versus 45 (27–58) (p = 0.023)]. Patients over 60 years were 23 times more likely to be FLT3 mutation-positive (p = 0.006). However, the number of FLT3 mutations did not correlate with age (r-Pearson: –0.244, p-value: 0.25). A total of 26 mutations were identified in the FLT3 gene with 2–16 FLT3 mutations in each FLT3 mutation-positive patient (mean: 5.6 mutations/patient). Tyrosine kinase domain (TKD) mutations in the FLT3 gene were detected in 58% of FLT3 mutation-positive patients. All FLT3 mutation-positive patients with a disease stage of pT2N1 or worse harbored at least one mutation in the TKD of FLT3.
Conclusions:
There is a wide spectrum and high frequency of FLT3 mutations in FTC. The precise role of FLT3 mutations in the genesis of FTC, as well as its potential role as a therapeutic target, requires further investigation.
Keywords
Introduction
Patients with differentiated thyroid cancer (DTC) may present with unfavorable prognostic factors. The five-year cancer-specific survival for patients presenting with distant metastases is only 40%1,2 as these patients frequently become refractory to radioiodine3,4 despite the advent of tyrosine kinase inhibitors (TKI). 5
To date, DTC risk stratification based on clinicopathological tumor characteristics remains an imperfect science. Recent research has focused on the identification of genetic markers, including BRAF and RAS mutations, as well as RET/PTC and PAX8/PPAR gamma rearrangements to improve risk stratification.2,5
New data are emerging for DTC from the use of genome-analyzing methods such as high-throughput next-generation sequencing (NGS), 6 yielding a so-called molecular signature 7 that may in the future become a clinically valuable tool for diagnosis, risk stratification, and personalized therapy.5,8 Furthermore, the definition of the spectrum and patterns of driver gene mutations in DTC may lead to the identification of targets for new treatments.
Although the Cancer Genome Atlas consortium has delivered new data on the genetic background of the papillary subvariety of DTC, 7 follicular thyroid cancer (FTC) has not yet been addressed. 9 The aim of the present study was to establish a mutational profile of FTC and to identify potential new mutations associated with FTC employing high-throughput NGS technology.
Material and methods
Ethical approval
The study was approved by the Bioethical Committee of the Poznań University of Medical Sciences (an approval no. 1061/15 from January 2015) and was conducted in accordance with the Declaration of Helsinki. Before surgery, patients gave generalized written informed consent for the use of their materials for scientific purposes and therefore the need for a specific written informed consent pertaining to the present study only was waived by the bioethical committee.
Study background
NGS sequencing was conducted using an Ion Personal Genome Machine Sequencer (Thermo Fisher Scientific; Indianapolis, USA) employing the Ion AmpliSeq Comprehensive Cancer Panel. The data obtained from genomic experiments were subjected to analysis using the dedicated software Variant Caller v5.2.1.38 (Thermo Fisher Scientific; Indianapolis, USA) and MutationTaster2 (Berlin Institute of Health, Berlin, Germany).
Patient characteristics
Index patient
The study was initiated based on clinical data obtained from an index patient, in whom the presence of mutations of the hot spot class III receptor fms-like tyrosine kinase-3 (FLT3; fetal liver kinase-2; human stem cell tyrosine kinase-1; CD135) was identified. This concerned a 29-year-old female patient diagnosed with FTC and metastases to the lymph nodes and lungs, who required two subsequent lymphadenectomies followed by three courses of radioiodine therapy (2009–2011). Due to unsatisfactory results, the patient was subsequently treated with lenvatinib from 2012–2015 and achieved total structural and partial biochemical remission. However, after 3 years, locoregional disease recurrence was found, and the drug was withdrawn. The patient was then re-operated on and received the last radioiodine therapy in 2015 and has been treated with sorafenib since 2016. Complete remission was not yet achieved at the last follow-up in June 2018.
In this index patient, DNA was isolated from a formalin-fixed, paraffin-embedded (FFPE) FTC specimen acquired at thyroidectomy in 2006. The specimen was re-reviewed to confirm the diagnosis and to indicate the most appropriate part for DNA sample collection.
Data collection
We analyzed 35 randomly selected patients [computer-aided simple random sampling (generated at stattrek.com)] at the tertiary care department of endocrinology at our university hospital, who were diagnosed with FTC according to the revised World Health Organization criteria from 2004, 10 between the years 2006 and 2018. Surgically obtained FTC specimens were subjected to further molecular analyses. The group consisted of 31 women and four men, all White, with a median age at diagnosis of 52 years (range 27–82 years). Only patients not already receiving treatment, which might affect the molecular signature of thyroid cancer at the time of diagnosis, nor suffering from any other endocrine disorders or cancer, were enrolled in the study. The single exclusion criterion was a patient’s medical records being incomplete.
Follow-up data on patients were collected up to July 2018.
Molecular studies and clinical analysis
Assessment of FLT3 mutational positivity was conducted with NGS sequencing. All specimens were reassessed by two independent board-certified pathologists to confirm the diagnosis and to indicate the most appropriate part for DNA sample collection, meaning those which most fulfilled FTC criteria such as vascular or capsular invasion. DNA was extracted with the High Pure FFPET DNA Isolation Kit (Roche Life Science, Indianapolis, USA) from FFPE FTC specimens obtained during thyroidectomy. 11 The Next Generation Sequencing Library was constructed with the Ion AmpliSeq™ Library Kit 2.0. (Life Technologies, Darmstadt, Germany). The template was prepared on the Ion OneTouch system (Thermo Fisher Scientific; Indianapolis, USA). Finally, DNA high-throughput sequencing was performed on the Ion PGM Sequencer (Thermo Fisher Scientific, Indianapolis, USA) with the Ion PGM™ Hi-Q™ Sequencing Kit (Life Technologies, Darmstadt, Germany) on the Ion 318™ sequencing chip (Life Technologies, Darmstadt, Germany).
An NGS Ion AmpliSeq Cancer Hotspot Panel v2 (Thermo Fisher Scientific, Indianapolis, USA) was used to study the coding regions and intronic flanking regions of the FLT3 gene. The analysis of other genes included in NGS Ion AmpliSeq Cancer Hotspot Panel v2, concerning the amount of data and the clarity of the presentation of results, is the subject of a separate report (submitted). Based on the costs, capacity, and analytical experience, in accordance with the guidelines for a comprehensive assessment of somatic mutation detection in cancer using whole-genome sequencing, 12 we determined a minimum of 30 × depth of sequencing coverage of each the tumor and normal genomes with paired reads on the order of 100–250 bp in length to identify tumor-specific somatic mutations.12,13
Data obtained were subjected to further analysis using the Variant caller v5.2.1.38 software (Thermo Fisher Scientific; Indianapolis, USA) and the MutationTaster2 algorithm. 14 Furthermore, to analyze a putative function of FLT3 mutations as driver mutations, four separate programs were employed: SIFT, 15 PolyPhen-2 (Polymorphism Phenotyping v2), 16 and MutationTaster2, 14 as well as FATHMM (Functional Analysis through Hidden Markov Models v2.3), which result in an index, calculated with a high-throughput web-server, which is able to predict the functional consequences of both coding variants, that is, nonsynonymous single nucleotide variants, and noncoding variants to distinguish between cancer-promoting/driver mutations and other germline polymorphisms.
The Catalogue of Somatic Mutations in Cancer (COSMIC), the dbSNP, 1000 Genome Project, ClinVar, and ExAC databases were checked for existing reports of the presence of particular FLT3 mutations. For the validation of mutations, we selected tissues bearing the most common mutations and performed a Sanger sequencing (Thermo Fisher Scientific; Indianapolis, USA). 17 Selected tissues were showing at least 20% tumor cell content to ensure a minimal threshold level for sensitivity using the di-deoxy method. The amplifications of selected targets were conducted using Clontech Titanium™ DNA Amplification Kit (Takara Bio Inc., Japan) and processed through 35 cycles (30s at 94°C, 30 s at 60°C, and 45s at 72°C). Primers were designed using the Primer3 Design Tool.18,19 The primers were as follows: exon 4 forward 5′- tctgaggggctgctaagag -3′, exon 4 reverse 5′- cgaaatgctgggattacagg -3′, exon 10 forward 5′- ccaccttggcttcacaaag -3′, exon 10 reverse 5′- aggatttctgcagcgagttc -3′, exon 13 forward 5′- tttccaaaagcacctgatcc -3′, exon 13 reverse 5′- ccttccctttcatccaagac -3′, exon 19 forward 5′- ccatcaccggtacctcctac -3′, exon 19 reverse 5′- ccctgaagctgcagaaaaac-3′. Capillary sequencing was conducted with the use of BigDye chemistry version 3.1, on an ABI 3130xl DNA Analyzer according to manufacturer instructions (Applied Biosystems, Foster City, CA). Sequence traces from tumor DNA samples were aligned to the genomic reference sequence of FLT3 gene (NG_007066_1) and analyzed using CodonCode Aligner software version 4.0.4 (http://www.codoncode.com).
Clinical data
Clinical records and results of histopathological examinations of the resected specimen following thyroidectomy were analyzed for all patients.
The following data were recorded from patient files: gender, age at diagnosis, tumor size, multifocality, capsular invasion without extra-thyroidal extension, presence of histopathological signs of chronic lymphocytic thyroiditis (presence of diffuse lymphocytic infiltrate, oxyphilic cells, and the formation of primary and secondary lymphoid follicles 20 ), and histopathological staging.
Tumors were classified as multifocal when two or more foci were found. In the case of multifocality, the size of a tumor was staged by the size of the largest focus. Staging procedures were performed according to the joint Union Contre le Cancer and American Joint Committee on Cancer Tumor, Node Metastasis (TNM) staging system (7th Edition). 21
For the present study, patients were defined as radioactive iodine (RAI) refractory if a cumulative activity of 22.2 GBq/600 mCi of RAI therapy were given without achieving complete remission.
Statistical analysis
The parameters were recorded and entered into a dedicated database. Descriptive analysis was used to summarize the collected data. To determine the normality of continuous variables, data were tested by the D’Agostino and Pearson omnibus normality test. Variables that were found to be normally distributed were expressed as means with respective standard deviations. Non-normal data were expressed as median and minimum-maximum values.
To compare differences between groups, the chi-square test or Fisher’s exact test (2 × 2 contingency table) were used as appropriate for categorical variables. Interval data were compared with the use of the Mann–Whitney U test as the data did not follow a normal distribution. Odds ratio (OR) and the 95% confidence interval (95% CI) were calculated using the group of FLT3 mutation-negative patients as the reference population. Correlation between the number of mutations in a single patient and their age was assessed with Pearson’s r correlation test.
A p-value <0.05 was regarded as significant. Statistical analyses were performed with StatSoft Statistica v10.0 and Analyse-It for Microsoft Excel v3.53.
Data availability
The datasets were submitted to the publicly accessible Catalogue of Somatic Mutations in Cancer (COSMIC) database.
Results
Index patient
The analysis revealed the presence of an FLT3 (fetal liver kinase-2; human stem cell) mutation on chromosome 13 – a heterozygous mutation c.1683A>G (p.L561L, COSM19740) in the FLT3 gene.
The mutation was not detected in the patient’s leucocyte DNA through NGS, confirming its somatic character. This particular mutation has not been detected in any other FTC specimen. Extension of the analysis using bioinformatic tools including SIFT, Polyphen-2, and MutationTaster2, 14 found further FLT3 mutations as described in Table 1. Fifteen further mutations were identified as potentially disease-causing, (i.e. driver), mutations. All have been previously published in the COSMIC database in acute myeloid leukemia (AML), although have not been previously reported in FTC. A total of 60% have been identified in acute lymphocytic leukemia (ALL) and a minority in myelodysplastic syndrome (MDS), lung adenocarcinoma, and gastric cancer. Two mutations may be sufficient to cause resistance to sorafenib (FLT3 D835H and D835Y), 22 quizartinib (D835V and D835Y), 23 and even one-to sunitinib (D835Y).24,25
The list of FLT3 mutations (Ensembl transcript ID: ENST0000241453 and Genbank transcript ID: NM_004119, UniProt number P36888) predicted as “disease-causing” by MutationTaster algorithm in CDS (alteration region).

AAE, amino acid exchange; DNA, Deoxyribonucleic acid; dbSNP, The Single Nucleotide Polymorphism Database; rs, reference SNP identifying number; COSMIC, The Catalogue of Somatic Mutations in Cancer; ID, identifier; FATHMM, Functional Analysis through Hidden Markov Models v2.3; AML, acute myeloid leukemia; ALL, acute lymphocytic leukemia; MDS, myelodysplastic syndrome; B ALL, B-cell acute lymphocytic leukemia
Study patients
FLT3 mutational status in the larger group of patients diagnosed with FTC enabled discrimination between FLT3 wild type patients (n = 11; 31%), without detectable mutation in the FLT3 gene, considered as ‘FLT3 negative’, and patients with at least one mutation in the FLT3 gene, considered ‘FLT3 positive’ (n = 24; 69%).
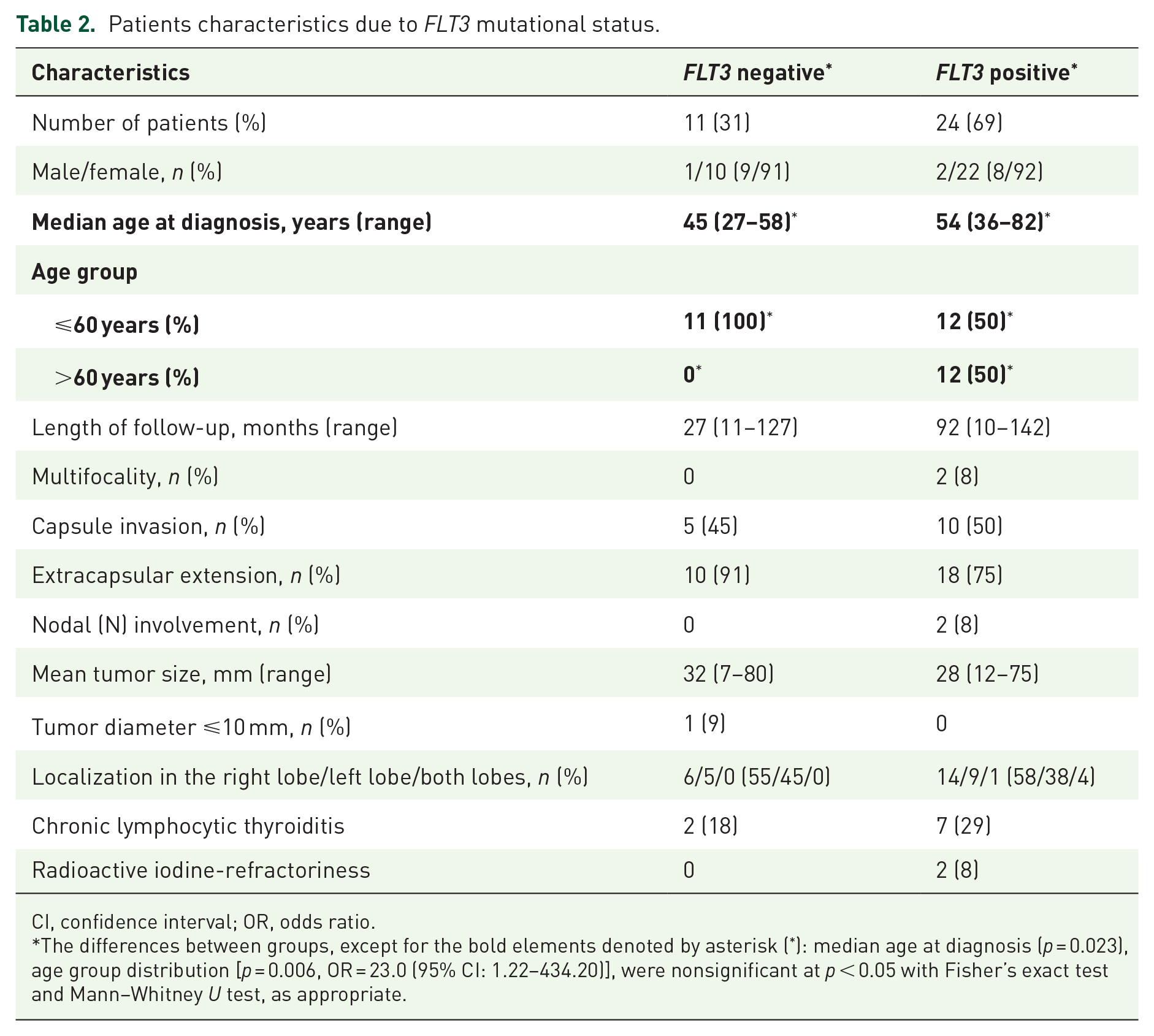
The demographic and clinical features of both groups are presented in Table 2. FLT3-positive patients were found to be significantly older than FLT3-negative patients with a median age at diagnosis of 54 (36–82) versus 45 (27–58) years (p = 0.023). Patients over 60 years were 23 times more likely to be FLT3 positive [p = 0.006, OR = 23.0 (95% CI: 1.22–434.20)]. There was no significant difference with regard to the distribution of sex, multifocality, capsule invasion, extracapsular extension, nodal involvement, mean tumor size, localization, or the presence of chronic lymphocytic thyroiditis.
Patients characteristics due to FLT3 mutational status.
CI, confidence interval; OR, odds ratio.
The differences between groups, except for the bold elements denoted by asterisk (*): median age at diagnosis (p = 0.023), age group distribution [p = 0.006, OR = 23.0 (95% CI: 1.22–434.20)], were nonsignificant at p < 0.05 with Fisher’s exact test and Mann–Whitney U test, as appropriate.
A total of 26 FLT3 gene mutations were identified with coverage of >30× (Table 3). The mean sequencing coverage of the target region across all samples was 60-fold. The most common mutations were single base exchanges in the form of missense substitutions. The most frequent mutations were V592A and Y599F in 10 patients, and G831E, R834Q, D835H, D835N, D835Y in nine patients (the last three mutations localized in the tyrosine kinase domain). In total, 2–16 FLT3 mutations were found in every FLT3-positive patient, with a mean of 5.6 mutations per patient, and protein kinase domain mutations were identified in the FLT3 gene in 14 patients (58%).
The list of FLT3 mutations with Ensembl transcript ID: ENST0000241453 and Genbank transcript ID: NM_004119 in peptide with UniProt number P36888 predicted as “disease-causing” by MutationTaster algorithm in CDS (alteration region).

AAE, amino acid exchange; DNA, Deoxyribonucleic acid; dbSNP, The Single Nucleotide Polymorphism Database; rs, reference SNP identifying number; COSMIC, The Catalogue of Somatic Mutations in Cancer; ID, identifier; FATHMM, Functional Analysis through Hidden Markov Models v2.3; AML, acute myeloid leukemia; ALL, acute lymphocytic leukemia; MDS, myelodysplastic syndrome; B ALL, B-cell acute lymphocytic leukemia; T ALL, T-cell acute lymphocytic leukemia
Quantification of the proportion of variant reads for a given mutation was made with variant allele frequency (VAF), which represents the percentage of tumor cells that harbor a specific mutation. The significantly highest mean VAF were observed for S451F (80.21%) and V592I (50.91%). The mean VAF for other mutations was below 20%.
The number of FLT3 mutations in a single patient did not correlate with age (r-Pearson: –0.244, p-value: 0.25).
The frequency of occurrence of FLT3 mutations in the tyrosine kinase domain was significantly higher in patients with a higher TNM stage, as all FLT3-positive patients with disease staging of pT2N1 and worse harbored at least one mutation in the tyrosine kinase domain (Figure 1). Two FLT3-positive patients were considered refractory to radioactive iodine; in contrast, this was not observed in any FLT3-negative patient.

FLT3 mutational spectrum observed in follicular thyroid cancers – distribution of acquired mutations in conjunction with the aggregation of the mutations into mutational categories: mutant (black), wild type (gray).
Discussion
This study demonstrates that the occurrence of FLT3 mutations in patients with FTC is common, as 69% percent of those patients examined harbored at least one mutation in the FLT3 gene. This variant has already been reported for the first time in FTC by our research team, 26 although had been previously known in AML, MDS, and hemangioblastoma.27–31
Data concerning the presence of FLT3 mutations in DTC cells are scarce. However, the application of novel techniques of genomic screening may enable detection of FLT3 mutations in a histopathological specimen from patients with DTC. In a recent study by Gerber and colleagues targeted NGS was employed to detect genetic mutations in poorly differentiated thyroid cancer. 32 Analysis of 25 samples revealed the presence of the mutated FLT3 gene in 16% of the analyzed specimens (in four patients). The reported mutations were of the missense type (G>A substitution in positions 1734, 1969, 2455, 2503, and 2515 and C>A substitution in position 2519 of cDNA). All detected mutations were previously reported in the COSMIC database as present in the hematopoietic and lymphoid tissue. Only substitutions in position 2503 of cDNA were common in our group of FTC patients and those reported by Gerber and colleagues.
In this study, a total of 26 mutations were found in the FLT3 gene, of which some were previously described in the dbSNP or ExAC databases. All mutations had a somatic nature and were predicted to be disease-causing by Polyphen-2 and MutationTaster2, and oncogenic. The majority of FLT3-positive patients (58%) harbor mutations in the FLT3 kinase domain (Figure 1). The primary tumor may exhibit significant clonal heterogeneity, and this diversity may be affected during disease progression. Cellular dynamics of tumor evolution and heterotypic interactions between tumor subpopulations may still be possible. All these factors may, of course, make targeting FTC even more difficult, but maybe still possible, as the majority of the mutations is seen in the tyrosine kinase domain.
To discuss clonal composition and the character of the mutations we used a concept of analyzing VAF for the mutations. As has already been proven in hematological malignancies, the allelic burden of somatic mutations may be successfully incorporated even into clinical practice for the diagnosis and prognosis of neoplasms. 33 According to the VAF, we could hypothesize on the role of the mutations, especially in those cases with more than one alteration: the mutation with the greatest VAF would likely be the driver mutation.34,35 We found that two mutations had a particular role in a possible natural history of FTC in our patients. S451F (localized in the extracellular domain) and V592I (in juxtamembrane domain) may be considered to be two oncogenic and driver mutations. Patients harbored one or both of these mutations. Analyzing their VAF, we may assume that their origin may be subclonal. A new finding of the possible particular role of those two mutations may be even more significant, taking into account their role as targets in the action of TKI, such as midostaurin. Further investigation with functional analysis is required to determine the functional role of all FLT3 mutations detected in this cohort of patients.
Although an association between FLT3 mutations and malignant disease, particularly in hematological malignancies, has been established, these findings are new for FTC. FLT3 is known to be involved in the regulation of proliferation, differentiation, and survival of hematopoietic and dendritic cells. 36 Its occurrence is linked mainly with AML patients, as about 30% harbor the FLT3 mutation, in which it is associated with reduced overall survival and increased risk of relapse compared with FLT3-negative AML patients. 37 FLT3 was reported to be the single most commonly mutated gene in AML by Yamamato and colleagues in 2001. 38
Among the mutations within an activation loop, the most commonly found ones in our group of FTC patients concerned asparagine residues (Asp835) mutations, similarly to AML, 39 MDS, and ALL patients. As is the case of FLT3-internal tandem duplication (ITD), they spontaneously activate tyrosine kinase activity and induce factor-independent proliferation of hematopoietic cell lines. 38 Although the role of FLT3 mutations in the transformation of myeloid cells is not yet known, it is highly probable that they provide a strong mitogenic and antiapoptotic signal. 40 Thus, these mutations may act synergistically with mutations blocking differentiation to induce AML and hypothetically, by analogy, FTC as well.
Not only kinase-activating mutations but also those located outside the juxtamembrane and kinase domains of FLT3, resulting in activated downstream signaling and conferred sensitivity to FLT3 inhibition, may reduce kinase activity and promote cancer induction.39,41 Among those were one of the most common point mutations in FLT3-positive patients, R834Q, which is expected to destabilize the autoinhibited kinase conformation and G831E that may stabilize FLT3 in the autoinhibited conformation and result in reduced kinase activity. The effect on cell function has also been described for the S451F (localized in the extracellular domain), Y572C and V592G (in the juxtamembrane domain), and R834Q (in the activation loop) mutations of FLT3. These results suggest that there are not only differences in signal transduction between various FLT3 mutation classes, that is, ITDs and activation loop mutations,42,43 but also between distinct alleles within a given mutation class, which may affect specific signaling pathways differently. 39
This study indicates that neoplasms of different primary sites may also share a common genetic background, as has been found in other studies of various forms of cancer 44 and introduces the potential for a novel, genetic-based rather than histology-based cancer classification as a basis for targeted treatment.
The number of patients studied is insufficient to establish whether the inclusion of FLT3 mutations may be used in the classification of FTC. However, the high rate of FLT3 mutations found indicates that FLT3 mutations are highly likely to play a role in the genesis of FTC. Further study, including an investigation into whether FLT3 positivity is associated with RAI-refractory disease, is required in a larger cohort.
Although precautions were taken to minimize potential bias, limitations in the study include the sample size (35 FTC patients) although this is mitigated by the precise inclusion criteria, which increases the reliability of the results at the cost of reducing the study sample size. Genotyping of the selected mutations is required in more FTC patients before making firm conclusions. To simplify the interpretation of NGS results the description of the genetic findings were focused on FLT3 mutations. However, Cancer Hotspot Panel v2, used for genetic analyses, included approximately 2800 COSMIC mutations from 50 oncogenes and tumor suppressor genes. Future work will include an analysis of other genetic mutations. Finally, the somatic type of the mutations has not been confirmed by NGS of peripheral DNA apart from the analysis in the first FLT3 patient treated with a TKI. However, the familial occurrence of the disease was taken into account for all patients to exclude evidence of a mutation germline type.
For confirmation of the mutations, we used capillary Sanger sequencing in selected samples (Figure 2). The method, however, has its limitations due to restricted sensitivity and its inability to perform parallel analysis of multiple targets, particularly in the case of cancer specimens. The limited sensitivity of Sanger sequencing of around 20% does not allow the investigation of sequence alterations detected by NGS at lower allele fractions. According to numerous recent reports, NGS platforms used in clinical and research settings are more accurate than Sanger sequencing.45–47

Sanger sequencing of commonly mutated codons of FLT3 gene in patients with follicular thyroid cancer.
An association between constitutive FLT3 activation and disease prognosis makes this protein an attractive target for therapeutic intervention. 48 The most common types of FLT3 mutations are ITD in the juxtamembrane domain and point mutations within the activation loop of FLT3,49,50 leading to increased FLT3 activation. 40 Furthermore, the study demonstrated that all FLT3-positive patients harbored at least one mutation in the FLT3 tyrosine kinase domain. Currently, there are numerous selective FLT3 inhibitors at various stages of clinical development, including sorafenib, lestaurtinib, tandutinib, quizartinib, sunitinib, midostaurin, gilteritinib, crenolanib, cabozantinib, Sel24-B489, G-749, AMG 925, TTT-3002, and FF-10101.51,52 These compounds are well tolerated at doses that achieve FLT3 inhibition and have shown moderate activity in AML patients with activating FLT3 mutations, warranting an investigation into their potential applications to the treatment of radioiodine-refractory, FLT3-positive FTC patients. Midostaurin and gilteritinib have been approved by FDA for treatment of AML with FLT3 mutations. 52
The potential association between FLT3 mutations and the occurrence of RAI-refractory disease and metastases in older FTC patients may be useful for prognostic stratification and individualization of treatment.
This study demonstrates the potential of NGS to inform the development of individualization of therapy in patients with metastatic, iodine-refractory DTC. The index patient with metastatic, refractory FTC harbored FLT3 D835H and FLT3 D835Y mutations, which caused resistance to sorafenib22,24 and may explain why the second line TKI treatment has not so far resulted in remission of the disease.
Conclusion
There is a wide spectrum and high frequency of FLT3 mutations in thyroid malignancy. The precise role of FLT3 mutations in the genesis of FTC, as well as its potential role as a therapeutic target for individualized therapy with specific kinase inhibitors warrants further investigation.
Footnotes
Acknowledgements
The authors would also like to thank Prof. Glenn Flux for English editing and his important support. We are grateful to all patients who agreed to take part in our study.
Conflict of interest statement
The authors declare that there is no conflict of interest.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was funded by PRELUDIUM Grant of Polish National Center for Science (grant number 2015/19/N/NZ5/02257 granted to MB), whom we would like to thank.
