Abstract
Background:
Ferroptosis plays a vital role in cancer development and treatment. The relationship between ferroptosis-related genes and breast cancer prognosis, as well as immunotherapy outcomes, remains unknown.
Objectives:
To evaluate the prognostic value of ferroptosis-related genes in breast cancer.
Methods:
We conducted differential expressions and prognostic analysis for ferroptosis-related genes on public databases and breast cancer patients in our center and analyzed their predictive value for immunotherapy of breast cancer patients.
Results:
We identified prognostic ferroptosis-related genes, constructed a nomogram, and validated key genes using patient data from our center. We also investigated ferroptosis-related genes significantly associated with immune infiltration and identified FTH1 as a promising biomarker for triple-negative breast cancer immunotherapy.
Conclusion:
Ferroptosis-related genes had potential prognostic value and predictive value for breast cancer immunotherapy.
Introduction
Female breast cancer is the most common cancer type worldwide, causing 2.3 million new cases and 690 thousand deaths each year. 1 Triple-negative breast cancer (TNBC) is defined as the absence of estrogen receptor, progesterone receptor, and human epidermal growth factor receptor-2, which comprises 15%–20% of all cases. 2 TNBC shows a more malignant biological behavior, higher recurrence rate, fewer treatment options, and worse prognosis compared to other subtypes.3,4 Due to few therapy options and relatively short overall survival (OS), more accurate prognostic factors and treatment options are warranted.
Ferroptosis, a novel form of regulated cell death driven by excessive lipid peroxidation, has been reported to reflect multiple pathways in cancer pathogenesis and development.1,5 Experimental molecules such as erastin and approved drugs such as sulfasalazine (for ulcerative colitis) could induce ferroptosis through inhibiting system XC-L-glutathione reduced-glutathione peroxidase 4 (GPX4) pathway. 6 A more recent study found that poly (ADP-ribose) polymerases inhibitor induces ferroptosis in ovarian cancer through inhibition of SLC7A11 (xCT) in a P53-dependent manner. 7 Furthermore, anti-programmed death ligand 1 (PD-L1) antibodies were reported to promote lipid peroxidation-dependent ferroptosis in tumor cells and had synergistic actions in tumor inhibition when combined with ferroptosis activators such as erastin. 8 However, few studies explored the prognostic and immunotherapy predictive value of ferroptosis-related genes in breast cancer patients.
Although chemotherapy remains the cornerstone in metastatic TNBC (mTNBC), immunotherapy has recently shown exciting results. A phase III IMpassion130 study enrolled 902 metastatic or inoperable TNBC patients and randomized them into atezolizumab or placebo plus nab-paclitaxel. The results showed significantly improved progression-free survival (PFS) in the atezolizumab group. 9 However, the phase III IMpassion131 study showed that atezolizumab combined with paclitaxel did not significantly improve the survival outcome in PD-L1-positive patients. 10 The KEYNOTE-355 study compared pembrolizumab or placebo plus chemotherapy in first-line treatment of mTNBC and indicated prolonged PFS in PD-L1-positive patients or patients with a combined positive score of greater than 10. 11 Although immunotherapy provides a great opportunity for patients, controversial results in different studies urge us to find more efficient biomarkers and prognostic factors to identify proper candidates for immunotherapy.
Thus, in this study, we explored the differential expression, prognostic value, and immunotherapy prediction of ferroptosis-related genes in breast cancer patients both in public databases and in our center. These findings could deepen our understanding of ferroptosis in breast cancer and uncover novel biomarkers for immunotherapy of breast cancer.
Methods
Gene expression, prognostic effect, and functional analysis in a public database
The mRNA expression data of normal and tumor tissues, as well as clinical outcomes of patients with breast cancer, were obtained from the GEO database: GSE93601 12 (https://www.ncbi.nlm.nih.gov/geo) and The Cancer Genome Atlas (TCGA) database: TCGA-BRCA (https://www.cancer.gov/tcga/). All data were normalized via log2-scale transformation to ensure standardization. Differential expression was analyzed by the “limma” package in R (version 4.0.2). Survival analysis was performed by “survival” package. Univariate and multivariate Cox regression models were used to identify the survival-related ferroptosis-related genes. A total of 149 ferroptosis-related genes, including drivers, suppressors, and markers, were obtained from the FerrDb database (http://www.zhounan.org/ferrdb/). 13 Functional analysis and network production were performed on Metascape Online (https://metascape.org/gp/index.html#/main/step1). 14
Nomogram construction
A nomogram was used to collect the prognostic factors and visualize the risk model for the calculation of the probability of 1, 3, and 5-year OS. 15 The calibration curve was mapped to evaluate the consistency between predicted and observed probability. The R packages “rms” and “survival” were used in the construction of the nomogram and calibration curves. In this nomogram, we could simply calculate the OS probability by adding up scores of each variable and locating this sum on the line of “total points.” Bootstrap self-sampling method and graphic calibration method were used to validate our model.
Human breast cancer tissue specimens
A set of tissue microarrays (TMAs) including 258 breast cancer patients was obtained from their first radical surgery with no prior systemic treatment from 2010 to 2020.
Patient data were collected from an electronic medical records system. Resected specimens were macroscopically examined to confirm the location and size of a tumor. For histology use, the specimens were fixed in 10% (v/v) formalin and processed for paraffin embedding. The Ethics Committee and Institutional Review Boards at Fudan University Shanghai Cancer Center approved this study for clinical investigation. All of the patients signed written informed consent forms before the study.
Immunohistochemistry
The primary antibodies SLC7A11/xCT (26864-1-AP; Proteintech, Wuhan, China, 1:200), ALOX12 (ab211506; Abcam, Cambridge UK, 1:50), GPX4 (67763-1-Ig; Proteintech, 1:100), and ACSL4 (22401-1-AP; Proteintech, 1:150) were used and incubated at 4°C overnight. TMA was scanned by Nikon E100 and processed through Nikon DS-U3 (Nikon, Tokyo, Japan) according to the manufacturer’s instructions.
Staining score was evaluated by stain intensity and the percent of positive cells, each patient was scored 0, 1, 2, or 3. Patients with 0 or 1 score were defined as low expression and 2 or 3 as high expression.
Immune infiltration analysis
RNAseq data were extracted from the TCGA-BRCA database and a correlation analysis was performed between FTH1 and immune infiltration matrix data. Using the ssGSEA algorithm provided in the R package GSVA1, the immune infiltration of 24 types of immune cells based on cloud data was performed. 16 Alternatively, the R package ESTIMATE was used to calculate the corresponding stromal and immune scores. 17 The analysis results were visualized using lollipop plots or violin plots with the ggplot2 package.
Immunotherapy cohort
A total of 12 patients with histologically confirmed mTNBC disease who used immunotherapy (PD-1/PD-L1 antibody)-based treatment as first-line treatment at Fudan University Shanghai Cancer Center were enrolled in our study.
Immunofluorescence
The immunofluorescence method is basically performed as before. 18 Briefly, after deparaffinization and antigen retrieval and blocking, incubate it overnight in primary antibody and 40 min in secondary antibody, washing, and TSA treatment before adding second primary antibody and second secondary antibody. Then stain it with DAPI before imaging.
Statistical analysis
The study results are presented as mean ± standard deviation for three independent experiments. Student’s t-test was used for comparison between groups. OS and recurrence-free survival (RFS) curves were conducted using Kaplan–Meier analyses and compared using the log-rank test. All statistical analyses were performed using the GraphPad Prism® 8.0 software (GraphPad Software, Boston, USA), R software (version 4.0.2), or SPSS (version 23.0, IBM, Chicago, USA). p < 0.05 (two-sided) was considered statistically significant. The reporting of this study conforms to the TRIPOD statement 19 (Supplemental File 3).
Results
Differentially expressed ferroptosis-related genes in breast cancer
We first screened differential genes of cancer tissue and normal tissue from both the TCGA-BRCA database and the GEO: GSE93601 database. As a result, 1265 dysregulated genes in the GEO database and 4966 dysregulated genes in the TCGA database were found (Figure 1(a)). We then intersected these dysregulated genes with ferroptosis-related genes obtained from the FerrDb database (Table S1) and showed them with a Venn diagram (Figure 1(b)). A total of 96 differentially expressed ferroptosis-related genes were identified (Table S2).

Differentially expressed ferroptosis-related genes in breast cancer. (a) Expression profiles of genes in normal and tumor samples in GSE93601. Genes were ranked according to their expression. The red color and blue color represent high and low expression. (b) Venn diagram of the differentially expressed genes and ferroptosis-related genes. (c, d) GO and KEGG analysis based on the Metascape Online. (e) Direct expression of differential genes using the TCGA-BRCA database.
To explore the relevant pathways of ferroptosis signatures in breast cancer, we performed Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis using Metascape Online (Figure 1(c) and (d)). The results of GO analysis showed that these differential genes were enriched in ferroptosis, response to stress, and nutrient levels. Similarly, the KEGG analysis revealed pathway enrichment in ferroptosis, response to stimuli, and nutrient levels. Furthermore, we explored the expression levels of key ferroptosis-related genes in the TCGA-BRCA database and found most ferroptosis suppressors (e.g., SLC7A11, FTH1) are upregulated in breast cancer tissue compared to normal tissue, while most ferroptosis promoters (e.g., ACSL4, ALOX12) are downregulated (Figure 1(e)). These findings indicated that ferroptosis might play an important role in breast cancer.
Prognostic value of ferroptosis-related genes in breast cancer
To further assess the prognostic value of ferroptosis-related genes in breast cancer patients, we conducted Cox regression analysis and showed in Kaplan–Meier method using mRNA expression data and patients’ survival information in the TCGA database. We found that six typical ferroptosis-related genes were associated with breast cancer prognosis: SLC7A11, TFRC, FTH1, ALOX12, HSBP1, and CBS (Figure 2(a)). Multivariate Cox regression revealed ALOX12 as an independent prognostic factor (adjusted hazard ratio (HR) 0.671, 95% confidence interval (CI) 0.486–0.927, p = 0.015). Moreover, since TNBC showed more severe biological behavior and poorer outcomes, we analyzed these genes in TNBC separately. The results confirmed the prognostic value of ferroptosis-related genes in TNBC patients (Figure 2(b)).

Ferroptosis-related genes predict the overall survival of breast cancer. (a) Kaplan–Meier plots show the ferroptosis-related genes with prognostic value in all breast cancer patients. (b) Kaplan–Meier plots show the ferroptosis-related genes with prognostic value in TNBC patients.
Construction of nomogram
Since multivariate analysis showed ALOX12 as a predictive factor, we further used a nomogram to combine ALOX12 with the basic characteristics of patients for survival prediction (Figure 3(a)).

Nomogram for predicting 1-year, 3-year, and 5-year OS for breast cancer patients. (a) To calculate survival probability, locate points of patient variables, add points, and locate the sum on the total points line. (b) Calibration curves for the probability of 3-year and 5-year OS.
We used two methods to validate our model. The concordance index (C-index) using the Bootstrap Self-sampling method, which could estimate the discrimination ability of the nomogram, was 0.730 (0.703–0.757). Furthermore, we performed graphic calibration method analysis and it also showed high accordance between predicted survival and actual survival both in 3-year OS and 5-year OS (Figure 3(b)).
Prognostic value of SLC7A11 and ALOX12 in our center
To further demonstrate the prognostic value of ferroptosis-related genes and their relations with adipocytes, we performed immunohistochemistry (IHC) tests for breast cancer patients in our center. A total of 258 breast cancer patients with tumor tissue array and complete medical records were enrolled and analyzed at Fudan University Shanghai Cancer Center (FUSCC). IHC staining was scored 0, 1, 2, and 3 according to stain intensity, and scores of 2 and 3 were defined as high expression. Baseline patient characteristics between two treatment groups divided by SLC7A11 and ALOX12 expression are summarized in Tables 1 and 2. No significant difference was observed in median age, T stage, and molecular types between high and low expression of both groups. ALOX12 high or SLC7A11 low patients had lower N stage and less vascular invasion. Survival analysis showed a significantly improved RFS in ALOX12-low patients compared to ALOX12-high patients in all patients (Figure 4(a)) and TNBC subtype (Figure 4(b)).
Baseline patient characteristics between two treatment groups divided by ALOX12 expression.

HER-2, human epidermal growth factor receptor-2; TNBC, triple-negative breast cancer.
Baseline patient characteristics between two treatment groups divided by SLC7A11 expression.

HER-2, human epidermal growth factor receptor-2; TNBC, triple-negative breast cancer.
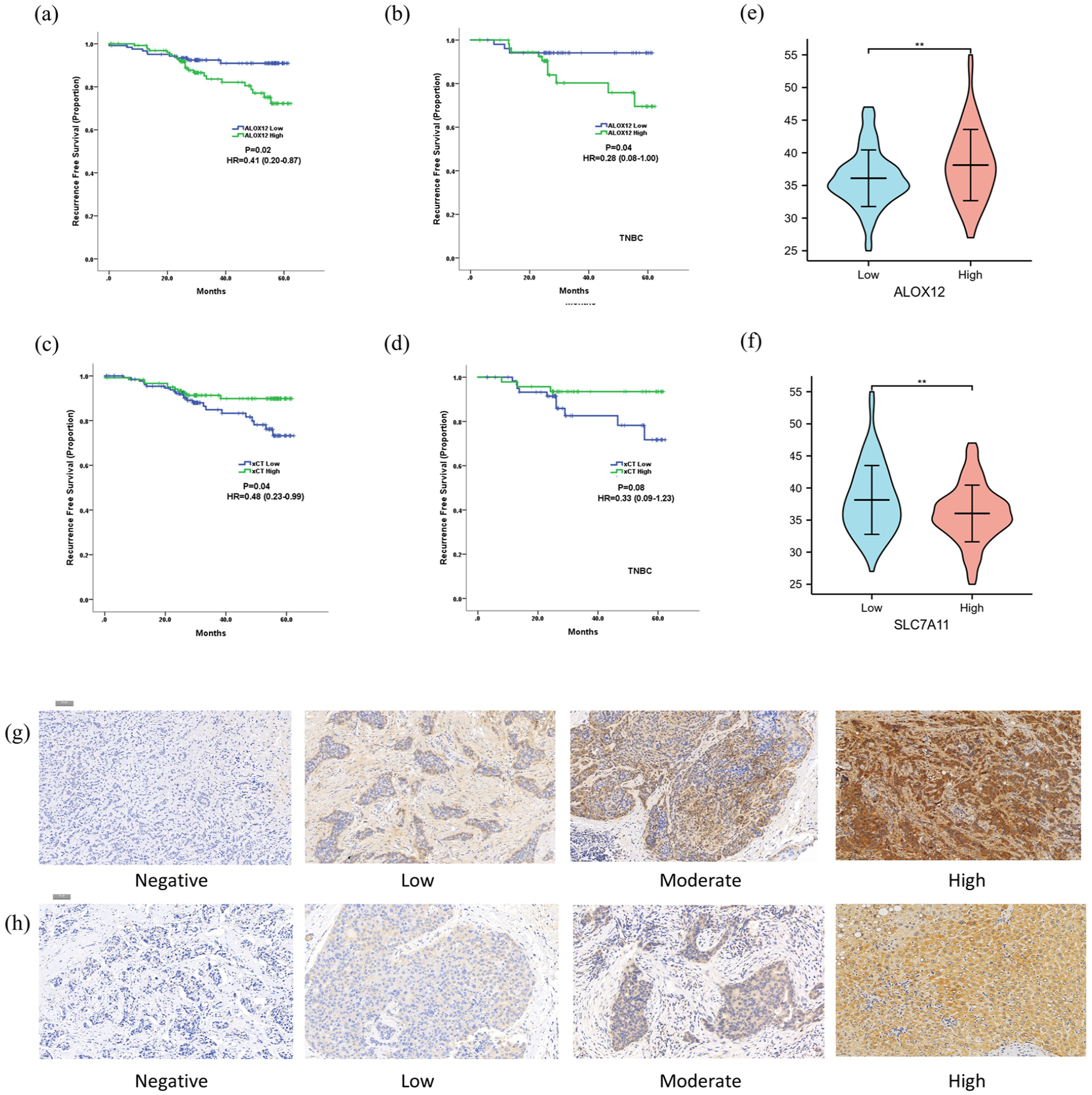
Prognostic value of SLC7A11 and ALOX12 in FUSCC. (a) Kaplan–Meier curves for RFS divided by ALOX12 expression in all patients. (b) Kaplan–Meier curves for RFS divided by ALOX12 expression in TNBC patients. (c) Kaplan–Meier curves for RFS divided by SLC7A11 expression in all patients. (d) Kaplan–Meier curves for RFS divided by SLC7A11 expression in TNBC patients. (e) Violin plot of BMI score divided by ALOX12 expression. (f) Violin plot of BMI score divided by SLC7A11 expression. (g) Representative IHC staining figures of ALOX12. (h) Representative IHC staining figures of SLC7A11.
Similarly, SLC7A11-high patients had a longer RFS compared to SLC7A11-low patients (Figure 4(c)), and a trend of predicting the TNBC subtype was observed (Figure 4(d)). Furthermore, we found that ALOX12-high and SLC7A11-low patients had significantly higher BMI scores (Figure 4(e) and (f)). Representative ALOX12 (Figure 4(g)) and SLC7A11 (Figure 4(h)) IHC figures were shown. These findings further suggested a meaningful prognostic value of SLC7A11 and ALOX12 and a close relation between fat tissue and ferroptosis in breast cancer patients.
FTH1 could predict immunotherapy outcomes of TNBC patients
We further evaluated ferroptosis-related genes and breast cancer immune infiltration.
We found that patients with high FTH1 expression had significantly higher levels of immune infiltration. The enriched immune cells included major immune effector cells such as NK cells and CD8+ T cells (Figure 5(a)). In addition, patients with high FTH1 expression had higher immune infiltration scores (Figure 5(b)). Subsequently, we included 12 TNBC patients from our center who received first-line immunotherapy for advanced disease and had available paraffin-embedded tissue sections. Immunofluorescence staining for FTH1 and PD-L1 was performed, with baseline information provided in Table 3. The results showed a significant positive correlation between FTH1 and PD-L1 expression (Figure 5(c)). Patients with high FTH1 expression had a significantly better median PFS compared to those with low FTH1 expression (12.4 vs 5.4 months, p = 0.005), and patients with high PD-L1 expression had a significantly better median PFS compared to those with low PD-L1 expression (12.4 vs 6.8 months, p = 0.016; Figure 5(d) and (e)). Multivariate analysis indicated that high FTH1 expression was an independent predictor of immunotherapy efficacy (HR = 0.052, 95% CI 0.006–0.478). Figure 5(f) shows immunofluorescence images of two typical patients, patient I had a PFS of 12 months and patient II had a PFS of 6 months.

Immunotherapy prediction value of FTH1. (a) Immuno-infiltration analysis of TCGA-BRCA database using ssGSEA model. (b) Immuno-infiltration analysis of TCGA-BRCA database using ESTIMATE model. (c) Relationship between FTH1 expression and PD-L1 expression in TNBC immunotherapy cohort of FUSCC. (d) Kaplan–Meier curves for PFS divided by FTH1 expression in the FUSCC cohort. (e) Kaplan–Meier curves for PFS divided by PD-L1 expression in the FUSCC cohort. (f) Representative immunotherapy patient immunofluorescence confocal images: Patient I, High expression of FTH1 and PD-L1, with good response to immunotherapy; Patient II: Low expression of FTH1 and PD-L1, with poor response to immunotherapy.
Baseline patient characteristics receiving immunotherapy in FUSCC divided by FTH1 expression.

DFI, Disease-Free Survival; FTH1, Ferritin Heavy Chain 1; FUSCC, Fudan University Shanghai Cancer Center; PD-L1, programmed death ligand 1.
Discussion
As a newly discovered form of programmed cell death, ferroptosis plays a more and more important role in tumor treatment and therapy resistance. However, cancer cells could use multiple ways to escape from ferroptosis. We previously found the protection value of adipocytes against breast cancer ferroptosis. 20 The present study further explored the prognostic value of ferroptosis-related genes on breast cancer, as well as the relationship between adipocytes and ferroptosis using public database and data from our center. Moreover, a novel combination strategy was figured out to overcome protection against breast cancer ferroptosis.
At first, breast cancer was considered less sensitive to ferroptosis in terms of limited ferroptosis inducers. 21 With the increasing knowledge of ferroptosis, more and more evidence indicates that ferroptosis participates in not only the inhibition of breast cancer growth but also the synergistic anticancer effect together with targeted therapy, especially for TNBC.22,23 Thus, investigation into ferroptosis protection could help us better understand the underlying mechanism of development, metastasis, and treatment response of breast cancer. We identify 96 ferroptosis-related genes differentially expressed in breast cancer and find six genes as predictors for breast cancer prognosis both in all patients and the TNBC subgroup, which on one hand adds evidence to support a link between ferroptosis and breast cancer and on the other hand give a hint for future studies.
ALOX12 catalyzes the production of lipid mediator which mediates cellular PUFA levels to regulate ferroptosis levels. 24 It is also reported that ALOX12 is required for a distinct ferroptosis pathway of p53-mediated tumor suppression. 25 We found ALOX12 as an independent predictor of OS in breast cancer patients and established a nomogram figure to visualize the risks for convenient use. The C-index of the nomogram is 0.7, which could be attributed to other genomic influences on prognosis not included in the nomogram. Similarly, ALOX12 was found to predict treatment response and prognosis of colorectal cancer. 26 Moreover, IHC staining in our center showed predictive value of ALOX12 and xCT in RFS of breast cancer patients. We also found that patients with different expressions of ALOX12 and xCT had differences in N staging and vascular invasion, further suggesting that these genes may be involved in the regulation of breast cancer invasion and metastasis. As far as we know, this is the first study reporting the prognostic value of ALOX12 and xCT both in medical centers and databases of breast cancer patients.
FTH1 is an important component of ferritin, responsible for storing cytoplasmic iron and exerting cellular antioxidant effects by sequestering oxidative active iron. 27 Both in vitro and in vivo experiments have confirmed that ferritin, as a key regulator of ferroptosis, can mediate cell sensitivity to ferroptosis.28,29 Our exploration suggests that FTH1 may be related to immune infiltration in breast cancer. We further proved FTH1 expression as an effective predictor for immunotherapy outcome of TNBC patients. The potential mechanism might be the upregulation of iron metabolism by FTH1 causing a reduction of oxidative stress and ferroptosis of the immune microenvironment among TNBC, thereby enhancing the effectiveness of immunotherapy associated with its role in iron metabolism and oxidative stress response. These findings suggested that FTH1 could be a promising biomarker for TNBC immunotherapy.
In conclusion, the present study discovered the prognostic value of ferroptosis-related genes in breast cancer patients and set up a nomogram model. Furthermore, we found FTH1 as a promising predictor for TNBC immunotherapy.
This study mostly focused on the prognostic and immune-prediction value of ferroptosis-related genes without more fundamental work. Thus, further explorations on the function and potential mechanisms are warranted in the future.
Conclusion
Ferroptosis-related genes like ALOX12 and SLC7A11 had potential prognostic value for breast cancer patients. Furthermore, FTH1 could predict immunotherapy outcomes for TNBC patients.
Supplemental Material
sj-pdf-1-tam-10.1177_17588359251322291 – Supplemental material for Ferroptosis-related genes as prognostic markers for survival and immunotherapy in triple-negative breast cancer: analysis of public databases and a single institution
Supplemental material, sj-pdf-1-tam-10.1177_17588359251322291 for Ferroptosis-related genes as prognostic markers for survival and immunotherapy in triple-negative breast cancer: analysis of public databases and a single institution by Yizhao Xie, Zhonghua Tao, Biyun Wang, Yannan Zhao, Xinyu Chen, Bin Li, Jinyan Wang, Guangliang Chen and Xichun Hu in Therapeutic Advances in Medical Oncology
Supplemental Material
sj-pdf-2-tam-10.1177_17588359251322291 – Supplemental material for Ferroptosis-related genes as prognostic markers for survival and immunotherapy in triple-negative breast cancer: analysis of public databases and a single institution
Supplemental material, sj-pdf-2-tam-10.1177_17588359251322291 for Ferroptosis-related genes as prognostic markers for survival and immunotherapy in triple-negative breast cancer: analysis of public databases and a single institution by Yizhao Xie, Zhonghua Tao, Biyun Wang, Yannan Zhao, Xinyu Chen, Bin Li, Jinyan Wang, Guangliang Chen and Xichun Hu in Therapeutic Advances in Medical Oncology
Supplemental Material
sj-pdf-3-tam-10.1177_17588359251322291 – Supplemental material for Ferroptosis-related genes as prognostic markers for survival and immunotherapy in triple-negative breast cancer: analysis of public databases and a single institution
Supplemental material, sj-pdf-3-tam-10.1177_17588359251322291 for Ferroptosis-related genes as prognostic markers for survival and immunotherapy in triple-negative breast cancer: analysis of public databases and a single institution by Yizhao Xie, Zhonghua Tao, Biyun Wang, Yannan Zhao, Xinyu Chen, Bin Li, Jinyan Wang, Guangliang Chen and Xichun Hu in Therapeutic Advances in Medical Oncology
Footnotes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
