Abstract
Background:
Patients with initially unresectable colorectal cancer liver metastases (IU-CRLM) might benefit from using an effective systemic treatment followed by resection of liver metastases but the curative success rate is quite low. Indeed, nearly one-third of patients exhibit early recurrence within the first 6 months after surgery, and these individuals often have poor overall survival.
Objectives:
This study aims to clarify the application value of serial circulating tumor DNA (ctDNA) analysis in predicting the clinical outcome of IU-CRLM patients following liver metastasectomy.
Design:
A retrospective study was conducted on a cohort of patients with IU-CRLM between February 2018 and April 2021.
Methods:
Plasma samples at different time points during CRLM treatment [baseline (BL), preoperation (PRE), postoperation (POST), end-of-treatment (EOT), and progressive disease (PD)] were retrospectively collected from patients with initially unresectable CRLM enrolled at the Sun Yat-sen University Cancer Center. Dynamic changes of SEPTIN 9 (SEPT9) and Neuropeptide Y (NPY) methylated circulating tumor DNA (MetctDNA) levels in serial plasma samples were detected using droplet-digital PCR (ddPCR).
Results:
SEPT9 and NPY genes were hypermethylated in colon cancer cell lines and tissues while no difference was observed between primary and metastatic tumors. Patients with MetctDNA positive at POST or EOT had significantly lower recurrence-free survival (RFS) compared to patients with MetctDNA negative at these time points [POST: Hazard ratio (HR) 9.44, 95% confidence interval (CI) 5.15–17.30, p < 0.001; EOT: HR 11.48, 95% CI 3.27–40.31, p < 0.001]. Multivariate analysis demonstrated that POST (OR 33.96, 95% CI 4.03–286.10, p = 0.001) and EOT (OR 18.36, 95% CI 1.14–295.71, p = 0.04) MetctDNA was an independent risk factor for early recurrence. Time-dependent receiver operating characteristic curve (T-ROC) analysis revealed that area under the curve (AUC) value was greatest at the relapse time point of 6 months post-intervention, with POST-AUC and EOT-AUC values of 0.74 (95% CI 0.66–0.81) and 0.73 (95% CI 0.53–0.94), respectively. Serial MetctDNA analysis showed that RFS was significantly lower in patients with no MetctDNA clearance compared with those with MetctDNA clearance (HR 26.05, 95% CI 4.92–137.81, p < 0.001).
Conclusion:
Our study confirmed that serial ctDNA analysis of NPY and SEPT9 gene methylation could effectively predict early recurrence in IU-CRLM patients, especially at POST and EOT.
Keywords
Introduction
Colorectal cancer (CRC) is the third most prevalent malignancy and the second most common cancer-related death worldwide. 1 More than half of CRC patients develop liver metastases during the course of their disease. 2 Notably, only 20–30% of CRC patients with limited liver metastases are candidates for curative surgery. 3 Factors such as extensive tumor burden, a high number of liver metastases, or poor anatomical position of lesions such as those close to critical biliary or vascular structures preclude resection. 4 In recent years, several studies have shown that patients with initially unresectable colorectal liver metastases (IU-CRLM) who respond to systemic chemotherapy may be converted to resectable and/or treatable lesions using a combination of radiofrequency ablation. Although the majority of patients are incurable, they often exhibit favorable long-term overall survival (OS).3,5 Around one-third of the patients had early recurrence within the first 6 months after surgery, especially patients with a high number of liver metastases.6,7 Although repeated local treatment of recurrent metastases might be beneficial, early recurrence has been reported as a poor prognostic factor for overall survival in patients with CRLM after hepatectomy.8,9
Carcinoembryonic antigen (CEA) and imaging techniques such as CT and MRI are commonly used in routine clinical detection of early recurrence. However, the sensitivity and specificity of these methods remain unsatisfactory. Circulating tumor DNA (ctDNA) has recently received increased attention as a promising biomarker. 10 It consists of a fragment of DNA released into the bloodstream by dead cancer cells that contain the same genetic and epigenetic changes as the cancer cells from which they originated. Previous assessment of the CRC epigenome has unveiled widespread aberration methylation in multiple driver genes. Advancements in understanding abnormal methylation have spurred the utilization of epigenetic changes as clinical biomarkers, with applications in the diagnosis, prognosis, and treatment of CRC. Of note, studies have shown that ctDNA is a promising tool for postoperative surveillance of stage I–III CRC.11–15 In particular, ctDNA assessment has also been reported as a strong indicator of minimal residual disease in patients with resectable CRLM.16–18 However, the value of ctDNA for predicting the survival of IU-CRLM patients remains undetermined. Our previous study demonstrated that ctDNA could predict recurrence in CRLM patients after hepatectomy. 18 In that study, we enrolled resectable and initially unresectable populations and utilized a 451-gene Next Generation Sequencing (NGS) panel-based approach to measure ctDNA. That strategy had several limitations, such as high cost, long detection time, and relatively low sensitivity and specificity. Therefore, the search for an easy and reproducible detection method for the early identification and management of IU-CRLM patients with recurrence after liver metastasectomy is of great clinical importance.
Aberrant methylation is an epigenetic change in the DNA that might be associated with the risk of tumorigenesis. The stability of the altered methylation makes them suitable biomarkers for early diagnosis, as well as predictors of prognosis and therapeutic efficacy in various malignant.19–22 Notably, the Food and Drug Administration previously approved a blood-based screening test using Septin 9 (SEPT9) methylation levels for average-risk adults aged 50 years or older who refused first-line screening modalities. 23 Although its specificity is inferior to conventional testing, blood-based screening garnered more participation and was preferred to fecal immunochemical testing, as evidenced by a small randomized controlled trial conducted in the United States.23,24 Besides, Neuropeptide Y (NPY) and other aberrant DNA methylation in the blood have been used as diagnostic and prognostic biomarkers in CRC patients.25–27 Furthermore, this tumor-agnostic method for ctDNA detection based on colorectal cancer-specific DNA methylation markers has some potential advantages, including quick turnaround time, no requirement to collect or analyze tumor tissues, and high cost-effectiveness.
Herein, we are the first to confirm the hypermethylation of SEPT9 and NPY genes in colon cell lines and tissues using droplet digital PCR (ddPCR) technology. Moreover, we offer more insight into the translational value of serial MetctDNA analysis in predicting early recurrence, especially in IU-CRLM patients after liver metastasectomy.
Methods
Cell lines and tissue specimens
Human colon adenocarcinoma cell lines (RKO, DLD1, HCT8, HCT15, HCT116, HT29, SW480, SW620, and LS174T) and human immortalized intestinal epithelial cell line (CCD841CON) were purchased from the Institute of Cell Biology of the Chinese Academy of Sciences (Shanghai, China). Cells were cultured in RPMI-1640 medium supplemented with 10% fetal bovine serum at a constant temperature of 37°C in a humidified chamber with a constant atmosphere of 5% CO2.
Formalin-fixed paraffin-embedded (FFPE) tissues of 92 cancer patients pathologically diagnosed with colorectal cancer were collected at Sun Yat-sen University Cancer Center. Methylation level was evaluated in the primary intestinal tissues of 61 patients (Stage I–IV), metastatic liver tissues of 31 patients, and their respective leukocyte samples.
Patient recruitment and blood sample acquisition
CRLM patients enrolled from February 2018 until April 2021 at Sun Yat-sen University Cancer Center in China were retrospectively recruited according to the following eligibility criteria: (1) pathologically and radiologically diagnosed with CRLM; (2) liver metastases were initially unresectable; (3) at least one blood sample was available from diagnosis to recurrence; and (4) Eastern Cooperative Oncology Group performance score of 0–1. As previously defined by the CELIM trial, 28 initially unresectable liver metastases are metastatic lesions ⩾5 and/or metastases considered technically unresectable by a multidisciplinary team according to one of the following criteria: (1) failure to perform R0 resection; (2) insufficient residual liver volume after resection; and (3) failure to preserve all three hepatic veins after resection, failure to ensure sufficient blood flow to and from the liver and bile ducts in the residual liver, and failure to preserve two adjacent liver segments. Patients with small (⩽1 cm) extrahepatic lesions that were not confirmed as metastatic lesions were eligible. In addition, we also collected 105 plasma samples from healthy individuals recruited between June 2019 and November 2019 at the Physical Examination Center, Sun Yat-sen University Cancer Center. Plasma was immediately separated from the blood samples by centrifugation at 1600g for 15 min (4°C) and stored at −80°C for DNA extraction. Preoperatively, patients received a median of four chemotherapy regimens (range: 3–8), and postoperatively, they underwent a median of two regimens (range: 1–6). Patients were assessed every 3 months postoperatively using imaging modalities such as CT and MRI, while physical examination and CEA monitoring were conducted at each follow-up. In addition, ‘no evidence of disease’ (NED) status was defined as the absence of detectable cancer assessed using imaging modalities and blood CEA following successful local treatment. All included patients and healthy controls (normal, N) provided written informed consent, and all study aspects were approved by the Sun Yat-sen University Cancer Center ethics committee (Approval number: SL-B2018-159-03).
DNA isolation and bisulfite conversion
DNA was isolated from colon adenocarcinoma cell lines, and DNA from leukocytes was purified using the QIAamp DNA Mini Kit (Qiagen, Inc., Hilden, Germany). DNA was isolated from tissues using the QIAamp DNA FFPE Tissue Kit (Qiagen, Inc., USA). Circulating free DNA (cfDNA) was extracted from 2 mL of plasma using the QIAamp Circulating Nucleic Acid Kit (Qiagen, Inc., USA) according to the manufacturer’s protocol. DNA bisulfite conversion was performed using the EZ DNA Methylation Kit (Zymo Research, Orange, CA, USA) according to the manufacturer’s instructions. Finally, 20 μL of modified DNA was obtained and immediately used for ddPCR or was stored at −20°C.
Methylated circulating tumor DNA detection and threshold determination
In this study, we focused on the clinically relevant biomarkers SEPT9 and NPY in colorectal cancer, a decision guided by practical considerations given the challenges in extensive methylation profiling. Before commencing the study, we found that SEPT9 and NPY exhibited the highest specificity and sensitivity in CRC specimens. The assessment of MetctDNA was performed using the QX200 Droplet Digital PCR System (Bio-Rad), based on previous SEPT9 29 and NPY 27 gene hypermethylation detection methods. In each sample, SEPT9 and NPY hypermethylation were measured using the unmethylated sequence of Glyceraldehyde-3-Phosphate Dehydrogenase (GAPDH) as a reference, and the ratio of methylated markers in each sample was determined. The sequences utilized are listed in Supplemental Table S1. The SEPT9/NPY/GAPDH multiplex ddPCR system was performed in two wells containing 6 µL of converted DNA per well for a total reaction system of 20 µL. The SEPT9/NPY/GAPDH assay was applied (Supplemental Table S1) and consisted of 10 μL of 2×ddPCR SuperMix (Bio-Rad), 100 nM Taqman®-labeled probe for GAPDH unmethylated sequences, as well as 100 nM FAM and 100 nM VIC-labeled probes to detect SEPT9 and NPY methylation sequences, respectively. Droplets were generated on a BioRad QX200 automated droplet generator. PCR was performed using Bio-Rad C1000. The thermal-cycling conditions were 95°C for 10 min (2°C/s ramp rate), 94°C for 30 s and 56°C for 1 min (45 cycles) (2°C/s ramp rate), 98°C for 10 min (2°C/s ramp rate), and maintaining the mixture at 4°C (1°C/s ramp rate). The droplets were quantified using the QX200 droplet reader from BioRad. Data were analyzed using QuantaSoft version 1.7.4 (BioRad, Biorad, Hercules, CA, USA).
Water was used as a negative control and universal hypermethylated DNA (Zymo Research, USA) as a positive control. The limit of blank (LOB) was calculated as described previously. 30 LOB was determined by the frequency of positive droplets measured in water control and healthy control (N = 105) DNA samples without hypermethylated ctDNA. Specifically, the LOBs for each target gene were LOB = 1 for SPET9 and LOB = 2 for NPY. Samples were further analyzed only when the number of positive droplets was higher than the LOB. All other samples below LOB were negative. As described above, the multiplex ddPCR system uses GAPDH unmethylated sequences as a reference for measuring amplifiable DNA in each sample. In this study, negative samples with less than 80 GAPDH genomes were censored to ensure sufficient sensitivity. MetctDNA levels were used to represent continuous variables and were defined as the fraction of MetctDNA and expressed as the percentage of SEPT9 and NPY methylated positive droplets in total circulating cell-free DNA (MetctDNA% = [MetctDNA/GAPDH] × 100). The methylation level of DNA extracted from tissues or cells (MetDNA) was measured in the same way as that of plasma samples. Finally, cfDNA extracted from healthy control plasma (N = 105) was analyzed to determine outlier values for methylation ratio. The observed median percentages of SEPT9 and NPY were 0.00% [interquartile range (IQR), 0.00–0.00%] and 0.00% (IQR, 0.00–0.00%), respectively. To assess dichotomous data, the median ratio of methylated genes, as well as the 25% percentile (Q1), 75% percentile (Q3), and IQR of the data, was calculated and used to determine a ‘positive threshold’ equivalent to ‘1.5IQR + Q3’ (0.00% and 0.00% for SEPT9 and NPY, respectively). The obtained MetctDNA % was then compared with the ‘threshold of positivity’ determined as described above. To be defined as MetctDNA-positive, both markers (SEPT9 and NPY) must be above that threshold, while MetctDNA-negative patients had either one marker or both below that threshold.
Statistical analysis
Median [interquartile range (IQR)] and frequency (percentage) were used to describe the clinicopathological characteristics of continuous and categorical variables, respectively. Differences in proportions of categorical variables were determined using the chi-squared test (or Fisher’s exact test, if applicable). Comparisons of continuous variables between two groups were examined using the Wilcoxon rank sum test. The Spearman correlation coefficient was utilized to detect the correlation between SEPT9 and NPY methylation levels in tissues. To facilitate the determination of the role of SEPT9 and NPY methylation-related biomarkers in the diagnosis of colorectal tissue, receiver operating characteristic curves (ROC) were utilized, and the area under the curve (AUC) was reported with 95% CI (Confidence interval). For all patients, postoperative samples were collected within 2 months after liver resection (termed POST), and end-of-treatment samples were the first samples collected within 1 month after completion of adjuvant chemotherapy (termed EOT). The primary outcome was recurrence-free survival (RFS), which is the time interval from the date of liver resection to the first radiological recurrence or the last follow-up date. Moreover, the median follow-up and IQR were also estimated. Time-dependent ROC (T-ROC) for RFS prediction was performed in POST and EOT settings to assess the relationship between the predictive efficiency of the MetctDNA status or CEA and time. Early recurrence risk factor analysis was performed using univariate and multivariate logistic regression analyses. The independent predictive performance of MetctDNA status was evaluated by univariate and multivariate analyses using Cox proportional hazards regression models. Hazard ratios (HRs), 95% CIs, and corresponding p values were calculated accordingly. In the univariate analysis, variables with p values <0.05 were further included in the subsequent multivariate analysis. Kaplan–Meier curves were used to calculate the survival rate, and comparisons at multiple time points were performed using the log-rank test. All statistical tests were two-sided, and a p value of less than 0.05 was considered statistically significant. All statistical analyses were performed using the R-3.6.3 platform (https://www.r-project.org).
Results
DNA methylation levels in colon cell lines and tissue samples
Figure 1 illustrates our study design and patient enrolment process. To determine the methylation level of SEPT9 and NPY in colon adenocarcinoma cell lines and tissues, we first measured the methylation levels of two genes in nine colon adenocarcinoma cell lines (RKO, DLD1, HCT8, HCT15, HCT116, HT29, SW480, SW620, and LS174T) and a human immortalized intestinal epithelial cell line (CCD841CON). The results showed that the methylation ratios of SEPT9 and NPY were significantly higher in colon cancer cell lines compared to the immortalized intestinal epithelial cell line [Figure 2(a)]. Next, we evaluated the methylation levels of SEPT9 and NPY in 61 paired primary intestinal tissues, 31 paired metastatic liver tissues, and corresponding leukocyte samples. Of note, methylation levels were significantly higher in tumor tissues compared to their respective leukocyte samples [p < 0.001; Figure 2(b)]. Meanwhile, no difference in DNA methylation levels was observed between primary intestinal tissues and liver metastases tissues [p > 0.05; Figure 2(c)]. Importantly, we observed a marked correlation between SEPT9 and NPY methylation levels [Figure 2(d)]. Individual data points are displayed as circles or triangles, with R = 0.42 and 0.68 for primary intestinal and liver metastasis tissues (both p < 0.001). To determine the role of SEPT9 and NPY methylation-related biomarkers in the diagnosis of colorectal cancer in tissues, we measured their respective AUCs. The AUC for NPY was 0.984 (95% CI 0.963–1), while the AUC for SPET9 was 0.982 (95% CI 0.961–1). More importantly, the combination of SEPT9 and NPY had an AUC of 1 [95% CI 1–1; Figure 2(e)]. At the same time, baseline or normal-setting SEPT9 and NPY MetctDNA% levels were not correlated with age and sex (Supplemental Figure S1).

Workflow of the study and patient enrolment. (a) Cell validation was performed by comparing nine colon cell lines and immortalized intestinal epithelial cell lines. Tissue validation was performed by comparing the primary intestinal tissues of 61 patients and the metastatic liver tissues of 31 patients matched to their leukocytes. Plasma MetctDNA assessment was performed by collecting a total of 212 blood samples, including 105 CRLM patients at diagnosis and 105 healthy controls. (b) Overview of blood collection at different time points for enrolled patients and study flow. Of the 181 patients with initial unresectable CRLM enrolled, the focus was on the 111 patients whose postoperative plasma samples were included in the analysis, as shown in the figure for the specific pairings. Of those only 42 patients had BL MetctDNA samples, while 20 patients had both BL and EOT MetctDNA samples. Meanwhile, 37 patients had POST as well as EOT MetctDNA samples. To assess the prognostic impact of postoperative and EOT MetctDNA on RFS in patients.

The methylation level of SEPT9 and NPY in colon adenocarcinoma cell lines and tissues. (a) Comparisons of MetDNA% of SEPT9 and NPY in the nine colon cell lines and immortalized intestinal epithelial cell line. (b) Comparisons of MetDNA% of SEPT9 and NPY in the primary intestinal tissues and metastatic liver tissues corresponding to leukocytes (p < 0.05). (c) Comparisons of MetDNA% of SEPT9 and NPY in the primary intestinal tissues and metastatic liver tissues. (d) The Spearman correlation coefficient presents the correlation between SEPT9 methylation level and NPY methylation level for primary intestinal and liver metastasis tissues (R = 0.42 and 0.68, both p < 0.001). (e) The ROC of NPY, SPET9, NPY + SPET9 for tissues.
Clinicopathological characteristics and assessment of Plasma DNA methylation levels
A total of 318 serial blood samples from 181 CRLM patients and plasma specimens from 105 healthy controls were retrospectively collected from our subjects between February 2018 and April 2021. Figure 1 also depicts the blood collection times, and Supplemental Table S2 details the clinical characteristics of patients included in the study. A total of 111 patients ultimately underwent liver metastasectomy and POST MetctDNA samples were collected accordingly (n = 111). Of those, only 42 patients had BL MetctDNA samples, while 20 patients had both BL and EOT MetctDNA samples. Meanwhile, 37 patients had POST as well as EOT MetctDNA samples. The follow-up endpoint of the 111 patients who received hepatectomy was in December 2022; starting from liver resection, the median follow-up time was 7.5 months (IQR, 3.8–19.0 months), and the median RFS was 7.5 months (95% CI 6.8–10.4 months). Among these patients, 85.6% (95 of 111) had developed tumor recurrence after hepatectomy, while local liver recurrence was observed in 70.5% (67 of 95) patients. The 6-month recurrence rate was 38.7% (43 of 111) and 88.4% (38 of 43) of those patients suffered from liver recurrence.
The ctDNA methylation levels of SEPT9 and NPY were assessed by ddPCR in 107 BL CRLM patients and 105 healthy controls blood samples. The SEPT9 and NPY MetctDNA% were significantly higher in BL patients compared to healthy controls [p < 0.001; Figure 3(a)]. The positive detection rate for BL samples was 95.3% (102/107), while only one of the healthy control samples was positive; the false-positive rate was 0.95% (1/105). Next, we analyzed the dynamic changes of SEPT9 and NPY MetctDNA% in a total of 318 samples at five time points, including BL, PRE, POST, EOT, and PD. The SEPT 9 and NPY MetctDNA% were significantly higher in BL compared to PRE [p < 0.001; Figure 3(a)]. The SEPT9 and NPY MetctDNA% in PD were higher than in the EOT groups [p < 0.001; Figure 3(a)]. The positive rate of MetctDNA detection gradually decreased with hepatectomy and postoperative chemotherapy [PRE 42.1%; POST 19.8%; EOT 10.8%; Figure 3(b)]. We found that 21 cases (84.0%) were MetctDNA positive at the time of PD [Figure 3(b)]. The SEPT9 and NPY POST MetctDNA% of patients who reached ‘no evidence of disease (NED)’ after surgery was significantly lower than that of non-NED patients [p < 0.001; Figure 3(c)].

Dynamic changes in MetctDNA levels during treatment. (a) Assessment of MetctDNA level at healthy controls (normal, N), BL, PRE, POST, EOT, and PD. p Values were calculated using the Wilcoxon rank sum test. (b) Positive rate of MetctDNA in five periods. (c) Comparison of NPY and SEPT9 methylation levels in NED and non-NED status.
The median duration from the date of surgery to POST blood collection was 41 days (IQR, 34–55 days). The median time from the completion of adjuvant chemotherapy to EOT blood collection was 1 day (IQR, 1–18 days). No significant differences were found in the percentage of MetctDNA-positive patients at different time points of POST and EOT assessment of MetctDNA (Supplemental Tables S3 and S4). Clinicopathologic characteristics and their association with POST and EOT MetctDNA status are shown in Table 1. Clinicopathologic variables significantly associated with POST MetctDNA positive, include the largest diameter of liver metastasis, elevated level of POST CEA, NED status, high clinical risk score, and recurrence (p < 0.05). Gender and NED status were significantly associated with EOT MetctDNA-positive status (p < 0.05).
Association between clinicopathological features and MetctDNA status in the postoperative and end-of-treatment setting.

Values are presented as median (IQR).
Synchronous liver metastases = time between diagnosis of primary colorectal tumor and metastatic disease less than 6 months.
The bold values indicate that the p-value is less than 0.05.
CEA, carcinoembryonic antigen; CRS, clinical risk score proposed by Fong; EOT, end-of-treatment; DFI, disease-free interval from primary tumor resection to liver metastases; IQR, interquartile range; LM, colorectal liver metastases; NED, no evidence of disease.
Prognostic significance of MetctDNA after resection
Next, we evaluated the prognostic value of MetctDNA in determining RFS at BL, PRE, POST, and EOT. Patients with MetctDNA positive at BL and PRE showed no significant trend in RFS compared to patients with MetctDNA negative (p > 0.05; Supplemental Figure S3). However, patients with POST or EOT MetctDNA positive had significantly lower RFS (median RFS of 1.75 months at POST and 4.34 months at EOT) compared to patients with MetctDNA-negative status (median RFS of 10.37 months at POST and 16.43 months at EOT) at these time points [POST: HR 9.44, 95% CI 5.15–17.30, p < 0.001; EOT: HR 11.48, 95% CI 3.27–40.31, p < 0.001; Figure 4(a) and (c)].
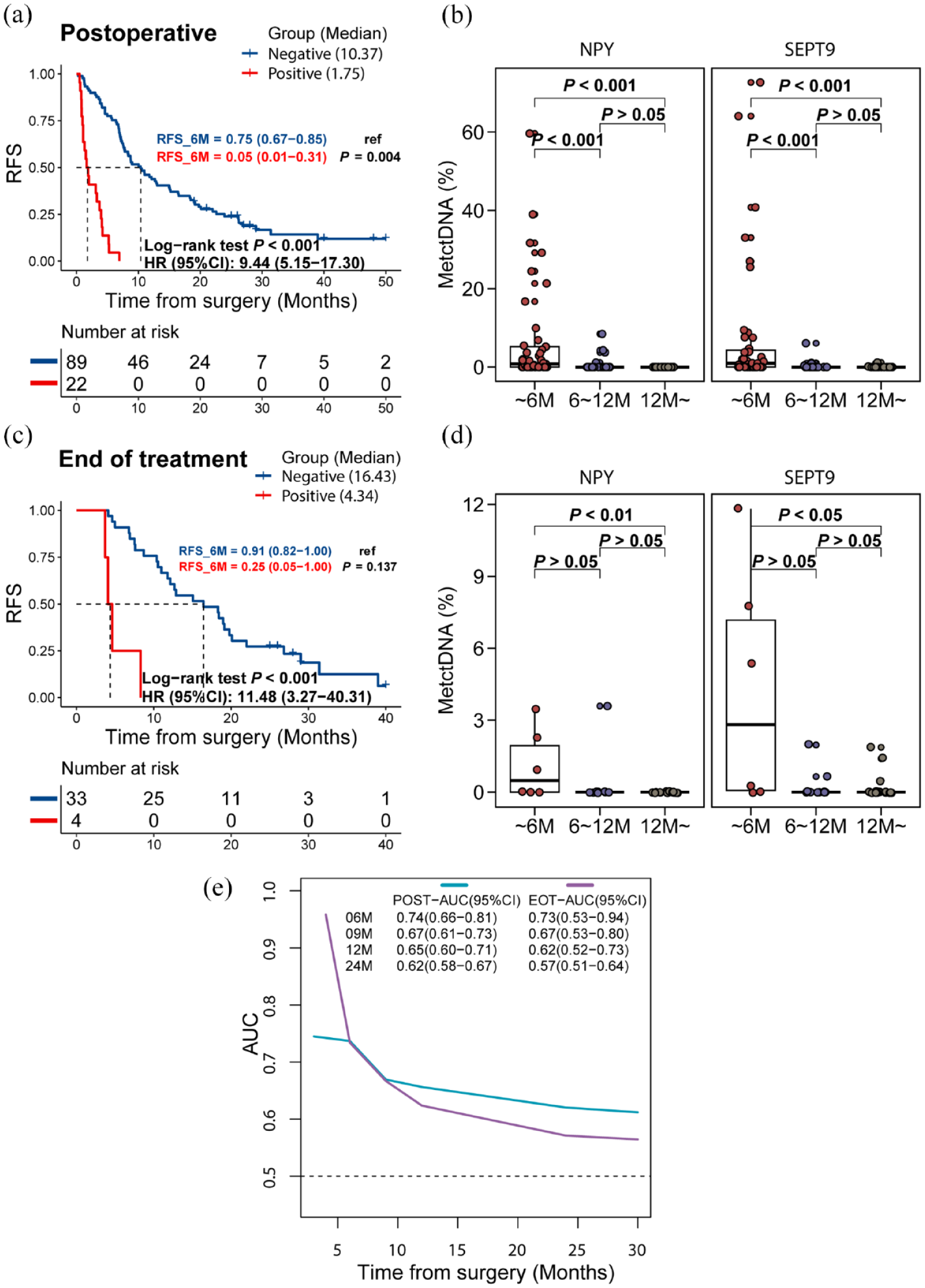
Prognostic significance of postoperative, and EOT MetctDNA. Kaplan–Meier survival analysis shows the probability of RFS by MetctDNA status of postoperation (a) and EOT (c). The SEPT and NPY MetctDNA % in the postoperative and EOT settings were higher in patients with a relapse time of fewer than 6 months than in those with a relapse time of 6–12 months and more than 12 months (b, d) (e) Time-dependent ROC of MetctDNA status for RFS prediction in postoperative and EOT settings. AUC at five time points was shown.
Univariate analysis in POST revealed that MetctDNA status, primary tumor location, elevated PRE CEA, elevated POST CEA, and NED status were significantly associated with RFS (p < 0.05; Table 2). Similarly, univariate analysis in EOT showed that MetctDNA status and elevated POST CEA and EOT CEA were significantly associated with RFS (p < 0.05; Table 2). In multivariate analysis, MetctDNA remained an independent prognostic marker for RFS following POST (HR 7.08, 95% CI 3.70–13.55, p < 0.001) and EOT (HR 39.87, 95% CI 3.44–461.37, p = 0.003; Table 2). On the other hand, CEA was not a significant independent predictor of RFS at POST or EOT, as shown in Table 2.
Univariate and multivariate analyses for recurrence-free survival.

Continuous variable.
The bold values indicate that the p-value is less than 0.05.
CEA, carcinoembryonic antigen; CI, confidence interval; DFI, disease-free interval from primary tumor resection to liver metastases; HR, hazard ratio; LM, liver metastases; NED, no evidence of disease; RFS, recurrence-free survival.
The recurrence rate was significantly higher in POST and EOT MetctDNA-positive patients than in MetctDNA-negative patients (Supplemental Figure S2A and S2B). T-ROC analysis was further performed to compare the predictive power of MetctDNA on RFS at different time points. The AUC of MetctDNA was largest at a relapse time of 6 months, with a POST-AUC and an EOT-AUC of 0.74 [95% CI 0.66–0.81; Figure 4(e)] and 0.73 [95% CI 0.53–0.94; Figure 4(e)], respectively. Meanwhile, the AUC of CEA at 6 months was lower, with an AUC of 0.67 (95% CI 0.56 to 0.77; Supplemental Figure S2C) and 0.67 (95% CI 0.42–0.91; Supplemental Figure S2D) at POST and EOT, respectively. Statistically significant variables (p < 0.05) screened from the univariate logistic regression analysis were included in the multivariate analysis. POST (OR 33.96, 95% CI 4.03 to 286.10, p = 0.001, Supplemental Table S5) and EOT (OR 18.36, 95% CI 1.14–295.71, p = 0.04; Supplemental Table S6) MetctDNAs were independent risk factors for early recurrence (6 months). Furthermore, in the POST and EOT settings, the level of SEPT9 and NPY MetctDNA% was higher in patients with a relapse time of less than 6 months compared to those with a relapse time of more than 12 months [p < 0.05; Figure 4(b) and (d)].
The prognostic analysis of BL, MetctDNA clearance, and serial MetctDNA
RFS estimates were stratified by BL NPY and SEPT9 MetctDNA% quartiles. Significant RFS differences between NPY and SEPT9 MetctDNA% quartiles were not observed at BL, although patients with the highest MetctDNA% quartile tended to have worse RFS (median RFS 6.8 months at SEPT9 and 7.2 months at NPY) than patients with the lowest MetctDNA% quartile (median RFS of 11 months and 12.1 months for SEPT9 and NPY, respectively) [NPY: HR 1.65, 95% CI 0.70 to 3.90, p = 0.250; SEPT9: 3.38, 95% CI 1.41 to 8.10, p = 0.006; Figure 5(a) and (b)]. The pairings in a longitudinal assessment of 28 patients based on serial BL-POST-EOT MetctDNA status revealed that only four patients had negative BL MetctDNA, with a median RFS of 8.4 months. Of the 24 BL MetctDNA-positive patients, 18 had persistent negative MetctDNA at POST and EOT, which we defined as MetctDNA clearance. Six patients had positive MetctDNA at POST or EOT, which we defined as no MetctDNA clearance. Patients with no clearance had significantly lower RFS compared to patients with clearance (HR 26.05, 95% CI 4.92 to 137.81, p < 0.001), with a median RFS of 2.3 and 12.4 months, respectively [Figure 5(c)]. Kaplan–Meier curves for RFS according to serial BL and POST MetctDNA status, serial BL, and EOT MetctDNA status are shown in Supplemental Figure S4. Based on serial BL-POST or BL-EOT MetctDNA status, patients who shifted from positive to negative MetctDNA had significantly higher RFS than those who remained positive after surgery (Supplemental Figure S4A and S4B). In all, 37 patients were stratified and further analyzed using a combination of POST and EOT MetctDNA status [Figure 5(d) and (e)]. Kaplan–Meier survival analysis showed that the difference between the four groups was statistically significant (p < 0.001). All patients in the negative to positive group (N = 3) relapsed at 9 months with a median RFS of 4.6 months. Of these, two had liver recurrences, while one had multiple lymph node metastases. The sole patient in the positive to positive group had an RFS of 3.7 months. Meanwhile, both patients with POST MetctDNA change from positive to negative relapsed at 9 months, and both had extrahepatic metastases (rib metastasis and single supraclavicular lymph node metastasis). The negative to negative group (N = 31) had a median RFS of 18.3 months, and only two patients relapsed within 6 months.

Baseline MetctDNA level, and the association of dynamic changes of MetctDNA with adjuvant chemotherapy and recurrence. RFS stratified by baseline NPY(a) and SEPT9(b) MetctDNA% quartiles. NPY cutoffs%: Q1 ⩽ 6.22, Q2 > 6.22 and ⩽ 22.06, Q3 > 22.06 and ⩽ 42.39, Q4 > 42.39. SEPT9 cutoffs%: Q1 ⩽ 17.61, Q2 > 17.61 and ⩽38.80, Q3 > 38.80 and ⩽61.02, Q4 > 61.02. (c) RFS according to baseline MetctDNA negative, MetctDNA clearance, and MetctDNA no clearance. (d) RFS according to serial postoperative and EOT MetctDNA status. (e) Sankey plot of MetctDNA dynamics and recurrence outcome for the 37 patients who had both serial POST and EOT MetctDNA samples available for analysis.
Discussion
In the present study, we conducted a dynamic ctDNA analysis of IU-CRLM patients using ddPCR to detect methylation levels of the NPY and SEPT9 genes. Importantly, we uncovered the translational potential of POST and EOT MetctDNA in predicting recurrence in IU-CRLM patients after liver resection.
A tumor-agnostic method was utilized for ctDNA detection of SEPT9 and NPY gene methylation based on colorectal cancer-specific DNA methylation markers reported in previous studies.27,29 Herein, we confirmed that SEPT9 and NPY genes were hypermethylated in colon cancer cell lines and CRC tumor tissues while no difference was observed between primary and metastatic tumors. In addition, due to the low ctDNA content, we detected the methylation of plasma ctDNA by utilizing the ddPCR method, which has many advantages, such as a fast turnaround, low cost, and good reproducibility. In addition, this approach does not require the collection or analysis of tumor tissues. Recently, Nadia Øgaard et al. reported using the tumor-agnostic methylation ddPCR test ‘TriMeth’ based on the three CRC-specific methylation markers C9orf50, CLIP4, and KCNQ5 and demonstrated the potential of serial ctDNA detection to guide postoperative resectable CRLM patient management. 17
Interestingly, Tie et al. previously used a tumor-informed personalized approach for ctDNA analysis, revealing that ctDNA decreased significantly following neoadjuvant chemotherapy and local treatment. Furthermore, patients with positive ctDNA at POST or EOT had significantly lower RFS and OS compared to ctDNA-negative patients at similar time points. 16 Consistent with our previous findings,17,18 the positive rate of SEPT9 and NPY MetctDNA% decreased gradually following preoperative chemotherapy, hepatectomy, and postoperative chemotherapy, while an increasing trend was observed in the presence of disease progression. Besides, the POST MetctDNA% of patients who reached NED after surgery was significantly lower than that of non-NED patients. Similarly, we found no significant correlation between BL and PRE MetctDNA status and RFS. Moreover, patients with MetctDNA positive at POST and EOT had poor outcomes, with an estimated 6-month RFS rate of 5.0% [95% CI 1–31%; Figure 4(a)] and 25.0% [95% CI 5–100%; Figure 4(c)] compared with 75.0% [95% CI 67–85%; Figure 4(a)] and 91.0% [95% CI 82–100%; Figure 4(c)] in those with MetctDNA negative at POST and EOT, respectively. Importantly, the 6-month relapse-free rate when both gene markers (SEPT9 and NPY positive) were used (5.0% (95 CI 1–31%) was superior to independent positive gene marker use (SEPT9 positive only: 34% (95% CI 22–52%) and NPY positive only: 17% (95% CI 7–37%). In our previous study, 18 79.4% (27 of 34 patients) of POST ctDNA-positive patients experienced disease recurrence, whereas, in the current study, 100% (22 of 22 patients) of such patients recurred within 9 months, with a median RFS of 1.75 months. This may be because the NGS panel of 451 genes used in the previous study defined samples as positive when the frequency of any one variant allele (VAF) was ⩾0.5%, bringing about false-positive results.
Recent studies using ctDNA detection approaches have reported recurrence risks of 30% to 43% in patients who were ctDNA negative after initially resectable CRLM resection.16–18 However, in our study, patients who were MetctDNA negative at POST and EOT still had a high likelihood of recurrence after 9 months, especially after more than 24 months, with 82.0% to 84.8% of patients relapsing. This may be due to the following reasons: First, Tie et al.’s and Nadia Øgaardet al.’s study included patients with initially resectable CRLM with a relatively good prognosis and a low recurrence rate after hepatectomy,16,17 whereas, in our study, we enrolled IU-CRLM patients, resulting in an overall higher recurrence rate. Second, it may be due to our study’s lack of continuous POST MetctDNA surveillance before radiological recurrence. Several studies showed that MetctDNA could be detected up to 3.1–8.0 months prior to radiographic detection of relapse.17,31,32 Therefore, our analysis leads us to conclude that MetctDNA testing at POST and EOT is an ideal predictor, particularly for early recurrence in IU-CRLM patients. According to T-ROC curve analysis, the 6-month post-treatment period yielded the largest AUC value, suggesting that this timeframe is optimal for analyzing POST and EOT MetctDNA to predict relapse in IU-CRLM patients. Notably, our results also revealed that POST and EOT SEPT9 and NPY MetctDNA levels were significantly higher in patients who relapsed before 6 months in contrast to patients who relapsed after 6 months, suggesting that elevated POST and EOT ctDNA methylation levels may serve as early indicators of a higher risk for relapse. In light of these observations, we posit that the 6-month post-intervention milestone serves as a critical juncture for assessing POST and EOT MetctDNA, offering a valuable window for predicting and potentially preventing early recurrence in IU-CRLM patients. The nuanced relationship between ctDNA methylation levels and the timing of relapse underscores the potential clinical utility of our findings, warranting further exploration and validation in larger cohorts for enhanced prognostic precision.
Furthermore, the serial MetctDNA analysis allowed us to explore the changes in MetctDNA with treatment and the correlation of these changes with outcomes. In the present study, 24 patients were eligible for sequential analysis of BL-POST-EOT MetctDNA status. Patients with no clearance after hepatectomy had significantly lower RFS than those with clearance. 83.3% (5 of 6) of patients with no clearance relapsed within 6 months, compared with 5.6% (1 of 18) of patients with clearance. In addition, our study focused on detecting the dynamic changes in MetctDNA during POST and EOT in 37 CRLM patients. The negative to negative group (N = 31) had a 6-month recurrence rate of 6.5% (2 of 31) with a median RFS of 18.3 months, while all three patients in the negative to positive group had recurrences before 9 months with a median RFS of 4.6 months. Meanwhile, the sole patient in the positive to positive group had an RFS of 3.7 months. However, two patients with extrahepatic metastases had POST MetctDNA change from positive to negative and an RFS of only 4.1 and 6.5 months, respectively. A previous study has reported that concordance rates of ctDNA-based liquid biopsy in mCRC patients depend significantly on the metastatic site. Therefore, caution is required when assessing MetctDNA levels in patients with extrahepatic metastases. 33 Together with the limited sample size, this might explain why MetctDNA conversion from positive to negative did not translate to a better RFS in this particular cohort, warranting further investigation for validation.
There is ongoing debate regarding the role of adjuvant chemotherapy in stage II colon cancer. The Dynamic randomized controlled phase II trial recently demonstrated for the first time that a ctDNA-guided approach could be used to determine whether patients with stage II CRC required adjuvant therapy. 34 Importantly, this study found that patients with detectable ctDNA benefit significantly from adjuvant therapy, while those with undetectable ctDNA are at a very low risk of recurrence and do not require further treatment. Nevertheless, it remains unclear whether ctDNA could guide adjuvant chemotherapy in CRC patients with liver metastases. Herein, we uncovered a correlation between poor RFS and CRLM patients who are MetctDNA positive at both POST and EOT. On the other hand, patients with negative MetctDNA at POST and EOT had a significantly better RFS and lower recurrence rate. Prospective studies with a larger sample size are needed to further validate whether ctDNA could be utilized to tailor perioperative and adjuvant decisions in CRLM patients. For instance, it remains to be determined if patients with postoperative negative ctDNA might avoid treatment-related toxicities by steering away from unnecessary adjuvant chemotherapy or whether positive ctDNA patients could instead benefit from treatment intensification, including biologics or hepatic intra-arterial chemotherapy to prevent disease recurrence.
Nevertheless, our study has several limitations. First, our study included a small sample of participants from a single center, reflected by wide confidence intervals in some of our outcomes. In addition, the small sample size may also give rise to deviations when further independent analyses were conducted, exemplified by the association of EOT MetctDNA-positive status with gender in our study. Therefore, further in-depth studies with a larger sample size are needed to substantiate our findings. Second, due to the retrospective nature of our study, not all CRLM patients had available serial plasma samples for MetctDNA analysis at various time points. Besides, this may give rise to inevitable selection bias; thus, randomized controlled trials are warranted for validation in the future. Third, the follow-up time after hepatectomy was not long enough, and the data collected for OS have not reached maturity. Fourthly, we lacked an external cohort to assess the predictive performance of our markers; thus, caution is warranted when interpreting our findings. In the future, prospective multicenter serial MetctDNA analysis studies are required to determine the clinical significance of MetctDNA in CRLM patients following hepatectomy.
Taken together, our study confirmed that SEPT9 and NPY genes are hypermethylated in colon cancer cell lines and tissues. Using ddPCR to detect the methylation levels of NPY and SEPT9 genes in the peripheral plasma of IU-CRLM patients is a feasible and reliable method. Importantly, we revealed that serial MetctDNA analysis, especially at POST and EOT, could effectively predict 6-month early recurrence in IU-CRLM patients.
Supplemental Material
sj-pdf-1-tam-10.1177_17588359241230752 – Supplemental material for Methylated ctDNA predicts early recurrence risk in patients undergoing resection of initially unresectable colorectal cancer liver metastases
Supplemental material, sj-pdf-1-tam-10.1177_17588359241230752 for Methylated ctDNA predicts early recurrence risk in patients undergoing resection of initially unresectable colorectal cancer liver metastases by Zhi-Gang Chen, Xiao-Meng Ji, Yu-Xia Xu, William Pat Fong, Xiao-Yun Liu, Jie-Ying Liang, Qiong Tan, Lei Wen, Yan-Yu Cai, De-Shen Wang and Yu-Hong Li in Therapeutic Advances in Medical Oncology
Supplemental Material
sj-pdf-10-tam-10.1177_17588359241230752 – Supplemental material for Methylated ctDNA predicts early recurrence risk in patients undergoing resection of initially unresectable colorectal cancer liver metastases
Supplemental material, sj-pdf-10-tam-10.1177_17588359241230752 for Methylated ctDNA predicts early recurrence risk in patients undergoing resection of initially unresectable colorectal cancer liver metastases by Zhi-Gang Chen, Xiao-Meng Ji, Yu-Xia Xu, William Pat Fong, Xiao-Yun Liu, Jie-Ying Liang, Qiong Tan, Lei Wen, Yan-Yu Cai, De-Shen Wang and Yu-Hong Li in Therapeutic Advances in Medical Oncology
Supplemental Material
sj-pdf-2-tam-10.1177_17588359241230752 – Supplemental material for Methylated ctDNA predicts early recurrence risk in patients undergoing resection of initially unresectable colorectal cancer liver metastases
Supplemental material, sj-pdf-2-tam-10.1177_17588359241230752 for Methylated ctDNA predicts early recurrence risk in patients undergoing resection of initially unresectable colorectal cancer liver metastases by Zhi-Gang Chen, Xiao-Meng Ji, Yu-Xia Xu, William Pat Fong, Xiao-Yun Liu, Jie-Ying Liang, Qiong Tan, Lei Wen, Yan-Yu Cai, De-Shen Wang and Yu-Hong Li in Therapeutic Advances in Medical Oncology
Supplemental Material
sj-pdf-3-tam-10.1177_17588359241230752 – Supplemental material for Methylated ctDNA predicts early recurrence risk in patients undergoing resection of initially unresectable colorectal cancer liver metastases
Supplemental material, sj-pdf-3-tam-10.1177_17588359241230752 for Methylated ctDNA predicts early recurrence risk in patients undergoing resection of initially unresectable colorectal cancer liver metastases by Zhi-Gang Chen, Xiao-Meng Ji, Yu-Xia Xu, William Pat Fong, Xiao-Yun Liu, Jie-Ying Liang, Qiong Tan, Lei Wen, Yan-Yu Cai, De-Shen Wang and Yu-Hong Li in Therapeutic Advances in Medical Oncology
Supplemental Material
sj-pdf-4-tam-10.1177_17588359241230752 – Supplemental material for Methylated ctDNA predicts early recurrence risk in patients undergoing resection of initially unresectable colorectal cancer liver metastases
Supplemental material, sj-pdf-4-tam-10.1177_17588359241230752 for Methylated ctDNA predicts early recurrence risk in patients undergoing resection of initially unresectable colorectal cancer liver metastases by Zhi-Gang Chen, Xiao-Meng Ji, Yu-Xia Xu, William Pat Fong, Xiao-Yun Liu, Jie-Ying Liang, Qiong Tan, Lei Wen, Yan-Yu Cai, De-Shen Wang and Yu-Hong Li in Therapeutic Advances in Medical Oncology
Supplemental Material
sj-pdf-5-tam-10.1177_17588359241230752 – Supplemental material for Methylated ctDNA predicts early recurrence risk in patients undergoing resection of initially unresectable colorectal cancer liver metastases
Supplemental material, sj-pdf-5-tam-10.1177_17588359241230752 for Methylated ctDNA predicts early recurrence risk in patients undergoing resection of initially unresectable colorectal cancer liver metastases by Zhi-Gang Chen, Xiao-Meng Ji, Yu-Xia Xu, William Pat Fong, Xiao-Yun Liu, Jie-Ying Liang, Qiong Tan, Lei Wen, Yan-Yu Cai, De-Shen Wang and Yu-Hong Li in Therapeutic Advances in Medical Oncology
Supplemental Material
sj-pdf-6-tam-10.1177_17588359241230752 – Supplemental material for Methylated ctDNA predicts early recurrence risk in patients undergoing resection of initially unresectable colorectal cancer liver metastases
Supplemental material, sj-pdf-6-tam-10.1177_17588359241230752 for Methylated ctDNA predicts early recurrence risk in patients undergoing resection of initially unresectable colorectal cancer liver metastases by Zhi-Gang Chen, Xiao-Meng Ji, Yu-Xia Xu, William Pat Fong, Xiao-Yun Liu, Jie-Ying Liang, Qiong Tan, Lei Wen, Yan-Yu Cai, De-Shen Wang and Yu-Hong Li in Therapeutic Advances in Medical Oncology
Supplemental Material
sj-pdf-7-tam-10.1177_17588359241230752 – Supplemental material for Methylated ctDNA predicts early recurrence risk in patients undergoing resection of initially unresectable colorectal cancer liver metastases
Supplemental material, sj-pdf-7-tam-10.1177_17588359241230752 for Methylated ctDNA predicts early recurrence risk in patients undergoing resection of initially unresectable colorectal cancer liver metastases by Zhi-Gang Chen, Xiao-Meng Ji, Yu-Xia Xu, William Pat Fong, Xiao-Yun Liu, Jie-Ying Liang, Qiong Tan, Lei Wen, Yan-Yu Cai, De-Shen Wang and Yu-Hong Li in Therapeutic Advances in Medical Oncology
Supplemental Material
sj-pdf-8-tam-10.1177_17588359241230752 – Supplemental material for Methylated ctDNA predicts early recurrence risk in patients undergoing resection of initially unresectable colorectal cancer liver metastases
Supplemental material, sj-pdf-8-tam-10.1177_17588359241230752 for Methylated ctDNA predicts early recurrence risk in patients undergoing resection of initially unresectable colorectal cancer liver metastases by Zhi-Gang Chen, Xiao-Meng Ji, Yu-Xia Xu, William Pat Fong, Xiao-Yun Liu, Jie-Ying Liang, Qiong Tan, Lei Wen, Yan-Yu Cai, De-Shen Wang and Yu-Hong Li in Therapeutic Advances in Medical Oncology
Supplemental Material
sj-pdf-9-tam-10.1177_17588359241230752 – Supplemental material for Methylated ctDNA predicts early recurrence risk in patients undergoing resection of initially unresectable colorectal cancer liver metastases
Supplemental material, sj-pdf-9-tam-10.1177_17588359241230752 for Methylated ctDNA predicts early recurrence risk in patients undergoing resection of initially unresectable colorectal cancer liver metastases by Zhi-Gang Chen, Xiao-Meng Ji, Yu-Xia Xu, William Pat Fong, Xiao-Yun Liu, Jie-Ying Liang, Qiong Tan, Lei Wen, Yan-Yu Cai, De-Shen Wang and Yu-Hong Li in Therapeutic Advances in Medical Oncology
Footnotes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
