Abstract
Background:
Recently, HER2-negative breast cancers have been reclassified by protein expression into ‘HER2-low’ and ‘HER2-zero’ subgroups, but the consideration of HER2-low breast cancer as a distinct biological subtype with differing prognoses remains controversial. By contrast, non-neutral ERBB2 copy number alteration (CNA) status is associated with inferior survival outcomes compared to ERBB2 CNA-neutral breast cancer, providing an alternative approach to classification.
Methods:
Here, we investigated the molecular landscape of non-metastatic HER2-negative BCs in relation to ERBB2 CNA status to elucidate biological differences. Molecular Taxonomy of Breast Cancer International Consortium (METABRIC) and The Cancer Genome Atlas (TCGA) TCGA-BRCA datasets (n = 1875) were analyzed.
Results:
Nearly two-fifths of the cohort harbored ERBB2 CNAs (39.4%), which were significantly enriched within hormone receptor-negative (56.1%) than within hormone receptor-positive BCs (35.5%; p < 0.0001). Globally, CNAs across the genome were significantly higher in ERBB2 non-neutral compared to neutral cohorts (p < 0.0001). Notably, genetic aberrations on chromosome 17 – BRCA1, NF1, TP53, MAP2K4, and NCOR1 – were widespread in the ERBB2 non-neutral cases. While chromosome 17q arm-level alterations were largely in tandem with ERBB2 CNA status, arm-level loss in chromosome 17p was prevalent regardless of ERBB2 gain, amplification, or loss. Differential gene expression analysis demonstrated that pathways involved in the cell cycle, proteasome, and DNA replication were upregulated in ERBB2 non-neutral cases.
Conclusion:
Classification of HER2-negative BCs according to ERBB2 CNA status reveals differences in the genomic landscape. The implications of concurrent aberrations in other genes on chromosome 17 merit further research in ERBB2 non-neutral BCs.
Background
The ERBB2 gene encodes for human epidermal growth factor receptor 2 (HER2), a transmembrane receptor tyrosine kinase that is overexpressed in 10–30% of invasive breast cancers. 1 Patients with HER2-positive breast cancers, as defined by high protein expression on immunohistochemistry (IHC) or gene amplification on in situ hybridization (ISH), derive significant benefit from HER2-directed targeted therapies.2,3 In recent years, a novel classification of HER2-negative breast cancers has emerged, defining a ‘HER2-low’ subtype based on low-moderate HER2 protein expression (IHC score of 1+ or 2+) and lack of gene amplification on ISH, 4 in contrast to HER2-zero (IHC score of 0) breast cancers. Importantly, patients with HER2-low breast cancers may benefit from novel HER2 antibody–drug conjugates (ADCs), with progression-free survival and overall survival improvements, demonstrated over standard chemotherapy in a recent phase III trial.5–7
However, it is contentious whether HER2-low breast cancers, as defined above, truly represent a distinct molecular subtype of breast cancer with unique biological characteristics.4,8–10 In particular, there are reproducibility issues in the scoring of HER2 protein expression on IHC, resulting in significant variability among pathologists and specialized centers.11,12 In addition, identification of the HER2 status may be complicated by intra-patient heterogeneity. 13 Consequently, studies investigating survival outcomes in HER2-low versus HER2-zero breast cancers have demonstrated mixed results,8,14–20 leading to uncertainty as to whether the current definition of HER2-low breast cancer has any biological or prognostic implications.
In a recent study, we demonstrated that HER2-low breast cancers conferred a superior prognosis compared to HER2-zero cases in the non-metastatic setting, but absolute differences were modest. Compared with HER2-zero tumors, HER2-low cases had significantly better relapse-free survival [hazard ratio (HR) 0.90, p < 0.001] and overall survival (HR 0.86, p < 0.001). 19 Specifically, improved survival outcomes were evident between patients with HER2 IHC 1+ and HER2-zero tumors, but not between those with HER2 IHC 2+ and HER2-zero tumors. In addition, we showed that ERBB2 non-neutral copy number alteration (CNA) status (as defined by the presence of loss or gain/amplification in ERBB2) was associated with worse relapse-free survival compared to ERBB2 neutral status in a combined Molecular Taxonomy of Breast Cancer International Consortium (METABRIC) and TCGA cohort of HER2-negative non-metastatic breast cancers (HR 1.39, p = 0.001). These findings suggest that the classification of traditionally HER2-negative breast cancers may be improved using information about ERBB2 CNA status.
Hence, in this paper, we examined the molecular landscape of non-metastatic HER2-negative breast cancers in relation to ERBB2 CNA status to further elucidate biological differences and to discuss potential clinical implications in this context.
Methods
Study cohort
Cases diagnosed with non-metastatic HER2-negative breast cancers in the Molecular Taxonomy of Breast Cancer International Consortium (METABRIC; n = 1192) and TCGA TCGA-BRCA (n = 683) datasets were extracted from cBioPortal for analysis.21–24 ERBB2 CNA, genomic mutational data including single nucleotide variants, small indels, and copy number alterations (CNA), as well as clinical and pathological details including age, grade, stage, and hormone receptor status were retrieved. Discrete ERBB2 CNA status was based on the Genomic Identification of Significant Targets in Cancer (GISTIC) method [− 2, loss of both copies; − 1, one copy loss; 0, neutral; 1, low-level gain (a few additional copies, often broad); 2, high-level amplification (more copies, often focal)]. 25
Bioinformatic analysis
Differential mutational and CNA analysis was conducted by grouping the patients into two different cohorts: neutral and non-neutral ERBB2 copy number. The entire cohort was first described with an oncoplot via maftools, 26 exhibiting 53 breast cancer-related genes that comprised variants with >25% of mean allele frequency. 27 Tumor mutation burden [i.e. the number of somatic coding variants per megabase (mt/Mb), with the capture size of 45 and 1 Mb, respectively, in TCGA and METABRIC datasets] was then calculated for each cohort. Furthermore, their association with ploidy status was investigated using Fisher’s exact test. AbsCN-seq 28 was used to estimate ploidy based on the copy number data from METABRIC as well as copy number and single nucleotide variant data from TCGA. The top 10 differentially altered genes, sorted by odds ratio (OR), from the 53 genes shown in oncoplot were presented in a co-barplot, and the number of patients exhibiting these 53 genes in each cohort was tabulated in a table, with the respective OR and false discovery rate (FDR). To investigate the effect of hormone receptor status, we repeated the analysis in hormone receptor-positive and hormone receptor-negative cohorts, respectively, and the ERBB2 non-neutral copy number cohort was further sub-categorized into gain/amplification (GISTIC = 1 or 2) and heterozygous loss (GISTIC = −1) to pinpoint the contributing sub-group in differentially altered genes.
Analysis of differentially expressed genes was performed in a similar approach. Since TCGA and METABRIC datasets were sequenced in different platforms (RNA-seq for TCGA and microarray for METABRIC), we employed DESeq2 29 and limma, 30 respectively, within TCGA and METABRIC datasets, to detect differentially expressed genes at FDR < 0.01 and abs(fold changes) > 1. Pathway enrichment analysis was then performed via Enrichr,31–33 using the KEGG database, to identify pathways that are significantly enriched in neutral and non-neutral cohorts, based on the upregulated genes that are shared between two datasets.
Statistical analysis
Comparisons of the frequencies of categorical variables were performed using Pearson’s chi-squared tests. Box–Whisker plots were used to represent continuous variables and Mann–Whitney–Wilcoxon tests with Bonferroni correction were used to evaluate potential associations. All statistical calculations were performed assuming a two-sided test with a significance level of 0.05 unless otherwise stated. All tests were performed using MedCalc for Windows version 19.0.4 (MedCalc Software, Ostend, Belgium).
Results
Patient cohort
A total of 1875 patients diagnosed with stage I–III HER2-negative (i.e. IHC score of 0–2+ and HER2-negative on ISH if HER2 2+) breast cancers were extracted from the METABRIC (n = 1192) and TCGA (n = 683) datasets. These include hormone receptor-positive (n = 1520) and hormone receptor-negative (n = 355) cases. Nearly two-fifths of the cases harbored ERBB2 CNAs or non-neutral ERBB2 status (n = 739; 39.4%), which included amplification (n = 57), gain (n = 242), and heterozygous loss (n = 440). The remaining were ERBB2 neutral (n = 1136; 60.6%).
In hormone receptor-negative breast cancer, heterozygous loss of ERBB2 was most commonly observed (44.5%), followed by neutral (43.9%), gain (9.6%), and amplification (2.0%). In hormone receptor-positive breast cancer, ERBB2 was most commonly copy number neutral (64.5%), while heterozygous loss (18.6%), gain (13.7%), and amplification (3.3%) were observed in the rest (Table 1, Supplemental Figure 1). Altogether, non-neutral ERBB2 CNA status was significantly more common within hormone receptor-negative (56.1%) than within hormone receptor-positive BCs (35.5%; p < 0.0001). We observed that there was a higher proportion of stage 1 cancer with neutral ERBB2 CNA status (374/557; 67.1%), compared to 59.5% (649/1090) of stage II tumors and 49.6% (113/228) of stage III disease. Notably, grade I breast cancers also had proportionately more ERBB2 neutral tumors (89/107; 83.2%), compared to 73.8% (360/488) of grade 2 tumors and 54.8% (304/555) of grade 3 tumors.
Characteristics of HER2-negative breast cancers from TCGA and METABRIC according to ERBB2 CNA status.

CNA, copy number alteration; HER2-, human epidermal growth factor receptor 2; METABRIC, Molecular Taxonomy of Breast Cancer International Consortium; TCGA, the cancer genome atlas.
Somatic mutation and copy number landscape
The list of 53 breast cancer-related genes is shown in Figure 1(a). The contribution of each variant type for the top 10 differentially altered genes, among these 53 genes, was exhibited in a co-barplot between the ERBB2 neutral and non-neutral cohorts. Alterations in BRCA1, NF1, TP53, MAP2K4, and NCOR1 were most prevalent in the ERBB2 non-neutral cohort [detected in ⩾85% of cases; Figure 1(b) and Supplemental Table 1]. ERBB2, BRCA1, and NF1 lie on chromosome 17q, while TP53, MAP2K4, and NCOR1 are located on chromosome 17p, suggesting a possible molecular pathologic event in ERBB2 non-neutral cases within classical HER2-negative breast cancers.
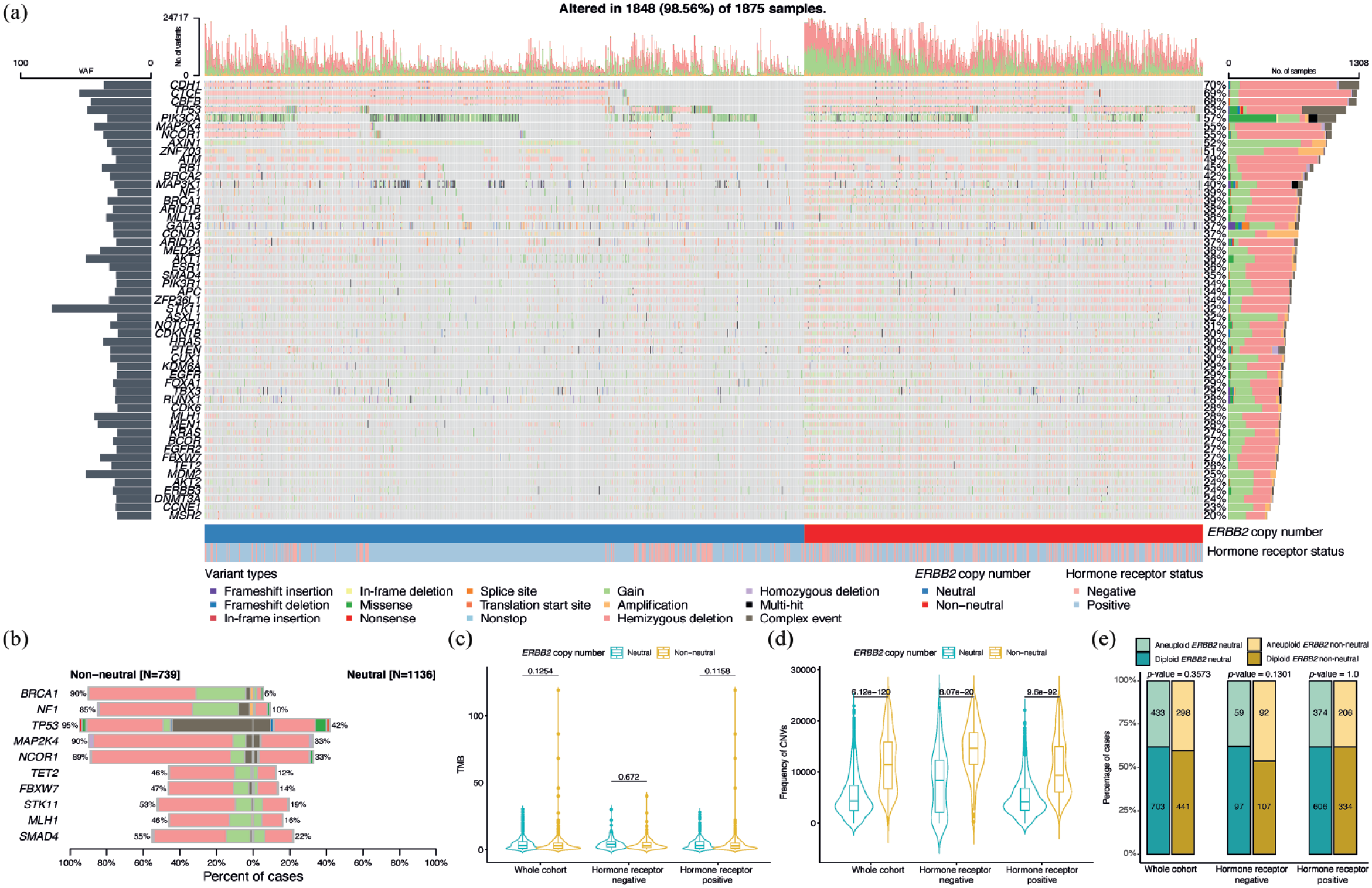
Genomic landscape of study cohort. Somatic mutational and copy number analysis was performed for 1875 cases from both METABRIC (n = 1192) and TCGA (n = 683) datasets. (a) An oncoplot is generated with the 53 breast cancer-related genes, which are comprised of variants with >25% of mean allele frequency. (b) The top 10 differentially altered genes from these 53 genes were shown in a co-barplot between the neutral and non-neutral cohorts. (c) Mann–Whitney–Wilcoxon tests, with Bonferroni correction, were performed for the TMB between patients harboring neutral and non-neutral ERBB2 copy numbers, for the whole cohort as well as within hormone receptor-negative and hormone receptor-positive cohorts, respectively. There is no significant difference in TMB between neutral and non-neutral copy numbers for all three comparisons (median of whole cohort and neutral: 3.00, whole cohort and non-neutral: 2.73, hormone receptor-negative and neutral: 3.89, hormone receptor-negative and non-neutral: 2.82, hormone receptor-positive and neutral: 3.00, hormone receptor-positive and non-neutral: 2.63). (d) As in (c), same statistical tests were performed on the frequency of CNA for all three different comparisons, and non-neutral cases have significantly more CNAs than those of neutral cases in whole cohort, hormone receptor-negative, and hormone receptor-positive cohorts (median of whole cohort and neutral: 4313, whole cohort and non-neutral: 11,360, hormone receptor-negative and neutral: 8350, hormone receptor-negative and non-neutral: 14,618, hormone receptor-positive and neutral: 4128, hormone receptor-positive and non-neutral: 9318). (e) An association analysis was performed between ERBB2 copy number and estimated ploidy from AbsCN-seq, based on the copy number data from METABRIC and a combination of copy number and single nucleotide variant data from TCGA. No significant association was detected between copy number and ploidy for all three comparisons (p values are annotated on top of bar chart).
In the whole cohort, the median TMB of ERBB2 neutral and non-neutral tumors was similar at 3.00 mt/Mb and 2.63 mt/Mb, respectively (p = 0.1254). TMB was also not significantly different within hormone receptor-positive (neutral, 3.00 and non-neutral, 2.63; p = 0.1158) and hormone receptor-negative cohorts [neutral, 3.89 and non-neutral, 2.82; p = 0.6720; Figure 1(c)]. Globally, CNAs across the entire genome were significantly higher in the ERBB2 non-neutral (whole cohort: 11,360; hormone receptor-negative cohort: 14,618; hormone receptor-positive cohort: 9318) compared to neutral (whole cohort: 4313; hormone receptor-negative cohort: 8350; hormone receptor-positive cohort: 4128) cohorts [p < 0.0001; Figure 1(d)]. An association analysis was performed between ERBB2 copy number and estimated ploidy from AbsCN-seq, based on the copy number data from METABRIC and a combination of copy number and single nucleotide variant data from TCGA. No significant association was detected between copy number and ploidy [Figure 1(e)].
Enrichment for alterations in seven genes (BRCA1, NF1, TP53, MAP2K4, NCOR1, TET2, and STK11) was demonstrated in both ERBB2 gain/amplification and heterozygous loss subgroups in the whole cohort, as well as in the hormone receptor-positive cohort (Supplemental Table 2 and Figure 2). Notably, BRCA1 and NF1 CNA were largely in congruence with ERBB2 CNA status, while TP53, MAP2K4, and NCOR1 showed frequent copy number loss regardless. In keeping with these findings from gene-level copy number analysis, arm-level loss in chromosome 17p was most prevalent regardless of ERBB2 gain, amplification, or loss. Whereas chromosome 17q arm-level alterations were largely in tandem with ERBB2 CNA status (Figure 3).

Genomic alteration landscape within ERBB2 CNA subgroups. Additional differential mutational and CNA analysis was performed among three groups of copy number: neutral (GISTIC = 0), gain/amplification (GISTIC = 1 or 2), and heterozygous loss (GISTIC = −1) for (a) the whole cohort, (b) hormone receptor-positive, and (c) hormone receptor-negative cohorts of HER2-negative breast cancers, as depicted in co-barplots. Similar to Figure 1(b), the top 10 differentially altered genes among the 53 genes were shown for each comparison. Seven genes (BRCA1, NF1, TP53, MAP2K4, NCOR1, TET2, and STK11) were shared among ERBB2 gain/amplification and heterozygous loss subgroups in the whole cohort, as well as in the hormone receptor-positive cohort.

Distribution of TCGA arm-level data with respect to ERBB2 copy number in each chromosome. In keeping with findings from gene-level CNA analysis, arm-level loss in chromosome 17p was most prevalent regardless of ERBB2 gain, amplification, or loss. Whereas, chromosome 17q arm-level alterations were largely in tandem with ERBB2 CNA. Only the TCGA dataset is included since no arm-level data are available for the METABRIC dataset.
Gene expression analysis
Differential expression analysis between ERBB2 copy number non-neutral and neutral cases revealed a total of 11,600 and 4612 differentially expressed genes respectively within TCGA and METABRIC datasets, at FDR < 0.01 and abs(fold changes) > 1 [Figure 4(a)]. Among these genes, 1704 (upregulated in non-neutral cases) and 964 (upregulated in neutral cases) were shared between these two datasets, and they were used in the pathway enrichment analysis (with the KEGG database). Specifically, pathways involved in the cell cycle, proteasome, and DNA replication were upregulated in ERBB2 non-neutral cases [Figure 4(b) and (c)]. Using the same filtering criteria of FDR < 0.01 and abs(fold changes) > 1, additional differential expression analysis was performed among three groups of ERBB2 CNA status: ERBB2 neutral (GISTIC = 0), gain/amplification (GISTIC = 1 or 2), and heterozygous loss (GISTIC = −1) for the whole cohort, hormone receptor-positive, and hormone receptor-negative cohorts of HER2-negative breast cancers from TCGA (Supplemental Table 3). Enrichment for cell cycle, proteasome, and DNA replication pathways was noted in both ERBB2 gain/amplification and heterozygous loss subgroups (Supplemental Figure 2A). These findings are mainly attributed to the hormone receptor-positive cohort (Supplemental Figure 2B and C). The METABRIC dataset was not utilized for this analysis since no differentially enriched pathways were significantly detected amongst the three subgroups, due to the smaller number of differentially expressed genes observed with the microarray sequencing platform.
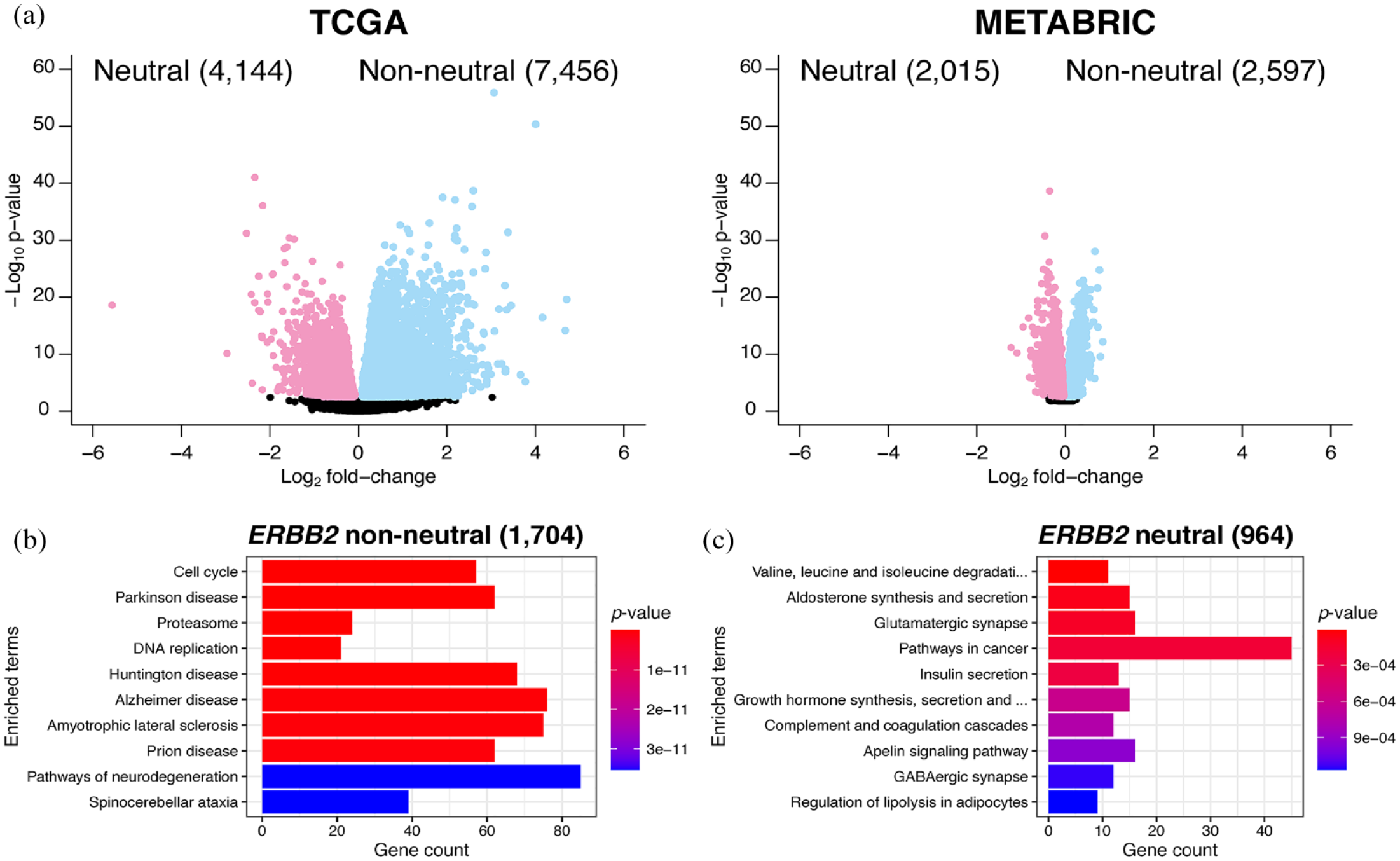
Differential expression analysis between ERBB2 copy number non-neutral and neutral cases for TCGA and METABRIC datasets. (a) Volcano plots showing fold changes and p values of differentially expressed genes in respective datasets, with light blue color representing genes passing the filter of FDR < 0.01 and fold changes >1 (i.e. genes that are upregulated in non-neutral cases), and light red color representing genes passing the filter of FDR <0.01 and fold-changes < (−1; i.e. genes that are upregulated in neutral cases). There are 7456 and 2597 genes that are, respectively, upregulated in TCGA and METABRIC datasets of non-neutral cases, as well as 4144 and 2015 genes in neutral cases; 1704 and 964 genes are shared between these two datasets for non-neutral and neutral cases, respectively. These genes are later used in the pathway enrichment analysis of (b) non-neutral and (c) neutral cases with the KEGG database.
Discussion
The traditional classification of breast cancers based on HER2 status is grounded on protein expression on IHC and/or gene copy number status on ISH.2,3 Approximately 60% of ‘HER2-negative’ breast cancers display low immunohistochemical expression of HER2, including in both hormone receptor-positive and hormone receptor-negative subtypes. Although patients with this group of breast cancers do not classically benefit from HER2-directed therapies, recent clinical trials demonstrated significant improvements in clinical outcomes with trastuzumab deruxtecan, an antibody–drug conjugate composed of an anti-HER2 antibody linked to a topoisomerase I inhibitor payload. 7 In our previous study, tumors harboring HER2 IHC scores of 1+ showed improved prognosis, as did those with ERBB2 CNA neutral status. 19 ERBB2 gene expression levels were correlated with both IHC and CNA scores in the TCGA-METABRIC cohort, with higher gene expression detected in patients with higher IHC and CNA scores, though no association with survival outcomes could be demonstrated based on RNA expression. These results highlight the complexities of subclassifying HER2-negative breast cancers and may imply new approaches for therapeutic intervention.
Current assays for the assessment of HER2 status rely largely on conventional IHC testing, along with ISH testing when deemed equivocal (2+). The accuracy of these scoring methods, particularly for HER2 IHC in the low (0 and 1+) range, has been questioned. Data assessment from College of American Pathologists surveys showed a poor agreement in the evaluation of HER2 IHC scores of 0 and 1+, while a study on 18 pathologists demonstrated a concordance of only 26% between 0 and 1+ scores. Alternative quantitative methods for improved disease classification and optimal patient selection for therapy are therefore urgently warranted. 11 Our results suggest that ERBB2 CNA estimation may offer a feasible approach for this purpose, although the ideal assay remains to be determined. Standard targeted FISH assays feature five ASCO-CAP groupings, with classic HER2-amplified breast cancer defined by HER2/CEP17 ratio ⩾2 and mean HER2 copy number ⩾4. Classic HER2 non-amplified breast cancer is defined by HER2/CEP17 ratio <2 and mean HER2 copy number <4, while the other four groups are defined as negative unless trumped by concurrent IHC scores. 34 The minimum cutoffs representing ‘HER2-low’ will require further studies.
Genome-wide assays such as array comparative genomic hybridization (aCGH), single nucleotide polymorphism arrays, and next-generation sequencing (NGS) methods offer yet additional options for obtaining ERBB2 CNA status, along with other potentially actionable genomic information.35–38 To our knowledge, data comparing contemporary IHC and FISH with various genome-wide assays are not available currently. In addition, the impact of intratumoral heterogeneity will need to be carefully assessed. The question of whether ERBB2 CNA status can predict benefit from novel HER2 ADCs also merits further investigation. ERBB2 heterozygous loss might be associated with upfront resistance to trastuzumab–deruxtecan based on biomarker analyses from the DAISY trial.39,40 Perhaps with the increasing adoption of NGS technologies in the clinical setting, an NGS-based diagnostic approach to ERBB2 CNA status assessment that complements conventional IHC/FISH testing might be feasible in the near future.
Breast cancer is known to have a generally low mutation burden, but is characterized by the high number of CNAs compared to other tumor types. 41 From our analysis, nearly two-fifths of HER2-negative non-metastatic breast cancers harbored ERBB2 CNAs, with heterozygous loss being most commonly observed. Non-neutral ERBB2 CNA status was significantly more prevalent within hormone receptor-negative as compared to hormone receptor-positive breast cancers (56.1% versus 35.5%, respectively) and was previously shown to be an independent prognostic factor for worse relapse-free survival. 19 Interestingly, non-neutral ERBB2 status was associated with a significantly higher genome-wide burden of CNAs, although there were no statistically significant differences in TMB and ploidy. In terms of specific genes, somatic alterations (particularly CNA) in genes on chromosome 17 – TP53, NF1, BRCA1, MAP2K4, and NCOR1 – were more prevalent in the ERBB2 non-neutral group. This observation is related to chromosomal arm-level events and is consistent with previous studies reporting a high frequency of 17p (short arm) losses but with complex combinations of gains and losses within 17q (long arm). 42 Furthermore, our results suggest that ERBB2 CNA, particularly heterozygous loss, is accompanied almost always by concurrent loss of major tumor suppressor genes on chromosome 17.43–46 The implications of these genes as potential contributors driving the inferior prognosis in this group of breast cancers warrant further investigation. In keeping with the association of ERBB2 CNA with worse patient survival outcomes, 19 analysis of differential gene expression revealed upregulated oncogenic pathways such as those involved in cell cycle signaling and DNA replication in the ERBB2 non-neutral group. Our findings are in keeping with a previous report demonstrating high somatic CNA burden with worse survival in patients with breast cancer, 47 as well as in prostate, endometrial, renal clear cell, thyroid, and colorectal cancer. 48 In other studies, higher somatic CNA burden was also associated with poorer responses to immunotherapy in melanoma,49,50 non-small-cell lung cancer, 51 gastrointestinal cancer, 52 and other metastatic cancers. 41
The current findings are limited by the availability of datasets used for the in silico analysis. We focused on non-metastatic, stage I–III HER2-negative breast cancers to follow up on our previous findings of inferior survival in ERBB2 non-neutral breast cancer in the non-metastatic setting. 19 Specific results pertaining to breast cancer subtypes, particularly hormone receptor-negative cases, will require validation in larger cohorts. Given that the new classification of HER2-low breast cancers impacts treatment decisions only in the metastatic setting currently, additional investigation in metastatic breast cancers may be required.
Conclusion
In conclusion, our work highlights the complexity of HER2-negative breast cancers in the context of their copy number status. Apart from differences in survival outcomes, the classification of HER2-negative breast cancers according to ERBB2 CNA status reveals distinct patterns of genomic aberrations especially in chromosome 17 genes. The biological and therapeutic implications of CNA classification merit further research.
Supplemental Material
sj-docx-1-tam-10.1177_17588359231206259 – Supplemental material for Classification of HER2-negative breast cancers by ERBB2 copy number alteration status reveals molecular differences associated with chromosome 17 gene aberrations
Supplemental material, sj-docx-1-tam-10.1177_17588359231206259 for Classification of HER2-negative breast cancers by ERBB2 copy number alteration status reveals molecular differences associated with chromosome 17 gene aberrations by Jui Wan Loh, Abner Herbert Lim, Jason Yongsheng Chan and Yoon-Sim Yap in Therapeutic Advances in Medical Oncology
Supplemental Material
sj-xlsx-2-tam-10.1177_17588359231206259 – Supplemental material for Classification of HER2-negative breast cancers by ERBB2 copy number alteration status reveals molecular differences associated with chromosome 17 gene aberrations
Supplemental material, sj-xlsx-2-tam-10.1177_17588359231206259 for Classification of HER2-negative breast cancers by ERBB2 copy number alteration status reveals molecular differences associated with chromosome 17 gene aberrations by Jui Wan Loh, Abner Herbert Lim, Jason Yongsheng Chan and Yoon-Sim Yap in Therapeutic Advances in Medical Oncology
Footnotes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
