Abstract
Background:
Circulating tumor DNA (ctDNA) has emerged as a potential biomarker for monitoring early non-small cell lung cancer (ENSCLC), particularly after radical surgery. However, the prognostic value of postoperative ctDNA is still being investigated due to the small sample size and heterogeneity of patients with ENSCLC in current trials. Moreover, the potential clinical utility of ctDNA assessment for administering adjuvant therapy (AT) in patients with ENSCLC is also an important area of active research.
Objectives:
We aimed to identify the prognostic value of postoperative ctDNA detection in ENSCLC patients with stages I–III.
Design:
This study type is a systematic review and meta-analysis.
Data sources and methods:
We conducted a search in the Cochrane Library, Embase, PubMed, and ScienceDirect for prospective or retrospective investigations involving patients with ENSCLC, gathering outcomes based on predefined end points. The literature review adhered to the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) guidelines, and the Newcastle–Ottawa scale was employed to carry out a quality evaluation of the included studies. The primary end point of the study was to evaluate the association of ctDNA status in two time points (within 1 month after surgery and long-term postoperative monitoring with more than 3 months) with relapse-free survival (RFS) and overall survival (OS). In addition, the study investigated the role of ctDNA in predicting the response to AT. The secondary end points of the study were to determine the impact of ctDNA on RFS and OS in different subgroups of ENSCLC patients based on pathological subtypes and TNM staging.
Results:
In total, 2149 studies were screened, and 11 studies met the inclusion criteria for the analysis. The presence of ctDNA within 1 month after surgery as well as long-term postoperative ctDNA were both associated with poorer RFS [hazard ratio (HR) = 4.43; 95% CI: 3.23–6.07 and HR = 7.99; 95% CI: 3.28–19.44, respectively] and worse OS (HR = 5.07; 95% CI: 2.80–9.19 and HR = 7.49; 95% CI: 3.42–16.43, respectively). Most subgroup analyses yielded similar results. Moreover, ctDNA-positive patients could acquire survival benefits from AT (HR = 0.30; 95% CI: 0.16–0.54), while ctDNA-negative patients that received AT did not show significant improvement in RFS (HR = 1.18; 95% CI: 0.67–2.09).
Conclusion:
The postoperative ctDNA assessment is a promising approach to stratify the risk of relapse and death in ENSCLC patients. Our data suggest that patients with negative ctDNA in the postoperative setting may not benefit from AT, which warrants further investigation. This finding, if validated in prospective trials with a larger sample size, could aid in better-individualized treatment for patients and avoid potential side effects of AT.
Registration:
This study was designed in accordance with PRISMA and registered with PROSPERO (CRD42022311615).
Keywords
Introduction
Lung cancer is a widespread malignancy globally, responsible for over 200,000 diagnoses and 150,000 fatalities annually.1,2 It can be categorized into non-small cell lung carcinoma (NSCLC) and small cell lung carcinoma, with the former constituting 80–85% of all lung carcinomas. 3 Although surgical intervention remains the favored treatment approach for early NSCLC, recurrence following curative treatment poses a considerable obstacle that may result in adverse outcomes due to minimal residual disease (MRD). 4
Although adjuvant chemotherapy has been shown to improve survival rates, the benefits are limited, with some studies indicating only a 5% increase in the 5-year overall survival (OS) rate for patients receiving postoperative adjuvant treatment. 5 Consequently, there is an urgent need for specific and sensitive markers that can effectively monitor postoperative recurrence of early nonsmall cell lung cancer (ENSCLC), which can help predict recurrence and prevent metastasis, ultimately leading to improved prognosis.
MRD is characterized by the presence of either isolated tumor cells or circulating tumor cells without any clinical or radiological evidence of metastasis. MRD remains in the body even after radical therapy has been performed to resect the primary tumor. 6 Studies have shown that detectable MRD is associated with occult residual disease and an increased risk of recurrence. 4 Circulating tumor DNA (ctDNA) refers to the nucleic acids derived from the tumor that is present in the cell-free DNA fraction of a patient’s blood plasma. 7 According to several studies, ctDNA has great clinical potential as a noninvasive biomarker to monitor MRD of cancers.3,6–8 Increasing evidence showed that positive ctDNA after curative treatment may predict a higher risk for relapse. Methods for detecting ctDNA include polymerase chain reaction (PCR)-based testing and next-generation sequencing (NGS). 7 However, there is currently no gold standard for the detection of ctDNA in ENSCLC.
Interestingly, previous studies have investigated the potential of ctDNA as a predictive marker in patients with resectable ENSCLC. Several prospective studies have reported a significant association between perioperative ctDNA positivity and shorter relapse-free survival (RFS) as well as poorer OS.9–12 However, a definitive conclusion regarding the clinical utility of ctDNA in ENSCLC cannot be drawn from these studies due to the small sample size and heterogeneity of patients. Furthermore, determining the optimal time point for postoperative ctDNA assessment remains a significant challenge. Gale et al. 8 reported that the detection of ctDNA within the landmark time frame of 2 weeks to 4 months after surgery was associated with shorter RFS, while detection within 1–3 days after surgery was not found to be associated with recurrence. In contrast, the DYNAMIC study by Chen et al. 13 found that undetectable ctDNA on the third day after surgery was correlated with longer RFS. Some studies have suggested that longitudinal monitoring after surgery may be beneficial for the early detection of relapse and enabling early intervention.8,12
In our work, we conducted a systematic review and meta-analysis of studies in ENSCLC patients after surgery, defining the prognostic role of ctDNA in ENSCLC patients and also exploring an optimal detection period with efficient utility and reliability.
Methods
Based on the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) guidelines, 14 a systematic review of the studies and meta-analysis of the postoperative ENSCLC patients was conducted to identify the relationship between postoperative ctDNA monitoring and survival outcomes, specifically RFS and OS. The study also determined the predictive role of ctDNA monitoring in the efficacy of adjuvant therapy (AT). This study was registered in the international prospective register of systematic reviews (PROSPERO 2022 CRD42022311615).
Data retrieval
We reviewed studies from electronic databases, including Cochrane Library, Embase, PubMed, and ScienceDirect. The search was conducted up to June 30, 2022, and detailed search strategy is listed in Supplemental File 1. Two authors, Kaibo Guo and Jiamin Lu, performed the systematic review independently and included published articles that met the criteria. In the case of any disagreements, another author, Song Zheng, was consulted to help resolve the issue.
Study selection
The studies included in this analysis satisfied the following criteria: (1) observational studies (prospective or retrospective), (2) studies including ENSCLC patients (I–III stage) who underwent radical resection, (3) studies recording postoperative ctDNA status as determined by NGS and containing corresponding outcome data such as RFS, disease-free survival (DFS), progression-free survival (PFS), or OS. Exclusion criteria included: (1) nonoperative patients or stage IV patients were excluded through reading full text or analyzing raw data and (2) the authors, clinical trial number, and institutions were examined to avoid repetition of studies.
The Newcastle–Ottawa scale (NOS) was employed to assess the quality of the studies included. Within the NOS, a total of nine points could be evaluated, encompassing patient selection (four points), outcome assessment (three points), and cohort comparability (two points). 15
Data extraction
The following information was collected from the manuscripts or raw data of the included studies: author names, country of origin, year of publication, study description, cancer subtype, stage distribution, treatments in the perioperative period, number of participants, the method for ctDNA analysis, ctDNA-positive criteria, survival outcomes, and median follow-up time. To provide a comprehensive estimate of RFS, we included studies that reported outcome measures such as RFS, DFS, and PFS.
In an effort to calculate the hazard ratio (HR) and corresponding 95% confidence interval based on the univariate and multivariate Cox regression analysis, we obtained all the feasible raw data. If the study did not include raw data, HR and 95% CI would be obtained by collecting the data from full text or extracting the information from the survival plot using Engauge Digitizer software.
Main outcomes
The primary end points of this study were to identify the significance of postoperative ctDNA status on RFS and OS for ENSCLC patients, as well as to explore the effectiveness of using postoperative ctDNA for predicting AT outcomes after surgery. This meta-analysis examined two modes of postoperative ctDNA monitoring: short-term monitoring (within 1 month after surgery) and long-term monitoring (more than 3 months after surgery). Long-term ctDNA monitoring refers to the final ctDNA result obtained during the monitoring period.
The secondary end points of this study were: (1) the effects of postoperative ctDNA on RFS and OS in different subgroups, including lung adenocarcinoma (LUAD) and non-lung adenocarcinoma (non-LUAD) patients. (2) The effects of postoperative ctDNA on RFS and OS in subgroups, including I–II-stage and III-stage patients with NSCLC.
Data synthesis
In all analyses, we conducted heterogeneity evaluation and reported the corresponding I 2 value. We pooled HRs by employing both fixed and random effects models, irrespective of the degree of heterogeneity. Typically, heterogeneity is considered when I 2 > 50% or p-value <0.05.
The fixed effects model used the inverse variance method to calculate the overall HR. In contrast, the random effects model incorporates heterogeneity by applying the DerSimonian–Laird method. When I 2 ⩽ 50%, it is advisable to use the fixed effects model to pool HRs. However, if I 2 > 50%, the random effects model may be a better choice. Further, if high heterogeneity was observed between included studies, sensitivity analysis was conducted to provide a comprehensive explanation of the results. p-Values for the pooled HRs were not reported. Funnel plot analysis and Egger’s test were performed to detect any potential publication bias. If publication bias was detected, the trim-and-fill method was conducted to normalize the outcomes. All analyses in this study were performed by R statistical software version 4.2.0 (R packages survival, survminer, meta).
Results
Literature screening
In total, 2149 articles were searched and obtained from the PubMed, Embase, Cochrane, and ScienceDirect databases, and the process of literature selection is illustrated in Figure 1. In total, we included 11 studies comprising 1104 patients with ENSCLC in our analysis.

Literature search and study selection according to PRISMA 2020 flow diagram for systematic reviews.
Study characteristics
All 11 studies included in the meta-analysis were prospective observational studies with postoperative ctDNA detection at multiple time points.6,8–13,16–19 Ten of these studies had ctDNA detection performed within 1 month after surgery,6,8–13,17–19 while eight studies performed long-term ctDNA detection after surgery.6,8,9,11,12,16,18,19 Four studies enrolled patients with postoperative positive ctDNA before AT,6,10,13,17 while five studies included patients with negative ctDNA status before AT.6,10,12,13,17 Of the included studies, three were conducted in the European population,8,12,16 while eight studies were conducted in the Asian population.6,9–11,13,17–19 Regarding the ENSCLC subtype, one study included only patients with LUAD, 18 while 10 studies enrolled patients with all ENSCLC subtypes.6,8–13,16,17,19 In terms of stage distribution, 10 studies included patients with stages I–III,6,8–13,16,17,19 while one study included patients with stage I. 18
Table 1 summarizes the characteristics of participants and interventions in the included studies, while Table 2 describes details on the ctDNA collections, survival end points characteristics, and median follow-up time. The definitions of RFS and OS in the included studies can be found in Supplemental Table 1. NOS-specific scores for each study can be found in Supplemental Table 2.
Study characteristics of the studies included in the systematic review and meta-analysis.

AT, adjuvant therapy; ChiCTR, Chinese Clinical Trial Registry; CT, chemotherapy; DRKS, Deutsches Register Klinischer Studien (German register of clinical trials); ICB, immune checkpoint blockade; ICI, immune checkpoint inhibitor; LUAD, lung adenocarcinoma; LUSC, lung squamous carcinoma; NA, not available; NAT, neoadjuvant therapy; NCT, national clinical trial; NOS, Newcastle–Ottawa scale, NSCLC, nonsmall cell lung cancer; RT, radiation therapy; TKI, tyrosine kinase inhibitor; TT, targeted therapy.
Results of survival and ctDNA collections characteristics.

ATG-seq, automated triple groom sequencing; cSMART, circulating single-molecule amplification and resequencing technology; ctDNA, circulating tumor DNA; DFS, disease-free survival; MAF, minor allele frequency; Multiplex-PCR, multiplex polymerase chain reaction; NA, not available, NGS, next-generation sequencing; OS, overall survival; PFS: progress-free survival, qPCR, quantitative real-time polymerase chain reaction; RFS, relapse-free survival; VAF, variant allele frequency.
Primary end points: Association of ctDNA levels with RFS and OS within 1 month after surgery
Ten studies (n = 941) collected data on postoperative ctDNA detection within 1 month and corresponding RFS,6,8–13,16,17,19 while five studies (n = 277) analyzed OS and postoperative ctDNA.8,9,11,13,16 Among the patients included in the RFS analysis, the presence of ctDNA within 1 month after surgery was identified in 128/941 (13.6%) patients, while in the OS analysis, ctDNA was detected in 59/277 (21.3%) patients. In summary, the presence of postoperative ctDNA within 1 month after surgery was significantly associated with shorter RFS (HR = 4.43; 95% CI: 3.23–6.07) as shown in Figure 2(a) and poorer OS (HR = 5.07; 95% CI: 2.80–9.19) as shown in Figure 2(b). These effects were observed consistently in both Asian and European populations. Due to low heterogeneity, the fixed effects model was used to estimate this analysis.
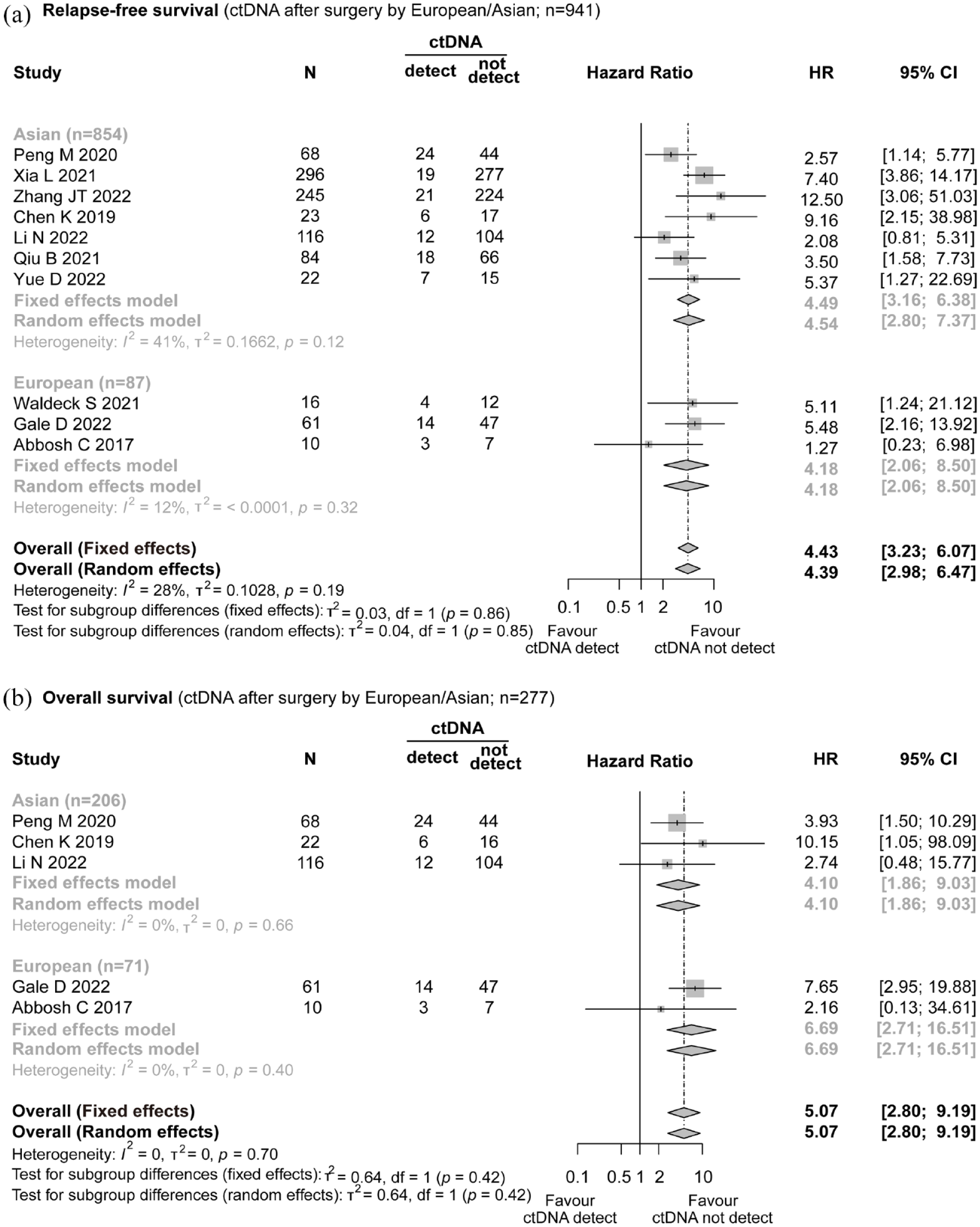
Forest plot illustrating the impact of ctDNA presence within 1 month after surgery on relapse-free survival and overall survival in patients with ENSCLC. (a) Relapse-free survival (ctDNA after surgery by European/Asian; n = 941). (b) Overall survival (ctDNA after surgery by European/Asian; n = 277).
Long-term postoperative ctDNA monitoring
Eight studies (n = 578) collected data on longitudinal ctDNA for more than 3 months and analyzed corresponding RFS,6,8,9,11,12,16,18,19 while three studies (n = 205) analyzed OS.8,9,11 Among the patients included in the RFS analysis, the presence of long-term postoperative ctDNA was identified in 189/578 (32.7%) patients, while in the OS analysis, ctDNA was detected in 51/205 (24.9%) patients. The presence of long-term postoperative ctDNA was significantly associated with poorer RFS (HR = 7.99; 95% CI: 3.28–19.44; as shown in Figure 3(a)) and OS (HR = 7.49; 95% CI: 3.42–16.43; as shown in Figure 3(b)). Due to high heterogeneity, the random effects model was used to estimate the effects, and a sensitivity analysis showed that the results were stable (Supplemental Figure 1).
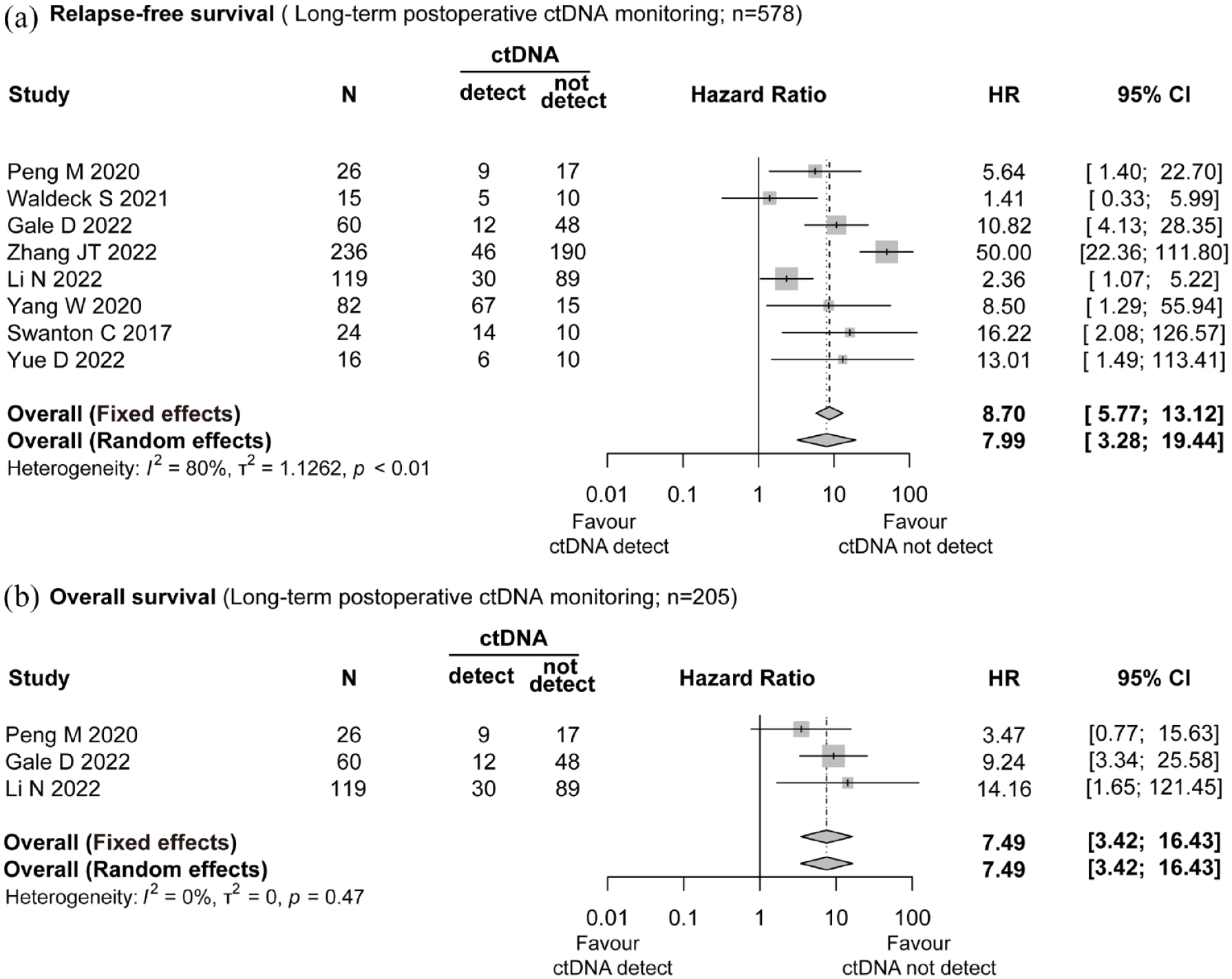
Forest plot illustrating the impact of long-term postoperative ctDNA monitoring on relapse-free survival and overall survival in patients with ENSCLC. (a) Relapse-free survival (long-term postoperative ctDNA monitoring; n = 578). (b) Overall survival (long-term postoperative ctDNA monitoring; n = 205).
Prognostic benefits of postoperative adjuvant therapy based on different ctDNA status
Four studies (n = 75) provided data on patients with postoperative positive ctDNA status within 1 month after surgery and who received AT,6,10,13,17 while five studies (n = 443) analyzed patients with postoperative negative ctDNA status within 1 month after surgery and who did not receive AT.6,10,12,13,17 Among patients with positive ctDNA, 28/43 (65.1%) patients who were treated with AT experienced recurrence, while 30/32 (93.8%) patients who were not treated with AT relapsed. In patients with negative ctDNA, the recurrence rate was 55/209 (26.3%) in the AT group, and 29/234 (12.4%) experienced relapse when the patients did not receive the AT.
Postoperative AT can significantly improve the RFS of patients with positive ctDNA (HR = 0.30; 95% CI: 0.16–0.54; as shown in Figure 4(a)); however, it may not provide the same benefit to patients with negative ctDNA (HR = 1.06; 95% CI: 0.59–1.91; as shown in Figure 4(b); trim-and-fill method, HR = 1.18; 95% CI: 0.67–2.09; Supplemental Figure 2). Due to insufficient data, it was not possible to conduct an analysis of the OS.

Forest plot illustrating the relationship between the efficacy of AT and (a) positive ctDNA and (b) negative ctDNA.
Secondary end points
The relationship between pathological types and ctDNA detection after surgery
The effects of postoperative ctDNA detection on the prognosis of ENSCLC patients with different pathological types are shown in Supplemental Table 3. Of the LUAD patients included in the analysis of RFS, 53/420 (12.6%) patients showed the presence of ctDNA within 1 month after surgery, and 30/114 (26.3%) patients showed long-term postoperative ctDNA detection. In non-LUAD patients, the detection rates were higher, with 38/136 (27.9%) patients showing the presence of ctDNA within 1 month after surgery and 26/51 (51.0%) patients showing long-term postoperative ctDNA detection. The OS analysis yielded similar results.
In patients with LUAD, the results were similar to primary end points. In this regard, the presence of postoperative ctDNA detection within 1 month was significantly associated with poorer RFS (HR = 6.3; 95% CI: 4.03–9.86) and OS (HR = 5.72; 95% CI: 2.54–12.88). Similarly, the existence of long-term postoperative ctDNA also led to worse RFS (HR = 5.28; 95% CI: 2.86–9.73) and OS (HR = 6.21; 95% CI: 2.42–15.94) for these patients.
For patients with non-LUAD, the presence of postoperative ctDNA detection within 1 month was also associated with a statistically significant worse RFS (HR = 3.56; 95% CI: 2.06–6.16) and OS (HR = 2.9; 95% CI: 1.10–7.64). Likewise, the occurrence of long-term postoperative ctDNA led to poorer RFS (HR = 4.1; 95% CI: 1.80–9.34) for these patients. The analysis of OS was not conducted in this part of the study due to the inclusion of only one study.
The relationship between different staging and postoperative ctDNA detection
The effects of postoperative ctDNA detection on the prognosis of ENSCLC patients with different staging are shown in Supplemental Table 4. In patients with stages I–II, ctDNA within 1 month after surgery and long-term postoperative ctDNA detection were 45/453 (9.9%) and 41/145 (28.3%), respectively. However, in patients with stage III, the detection rates were higher, with 41/89 (46.1%) and 14/17 (82.4%) being positive for ctDNA within 1 month after surgery and long-term postoperative ctDNA, respectively. In the analysis of OS, patients with stage III had higher detection rates of short-term or long-term postoperative ctDNA compared to those with stages I–II.
In patients with stages I–II, the presence of postoperative ctDNA detection within 1 month was correlated with a statistically significant poorer RFS (HR = 6.12; 95% CI: 2.73–13.71) and OS (HR = 5.12; 95% CI: 2.29–11.46). Similarly, presence of long-term postoperative ctDNA resulted in poorer RFS (HR = 6.91; 95% CI: 3.69–12.94) and OS (HR = 10.14; 95% CI: 3.52–29.22) for these patients. Thus, patients with stages I–II showed similar results to the findings in the previous sections.
For patients with stage III, the presence of postoperative ctDNA detection within 1 month was also associated with a statistically significantly worse RFS (HR = 2.66; 95% CI: 1.58–4.46) and OS (HR = 5.26; 95% CI: 2.34–11.80). However, the existence of long-term postoperative ctDNA did not result in poorer RFS (HR = 1.81; 95% CI: 0.75–4.39) and OS (HR = 2.89; 95% CI: 0.55–15.16) outcomes for these patients.
Quality evaluation and risk of bias assessment
According to NOS, all included studies scored between 6 and 9. Risks of publication bias were presented as funnel plots in Figure 5 and Supplemental Figure 3. Results indicated some potential publication bias in the RFS analysis of the therapeutic effects of AT on patients with negative ctDNA (p = 0.03). Accordingly, we performed the trim-and-fill method and found that the conclusion is robust, as shown in Supplemental Figure 4.

Funnel plots demonstrating the publication bias in (a andb) ctDNA presence within 1 month after surgery and (c and d) long-term postoperative ctDNA monitoring.
Discussion
Although radical surgery can improve outcomes of ENSCLC patients with relatively better survival rates, some of them still experience relapse. 4 Therefore, adjuvant therapy is commonly administered to all postoperative patients to eliminate residual tumors and reduce the risk of cancer recurrence. 20 This underscores the importance of identifying additional novel biomarkers that can more accurately predict the risk of recurrence and aid in the decision to use adjuvant therapy. 21
Our meta-analysis focused on ENSCLC patients who underwent radical surgery and highlights that postoperative ctDNA detection has a promising potential role as a prognostic biomarker for these patients. The presence of ctDNA within 1 month after surgery is strongly associated with an increased risk of relapse in patients with ENSCLC, regardless of the ethnicity of the participants. Subgroup analysis based on ENSCLC subtype or TNM staging was performed for the included patients, and the results being consistent further supports the reliability of the finding. This suggests that ctDNA detection might be a universal biomarker for predicting relapse in ENSCLC patients.
Significantly, the detection rates of ctDNA were reduced in patients with LUAD or stages I–II, and the presence of positive ctDNA had a more substantial influence on these patients’ RFS. This could be attributed to two factors: firstly, the participants with LUAD or stages I–II represented a large proportion of the included studies. Secondly, the non-LUAD group primarily consisted of squamous cell carcinoma, and the release of ctDNA may be associated with a higher prevalence of intratumoral necrosis.16,22 Along the same line, the lower release of ctDNA in LUAD may be related to its histopathological characteristics, such as the lower incidence of necrosis and more solid tumor structure compared to squamous cell carcinoma. Additionally, tumors with stages I–II are generally considered to have less aggressive biology compared to stage III tumors. 23
The efficacy of AT in ENSCLC patients remains uncertain, highlighting the need for further research in this area. 24 To investigate this, we analyzed the efficacy of AT in two subgroups of patients: ctDNA-positive and ctDNA-negative patients within 1 month of surgery. Interestingly, our results revealed no significant correlation between AT usage and improved RFS in ctDNA-negative patients. On the contrary, ctDNA-positive patients who did not receive AT exhibited worse RFS, indicating that AT may be beneficial for this patient population. The limited sample size of ctDNA-positive patients (75 positive patients in four studies) in the studies included in this meta-analysis is a potential limitation that should be addressed by further research.
In our analysis, long-term detection of ctDNA after surgery demonstrated greater clinical significance compared to short-term detection.6,8,9,11,12,16,18,19 Although the studies exhibited some heterogeneity, sensitivity analysis revealed consistent and robust results. Further, the association between long-term detection of ctDNA and poor RFS was consistent regardless of histological subtypes (LUAD versus non-LUAD). This suggests that ctDNA detection may be a useful prognostic biomarker for all ENSCLC patients, regardless of tumor type. Furthermore, long-term detection of positive ctDNA was linked to worse RFS in patients with the I–II stage, while the association with III-stage patients was not statistically significant. Nevertheless, it is important to note that the III-stage patient subgroup included only 31 patients, and further studies are necessary to confirm this correlation.
Our meta-analysis comprehensively analyzed several studies investigating the correlation between ctDNA and OS in ENSCLC patients.8,9,11–13,16 Our pooled analysis revealed that two detection periods, including postoperative ctDNA within 1 month after surgery and long-term postoperative ctDNA, were both strongly correlated with the risk of relapse as well as worse OS. These findings indicate that ctDNA detection may provide valuable information for the disease management of patients with ENSCLC who have undergone radical surgery by enabling more personalized and effective treatment approaches.
Nevertheless, this meta-analysis has some limitations, mainly related to the potential heterogeneity among the included studies. Firstly, although all studies used next-generation sequencing (NGS) technology to assess ctDNA status, variations existed in the reference gene panel and detection methods among the studies. For instance, one study used a 769 gene NGS panel, 10 while another study employed a 127 gene cSMART (NGS & Multiplex-PCR) panel. 11 The impact of the detection and analysis method on the prognostic roles of ctDNA is still unclear, emphasizing the need for a coordinated effort to establish a standardized testing protocol. Secondly, we considered ctDNA as a binary variable (detected/undetected) for analysis because this information was most easily available in most included studies. The use of a binary variable for ctDNA detection could have potentially led to the loss of information. The majority of studies used variant allele frequency (VAF) or minor allele frequency (MAF) to distinguish ctDNA status, and the threshold value for defining ctDNA positivity varied across different studies. Furthermore, we were unable to account for the relative amount of ctDNA during postoperative follow-up, which may have influenced patient outcomes. Thirdly, the timing and frequency of ctDNA assessment varied among the studies included in our meta-analysis, which may have introduced some heterogeneity in our results. We focused on two distinct time points for ctDNA monitoring: within 1 month after surgery and more than 3 months after surgery. We noted that the latter group was more heterogeneous, as it included studies with varying follow-up times. In addition, the lack of information on adjuvant therapy may affect the prognostic value of ctDNA in patients with long-term surveillance. Fourthly, we observed a lack of harmonization in the end points among the included studies. To address this, we used a composite end point RFS to summarize data on RFS, DFS, and PFS from each study (Table 2 and Supplemental Table 1) in the analysis of outcomes. However, we acknowledge that while these concepts are closely related, they are not completely identical. Finally, it should be noted that the sample size of our study was relatively small in two specific areas. Firstly, when studying the effects of adjuvant therapy in ctDNA-positive patients and secondly, when exploring the impact of ctDNA on ENSCLC patients with subgroups, particularly those in stage III. As a result, further multicenter studies with larger sample sizes are necessary to confirm the findings in these areas.
In conclusion, this work suggests that assessing ctDNA, either within 1 month after surgery or in long-term postoperative follow-up, may be a valuable tool to identify the risk of relapse and death in patients with ENSCLC who have undergone radical surgery. As for tailoring the treatment of patients, our findings suggest that patients with negative ctDNA in the postoperative setting may not benefit from adjuvant therapy. However, patients with positive ctDNA need to actively undergo adjuvant therapy, as it could potentially improve patient outcomes. Despite several limitations mainly derived from the heterogeneity of the included studies, this meta-analysis highlights the application of ctDNA as a tool to guide treatment in postoperative patients with ENSCLC. However, further research is needed to validate and refine the findings of this meta-analysis in prospective clinical trials.
Supplemental Material
sj-docx-1-tam-10.1177_17588359231177008 – Supplemental material for Prognostic value of postoperative ctDNA detection in patients with early non-small cell lung cancer: a systematic review and meta-analysis
Supplemental material, sj-docx-1-tam-10.1177_17588359231177008 for Prognostic value of postoperative ctDNA detection in patients with early non-small cell lung cancer: a systematic review and meta-analysis by Kaibo Guo, Jiamin Lu, Yidan Lou and Song Zheng in Therapeutic Advances in Medical Oncology
Supplemental Material
sj-tif-3-tam-10.1177_17588359231177008 – Supplemental material for Prognostic value of postoperative ctDNA detection in patients with early non-small cell lung cancer: a systematic review and meta-analysis
Supplemental material, sj-tif-3-tam-10.1177_17588359231177008 for Prognostic value of postoperative ctDNA detection in patients with early non-small cell lung cancer: a systematic review and meta-analysis by Kaibo Guo, Jiamin Lu, Yidan Lou and Song Zheng in Therapeutic Advances in Medical Oncology
Supplemental Material
sj-tif-4-tam-10.1177_17588359231177008 – Supplemental material for Prognostic value of postoperative ctDNA detection in patients with early non-small cell lung cancer: a systematic review and meta-analysis
Supplemental material, sj-tif-4-tam-10.1177_17588359231177008 for Prognostic value of postoperative ctDNA detection in patients with early non-small cell lung cancer: a systematic review and meta-analysis by Kaibo Guo, Jiamin Lu, Yidan Lou and Song Zheng in Therapeutic Advances in Medical Oncology
Supplemental Material
sj-tif-5-tam-10.1177_17588359231177008 – Supplemental material for Prognostic value of postoperative ctDNA detection in patients with early non-small cell lung cancer: a systematic review and meta-analysis
Supplemental material, sj-tif-5-tam-10.1177_17588359231177008 for Prognostic value of postoperative ctDNA detection in patients with early non-small cell lung cancer: a systematic review and meta-analysis by Kaibo Guo, Jiamin Lu, Yidan Lou and Song Zheng in Therapeutic Advances in Medical Oncology
Supplemental Material
sj-tif-6-tam-10.1177_17588359231177008 – Supplemental material for Prognostic value of postoperative ctDNA detection in patients with early non-small cell lung cancer: a systematic review and meta-analysis
Supplemental material, sj-tif-6-tam-10.1177_17588359231177008 for Prognostic value of postoperative ctDNA detection in patients with early non-small cell lung cancer: a systematic review and meta-analysis by Kaibo Guo, Jiamin Lu, Yidan Lou and Song Zheng in Therapeutic Advances in Medical Oncology
Supplemental Material
sj-xls-2-tam-10.1177_17588359231177008 – Supplemental material for Prognostic value of postoperative ctDNA detection in patients with early non-small cell lung cancer: a systematic review and meta-analysis
Supplemental material, sj-xls-2-tam-10.1177_17588359231177008 for Prognostic value of postoperative ctDNA detection in patients with early non-small cell lung cancer: a systematic review and meta-analysis by Kaibo Guo, Jiamin Lu, Yidan Lou and Song Zheng in Therapeutic Advances in Medical Oncology
Footnotes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
