Abstract
Lung cancer has a high morbidity and mortality rate, and affected patients have a poor prognosis and low survival. The therapeutic approaches for lung cancer treatment, including surgery, radiotherapy, and chemotherapy, are not completely effective, due to late diagnosis. Although the identification of genetic drivers has contributed to the improvement of lung cancer clinical management, the discovery of new diagnostic and prognostic tools remains a critical issue. Liquid biopsy (LB) represents a minimally invasive approach and practical alternative source to investigate tumor-derived alterations and to facilitate the selection of targeted therapies. LB allows for the testing of different analytes such as circulating tumor cells, extracellular vesicles (EVs), tumor-educated platelets, and cell-free nucleic acids including DNAs, RNAs, and noncoding RNAs (ncRNAs). Several regulatory factors control the key cellular oncogenic pathways involved in cancers. ncRNAs have a wide range of regulatory effects in lung cancers. This review focuses on emerging regulatory ncRNAs, freely circulating in body fluids or shuttled by EVs, such as circular-RNAs, small nucleolar-RNAs, small nuclear-RNAs, and piwi-RNAs, as new biomarkers for early detection, prognosis, and monitoring of therapeutic strategy of lung cancer treatment.
Introduction
Lung cancer (LC) is one of the primary causes of mortality worldwide, with about 2,200,000 new cases and around 1,800,000 deaths estimated in the world in 2020 by the Global Cancer Observatory (https://gco.iarc.fr). LC originates from basal epithelial cells of the lung and is classified into two main categories: non-small-cell lung cancer (NSCLC) and small cell lung cancer (SCLC). 1 NSCLC is the most prominent form of LC; about 85% of all LC are NSCLC, this subtype represents one of the most frequent causes of cancer-related death.2,3 The prognoses and therapeutic strategies for NSCLC depend on the disease stage at diagnosis. While the surgery remains the standard of care for NSCLC, 4 it is often unsuccessful in patients with metastasis. Unfortunately, NSCLC patients are mostly diagnosed at advanced stages since they are frequently asymptomatic at early stages;5,6 thus, finding new biomarkers for early diagnosis is an urgent need.
The understanding of NSCLC pathogenesis has improved through the identification of activating mutations and amplifications of oncogenes, including Kirsten rat sarcoma virus (KRAS),7,8 epidermal growth factor receptor (EGFR), 9 and inactivating mutations in tumor suppressive genes, such as p53. 10 Nowadays, it is estimated that up to 69% of advanced NSCLC patients carry druggable genetic alterations in different genes, such as EGFR, KRAS, anaplastic lymphoma kinase (ALK), human epidermal growth factor receptor 2 (HER2), c-Ros oncogene 1 (ROS1), or V-raf murine sarcoma oncogene homolog B1 (BRAF). 11 The list of exploitable targets is rapidly growing and includes eight different genes with approved targeted therapies, including EGFR, ALK, ROS1, MET, BRAF, KRAS, NTRK, and RET. 12 Despite initial responses to targeted therapies, the clinical benefit of these agents is typically limited and virtually all patients progress through the acquisition of multiple on-target and/or off-target resistance mechanisms. 13 Therefore, it is important to identify new therapeutic targets for NSCLC. 14 In the era of precision medicine, NSCLC clinical management can benefit from a new tool, liquid biopsy (LB). 15
LB is a minimally invasive procedure, alternative, or complementary to tissue biopsy, and utilizes body fluids, such as blood, plasma, serum, saliva, or urine to collect elements mirroring the patient-specific disease state. 12 LB allows the analysis of circulating biomarkers that reflect systemic tumor burden and represent intra-tumoral heterogeneity. LB includes the analysis of circulating tumor cells (CTC), extracellular vesicles (EVs), tumor-educated platelets (TEPs), and cell-free nucleic acids (cf-NAs) such as circulating tumor DNA (ctDNA) and circulating tumor RNA (ctRNA). The combination of circulating factors transported in the bloodstream has been referred to as ‘tumor circulome’. In the near future, the diagnosis based on cf-NAs could represent a new route in personalized medicine.16,17 Currently, CTCs and ctDNA are the only components of LB approved in clinical practice by the Food and Drug Administration, as biomarkers for diagnosis, prognosis, and monitoring treatment response.
LB is an important source of cf-NAs and provides information about cancer cell mutation profiles, while cell-free noncoding RNAs (ncRNAs) are promising biomarker candidates in the diagnosis and prognosis of cancer. 14 The clinical application of LB technology in LC, includes early diagnosis, personalized treatment, prognosis prediction, longitudinal monitoring for cancer progression, and therapeutic response. LB is also considered a promising method for the identification of patients with a high risk of disease progression after curative surgery. 18
The analysis of CTCs in patients with early NSCLC reflects their value in early diagnosis. CTC amount increases significantly in most patients with tumor progression. 19 CTCs could be used as an additional method to detect ALK gene rearrangement when appropriate tissue biopsy samples could not be obtained. 20 Moreover, CTCs are used to supplement the tissue-based EGFR mutation detection in LC and to guide the precision treatment of EGFR. 21 LB has also entered in clinical practice as ctDNA-based tumor genotyping for the management of advanced-stage LC. 22 It was reported that the plasma ctDNA levels, in NSCLC patients, are higher than in subjects with chronic respiratory inflammation or healthy individuals. Collecting ctDNA from blood is a non-invasive method, with high clinical potential, that can be repeated over time, contributing to the identification of small residual diseases or recurrence rates. 23
In advanced NSCLC, the use of ctDNA, which had previously been limited to EGFR mutation detection in patients with inadequate tissue for tumor genotyping and/or for the identification of EGFR, T790M-resistant mutation, has now been extended to other genetic aberrations. Thanks to the use of plasma next-generation sequencing (NGS), this analysis, termed the ‘plasma-first’ approach, is moved to a treatment-naïve setting. 3
Recently, it was also developed a new integrated genomic strategy for early-stage LC detection using a ctDNA-based machine-learning platform, named LC likelihood in plasma. This study introduces improvements to Cancer Personalized Profiling by deep Sequencing ctDNA analysis that facilitates screening applications. 24 Nowadays, the European Society for Medical Oncology and the International Society for Study of LC have recommended the use of a multigene NGS approach in the molecular evaluation of advanced-stage NSCLC patients. 3 Although LB is a growing field in cancer management, some challenges to the sensitivity of CTCs and ctDNA in clinical practice remain. The percentage of ctDNA is often as low as 1% and the number of CTCs is limited. EVs, ctRNA, and TEPs are considered novel tumor circulome elements with great potential at any stage of cancer for adequate clinical management. 25 EVs could overcome some limits of LB; they have the advantages to be abundant in blood samples of cancer patients, stable in biofluids, and protect cf-NAs from degradation. Although EVs isolation and quantification are challenging and need standardization, the analysis of the different biological components of LB could be used to explore complementary aspects to illustrate the molecular profiles of LC comprehensively. 3
This review focuses on circulating-free or EVs containing emerging regulatory ncRNAs, such as circular-RNAs (CircRNAs), small nucleolar-RNAs (snoRNAs), small nuclear-RNAs (snRNAs), and piwi-interacting RNAs (piRNAs), in LC. These ncRNAs are essential in maintaining the spatial–temporal architecture of transcriptional and translational programs under malignant conditions and have gained the progressive attention of the scientific community. In addition, we discuss the potential roles of EVs and ncRNAs as biomarkers in LC LB (Figure 1).

LB as a tool for LC management. Schematic representation of LB and EV components, focusing on emerging ncRNAs described in EVs.
Extracellular vesicles
EVs are nanoscale membrane particles, released by all eukaryotic and prokaryotic cytotypes, in physiological and pathological conditions.26–28 EVs carry proteins, metabolites, lipids, and nucleic acids, including DNA fragments and RNAs that can induce phenotypic reprogramming of target cells.29–31 Based on their biogenesis and size, EVs are classified into two main classes: exosomes and microvesicles (MVs; Figure 2). 32
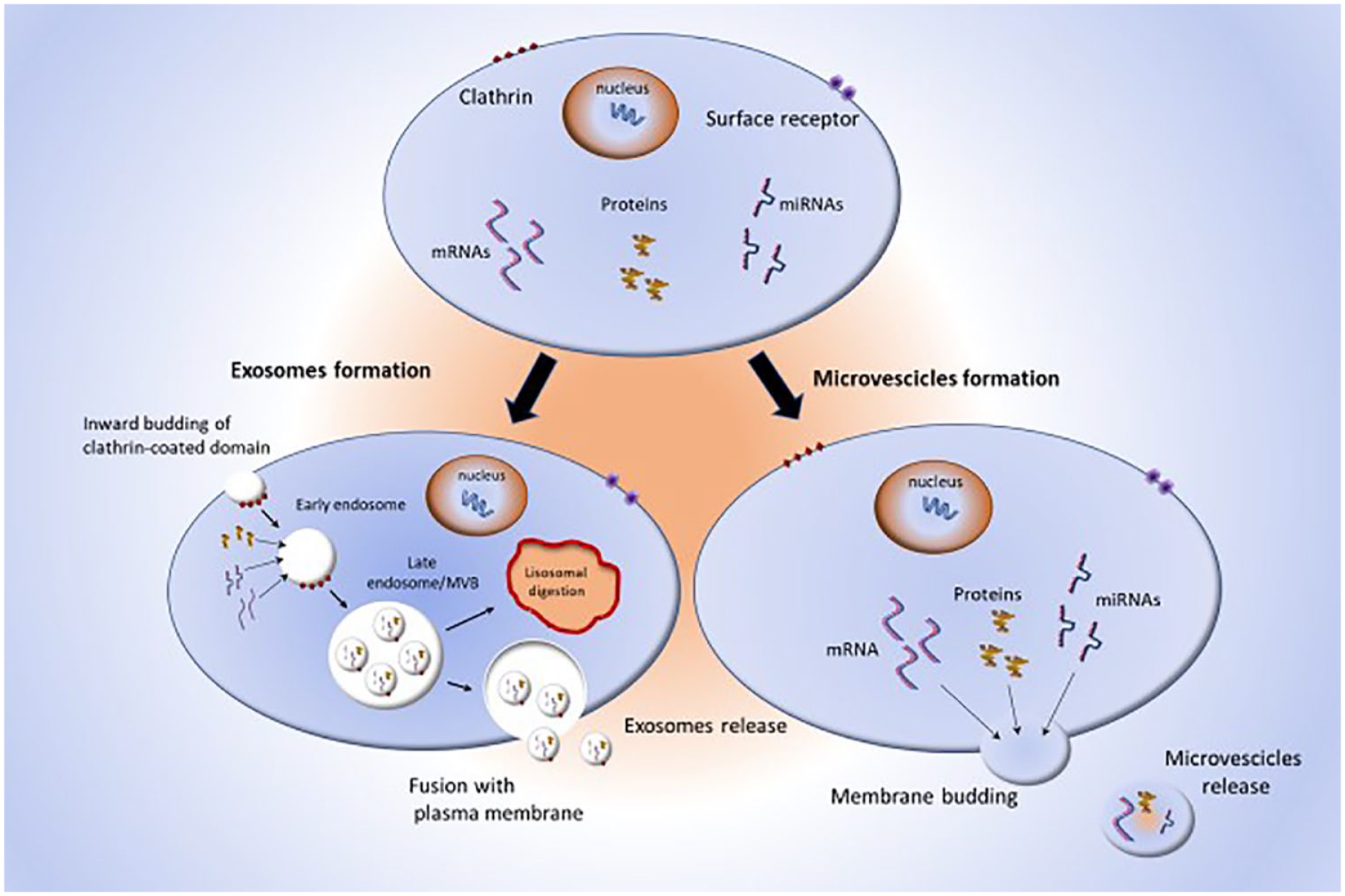
Schematic representation of EV biogenesis. Donor cells release exosomes and MVs with different mechanisms.
Exosomes are lipid bilayer nanovesicles of endosomal origin, with a diameter of 30–150 nm instead, MVs have a diameter of 200–1000 nm and are shed directly from discrete microdomains of the plasma membrane, enriched in phosphatidylserine, cholesterol, and lipid rafts. 33
EVs can be collected from various biofluids, such as serum, plasma, saliva, breast milk, nasal secretions, urine, semen, and pathological effusions.6,29,34,35 Several mechanisms of EV release have been described, that involve different machineries such as ESCRT complex, tetraspanins, sphingomyelinases, redistribution of phospholipids, and depolymerization of cytoskeletal actin. 26
EVs can be internalized by target cells through receptor-ligand binding, direct fusion with plasma membranes, phagocytosis, micropinocytosis, actin polymerization, and filopodia extension. It has also been reported that EVs can be internalized by target cells as intact vesicles surfing on filopodia.27,36–38 EVs encapsulate their cargo in a phospholipidic bilayer, providing high stability, a long half-life, and resistance to degradation. Their stability is attributed to the lipid components of their membranes, which are enriched in cholesterol, phosphatidylserine, glycosphingolipids, sphingomyelin, and annexin, as compared to the cellular plasma membrane. 39
EVs can travel long distances and deliver their cargo to specific cell types through specific ligand–receptor interactions. 39
Another intriguing feature of EVs is their potential to cross tissue barriers. Several studies showed their ability to cross the blood–brain barrier. EVs’ stability, addressability, and barrier penetration make them encouraging therapeutic delivery devices.40,41
EV cargos, transported in directed ways to target cells, can act in autocrine, paracrine, and systemic manners. EVs regulate biological processes and crosstalk between cells, playing an important role in cancer.34,42–44 Several reports demonstrate the key role of EVs in cancer progression, 45 premetastatic niche formation, 46 metastasis,47,48 and drug resistance.49,50 The potential for EVs as biomarkers in LC early detection and progression has been reported as well. EVs have been identified as good biomarkers for monitoring response to LC therapies. 26
Since anticancer therapies alter the amount of EVs in biofluids, the dynamic analysis of circulating EVs and their contents can provide real-time information on therapeutic responses.
51
EVs promise to be a next-generation diagnostic and therapeutic tool in LC, to monitor immune-checkpoint inhibitors (ICIs) therapy with programmed cell death protein 1 (PD-1) or programmed cell death-ligand 1 (PD-L1) antibodies, a standard treatment for advanced NSCLC. It was reported that specific microRNAs (miRNAs) in plasma EVs are differentially expressed between responders and non-responders’ patients to ICIs and have potential as predictive biomarkers for anti–PD-1/PD-L1 treatment response.
52
Recently, de Miguel-Perez
As aforementioned, EVs are natural intercellular shuttles for NAs such as DNA, and RNAs including ncRNAs.54,55 The DNA packaging into EVs was significantly higher in cancer-derived EVs compared to EVs from noncancer cells. In LC, the combination of RNA/DNA contained in exosomes (exoRNA/DNA) and ctDNA for T790M detection has higher sensitivity and specificity compared to ctDNA alone. 56
Recently, Park and colleagues demonstrated that also EVs derived from bronchial washing (BW-EVs) could be used for accurate and frequent genotyping of EGFR mutation in patients with NSCLC. BW-EVs are useful for the early detection of actionable mutations, for the selection of personalized therapy, and for monitoring disease progression. The well-known T790M mutation seems to be detected from BW-EV-DNA with a superior detection rate to plasma-derived ctDNA or tissue biopsy. 57 Although the ‘gold standard’ source for LB testing on EGFR is represented by ctDNA, recovered from plasma, recent studies indicate that EVs are a good source of RNA to identify EGFR mutations. EV-RNA and ctDNA can function as independent biological sources in LB of NSCLC patients providing a complementary informative set of tumor dynamics. While ctDNA is predominantly released as a consequence of the apoptotic and necrotic process, EVs are released from viable cells to transmit information to other cells providing a complex cargo available for molecular analysis. 58 Moreover, it was demonstrated that EVs carry mutated EGFR as mRNA and protein, mirroring the disease status in metastatic NSCLC. Sensitizing (exon 19 deletion, L858R) and resistance (T790M) mutations were quantified in EV-RNA. The comparison of mutation detection between EV-RNA and ctDNA using digital droplet PCR (ddPCR) indicates that EVs have a better detection rate for exon 19 deletions and L858R point mutation. 59 These reports suggest that EV-RNA provides a new tool appropriate for use in clinical practice to investigate the dynamics of common driver EGFR mutations in NSCLC patients receiving TKIs.
Noncoding RNAs
ncRNAs are molecules of RNA with no protein translation potential, involved in physiological and pathological processes. 60 Data from genome-wide transcriptional analysis in humans have shown that the amount of protein-coding transcripts account for approximately 2% of the entire genome, while ncRNAs represent about 98% of all genomic output.29,61 Interestingly, it has been reported that the proportion of noncoding regions in the genome increases according to the complexity of organisms, suggesting an important role for these sequences in the physiology and development of organisms.62,63 Studies on these non-protein-coding RNAs have received a lot of interest in many fields, especially in cancer, leading to new hypotheses about cancer biology. 64 ncRNAs can be classified as housekeeping and regulatory. Housekeeping ncRNAs are abundantly and ubiquitously expressed in cells, and primarily regulate generic cellular functions. While regulatory ncRNAs are usually considered key regulatory RNA molecules, function as modulators of gene expression at epigenetic, transcriptional, and post-transcriptional levels.65–67
Based on their molecular size, all regulatory ncRNAs are subclassified into small ncRNAs, with transcripts shorter than 200 nucleotides, including miRNAs, snRNAs, snoRNAs, and piRNAs and large ncRNAs, with transcripts longer than 200 nucleotides, that include CircRNAs. 68
Despite comprising less than 1% of total cellular ncRNA content, regulatory ncRNAs play crucial roles in transcription, post-transcriptional mechanisms, and translation. Regulatory ncRNA-mediated gene silencing constitutes one important type of epigenetic alteration and has been implicated in several cases of human carcinogenesis. Regulatory ncRNAs (Figure 3) modulate gene expression through various mechanisms.69,70 ncRNAs can influence lung tumorigenesis, and they play an important role in premetastatic niche formation and metastasis of NSCLC. 71 Since ncRNAs circulate in biofluid freely or encapsulated in EVs, they are intriguing as potential biomarkers and therapeutic targets in LC LB scenario.
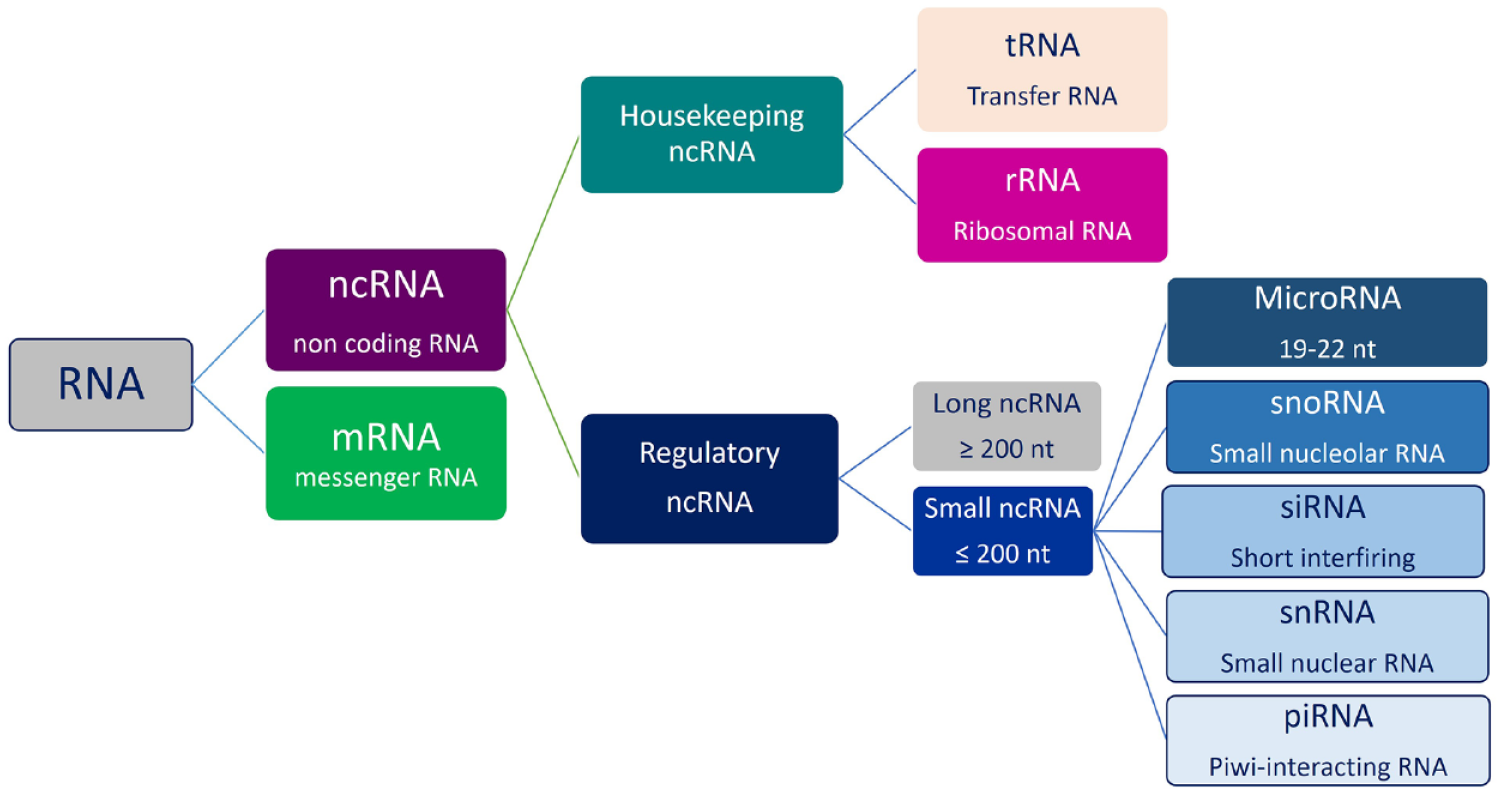
RNA classification. RNAs are divided into two main classes: mRNAs and ncRNAs. Each category has different components.
In recent years, a variety of regulatory ncRNAs have been identified, as key modulators of gene expression in different cellular pathways and systems.68,69 Recent findings changed the description of ncRNAs from ‘junk’ transcriptional products to functional regulatory molecules that mediate key processes such as gene expression, chromatin remodeling, transcription, post-transcriptional modifications, and signal transduction. 70 The tRNA-derived fragments (tRFs)-based or RNA-related fragments (rRFs)-based therapies might become useful to offer new therapeutic options. 72
In the classical view, housekeeping ncRNAs, involved in the maintenance of normal cell functionalities, include transfer RNAs (tRNAs) and ribosome RNAs (rRNAs), which are important in protein translation. This viewpoint was challenged by recent findings on the dysregulated expression of tRNAs in several cancers, including LC. 54 The tRNAs are cleaved by angiogenin in tRNA-derived stress-induced RNAs (tiRNAs), which can be further processed by Dicer into tRFs. In LC tissue, tRNAs-Leu and tRNAs-Val are overexpressed, respectively, in 37% and 26% of samples. 73 The tRF-Leu-CAG is upregulated in LC and involved in cell proliferation and cell cycle progression, interacting with AURKA protein. 74 The tRF-Leu-CAG could be a new diagnostic marker and potential therapeutic target in NSCLC. Specific blockade of some tRFs, such as tRF-Leu-CAG, may have considerable clinical application in suppressing LC cell proliferation and cell cycle progression. 73 Recently, it was reported that EVs shuttle tRFs with effects on various cellular processes; the dysregulation of EV-tRFs has been associated with cancer progression and they are potential novel biomarkers for cancer diagnosis. 75
The rRNAs, like tRNAs, can be cleaved into small ribosomal rRFs. The function of rRFs is not limited to gene silencing, their structure also plays a key role in rRNA stability. 72 The rRNAs are abundant in EVs released by different cancer cell lines; a high-throughput study on human breast cancer cell lines showed that over 80% of RNAs contained in EVs are rRNAs. 76 The rRNAs were also detected in EVs collected by different body fluids such as the serum, plasma, urine, and saliva. 77
Circular-RNAs
CircRNAs have been identified as a new class of ncRNA with high regulatory potential. 78 CircRNAs contained in blood, either free or encapsulated within EVs, have several advantages over canonical linear RNAs as cancer biomarkers. 79
CircRNAs, which are found in a large amount within tissues, cells, and body fluids, are aberrantly expressed in cancer tissues and regulate tumor progression. Since they are expressed in stage-specific manners, several studies have shown their potential as helpful diagnostic and prognostic biomarkers for cancers. 80
CircRNAs are generated for back-splicing processing from linear pre-messenger RNAs when the 3′ and 5′ ends are ligated to form a continuous loop and covalently closed. 79 CircRNAs were first discovered in 1990 when observing that exons of a tumor suppressor gene after their splicing were joined in a different order than their genomic sequence. 81 CircRNAs are highly resistant to RNAse activity because of the lack of 5′ and 3′ ends, 82 being more stable and having longer half-lives than canonical linear isoforms. In 2015, CircRNAs were described in EVs for the first time, when high-throughput technologies such as genome-wide RNA-seq analyses showed that CircRNAs were enriched in EVs in comparison with parental cells. CircRNAs contained in EVs may be controlled by alteration of associated miRNA levels in parental cells, transferring the biological activity to target cells.83,84
The principal functions of CircRNAs are miRNA inhibition, interaction with RNA-binding proteins, and regulation of parental genes. Particularly, CircRNAs act as miRNA sponges protecting target genes from repression by miRNAs. Notably, one CircRNA can sponge different miRNAs, establishing an intricate and precise regulatory network.85,86
CircRNAs are emerging candidates as biomarkers for diagnosis, prognosis, and therapeutic response of NSCLC in LB.
86
CircRNAs are involved in various aspects of LC progression such as cell growth, proliferation, migration, metastasis, drug resistance, and immune escape as summarized in Figure 4. Recently, it was reported that ciRS-7 is associated with NSCLC development. CiRS-7, also called CDR1as, is one of the earliest discovered CircRNAs originating from the back-splicing of CDR1 gene.
87
This CircRNA has about 70 miRNA-binding sites and acts as a sponge for miR-7. CiRS-7 is overexpressed in tissues and cell lines of NSCLC and may promote cancer cell proliferation

CircRNAs are involved in different steps of LC progression such as cell growth, migration, metastasis, drug resistance, and immune escape.
Another CircRNA overexpressed in NSCLC tissues compared to adjacent tissues is has_circ_0014130 (circPIP5-K1A), which acts as a sponge for miR-600 and inhibits its activity, upregulating hypoxia-inducible factor 1α, which is involved in tumor proliferation and metastasis.89
The proto-oncogenic CircRNA (circ-PRKCI) originates from the 3q26.2 amplicon, one of the most frequent genomic aberrations in cancer. The increased expression of Circ-PRKCI in lung tissues, caused by amplification of 3q26.2 locus, induces tumorigenesis and progression of LC. Circ-PRKCI functions as a sponge for miR-545 and miR-589 and annuls their suppression of pro-tumorigenic transcription factor
In addition, circMAN2B2 is a product of MAN2B2 mRNA splicing and promotes FOXK1 expression by sponging miR-1275 in LC cells, inducing LC progression. 91 FLI1 exonic CircRNA (FECR) is considered an oncogenic driver that helps tumor metastasis by modulating the miR584-ROCK1 pathway. Notably, FECR1 is contained in exosomes collected in the serum of LC patients. Exosomal FECR1 has been associated with poor tumor survival and clinical response to chemotherapy and is useful as a cancer prognostic factor and as a potential biomarker to track LC progression. 91
Recently a novel CircRNA-103809/miR-377-3p/GOT1 axis that contributes to cisplatin resistance in NSCLC
Moreover, it has been observed that has-CircRNA-002178 can enhance PD-L1 expression in cancer cells inducing T-cell exhaustion
Furthermore, Circ-CPA4 is highly expressed in NSCLC relative to normal human bronchial epithelial cells. Circ-CPA4 regulates cell growth, migration, stemness, and drug resistance in NSCLC cells and is involved in CD8+ T-cell inactivation,
Recently, it was demonstrated that CircRNA-102481 is overexpressed in exosomes collected from serum in EGFR-TKIs-resistant patients. The silencing of this CircRNA inhibits EGFR-TKIs-resistant NSCLC cell proliferation and induces apoptosis. CircRNA-102481 overexpression could promote EGFR-TKIs-sensitive NSCLC cell proliferation and inhibit cell apoptosis, which suggests that CircRNA-102481 may contribute to EGFR-TKIs resistance in NSCLC.
96
Another CircRNA highly expressed in NSCLC cells is circSATB2, which positively regulates fascin homolog 1, actin-bundling protein 1 (FSCN1) expression
The specificity and stability of CircRNAs and the capacity of EVs to interact with target cells while remaining stable in the bloodstream, make CircRNAs shuttled by EVs very attractive as biomarkers for LC early detection. 97
Emerging evidence demonstrates that CircRNAs are aberrantly expressed in progressive degenerative lung diseases such as chronic obstructive pulmonary disease (COPD), in which cigarette smoke is considered a crucial risk factor. Recently, it was reported that circ-RBMS1 derived from the RBMS1 gene was higher in COPD patients, and cigarette smoke increased circ-RBMS1 expression in a dose-dependent manner. Circ-RBMS1 directly targeted miR-197-3p and this miRNA targeted FBXO11.
Furthermore, it was reported that circ-FASRA in plasma has diagnostic value for NSCLC and can be used as a biomarker for the detection of non-invasive NSCLC. With the development of second-generation sequencing technology, screening CircRNA differentially expressed in body fluid samples of LC patients and healthy controls can be used to find a new biomarker for diagnosis and prognosis and as a potential therapeutic target. 100
Small nuclear RNA
Several diseases such as cancer and chronic lung disease have been associated with snRNAs alterations, acting on the cellular transcriptome. Each snRNA, such as U1, and U2 (the nomenclature arises from high uridine content), is complexed with small nuclear ribonucleoproteins to exert their functions. 70 The snRNAs act in spliceosome complexes, recognizing 5′ and 3′intron/exon boundaries during splicing of introns from pre-messenger RNA transcripts. 101
snRNAs (<200 nucleotides) include a small group of non-polyadenylated, noncoding transcripts that act in the nucleoplasm. snRNAs are divided into two classes, Sm and Lsm class RNAs, based on common sequence features and protein cofactors. Sm-class RNAs are characterized by a 5′-trimethylguanosine cap, a 3′ stem-loop, and a Sm-site that consists of a binding site for a group of seven Sm proteins, forming a hetero-heptameric ring structure. Lsm-class RNAs consist of a mono-methyl-phosphate cap and a 3′ stem-loop, ending in a stretch of uridines that form the binding site for a hetero-heptameric ring of Lsm proteins. The principal components of Sm-class RNAs are U1, U2, U4, U4atac, U5, U7, U11, and U12, whereas the more studied Sm-class RNAs are U6 and U6atac. 102
The high stability of snRNAs in biological samples makes them novel diagnostic biomarkers, supporting prognostic or predictive indicators, and tools to monitor treatment in cancer patients. 103
Fragments of U2 snRNA (called RNU2-1f) may be collected in sera of patients with pancreatic, colorectal, and ovarian cancers and cerebrospinal fluid of patients with primary central nervous system lymphoma.104–106 RNU2-1f has also been found in the serum of patients with metastatic melanoma. 107 High levels of this snRNA have been associated with tumor progression and after surgical removal of pancreatic and colorectal tumors, RNU2-1f levels have been shown to decrease. 101 Furthermore, RNU2 is highly expressed in lung tissue and efficiently exported into circulation, elevated levels of RNU2-1f have been detected in the serum of LC patients. RNU2-1f may be a potential biomarker for LC patients. 108 LC snRNAs can regulate alternative splicing to drive genetic and neoplastic disease and alter mRNA profile in TEPs. Platelets are ‘educated’ by their tumor environment, containing a dynamic variety of RNA subsets, including snRNAs. RNA profiles of TEPs allow for the distinction between cancer patients and healthy controls. For instance, RNA profiles of TEPs can be utilized to predict oncogenic status, such as MET or HER2 positivity, and reveal gene mutations in KRAS or EGFR. In addition, U1, U2, and U5 are significantly downregulated in TEP of LC patients compared with healthy controls. TEP U1, U2, and U5 levels can be decreased in LC patients, and their downregulation has been correlated with LC progression. Moreover, TEP U1, U2, and U5 levels may be directly correlated with paired exosomes and TEP from treated patients but not from untreated patients. U1, and U5 but not U2 in platelets can be elevated by exosomes released from apoptotic cells. 109
Small nucleolar RNA
snoRNAs are a group of intron-encoded ncRNAs mainly accumulated in nucleoli that consist of 60–300 nucleotides. snoRNAs are grouped into two families called box C/D snoRNAs (SNORDs) and box H/ACA snoRNAs (SNORAs). 110
snoRNAs are involved in post-transcriptional modification and maturation of ribosomal RNAs, snRNAs, and other RNAs. 111 snoRNAs are involved in several physiological and pathological processes. In addition, snoRNAs have oncogenic or tumor-suppressive functions in different cancers activating invasion, metastasis, angiogenesis, and sustained proliferative signaling or increasing growth suppressors and cell death. Some reports suggest that snoRNAs are associated with p53 regulation. P53 is a well-known tumor suppressor that responds to cellular stresses to regulate the expression of target genes involved in cell cycle arrest, apoptosis, and DNA repair. 112 Recently, it was reported that snoRNAs are associated with p53 pathway. In particular, SnoRNA42 is overexpressed in NSCLC and has an oncogenic role by affecting p53 expression. 113
Moreover, SNORD78 is upregulated in LC and tumor-initiating cells of LC, suggesting that it might play a role in lung tumorigenesis. SNORD78 is also upregulated in cancer stem cells in NSCLC and is essential for the self-renewal of cancer stem cells in NSCLC. 114
It was reported that SNORD46 acts as an oncogene in LC.
snoRNAs can predict LC progression from the initial stages. The overexpression of SNORD28, SNORA21, SNORA47, SNORD66, SNORA78, and SNORA68 leads to worse overall survival in LC patients and these snoRNAs are differentially expressed between lung tumors of stage I and normal tissue. 116 In NSCLC patients, SNORD33, SNORA42, SNORD66, and SNORD78 are overexpressed. Moreover, snoRNAs can be considered LC biomarkers in combination with other miRNAs. 117 A panel of biomarkers for LC, comprising miR-21, miR-32, and miR-210 and SNORD66, SNORD78, collected from sputum, was developed to be used as a potential tool for a non-invasive LC diagnosis. It was reported that the combined use of miRNAs and snoRNAs demonstrated higher sensitivity and specificity compared with a single type of ncRNA biomarkers, offering a new approach for LC early detection. 118
Interestingly, it was reported that five snoRNAs: SNORA14B, SNORA18, SNORA25, SNORA74A, and SNORD22 were encapsulated in exosomes isolated from the serum of pancreatic cancer patients and conditioned medium of pancreatic cell lines. SNORA74A and SNORA25 are highly expressed in the early stages of pancreatic cancer in comparison with healthy controls. 119 These data indicate that exosomal snoRNAs are useful in the diagnosis of cancer; further studies are needed to confirm their role in LC.
Piwi-interacting RNAs
P-element-induced wimpy testis piRNAs are a new investigated class of small ncRNAs. 120 piRNAs have been discovered in male gonadal cells, with the central role of protecting germinal cells from transposable elements (TEs), especially of viral origin, and are germline specific. 121 piRNAs are small ncRNAs of 26–31 nt; they have a 2′-O-methylation at the 3′ end as a distinctive and exclusive feature of all piRNAs. The precursors of piRNAs are single-stranded transcripts without secondary hairpin structures.122,123 These precursors are generated from precise genomic locations with repetitive elements with a Dicer-independent process. The nascent piRNAs require further post-transcriptional modifications to become mature piRNAs. The biogenesis of piRNAs includes two steps: a primary and secondary amplification cycle referred to as a ‘ping-pong cycle’ in which piRNAs are bound to piwi proteins. 124 piRNA biogenesis is an adaptive process that silences active transposons with sequences complementary to piRNA cluster transcripts. piRNAs control and silence TEs to protect the genome since uncontrolled TE expression may lead to a loss of genome integrity. 125 Recently, it was reported that piRNAs are widely expressed in somatic cells and human cancer cells. About 30,000 piRNAs were found in the human genome and recent studies have suggested that piRNAs play a role in human cancer pathogenesis. Cancer, stem, and germ cells share key biological characteristics such as the ability for self-renewal and rapid proliferation. Although piRNAs were first described as important regulators in maintaining germline stem cells, it is conceivable that rapidly dividing cancer cells might adopt and utilize self-renewal machinery like germ cells. Recently, a growing number of studies have revealed the role of piRNAs in cancers, introducing a new biological concept in which mechanisms of piRNA-mediated gene regulation specific to germline cells also have oncogenic and tumor suppressive roles. 126
piRNAs are expressed in a tissue-specific manner in several human tissues, regulating important signaling pathways at the transcriptional or post-transcriptional level. In malignant cells, piRNAs participate in the epigenetic regulation of DNA 123 and are essential for maintaining cancer stemness. 127 The aberrant expression of piRNAs and PIWI proteins in cancers might be used as new biomarkers and therapeutic targets for tumor diagnosis and treatment. It was suggested that piRNA-54265 can be used as a biomarker for the early detection and clinical monitoring of colorectal cancer. piRNA-36712 has been described as a novel tumor suppressor and might be a breast cancer prognostic predictor. In addition, piR-823 has been found in cancer cells and plasma, has a role in regulating tumor cell growth, and its expression fluctuates in many cancers, including gastric cancer and multiple myeloma. 128 In colorectal cancer, piR-823 downregulation increases cell apoptosis by inducing transcription factor HSF1, an apoptosis activator, and inhibits cell proliferation. 129 piRNA-54265 is highly expressed in colorectal cancer tissue and serum; this piRNA activates STAT3 signaling, thereby inducing cancer progression. 130 In multiple myeloma, piRNA-823 silencing induces the expression of apoptosis-related genes, modulating DNA methylation. 131 Another piRNA, piR-651, is downregulated in patients with Hodgkin lymphoma relative to healthy controls. Furthermore, low levels of piR-651 correlate with poor prognosis in Hodgkin lymphoma patients. 132 piRNAs are not easily degradable and can move across the plasma membrane. This feature suggests that piRNAs can easily be detected in body fluids, and recent studies have indicated that piRNAs, contained in EVs released by cancer cells, remain stable in body fluids. Peng and colleagues have demonstrated that piRNAs are contained in EVs from the urine of patients with prostate cancer. The expression of piR349843, piR382289, piR158533, and piR002468 in urinary EVs is increased in prostate cancer patients relative to healthy controls. 133
Emerging findings suggest that piRNAs may be considered potential biomarkers for LC diagnosis. It has been shown that piR-hsa-211106 can inhibit the progression of lung adenocarcinomas enhancing chemotherapy sensitivity, suggesting that piR-hsa-211106 is a potential therapeutic target for LC. 134 piRNA expression profiles of lung adenocarcinoma tissues and adjacent normal tissues have identified ten piRNAs overexpressed in tumor tissues. Among these, piR-hsa-26925 and piR-hsa-5444 are upregulated in EVs collected from the serum of patients with lung adenocarcinoma. 135 These findings indicate that LB can allow for the detection of both free piRNAs and piRNAs contained within EVs. Moreover, piRNAs may become therapeutic and diagnostic tools for various cancer types, including LC.
Conclusion and perspectives
LB is a promising diagnostic tool with several advantages over conventional invasive methods. LB is useful to identify prognostic, diagnostic, and predictive biomarkers in LC not only for its minimal invasiveness, which allows repeating the biopsy within the scope of tumor surveillance and yields genetic information about cancer, considering the tumor heterogeneity and the presence of subclones. LB represents a source of biomarkers including EVs, ctDNA, CTC, and cf-NAs such as ncRNAs. EVs appear to be ideal for LB thanks to the stability of their membrane and their ability to travel in body fluids, at long distances.
Several ncRNAs have been detected in biofluids. The origin of these molecules might be passively released by dead cells or active secretion
Emerging ncRNA can be useful for their diagnostic and prognostic potential (Figure 5). The structural stability of CircRNAs may have interesting applications for designing drugs that can be delivered free or encapsulated in EVs. 137 In addition, EVs containing CircRNAs are internalized by recipient cells, where they affect post-transcriptional regulation of gene expression and can be exploited for their therapeutic potential. 138 Some CircRNAs are found to be significantly associated with LC. Many clinical results have demonstrated that CircRNAs can be considered potential biomarkers. Nevertheless, the sample quantities of these studies are limited, and the precise regulatory mechanism of CircRNAs in LC should be better defined. 139

ncRNAs as diagnosis and prognostic biomarkers in LC.
The aberrant expression of piRNAs has been associated with cancer progression and metastasis in different cancer types, suggesting their role in cancers as oncogenes or tumor suppressors. Although they can be detected in the blood of cancer patients, few reports have been published about piRNAs contained in EVs. 140
Although snoRNAs localize primarily to the nucleus and have previously been considered housekeeping genes, accumulating evidence indicates that snoRNAs have oncogenic roles. Since snoRNAs are abundant and easily detectable in cancers and blood and are functionally relevant in oncogenesis, they may become an important target for cancer therapy. 141 Moreover, snoRNAs contained in EVs can be considered novel biomarkers for LC diagnosis. 142 snoRNAs might become pivotal elements to improve the knowledge of LC and relevant multimodal tools to ameliorate cancer patient management from their diagnosis to their treatment. 143
While the evaluation of actionable mutations in LB has a confirmed clinical value, the use of epigenetic alterations, with few exceptions, has not reached clinical practice yet. Among the different fields where epigenetic factors may play a role as biomarkers, screening and diagnosis are the areas closer to the clinic. 144
Since the functions of many ncRNAs are not completely known and EVs shuttle a myriad of biomolecules, EV language remains difficult to decode. Understanding the complex networks of interactions coordinated by ncRNAs would offer a unique chance to design better therapeutic options. Further studies are needed to confirm the potential role of ncRNAs freely circulating in body fluids or contained in EVs as biomarkers in the field of precision oncology.
Currently, few clinical trials (precisely 13) on EVs in LC management have been proposed (www.clinicaltrials.gov), but the number of clinical studies on EV-ncRNAs (as reported in Table 1) in this field growing exponentially.
Clinical studies on ncRNAs as diagnosis and prognostic biomarkers in LC.

AUC, are under the curve; CircRNAs, circular RNAs; ncRNAs, noncoding RNAs; NSCLC, non-small-cell lung cancer; piRNAs, piwi-interacting RNAs; snoRNA, small nucleolar RNA; snRNA, small nuclear RNA; TEP, tumor-educated platelet.
The combined analysis of the different components of LB, including the ncRNAs contained in EVs, may potentially help to comprehend the dynamics of molecular alterations and support the clinical decisions in LC management. Overall, while this field may seem to be in infancy, the evidence reported in this review indicates the potential of ncRNAs as rising stars biomarkers and therapeutic targets for precision oncological treatments in LC patients.
