Abstract
The treatment of metastatic renal cell carcinoma has significantly evolved in recent years, particularly with the advent of novel immune checkpoint inhibitors (ICI). Despite the striking benefits observed on a population level, outcomes vary and some patients do not respond to ICI-based regimens, ultimately require salvage therapies. An ever deeper understanding of the disease biology mediated by the development of multiple high-throughput molecular omics has led to significant progress in biomarkers discovery. But despite growing insights into the molecular underpinnings of the tumor microenvironment, biomarkers have not been integrated successfully into clinical practice. In this review, we discuss some of the novel emerging predictive biomarkers to ICIs in metastatic renal cell carcinoma.
Introduction
Renal cell carcinoma (RCC) is estimated to be the sixth and eighth most frequently diagnosed cancer in men and women, respectively, in 2021 in the United States. 1 Globally, RCC poses a significant healthcare burden with >400,000 new cases and >175,000 attributable deaths annually. 2 Over 20% of patients diagnosed with RCC present with synchronous metastases and 30% of patients who are treated with nephrectomy in a curative intent subsequently develop recurrent metastatic disease.3,4 RCC comprises a diverse group of malignancies with increasingly defined genomic features and clinical outcomes.5,6 Histologically, it is broadly divided into clear cell RCC (ccRCC) which accounts for approximately 75% of cases and non-clear cell RCC (nccRCC) comprising the remaining 25% of cases. 7
A landmark breakthrough discovery in the biology of ccRCC was achieved when it was recognized that the majority of sporadic ccRCC cases are driven by inactivating mutations or methylation of von Hippel-Lindau (VHL), leading to accumulation of hypoxia-inducible factor (HIF) which in turn leads to activation of pathways that promote angiogenesis such as the vascular endothelial growth factor (VEGF) and platelet-derived growth factor (PDGF).8,9 Furthermore, RCC is known to be one of the most immune-responsive solid tumors with a frequent high infiltration of several immune cells. 10 As our understanding of the disease biology and molecular interplay in RCC improved,11,12 this led to an expansion of the therapeutic landscape of metastatic RCC over the last two decades, which resulted in a plethora of FDA-approved drugs, including therapies targeting tyrosine kinase inhibitors with activity against the VEGF receptor, antibodies directed against VEGF, small molecule inhibitors of mammalian target of rapamycin (mTOR), and more recently, immune checkpoint inhibitors (ICIs) targeting the PD-1/PDL-1 and CTLA-4 axis.13,14 As per contemporary guidelines and based on recent randomized phase III studies, the current standard first-line treatment for patients with metastatic RCC consist of dual immune checkpoint inhibition with anti-PD1 plus anti-CTLA-4 ICIs or the combination of anti-VEGF receptor TKI (VEGF-TKI) plus anti-PD-1 ICI. 15
Relevantly, despite these advances in immunotherapy in RCC, hitherto, the role of biomarkers was limited on the pivotal trials for these newly approved standards and few metrics help predict treatment responses, durable benefit or early disease progression.16,17 Currently, survival probability can be best estimated using prognostic risk models, including the Memorial Sloan Kettering Cancer Center (MSKCC) risk score, 18 developed during the cytokine era, and the International Metastatic RCC Database Consortium (IMDC) risk score, 19 developed in the targeted therapy era (Table 1). Both models are computed based on clinical and laboratory variables; however, their applicability and accuracy in the current immunotherapy era may be compromised as ICI now represent the backbone therapy in metastatic RCC. 20
Prognostic models for metastatic RCC.

IMDC, International Metastatic RCC Database Consortium; KPS, Karnofsky performance status; MSKCC, Memorial Sloan Kettering Cancer Center; RCC, renal cell carcinoma.
Biomarkers discovery in RCC has been challenging due to multiple factors, including the diverse molecular and genomic heterogeneity of RCC and the fact that the driving genomic events in RCC are primarily due to loss of function events rather than actionable gain of function mutations.21–23 Herein, we will review some of the emerging data in the context of ICI therapy for metastatic disease, including circulating and tissue-based biomarkers.
Programmed death-ligand 1 expression
Program death ligand-1 (PD-L1) is the ligand for PD-1, a key immune checkpoint receptor expressed by activated T cells that has been successfully targeted by multiple checkpoint inhibitors. 24 High PD-L1 expression by immunohistochemistry (IHC) on tumor cells is a well-known adverse prognostic factor in patients with metastatic RCC treated with TKI therapy and is associated with aggressive features of RCC, such as a high nuclear grade, lymph node involvement, and metastatic disease.25,26 In the immunotherapy era, PD-L1 expression by IHC is by far the most widely studied biomarker; 27 however, unlike other solid tumors like non-small cell lung cancer where PD-L1 expression is incorporated in treatment decisions, 28 the role of PD-L1 expression in directing kidney cancer treatment is still debatable, and currently it is not used in our treatment decision making. 15 Widely acknowledged limitations of PD-L1 IHC include the spatial heterogeneity of PD-L1 expression within the biopsied lesion,29,30 the expression of PD-L1 in a wide range of cells in the tumor microenvironment, 31 and the lack of standardization with different antibody tests and methods to evaluate PD-L1 expression (tumor cells, immune cells, or both). 32
The phase III CheckMate 025 clinical trial compared nivolumab with everolimus in patients with previously treated metastatic ccRCC and demonstrated overall survival (OS) benefit for patients treated with nivolumab compared with patients treated with everolimus [25.0 versus 19.6 months; hazard ratio (HR) = 0.73, 98.5% confidence interval (CI) = 0.57–0.93] irrespective of PD-L1 expression. 33 In the subgroup analysis of the study, an inferior OS was observed in patients with PD-L1 expression ⩾1% compared with those with PD-L1 expression <1%. The phase III Checkmate 214 clinical trial investigated the combination of nivolumab plus ipilimumab with sunitinib in previously untreated metastatic ccRCC, the study led to a significant prolongation of OS and progression free survival (PFS), as well as higher objective response rates (ORR) in patients treated with the combination. 34 In an exploratory analysis, OS and ORR favored nivolumab plus ipilimumab irrespective of PD-L1 expression; however, the benefit was more pronounced in the PD-L1 positive group. In particular, the ORR to the combination was higher in the PD-L1-positive group (ORR among patients with <1% PD-L1 37% versus 58% among patients with ⩾1% PD-L1), suggesting that PD-L1 expression on tumor cells is not only prognostic but also predictive for this particular regimen.
With regard to ICI plus VEGF-TKI combinations, several combinations reported significant prolongation of PFS, OS, and likelihood of response irrespective of PD-L1 expression when compared with sunitinib including pembrolizumab plus axitinib in the KEYNOTE-426 phase III clinical trial, 35 nivolumab plus cabozantinib in the CheckMate 9ER phase III clinical trial, 36 and pembrolizumab plus lenvatinib in the CLEAR phase III clinical trial. 37 Given the broad efficacy (and superiority over TKI monotherapy) observed with all approved combinations, regardless of PD-L1 expression levels, it has neither been approved nor embraced as a clinical biomarker in this disease.
Genomic alterations
Large-scale sequencing efforts have studied the association between clinical outcomes to ICI and the genomic landscape of clear cell RCC, including genomic and copy number alterations. In several solid tumors such as non-small cell lung cancer (NSCLC) and urothelial carcinoma (UC), it was demonstrated that tumor mutational load is associated with improved clinical outcomes to tumor mutational load.38,39 This stems from the hypothesis that in tumors with a high mutational load, there is an increased production of neoantigens, which subsequently stimulates the anti-tumor immune systemic response.40,41 In 2020, a tumor-agnostic approval was announced by the FDA for pembrolizumab in patients with tissue tumor mutational burden–high (TMB-H; ⩾10 mutations/megabase), who have progressed following prior treatment and have no satisfactory alternative treatment options. 42
However, RCC is known to exhibit low tumor mutational load with a median of 1.1 mutations per megabase,43,44 and correlation has not been demonstrated with response to ICI based on several retrospective and prospective studies.45–48 This lack of association extends to exploratory analyses of frontline combination regimens, in that TMB did not correlate with outcomes on CheckMate 214 (ipilimumab plus nivolumab), 49 JAVELIN renal 101 study (axitinib plus avelumab), 50 or KEYNOTE 146 (pembrolizumab plus lenvatinib). 51
The mammalian SWItch/Sucrose Non-Fermentable complex (mSWI/SNF) is a chromatin remodeling complex that has recently been implicated as a potential biomarker for immunotherapy, in particular the polybromo and BRG-1 associated factors (PBAF) complex; this includes the genes ARID2, PBRM1, and BRD7. 52 While early reports suggested that LOF alterations in PBRM1 might be predictive of benefit from ICI monotherapy with nivolumab,53,54 this could not be confirmed in other retrospective 55 and prospective datasets.48–50
ARID1A is another member of the mSWI/SNF, which has been studied as a biomarker of clinical benefit to ICI.56,57 Correlative analyses from the phase III IMmotion 151 study demonstrated that the presence of loss of function mutations in ARID1A conferred significant improved PFS treated with atezolizumab plus bevacizumab versus sunitinib (ARID1A altered versus wild type, HR = 0.50; 95% CI: 0.26–0.96; median PFS: 20.7 versus 6.8 months). 58
DNA damage and repair (DDR) gene alterations are associated with improved outcomes to ICI such as in other solid tumor malignancies.59,60 In RCC, analysis of a select panel of key DDR gene alterations (34 genes) across 229 patients showed that deleterious DDR gene alterations were frequent (19%). 61 The presence of DDR gene alteration was associated with improved OS to IO treatment and not associated with outcome in TKI-treated patients. Another study of patients with metastatic RCC (n = 34) treated with ICI showed that patients with disease control displayed enrichment of mutations in DNA repair genes. 45
The significance of copy number alterations was tested in a pooled analysis from three clinical trials (CheckMate 025, 009, 010). Overall, copy number burden did not affect survival; however, loss of 9p21.3 was associated with worse OS following ICI treatment (p = 0.011). 47 Similar associations have been demonstrated in other diseases. 62
Human endogenous retroviruses (ERVs) are remnants of exogenous retroviruses that have integrated into the human genome throughout evolution. 63 In a case series of patients with metastatic ccRCC who received hematopoietic stem cell transplant, a CD8+ T-cell clone was isolated from a long-term responding patient and was found to have recognized a tumor antigen encoded by human endogenous retrovirus type-E (ERV-E). 64 Several studies since have reported that ERVs are aberrantly expressed in ccRCC. A pan-cancer TCGA analysis identified the prevalence of potentially immunogenic ERVs across solid tumors, notably with highest expression in ccRCC, breast cancer, colon cancer, and head and neck squamous cell cancer. 65 The authors found that expression of ERV 3-2, the most prevalent ERV, was enriched in ICI responders compared with non-responders in 24 patients with metastatic ccRCC. Similar findings have since been reported for two additional ERVs (ERV2282 and EVR3382), which were weakly potentially associated with clinical outcomes in a separate anti-PD-1-treated cohort. 47 However, in a recent biomarker analysis of patients treated with metastatic RCC treated on the phase III CheckMate-025 clinical trial, none of the previously identified ERVs were found to be associated with clinical outcomes in the ICI arm. 47
Gene expression signatures
While tumor genomics has proven to be of limited use in predicting benefit from ICI therapies, expression analyses on the RNA level are increasingly helpful to gain insights into the complexities of the immune microenvironment and hold promise for RCC biomarker development in the context of immunotherapies.
Gene expression profiling (GEP) of bulk RNA has helped delineate different molecular subtypes of conventional RCC which were evaluated in several clinical trials (Table 2). Exploratory analyses from the phase II IMmotion150 study (atezolizumab plus bevacizumab versus atezolizumab versus sunitinib) identified three unique signatures including angiogenesis, pre-existing T effector immune activation, and myeloid inflammation. 48 Tumors with a high versus low angiogenesis signature were found to derive added benefit from sunitinib monotherapy, while such associations were not seen on the other treatment arms. Similarly, high versus low T-effector signature correlated with improved PFS and ORR for patients receiving atezolizumab plus bevacizumab and a trend for the same with atezolizumab alone, but this was not the case in patients treated on the sunitinib arm. Finally, in patients with T-effector high tumors and with a concurrent increase in myeloid infiltration, clear benefit with the addition of bevacizumab was observed, while the combination did not appear to add much in patients with tumors that were T effector high/myeloid low. 48
Emerging novel gene expression signatures in metastatic RCC.
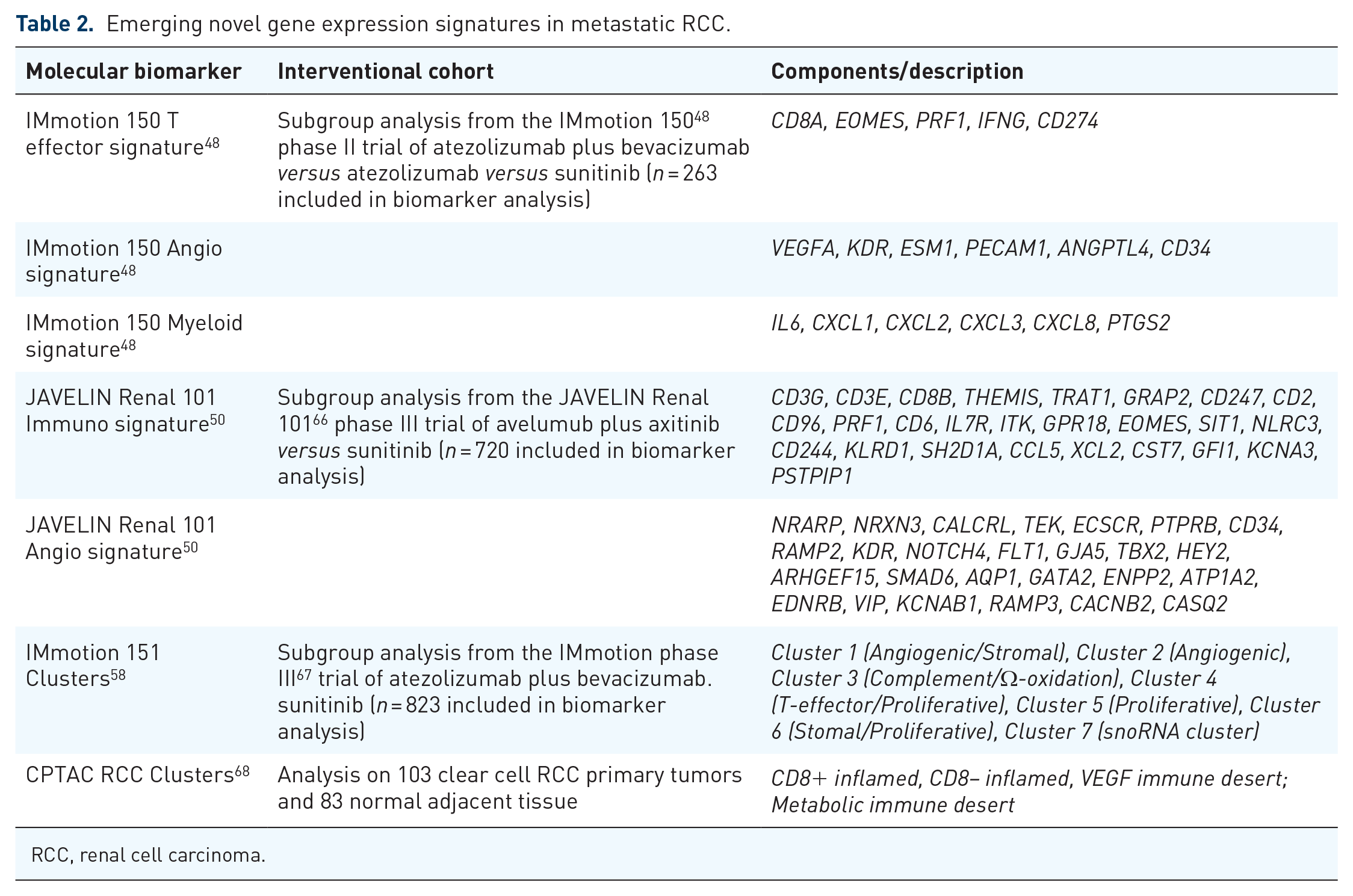
RCC, renal cell carcinoma.
A similar approach was conducted in the phase III clinical trial JAVELIN Renal 101 (avelumab plus axitinib versus sunitinib). 66 In this study, the investigators developed a novel immune GEP signature referred to as ‘JAVELIN Renal 101 Immuno’, which is derived from 26 genes involved in T-cell receptor signaling, proliferation, NK cell mediated cytotoxicity, chemokines, and other immune response genes. 50 In the avelumab plus axitinib arm, patients with high expression of this signature had improved median PFS of 15.2 months versus 9.8 months in those with lower expression (HR = 0.60, 95% CI = 0.44–0.83; p = 0.0019); there was no impact of this signature in the sunitinib arm. The authors also proposed a distinct GEP signature of 26 mostly angiogenic genes that could predict outcomes in the sunitinib arm referred to as ‘JAVELIN Renal 101 Angio’. In this study, Weighted Gene Coexpression Network Analysis was used to identify the Javelin Renal 101 transcriptomic signatures. 67
Different authors subsequently applied GEP signatures from IMmotion 150 and JAVELIN Renal 101 studies to the biomarker cohort of Checkmate 214 study (nivolumab plus ipilimumab versus sunitinib), 34 and no correlation with benefit from nivolumab plus ipilimumab was observed. 49 The investigators also conducted a hallmark gene set enrichment analysis as part of the biomarker analyses in Checkmate 214 and found that tumor necrosis factor alpha (TNF alpha) and epithelial-mesenchymal transition (EMT) gene sets expression was higher among patients with PFS >18 months treated with nivolumab plus ipilimumab. In a separate effort, authors assessed IMmotion 150 and JAVELIN Renal 101 GEP signatures in a pooled analysis from three nivolumab monotherapy trials and could not confirm an association between the signatures and clinical outcomes. 47
Investigators on the IMmotion 151 phase III study58,68 (atezolizumab plus bevacizumab versus sunitinib), using bulk RNA sequencing and targeted mutation sequencing by utilizing the non-negative matrix factorization method, 69 proposed an integrated molecular classification of metastatic ccRCC with delineation of seven transcriptionally defined and genomically annotated, biologically distinct categories of tumors. Treatment effects on the trial were then put in the context of these multi-omics subgroups. The authors broadly grouped patients into clusters with angiogenic phenotypes (clusters 1 and 2) and proliferative phenotypes (clusters 4–6). Clusters 1 and 2 are highly angiogenic, enriched with PBRM1 and KDM5 C mutation, and they were associated with improved outcomes to sunitinib. Cluster 3 had a high expression of cell cycle and complement cascade genes, as well as genes associated with the cytochrome P450 family, which is involved in omega oxidation. This cluster was also associated with low expression of angiogenesis and immune-related genes with relatively poor clinical outcomes regardless of the type of treatment received. Cluster 4 was enriched with T-effector gene signatures with improved outcomes to ICI and characterized as highly immunogenic with an expression of immune-related genes, high PD-L1 expression, and immune cell infiltration. Cluster 5 displayed a low rate of VHL mutation and included sarcomatoid dedifferentiation as well as 15 patients who were found to be of MiT family translocation RCC (tRCC). Cluster 6 was associated with prominent stroma and myeloid signatures. Overall, Clusters 5 and 6 were associated with improved outcomes to ICI but less pronounced compared with Cluster 4. Cluster 7 was a small cohort representing 3% of all tumors which is characterized by high expression of small nucleolar RNAs (snoRNAs) and low mutation rates.
Investigators with the Clinical Proteomic Tumor Analysis Consortium (CPTAC) recently reported comprehensive proteogenomic analyses of 103 ccRCC tumors and 83 samples of normal adjacent tissue. 70 Previously described GES were then applied. Based on the combined analyses of RNA, proteome, and phospho-proteome, four immune-based ccRCC were identified, discriminated by relative immune and stromal infiltration including (1)CD8+ inflamed; characterized by high CD8+ T cell infiltration, increased expression of the immune evasion markers PD1/PD-L/1PD-L2/CTLA4, high BAP1 mutations, and interferon-gamma signaling. This cluster was also strongly associated with elevated IMmotion 150 T effector signature; (2) CD8− inflamed; characterized by enriched signatures of platelet degranulation, EMT, complement, and coagulation cascades; (3) VEGF immune desert, characterized by enriched signatures of angiogenesis, notch signaling, and Rap1 signaling. Notably, IMmotion 150 angiogenesis signature was enriched in this cluster. (4) Metabolic immune desert, which displayed elevated MYC signatures, mTOR signaling, and a unique metabolic profile that included elevated mitochondrial, oxidative phosphorylation, and glycolysis protein. This cluster also displayed low immune, stromal, and microenvironment scores. These clusters may have an implication on systemic treatment outcomes and warrant further investigation.
Subsequent analyses put transcriptomic findings in context with similar deconvolution analysis across a large cohort of immune-responsive solid tumors including UC, non-small cell lung cancer, and RCC. 71 Relevant differences were noted across indications, both in terms of the immune microenvironment per se and the applicability of signatures. Different predictive models were built utilizing multiple machine learning approaches, and these failed to identify a unifying transcriptional signature predictive of response and showed poor performances in most cases. This highlights the important difference between the associations with better survival outcomes and predictive potential; the latter is an essential requirement for the application of these signatures in clinical practice.
Furthermore, novel sequencing strategies are approaching, particularly single cell RNA sequencing (scRNAseq). These are shedding lights on which cellular compartments are contributing to these expression signatures and help understand the dynamics of the immune microenvironment in much greater detail.72–74
Notable opportunities but also challenges lie in the task of integrating these various RNA-based signatures, making them available to the scientific community at large, and ideally simplifying them to the extent that molecular selection can be prospectively applied and integrated into innovative clinical trial concepts.
Peripheral blood biomarkers
Investigators have evaluated the impact of readily available, routinely used but also that of novel peripheral blood biomarkers in the context of ICI and metastatic RCC. Several authors have explored complete blood cell counts, particularly the distribution of leukocyte subsets in this regard. In a retrospective cohort of 90 patients receiving nivolumab therapy, the risk of progressive disease was increased with a higher baseline neutrophil-to-lymphocyte ratio (NLR) (HR = 1.86, 95% CI, 1.05–3.29), whereas a higher baseline eosinophil count was associated with a lower risk of progression (HR, 0.54; 95% CI, 0.30–0.98; p = 0.042). 75 In a similar analysis, a retrospective review of 142 patients treated with ICI suggested that early decline of NLR at 6 weeks after the initiation of ICI was associated with significantly improved outcomes. 76 In a cohort of 111 patients with metastatic ccRCC treated with nivolumab plus ipilimumab, low baseline neutrophil-to-eosinophil ratio (NER) was associated with significant improvement in PFS, OS, and ORR. However, low baseline NLR was only associated with significant improvement in OS but not in PFS or ORR. 77 In the JAVELIN Renal 101 study, low pretreatment NLR was associated with improved OS and a trend toward improved PFS in patients treated with avelumab plus axitinib. 78
C-reactive protein (CRP) is a known peripheral serum biomarker of inflammation, and prior studies showed that high baseline CRP is associated with worse OS in patients treated with systemic therapies, including VEGF-TKI and nivolumab.79,80 A recent retrospective study identified that a significant decline in CRP within 3 months of initiation of ICI is associated with improved ORR and PFS, suggestive that CRP kinetics could help to identify early disease progression. 81
Circulating cytokines have been studied in kidney cancer as a potential biomarker of systemic therapy, including with targeted therapies and recently with ICI.82,83 A recent prospective correlative study examined the association between distinct pre and on-treatment circulating cytokines with clinical outcomes in 56 patients with metastatic RCC treated ICI or VEGF-TKI. 84 Pretreatment cytokine levels did not have an impact on clinical outcomes from ICI in this cohort, but the investigators found that patients who derived clinical benefit (defined as PR, CR, or SD at 6 months) had higher relative increases in levels of IFN-gamma and IL-12 from baseline to 1 month on treatment.
In a pooled analysis from multiple randomized trials representing 1445 patients with metastatic UC or metastatic RCC treated with atezolizumab, 85 high baseline levels of IL-8 in plasma, peripheral blood mononuclear cells, and in tumors were associated with decreased efficacy of atezolizumab in patients with metastatic UC and metastatic RCC. In a phase II study of atezolizumab plus bevacizumab in patients with advanced nccRCC, circulating cytokines were collected at baseline and on treatment. 86 Results showed that high baseline levels of specific immunosuppressive cytokines (IL1α, IL6, CCL4, and IL13) were associated with inferior PFS and OS. A decline in these markers on-therapy was not predictive of outcomes.
Human leukocyte antigen diversity
Human leukocyte antigen (HLA) genotyping plays a critical role in immunogenic antigens presentation and could impact outcomes to ICI. Prior studies have demonstrated that T-cell receptor (TCR) clonality is associated with HLA class I genetic variability and evolutionary divergence, which in turn are associated with ICI response in melanoma. 87 In RCC, recent biomarker effort from the phase Ib/2 KEYNOTE 146 of pembrolizumab plus lenvatinib in metastatic ccRCC identified that high HLA-I evolutionary divergence, which is a measure of functional divergence between the two HLA alleles, was strongly predictive of longer PFS, as well as longer duration of response, 88 while zygosity at HLA-class I loci was not. Furthermore, analysis from JAVELIN Renal 101 showed impact of HLA typing on treatment outcomes, in particular, five HLA alleles (A∗01:01, A∗03:01, B∗40:02, B∗57:01, and C∗06:02) were associated with treatment arm-specific differences in PFS. 50 No correlation with survival was seen in a third report which investigated HLA class I heterozygosity in nivolumab monotherapy-treated patients pooled from three separate trials. 47
Microbiome
In recent years, the microbiome has emerged as a key component of the host immune system and has been increasingly recognized for its role in cancer biology. 89 This extends to its interplay with cancer therapies, and several studies have pointed to an association between the gut microbiome and outcomes to ICI therapy in solid tumors, highlighting the significance of microbial diversity but also specific bacterial species in this context.90,91
A previously reported prospective observational study collected stool from 31 patients with metastatic RCC prior to initiating ICI therapy (including nivolumab monotherapy and nivolumab plus ipilimumab combination). 92 Gut microbiota composition was assessed using whole-genome shotgun metagenomic sequencing. Clinical benefit [defined as partial response (PR), complete response (CR), or stable disease (SD) > 4 months] was associated with greater alpha diversity and the relative abundance of Akkermansia spp. In a separate study, investigators analyzed the gut microbiome from stool samples obtained from 69 advanced RCC patients who were treated with nivolumab as part of the phase II GETUG-AFU 26 NIVOREN. 93 Their work identified that recent antibiotic use reduced the ORR to nivolumab (from 28% to 9%, p < 0.03), which the authors speculated may have been related to the effect of antibiotics on modulating the gut microbiome to a less favorable ‘gut immune-environment’.
Building on the knowledge of the gut microbiome’s role in modulating outcomes to ICI, a recent phase II study investigated the use of nivolumab plus ipilimumab with or without the addition of CBM-588 in patients with metastatic ccRCC. 94 CBM-588 is a probiotic comprised of Clostridium butyricum, which was shown to augment the activity of ICI in non-small cell lung cancer through the modulation of gut microbiota. 95 In this study, among 29 evaluable patients with metastatic ccRCC, the primary endpoint of detectable change in gut microbiome was not reached, however, the ORR was significantly higher among patients receiving nivolumab plus ipilimumab plus CBM-588 compared with those receiving nivolumab plus ipilimumab (59% versus 11%; p = 0.024).
Biomarker-driven clinical trials
With the considerable progress in molecular diagnostics, biomarkers-driven clinical trials have gained interest in recent years. The BIONIKK trial 96 was a phase II biomarker-driven clinical trial evaluating nivolumab versus nivolumab plus ipilimumab versus VEGF-TKI (sunitinib or pazopanib) with treatment assignment per baseline molecular strata. The design of the study was informed by transcriptomic analysis conducted by Beuselinck et al. 97 based on the tumor microenvironment features of 53 primary ccRCC tumors from patients who were treated with sunitinib in the first-line setting. They identified four distinct ccRCC clusters (ccrcc1 to 4); ccrcc1 ‘immune-low’ and ccrcc4 ‘immune-high’ tumors were associated with worse outcomes to sunitinib. In contrast, ccrcc2 ‘angio-high’ and ccrcc3 ‘normal-like’ tumors were associated with improved outcomes to sunitinib. According to ccrcc1-4 clusters, patients were randomized to nivolumab or nivolumab plus ipilimumab or VEGF-TKI (sunitinib or pazopanib) with the primary endpoint of ORR. Patients within ccrcc 2 ‘angio-high’ cluster achieved high responses with VEGF-TKI as expected given the enriched angiogenic signatures in this group. ORR was comparable between nivolumab and nivolumab plus ipilimumab in the ‘immune-high’ ccrcc4 group; however, ORR for nivolumab plus ipilimumab was numerically higher than nivolumab alone in the ccrcc1 group ‘immune-low’. The study highlights the need for adequate sample sizes in multi-arm biomarker directed trials, but <1 month turnaround of RNA-based profiling demonstrates that expression-based treatment assignment can be achieved and hence sets the bar for future investigations. Table 3 summarizes some of the ongoing biomarker-driven clinical trials in clear cell RCC.
Select ongoing biomarker-driven clinical trials in metastatic RCC.
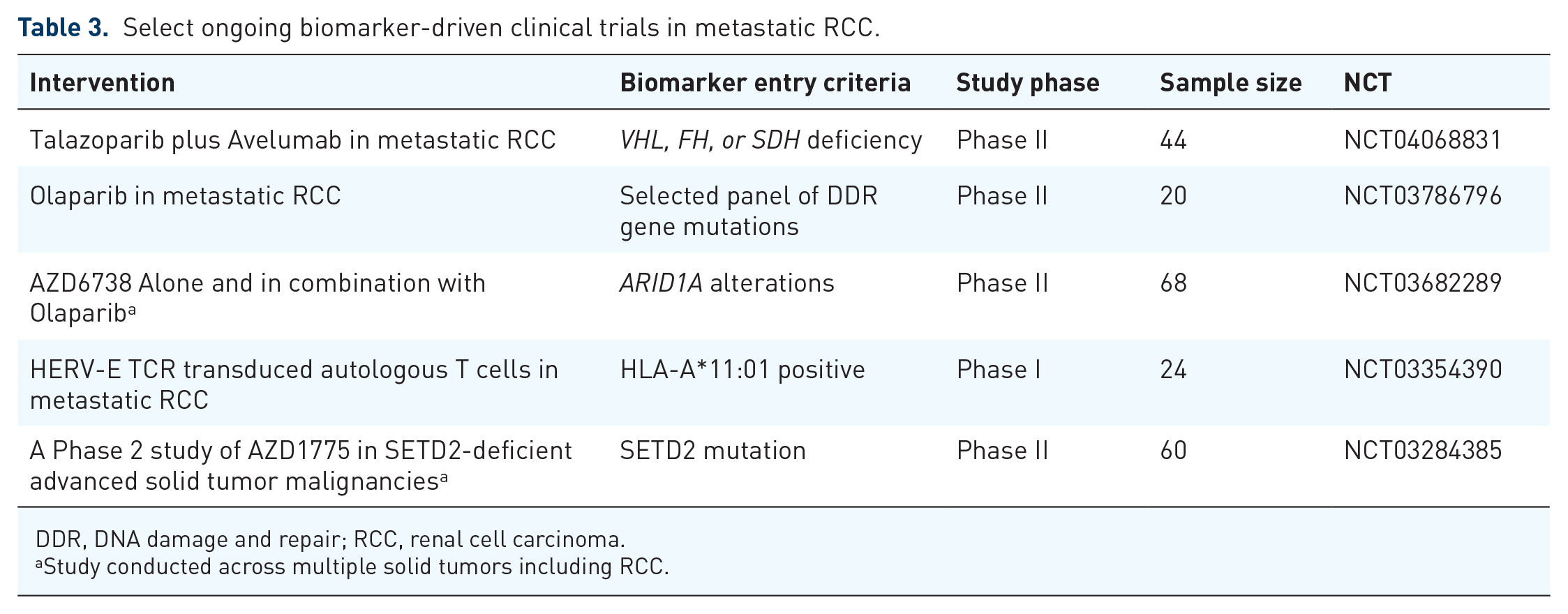
DDR, DNA damage and repair; RCC, renal cell carcinoma.
Study conducted across multiple solid tumors including RCC.
Conclusion
With the tremendous expansion in therapeutic armamentarium of kidney cancer, the fast pace of immunotherapeutic innovation, and rapidly evolving methodology to molecularly annotate patient-derived specimens, there is an unprecedented need to develop clinically applicable biomarkers. Novel biomarker discovery endeavors should focus on incorporating different types of biomarkers (clinical, genomic, transcriptomic, blood-based variables) in an effort to build a composite ‘score’ that would help to assess more comprehensively the molecular characteristics of the tumor and the corresponding response to ICIs. The emphasis of the field must be on delineation of rational molecular endpoints which can inform the design of large registries and ultimately that of innovative, biomarker-driven clinical trials. For this bright future in kidney cancer biomarkers discovery to materialize and have the clinical impact we seek, we need to learn from prior failures and change the culture to pursue biomarker discovery early, as an integral part of all prospective efforts of developing novel therapeutic strategies.
Footnotes
Author contributions
YG: Concept, manuscript writing and apporoval
MHV: Concept, manuscript writing and apporoval
Conflict of interest statement
The authors declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: M.H.V. reports receiving commercial research grants from Bristol-Myers Squibb, Pfizer, and Genentech/Roche; honoraria from Novartis, BMS travel/accommodation from Astra Zeneca, Eisai, Novartis, and Takeda; consultant/advisory board member for Alexion Pharmaceuticals, Aveo, Bayer, Calithera Biosciences, Corvus Pharmaceuticals, Exelixis, Eisai, GlaxoSmithKline, Merck, and Natera; Onquality Pharmaceuticals; Novartis; and Pfizer. Y.G. is consultant/advisory board member for Exelixis and Aveo.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
