Abstract
Background:
EGFR/c-Met activation/amplification and co-expression, mTOR upregulation/activation, and Akt/Wnt signaling upregulation have been individually associated with more aggressive disease and characterized as potential prognostic markers for lung cancer patients.
Methods:
Tumors obtained from 109 participants with stage I–IV non-small cell lung cancer (NSCLC) were studied for EGFR/c-Met co-localization as well as for total and active forms of EGFR, c-Met, mTOR, S6K, beta-catenin, and Axin2. Slides were graded by two independent blinded pathologists using a validated scoring system. Protein expression profile correlations were assessed using Pearson correlation and Spearman’s rho. Prognosis was assessed using Kaplan–Meier analysis.
Results:
Protein expression profile analysis revealed significant correlations between EGFR/p-EGFR (
Conclusions:
These findings suggest that increased EGFR/c-Met signaling is correlated with upregulated mTOR/S6K signaling, which may in turn be associated with shorter median survival in late-stage NSCLC.
Introduction
Lung cancer is the leading cause of cancer mortality worldwide. In 2019, a projected 142,670 people in the United States died from lung cancer, more deaths than those attributed to colon, breast, prostate, and ovarian cancers combined. 1 Among diagnosed lung cancer cases, 80–90% are classified as non-small cell lung cancer (NSCLC), the form of lung cancer investigated in this study. 2 Approximately 75% of NSCLC cases are locally advanced or metastatic (stage III–IV) at the time of diagnosis, obviating the role of surgery. 3 Over the past decade, the development of targeted tyrosine kinase inhibitor (TKI) therapies and guideline-directed screening for prognostic and predictive molecular markers has led to a significant improvement in progression-free survival. Specifically, routine testing for EGFR, ALK, ROS1, BRAF, MET, ERBB2 (HER2), KRAS, and RET is recommended by the International Association for the Study of Lung Cancer (IASLC) and the Association for Molecular Pathology.4–16 Nevertheless, NSCLC-attributed mortality rates remain high.1,17 In addition, studies have elucidated the role of upregulation of molecular signaling pathways, including EGFR, c-Met, mTOR, and Akt/Wnt, in the pathogenesis of NSCLC.18,19
c-Met, also known as the hepatocyte growth factor receptor (HGFR), is a membrane-bound Receptor Tyrosine Kinase (RTK) that is coded for by the MET gene. When amplified/activated, c-Met plays a role in tumor growth, angiogenesis, and metastasis.20,21 In addition, amplified c-Met expression has been correlated with poor prognosis in other tumor types, including liver, gastric, breast, and brain malignancies. 21 Activation of c-Met signaling has also been associated with upregulation of a variety of downstream signaling pathways, including the RAS/MAPK, PI3K/Akt, and Wnt pathways.22–24 In NSCLC, co-localization of c-Met and epidermal growth factor receptor (EGFR) has been shown to have a synergistic effect on cell proliferation, downstream activation of signal transduction, and may indicate a more aggressive tumor phenotype that results in worse prognosis. 25
The mammalian target of rapamycin/S6 protein kinase (mTOR/S6K) pathway is a key intracellular regulator of cell growth, survival, migration, and invasion in NSCLC.26,27 Increased mTOR/S6K activity has been associated with poor clinical prognosis in early-stage NSCLC.18,19,26,28 In both pre-clinical studies and clinical trials, the inhibition of mTOR has demonstrated potential benefit when employed to treat NSCLC.29,30 Clinical trials have also been conducted to test the efficacy of mTOR inhibitors in combination therapy. Therefore, the study of multiple target proteins found in tumors for use in combination could help improve patient prognosis. 31
The Wnt pathway proteins consist of highly conserved glycoproteins that induce downstream signaling upon binding to the activated Frizzled transmembrane receptor, a mechanism of action which stabilizes and inhibits the degradation of beta-catenin.
32
Active beta-catenin is the nuclear, signaling form of beta-catenin that is transcriptionally active and unphosphorylated at serine 37 and threonine 41.33,34 Stabilization of beta-catenin leads to transcriptional activation of genes involved in cellular proliferation.32,35 Axin2, a negative regulator of beta-catenin, functions by facilitating the phosphorylation of beta-catenin and thereby tagging it for ubiquitin-mediated destruction.
32
Various studies have implicated increased nuclear beta-catenin in both the oncogenesis and pathogenesis of NSCLC, particularly in lung tumors with EGFR mutations.35,36 In addition,
Correlations in protein expression and activation patterns between EGFR, c-Met, EGFR/c-Met co-localization, mTOR, S6K, beta-catenin, and Axin2 in lung tumors are not well established. Currently, few studies have investigated the roles of the Wnt pathway; Axin2, a negative regulator of the Wnt pathway; and EGFR/c-Met co-expression on activation of the downstream mTOR pathway. Furthermore, the fundamental role of mTOR in late-stage NSCLC prognosis is not clearly defined. In response to these deficiencies, this novel study aimed to evaluate correlations in protein expression and activation patterns between EGFR, c-Met, EGFR/c-Met co-localization, mTOR, S6K, beta-catenin, and Axin2 to determine if these signaling pathways were concomitantly upregulated
Materials and methods
Case selection, tissue acquisition, and exclusion criteria
All patients at participating institutions with a diagnosis of stage I–IV NSCLC were considered for participation in the study. Samples were obtained in accordance with approval from the University of Illinois College of Medicine at Rockford Institutional Review Board (IRB) under approval number 351597-11. Written informed consent was obtained from all patients prior to their inclusion in the study. This consent form emphasized how participation in the study was voluntary and all tissue samples were de-identified to ensure patient confidentiality. Patients were excluded from participating if they declined to consent to the use of their tissue. A total of 109 patients was identified. All samples used in this study were collected from Winnebago County in Illinois, which has a 14% higher age-adjusted mortality rate (2016) due to lung cancer compared with the national rate and a 20% higher average annual age-adjusted incidence rate (2011–2015) of lung cancer compared with the national rate.37,38
Tissue de-identification and study blinding
A research coordinator not involved in staining, grading, or data analysis was responsible for de-identifying the tissue blocks and labeling them with randomly generated numerical study codes. All investigators involved in processing, staining, and grading of the tissues were blinded to all patient data and outcomes until all cases were scored and re-associated with the de-identified outcome data and clinical treatment for statistical analysis.
Tissue preparation
Tissues were fixed in 10% neutral buffered formalin for a minimum of 6 h and dehydrated in increasing Ethanol (EtOH) concentrations, followed by infiltration with xylene and paraffin. Paraffin blocks were prepared by being embedded in hot wax and then cooled on a cold plate. Fresh slides were sectioned from the coded, paraffin-embedded tissue blocks using a microtome to cut 4-micron thick sections, which were mounted on glass slides.
Immunohistochemistry staining protocol
In brief, protein expression of interest was examined retrospectively in NSCLC patients through immunohistochemistry (IHC) using formalin-fixed paraffin embedded tumor samples and individual probes, as described earlier. 39 Samples were double-stained during assessment of co-localization. Positive and negative staining controls used for each protein marker are listed in Supplemental Data Table S1. Positive control tissues used were known to express the targeted protein while negative controls were obtained through omission of primary antibody on a positive control tissue.
Reagents for IHC staining
Avidin/biotin blocking kit (# SP-2001), normal horse serum (# S-2000), normal goat serum (# S-1000), VECTASTAIN Elite ABC kit (# PK-6100), VECTASTAIN ABC-AP kit (# AK-5000), ImmPACT DAB peroxidase substrate (# SK-4105), and VECTOR red alkaline phosphatase substrate kit (# SK-5100) were obtained from Vector Laboratories (Burlingame, CA, USA). Hematoxylin was obtained from Sigma-Aldrich (St Louis, MO, USA). Permount was obtained from Fisher Scientific (Pittsburgh, PA, USA). Antibodies used were Horseradish Peroxidase (HRP)-conjugated anti-EGFR (1:2000 dilution) Thermo Fisher Scientific (Rockford, IL, USA) cat.# PA1-1110S-HRP, Anti-p-c-Met (1:1000 dilution) Life Technologies (Grand Island, NY, USA) cat.# 3077S, biotinylated anti-mouse IgG (1:250 dilution) Vector Laboratories (Burlingame, CA, USA) cat.# BA-2000, and biotinylated anti-rabbit IgG (1:250 dilution) Vector Laboratories (Burlingame, CA, USA) cat.# BA-1000. Remaining antibodies obtained from Cell Signaling Technology (Danvers, MA, USA) were the following: Anti-p-EGFR (1:3200 dilution) cat.# 2236S, Anti-c-Met (1:300 dilution) cat.# 8198P, Anti-mTOR (1:50 dilution) cat.# 2983, Anti-p-mTOR (1:100 dilution) cat.# 2976S, Anti-p70 S6K (1:400 dilution) CST cat.# 2708S, and Anti-pp70 S6K (1:150 dilution) cat.# 9234S. All products were stored and used according to manufacturers’ instructions.
IHC staining of control tissues
Positive control tissues were known to express the target protein, while negative controls omitted primary antibody on positive control tissue. In addition, cells in the normal areas of the lung which do not stain were also used as an internal negative control. Positive controls were placenta for EGFR, EGF-treated H2170 cells for p-EGFR, liver/kidney for c-Met, HGF-treated H2170 cells for p-c-Met, validated NSCLC tissue for mTOR, EGF-treated H2170 cells for p-mTOR, liver/spleen/validated NSCLC tissue for S6K, and HGF-treated H2170 cells for p-S6K. H2170 cell line protein expression patterns were verified by western blotting, as described earlier. 18
Grading of stained slides
Staining intensity was graded by two blinded pathologists on a 0–3 scale, with 0 = no staining, 1 = weak staining, 2 = moderate staining, and 3 = strong staining. The grading pathologist also recorded the percentage area of tumor cells staining positively for each corresponding intensity score. The overall graded score was obtained by multiplying the intensity scores (from 0 to 3) by the corresponding percentage of positively stained cells (from 0% to 100%) with a scale ranging from 0 to 300. The equation for obtaining the grading score is as follows: %Tumor Grade 3*3 + %Tumor Grade 2*2 + %Tumor Grade 1*1 + %Tumor Grade 0*0, as described earlier. 40
Statistical analysis
Two-tailed
Results
Demographics
Table 1 presents the demographics information for tissues obtained from the 109 participants in this study. The table includes information on patient sex, race, smoking status, smoking history, histology, cancer stage at diagnosis, and treatment received (Table 1). When comparing demographic characteristics between high and low expression groups for each protein of interest, we found several significant differences. Late-stage (stage IIIb–IV) patients were more likely than early-stage (stage I–IIIa) patients to show high expression of EGFR (
Demographics table for NSCLC patients.
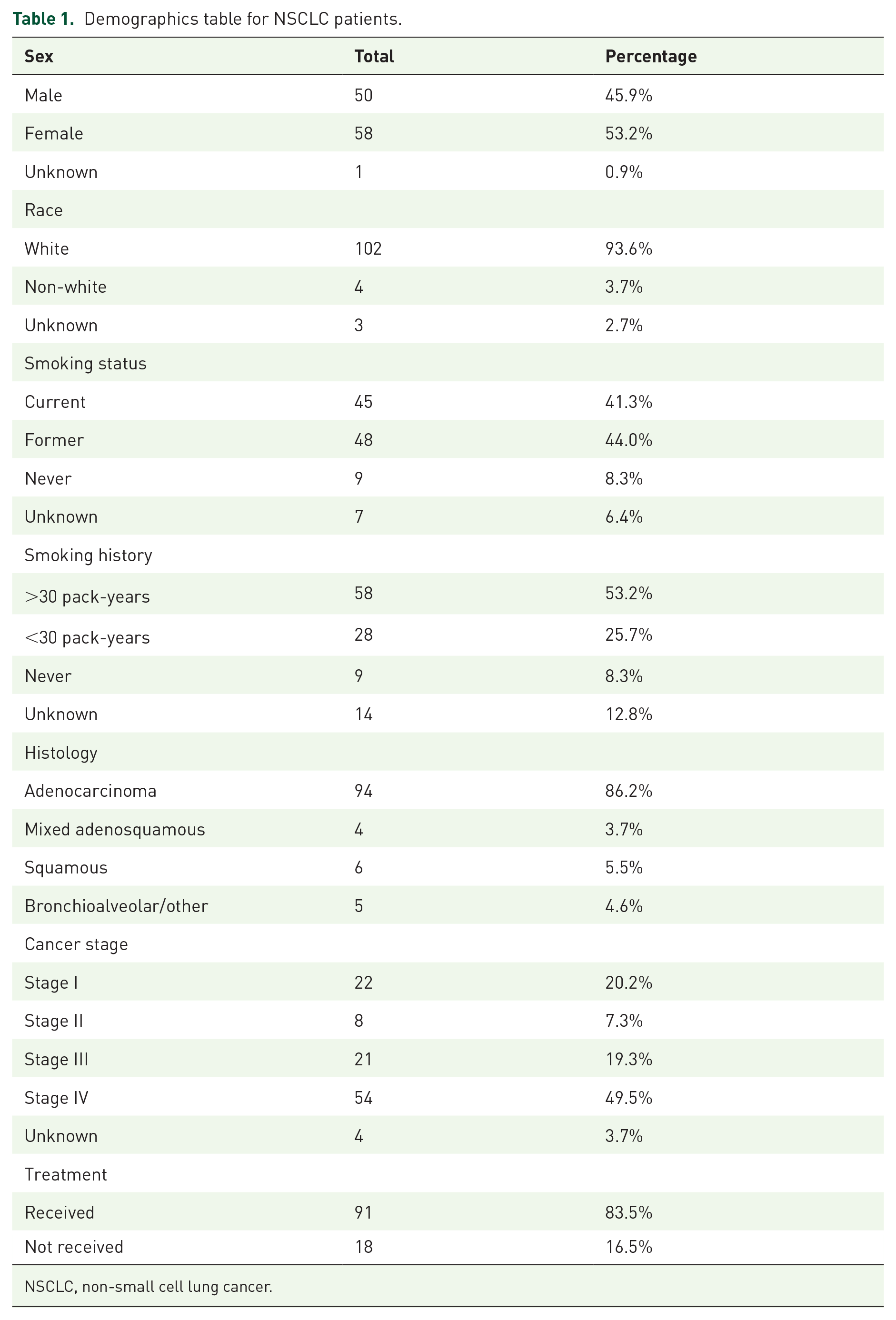
NSCLC, non-small cell lung cancer.
Sample distribution
Protein expression/activity levels were not normally distributed, as indicated by both the Kolmogirov–Smirnov and the Shapiro–Wilk tests.
Protein expression patterns and profile correlations
In general, most of the proteins were expressed over a broad range, from a minimum of 0 to a maximum of 300. Total beta-catenin was more heavily expressed in all samples, with a minimum expression of 100 and maximum of 300. p-c-Met and p-mTOR had a minimum expression of 0 and a maximum expression of 280. Mean expression was higher for S6K and p-S6K than mTOR and p-mTOR, a possible result of signal amplification and the capacity of one molecule of p-mTOR to activate multiple p-S6Ks. Standard deviation was lower on average for the total proteins, with larger standard deviations seen in activated/phosphorylated proteins. S6K and p-S6K demonstrated relatively higher levels of expression with the lowest variation. As S6K was the furthest downstream marker assayed in this study, this result might represent a confluence of various cell signaling pathways converging on this signaling protein (Figure 1).

Immunohistochemistry of different biomarkers in tissue biopsies of patients with NSCLC. (a) EGFR and p-EGFR. (b) mTOR and p-mTOR. (c) c-Met and p-c-Met. (d) S6K and p-S6K. (e) Double staining for EGFR/c-Met and p-EGFR/p-c-Met. EGFR/p-EGFR is brown, c-Met/p-c-Met is red. (f) Total beta-catenin and double staining of active beta-catenin/Axin2. Beta-catenin is brown and Axin2 is red. NSCLC, non-small cell lung cancer.
We observed that EGFR expression was correlated with downstream mTOR expression and p-S6K activity (
Total beta-catenin was correlated with total and active EGFR and S6K as well as total Axin2 (all
Spearman and Pearson correlations between selected biomarkers.
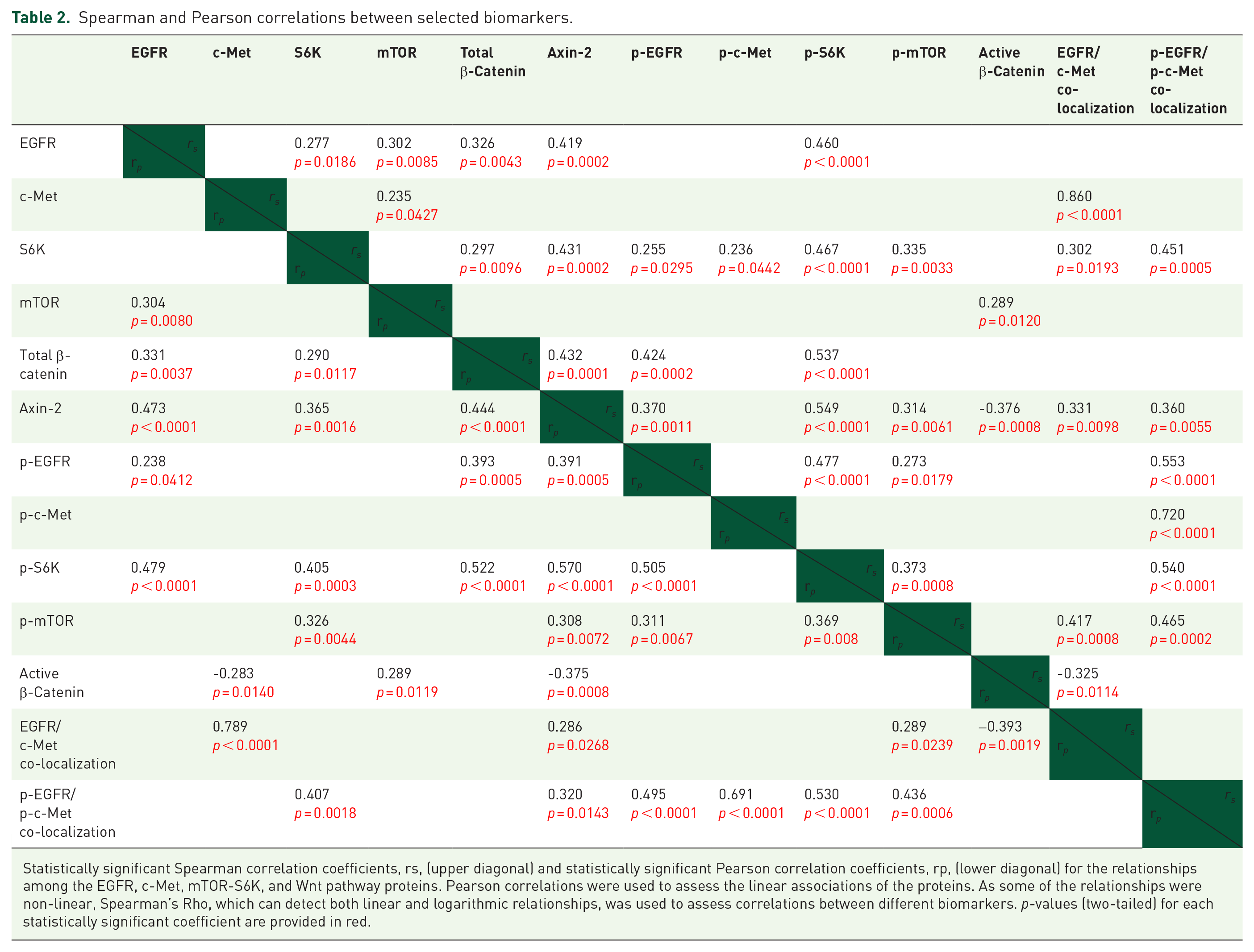
Statistically significant Spearman correlation coefficients, rs, (upper diagonal) and statistically significant Pearson correlation coefficients, rp, (lower diagonal) for the relationships among the EGFR, c-Met, mTOR-S6K, and Wnt pathway proteins. Pearson correlations were used to assess the linear associations of the proteins. As some of the relationships were non-linear, Spearman’s Rho, which can detect both linear and logarithmic relationships, was used to assess correlations between different biomarkers.
Clinical outcomes and protein expression in patients not stratified for cancer stage
KM analysis comparing high and low expression groups revealed that the elevation in expression and activation of several proteins at the time of diagnosis was associated with significantly worse patient prognosis. In particular, for the following proteins, high expression was associated with shorter median survival compared with low expression: EGFR (13.0 months

Kaplan–Meier survival analyses not stratified by cancer staging demonstrated that increased expression levels of total S6K, total beta-catenin, Axin2, p-mTOR, p-S6K, EGFR/c-Met co-expression, and EGFR were significantly associated with shorter median survival. Median expression level of each biomarker was chosen as the cut-off between the high and low expression groups. Equality of survival distributions for the two expression groups were studied using the log rank test.
In summary, amplified expression of EGFR and elevated co-localization of EGFR/c-Met, along with amplified expression and activation of the mTOR-S6K pathway at the time of diagnosis, were associated with worse prognosis. Increased total beta-catenin expression was also associated with worse prognosis. Axin2, which may serve as an indicator of downregulated active beta-catenin, was similarly associated with a worse prognosis.
Early versus late-stage disease analysis
In response to the concurrent observations that patients with high expression levels were more likely to be diagnosed at late stages and less likely to undergo surgical resection, we performed a KM analysis of the survival data. To do this analysis, we stratified survival into early (I–IIIa) and late (IIIb–IV) disease categories, depending on stage at the time of diagnosis. Patients with late-stage disease who had high levels of mTOR and p-mTOR protein expression had shorter median survival compared with those with late-stage disease and lower levels of protein expression (mTOR: 6.7

Kaplan–Meier survival analyses stratified by stage IIIB–IV NSCLC demonstrated that increased co-expression of EGFR/c-Met and increased expression of p-mTOR and total mTOR were significantly associated with shorter median survival. Median expression level of each biomarker was chosen as the cut-off between the high and low expression groups. Equality of survival distributions for the two expression groups were studied using the log rank test.
In summary, after controlling for stage of disease at diagnosis, high levels of mTOR expression, p-mTOR activation, and EGFR/c-Met co-localization were associated with a significantly shorter median survival compared with those with lower expression/activation/co-localization.
Discussion
In general, the diagnosis of NSCLC correlates with poor prognosis. 42 The current availabilities of targeted TKI therapy and guideline-directed screening for molecular markers have led to a significant improvement in progression-free survival.9,10,13,14 Despite an advanced understanding of NSCLC tumor biology and treatment, NSCLC-attributed mortality rates remain high.1,17 This observation has prompted further investigation into the tumor biology of NSCLC, and multiple mechanisms which appear to play a role in NSCLC pathogenesis and resistance to TKIs have been elucidated here.4,6–8,12,15,16
We observed that EGFR and p-EGFR expression were correlated with downstream expression and activity of mTOR-S6K and Axin2. Interestingly, p-EGFR/p-c-Met co-localization was strongly correlated with increased mTOR-S6K activation. This implies that signaling through co-localized activated EGFR/c-Met may be a potent activator of the mTOR-S6K pathway.18,19 Total beta-catenin was correlated with various proteins in the EGFR and mTOR-S6K pathways in addition to Axin2, a negative regulator of the Wnt signaling pathway.
41
Finally, strong correlations of Axin2 with the majority of total and active EGFR-mTOR-S6K signaling proteins may imply that activation of this signaling axis may lead to downregulation of Wnt signaling through the negative regulatory effects of Axin2 and by promotion of phosphorylation and degradation of beta-catenin. These associations suggest that
Furthermore, studies suggest that the mTOR pathway may play a key role in NSCLC pathogenesis.12,15 Increased downstream mTOR activity has been associated with EGFR, PI3K/Akt, and c-Met as well as various other signaling proteins (Figure 4).6,15,43 This proposed mechanism is consistent with data published from the The Cancer Genome Atlas (TCGA) database evaluating tumors across multiple cancer types, including more than 500 lung adenocarcinomas, which suggests that EGFR, PI3K, and Wnt pathways play a significant role in oncogenic signaling utilizing crosstalk through positive and negative regulators, including Axin2 and GSK-3. 44 Our previous mechanistic studies in H2170 and H358 cell lines made resistant to EGFR and c-MET tyrosine kinase inhibitors found 2–8-fold upregulation of mTOR (p-S6K, and p-mTOR) and Wnt (active beta-catenin) signaling proteins. 18 Additional studies supporting our proposed mechanism were completed in H2170 and H1975 TKI-resistant cell lines and found key Wnt and mTOR proteins to be differentially modulated, including upregulation of active beta-catenin and p-GSK-3β in H2170 cells. 19 In H1975 cells combination treatment with an mTOR inhibitor (everolimus) and erlotinib resulted in synergistic cell growth inhibition. 19 In clinical studies, patients with early-stage disease, high mTOR activity, and histological evidence of angioinvasion were reported to have poor prognosis. 26 Here, we observed that patients across all stages of NSCLC who expressed high levels of EGFR, mTOR, p-mTOR, S6K, p-S6K, total beta-catenin, Axin2, and EGFR/c-Met co-localization at the time of diagnosis had a significantly worse prognosis compared with those with low expression levels. Alternatively, those with higher expression and activation of the previously mentioned proteins had an increased likelihood of advanced disease diagnosis and were less likely to undergo surgical resection, two potential contributions to worse prognosis. Even after controlling for stage at diagnosis, patients with elevated levels of mTOR and p-mTOR and co-localized EGFR/c-Met were still found to have a worse prognosis.

A proposed mechanism for the EGFR/c-Met-mTOR/S6K signaling axis in NSCLC. Both EGFR and c-Met expression can lead to activation of the PI3K, mTOR, and MAPK pathways, resulting in increased cell proliferation, survival, migration, and angiogenesis through downstream targets such as Cyclin D1, p21, Bcl-2, wee-1, c-fos, c-jun and fosL1.45–49 In addition, Wnt pathway activation leads to the recruitment of Axin2, CKI, and GSK-3, allowing for nuclear accumulation of active beta-catenin and enhanced transcription. Finally, it is shown that upregulation of Axin2, a negative regulator of the Wnt signaling pathway, can bind to GSK3 and lead to phosphorylation and degradation of beta-catenin. NSCLC, non-small cell lung cancer.
The association of mTOR activity with poor prognosis may be due to a variety of mechanisms. The mTOR protein itself is a well-characterized mitogenic signaling protein that regulates downstream molecules such as cyclin D, an integral player in cell cycle progression at the G1-S checkpoint. 50 Therefore, no matter the stage, a NSCLC tumor expressing increased levels of total and active mTOR may be inherently more capable of proliferation and thus be more clinically aggressive. Conversely, it may be possible that amplified protein expression or activation is a mere reflection of disease that has already advanced. The mTOR pathway also lies downstream from several proteins involved in proliferation and survival, as well as other mediator proteins, which utilize mTOR signaling to achieve their effects.51,52 This suggests that mTOR activity, specifically at the time of diagnosis, may reflect the downstream summation of multiple upregulated signaling pathways. If true, this could explain why some studies have found no correlation between increased mTOR and prognosis, while others have.53,54 It is also a possibility that upstream, activating mediators of the mTOR pathway drive disease phenotypes by activating alternative or complementary pathways. By way of this reasoning, perhaps elevated EGFR/c-Met co-localization leading to upregulated mTOR signaling is associated with worse prognosis at the time of diagnosis, whereas alternative upstream pathways leading to mTOR may not necessarily be so. Currently, few studies have evaluated whether, in the presence of increased mTOR-S6K, variations in signaling proteins upstream of mTOR-S6K are better predictors of prognosis. Our study has investigated this question.
As seen in the results of this study, EGFR expression and activation, when co-expressed with c-Met, demonstrated a statistically significant positive correlation with mTOR activation. This suggests that the mTOR pathway acts as a downstream mediator through which EGFR signaling plays a role in NSCLC pathogenesis.6,7,12,15 This model is further supported by the observation that expression of p-S6K, a downstream mTOR pathway protein, is positively correlated with EGFR expression and activation. It is also notable that studies have observed EGFR-c-MET heterodimerization, 55 co-precipitation 56 and also that both c-Met (61%) and EGFR (80%) are highly expressed in NSCLC tumors.39,57 Earlier studies have also shown a synergistic effect on inhibition of proliferation in the presence of EGFR and c-MET inhibitors leading to synergistic inhibition of downstream signaling. 25 Therefore, double IHC staining showing the co-localization of p-EGFR and p-c-MET in patient tumors is suggestive of crosstalk and increased activation of both receptor tyrosine kinases. In addition, the correlation of increased p-EGFR/p-c-MET with upregulation of downstream p-mTOR and p-S6K in human tissues offers clinical data to support our prior mechanistic pre-clinical studies reporting that EGFR and c-MET may signal through mTOR/S6K to mediate their oncogenic potential. 18
Additional studies should be done to characterize mTOR expression profiles completely in a larger NSCLC cohort and determine their clinical relevance. More studies are also needed to validate prospectively whether mTOR and p-mTOR testing in advanced NSCLC may be a viable prognostic biomarker for clinicians at diagnosis. Furthermore, as targeted mTOR inhibitors become available for the standard of care treatment of NSCLC, future studies may elucidate whether testing for mTOR and p-mTOR may provide a level of personalized medicine through predicting which patients are likely to benefit from mTOR inhibitor therapy. Finally, among the biomarkers investigated, molecular alterations have been found in EGFR, c-MET, Axin-2, and beta-catenin.58–61 In the future, it would be interesting to study if molecular alterations contribute to modulation in expression of these biomarkers.
Conclusion
We conclude that EGFR/c-Met co-expression and the correlated activation of the mTOR pathway represents a possible mechanism for a more aggressive NSCLC tumor phenotype and may be clinically associated with shorter median survival in late-stage NSCLC patients (Figure 3). In addition, Axin2 may also be a predictive biomarker for lung cancer prognosis.
Supplemental Material
Supplementary_Data_Final – Supplemental material for EGFR/c-Met and mTOR signaling are predictors of survival in non-small cell lung cancer
Supplemental material, Supplementary_Data_Final for EGFR/c-Met and mTOR signaling are predictors of survival in non-small cell lung cancer by Zachary D Crees, Caleb Shearrow, Leo Lin, Jennifer Girard, Kavin Arasi, Aayush Bhoraskar, Joseph Berei, Adam Eckburg, Austin D. Anderson, Christian Garcia, Ariana Munger, Sunil Palani, Thomas J Smith, Shylendra B Sreenivassappa, Connie Vitali, Odile David and Neelu Puri in Therapeutic Advances in Medical Oncology
Footnotes
Conflict of interest statement
The authors declare that there is no conflict of interest.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Research reported in this publication was supported by the National Cancer Institute of the National Institutes of Health under award number R21CA158965-01A1(![]() ) and Community Grant from the Community Foundation of Northern Illinois (Grant ID # I3736) to Neelu Puri. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
) and Community Grant from the Community Foundation of Northern Illinois (Grant ID # I3736) to Neelu Puri. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
