Abstract
Background:
Emerging evidence indicates that gut inflammatory and immune response play a key role in the pathophysiology of stroke and may become a promising therapeutic target. However, the specific role of the microbiota-gut-brain axis in poststroke aphasia (PSA) patients remains unclear.
Objectives:
The aim of this study was to investigate the relationships among the gut microbiota, neuroendocrine-immune network, brain network properties, and language function in patients with PSA.
Design:
This is a cross-sectional, observational, monocentric study.
Methods:
This study enrolled 15 PSA patients, 10 non-PSA patients, and 15 healthy controls (HCs). All subjects underwent stool microbiota analysis, blood inflammatory cytokines assessment, and brain-gut peptide examination. PSA patients and HCs underwent additional resting-state functional MRI (rs-fMRI) brain scans. The rs-fMRI data were utilized to create whole-brain connectivity maps, and graph theory was employed to characterize the network topological properties. Analysis of variance and the Kruskal–Wallis test were used for comparisons among the three groups. Correlation analyses were subsequently conducted to explore relationships among factors showing significant group differences.
Results:
Compared with non-PSA patients and HCs, PSA patients displayed alterations in the gut microbiota composition, increased systemic inflammation, changes in brain-gut peptides, and had worse language performance. Graph theoretical analysis revealed that PSA patients exhibited small-world topology. Furthermore, nodal measures in brain network analysis showed activation of homologous speech areas in the right hemisphere, while the nodal properties of brain regions near the lesion in the left hemisphere decreased in patients with PSA compared with HCs.
Conclusion:
The present study revealed, for the first time, that an imbalance in gut microbiota was accompanied by the neuroendocrine-immune network disorder and abnormal changes in the brain network in PSA patients.
Plain language summary
Aphasia is one of the most common complications of stroke, affecting approximately one-third of stroke survivors. Poststroke aphasia (PSA) is associated with more severe strokes, higher mortality, slower functional recovery, and greater healthcare costs compared to stroke patients without PSA. The unique characteristics of human language, which cannot be fully replicated in animal models, present a significant barrier to exploring the pathogenesis of PSA. Consequently, understanding the underlying neural pathways and identifying novel therapeutic targets is critically important. Recent evidence highlights the bidirectional communication between the gut microbiota and the brain, referred to as the microbiota-gut-brain axis. Our prior studies have demonstrated gut microbiota imbalances in patients with acute ischemic stroke, which significantly influence outcomes and prognosis. Considering the marked differences in disease severity and long-term prognosis between PSA patients and non-PSA patients, we hypothesized that variations in the microbiota-gut-brain axis might be linked to language function. To test this hypothesis, we employed functional magnetic resonance imaging (fMRI), 16S rDNA sequencing and enzyme-linked immunosorbent assay (ELISA) to explore differences in gut microbiota composition, neuroendocrine-immune network (NEI) network indicators, and brain network topology among PSA patients, non-PSA patients, and healthy controls (HCs). Our findings revealed that PSA patients, compared to non-PSA patients and HCs, exhibited gut microbiota dysbiosis, increased inflammatory responses, abnormal secretion of brain-gut peptides, and early activation of homologous language-related regions in the right hemisphere. These results provide new insights into the role of the gut microbiota in language recovery in PSA and highlight the gut microbiota as a promising therapeutic target for this condition.
Background
As one of the most common complications of stroke,1,2 approximately one-third of stroke survivors suffer from aphasia. 3 The presence of aphasia is a predictor of poor rehabilitation outcomes. 4 The presence of poststroke aphasia (PSA) is associated with more severe strokes, greater mortality, a decreased rate of functional recovery, and greater healthcare costs. 3 Due to the unique characteristics of human language, which are absent in animals, suitable animal models are currently unavailable for investigating this condition. This limitation has become a significant bottleneck in the investigation of the pathogenesis of PSA. Hence, an understanding of potential neural pathways and the identification of new therapeutic targets are vital.
Recent studies have shown that up to 50% of patients experience gastrointestinal complications (including constipation, dysphagia, and gastrointestinal bleeding) after a stroke.5–7 Gastrointestinal complications following a stroke can significantly impact patient outcomes, thus leading to delayed recovery, increased mortality rates, and the progression of neurological deficits.8–10 Accumulating evidence over the past decades has increasingly highlighted the significant role of poststroke changes in the microbiota-gut-brain axis (MGBA) in contributing to stroke pathology and as potential targets for future therapies. 11 The gut and central nervous system (CNS) are intimately connected through this axis, and it involves a complex neuroendocrine-immune (NEI) network, which allows for dynamic, bidirectional communication, and interaction. Ischemic stroke can induce local neuroinflammation and change peripheral immune homeostasis, which may result in dysfunction of the intestinal barrier, such as the disruption of tight junction proteins, 12 dysregulation of the sympathetic nervous system, 13 and impairment of gut motility, 10 thus leading to gut microbiota dysbiosis. 14 As a result, changes in the gut microbiota may represent one of the risk factors for ischemic stroke and determine stroke outcomes. An increasing number of studies have shown that gut dysbiosis exacerbates stroke severity, which can affect the prognosis after a stroke via various mechanisms, including microflora migration, intestinal bacterial metabolites, and immune regulation,15,16 which gradually improve with the recovery of stroke. 17 Additionally, gut dysbiosis may accelerate the progression of cognitive impairment. Language is unique to humans and is an important aspect of the cognitive system. Recent research has revealed that the inflammatory and metabolic pathways that are activated by gut microbial products can affect cognition, learning, memory, and other functions of the brain.18–20 In summary, existing studies have confirmed that gut dysbiosis is related to the onset of stroke and that the gut microbiota is rapidly dysregulated in acute stroke, which further exacerbates stroke. However, few studies have investigated changes in the gut microbiota in the subacute phase of stroke, especially regarding how the gut affects the brain in patients with PSA. Furthermore, few studies have explored the recovery of language after stroke from the perspective of the MGBA. When considering the potential effects of the microbiota on the brain that have been reported in recent studies, we hypothesized that the gut microbiota may cause CNS disorders via the NEI network in PSA patients.
Neuroplasticity is a constant and adaptive process that changes in response to intrinsic or extrinsic stimuli, the environment, and cerebral insults. This ability not only enables adaptive changes in action, perception, and cognition in the brain but also endows the brain with considerable potential for neuroplasticity during development and skill learning, as well as after cerebral injury. 21 The human brain has a complex topological structure that can be altered by many cognitive diseases. Language is a cognitive function of large-scale brain network cooperation, and local damage can cause global network dysfunction. Changes in functional connectivity in different brain regions after stroke are related to the development of and recovery from aphasia after stroke. Therefore, the investigation of the topological properties of the human brain is expected to reveal its structure and functional mechanism. The brain network is a complex system; however, we can reveal its rules of connection, as well as the processing and transmission mechanisms of information inside of the brain, by constructing functional connections and combining graph theory analysis methods. 22 As a result, we can further investigate and interpret the neuroplasticity mechanisms underlying language recovery in PSA patients.
In the present study, we employed a graph theory-based approach to systematically explore alterations in the brain networks of PSA patients. Moreover, we sought to further elucidate the underlying relationships among the microbiota, NEI network, brain network, and language function in patients with PSA. Given the notable disparities in disease severity and long-term prognosis between PSA and non-PSA stroke patients, we assumed that dissimilarities may exist in the microbiota and NEI networks of these two groups, which may be linked to language function. Thus, our objective was to investigate the reciprocal relationship between the gut microbiota and the brain in PSA patients with the aim of increasing the understanding of the interplay between PSA and gut health. A systematic flowchart of the study design is shown in Figure 1.
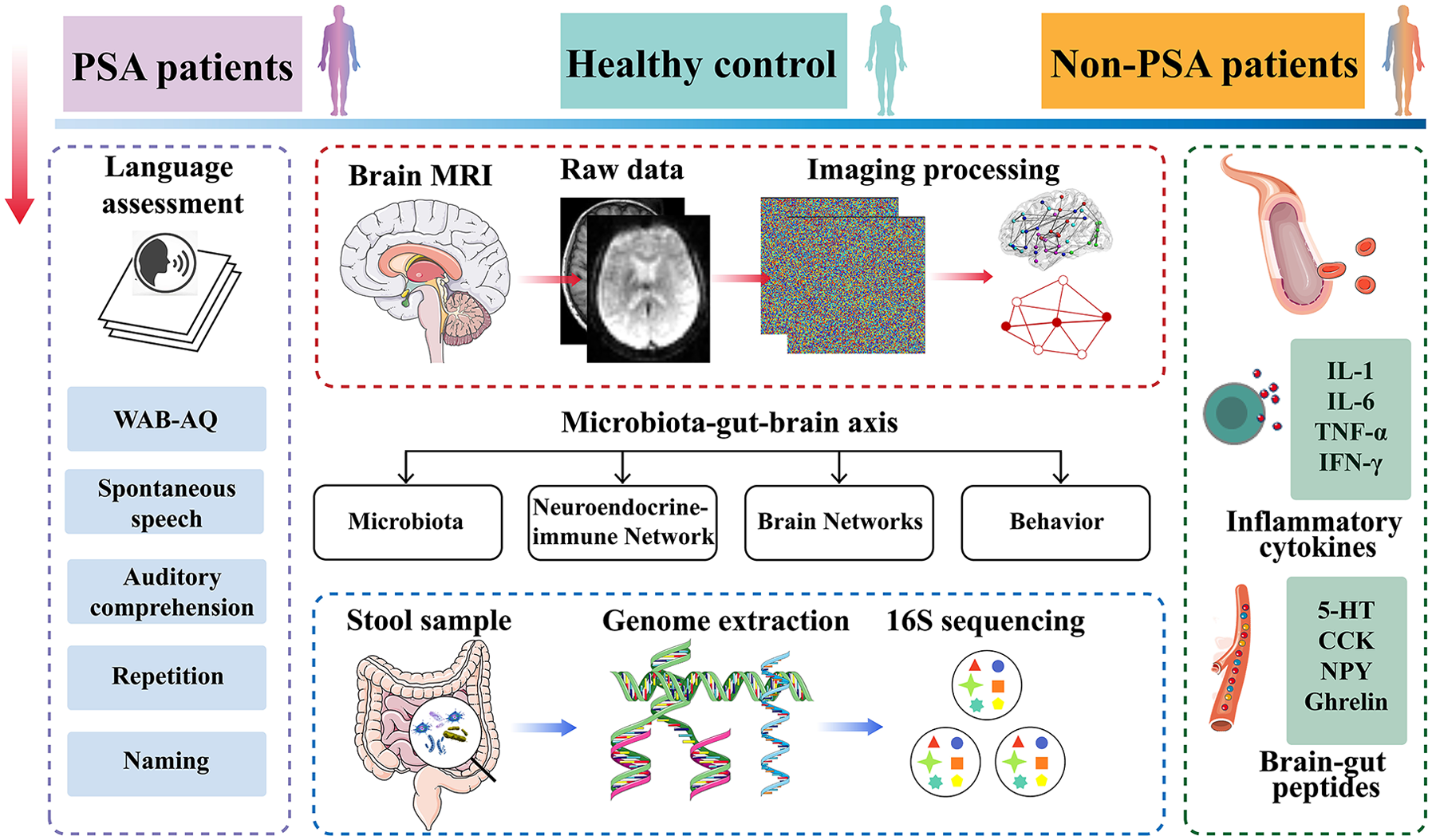
Flowchart of the study design.
Materials and methods
Participants
This study was approved by the Ethics Committee of Dongzhimen Hospital Affiliated to Beijing University of Chinese Medicine. Between December 10, 2020 and January 20, 2022, 15 PSA patients and 10 non-PSA stroke patients were enrolled from the Department of Neurology, Dongzhimen Hospital, Beijing University of Chinese Medicine; additionally, 15 healthy controls (HCs) were also recruited. PSA patients who met the following criteria were included: (1) were diagnosed with ischemic stroke via computed tomography or MRI methods at 14–30 days after symptom onset; (2) were aged between 18 and 80 years, and their native language were Chinese and right-handed; (3) at least a primary school and above education level with no serious heart, liver, or kidney diseases; (4) were fully conscious and without cognitive impairment; (5) had normal language function before stroke onset and dominant language dysfunction after stroke with mild limb dysfunction; (6) had a specific aphasia syndrome diagnosed as nonfluent aphasia after stroke via the Western Aphasia Battery-Chinese version (WAB-C); (7) scored between 2–4 on the Boston Diagnostic Aphasia Examination; and (8) able to cooperate for the 30-min MRI examination. The exclusion criteria were as follows: (1) severe hearing impairment, dysopia, or dysarthria; (2) language dysfunction caused by congenital or childhood diseases; (3) received pacemaker surgery, coronary intervention, or coronary artery bypass surgery or had other metal products found in the body; (4) history of brain tumor, brain trauma, or neuropsychiatric disorders; (5) use of any antibiotics, probiotics, or antimicrobial preparations in the previous 2 weeks before the start of the study; and (6) diagnosis of immune-mediated inflammatory disease, such as Crohn’s disease or ulcerative colitis. The inclusion criteria for the HCs were as follows: (1) closely matched age and educational levels with the PSA patient cohort; (2) right-handedness; (3) could undergo MRI examination and language assessments; and (4) did not use any antibiotics, probiotics, or antimicrobial preparations in the previous 2 weeks before the start of the study. By applying the inclusion and exclusion criteria for patients with PSA, we included non-PSA stroke patients who were matched with PSA patients for the size and location of the lesion. We present the following article in accordance with the Strengthening the Reporting of Observational Studies in Epidemiology (STROBE) reporting checklist (Supplemental Material).
Blood sample collection and analysis
Venous blood samples were collected from all of the participants after overnight fasting, placed at room temperature for stratification, and centrifuged at 3000 rpm for 10 min. Serum samples were collected, frozen at −80°C, and subsequently detected by enzyme-linked immunosorbent assay (ELISA). The inflammatory cytokines that were analyzed included interleukin-1 (IL-1) (pg/mL), interleukin-6 (IL-6) (pg/mL), tumor necrosis factor-alpha (TNF-α) (pg/mL), and interferon-gamma (IFN-γ) (pg/mL). The brain-gut peptides included 5-hydroxytryptophan (5-HT) (ng/mL), cholecystokinin (CCK) (pg/mL), ghrelin (pg/mL), and neuropeptide Y (NPY) (ng/L).
Stool sample collection and gut microbiota analysis
Stool samples were collected from each participant on the same day and promptly transferred them to a laboratory freezer set at −80°C for cryopreservation. DNA extraction was performed via the QIAamp® PowerFecal® DNA Kit (Qiagen, Hilden, Germany), and the DNA samples were quantified and validated. The V3–V4 hypervariable region of the 16S rRNA gene was selected as the target fragment for amplification and sequencing. To construct a polymerase chain reaction (PCR)-based 16S rDNA amplicon library for sequencing, PCR enrichment of the V4 hypervariable region of the 16S rDNA was performed with the forward primer 341F (5′-CCTACGGGNGGCWGCAG-3′) and reverse primer 806R (5′-GGACTACHVGGGTWTCTAAT-3′). The qualified amplicon mixture was then sequenced on the MiSeq platform via the PE250 sequencing strategy. Prior to 16S rDNA data analysis, the raw reads were processed to filter out adaptors, low-quality sequences, and ambiguous bases. Paired-end reads were subsequently assembled into tags using the Fast Length Adjustment of Short Reads (FLASH) program. The effective tags were clustered into operational taxonomic units (OTUs) with a cutoff value of 97% using UPARSE software v9.1.13 (https://www.drive5.com/uparse/), and chimeric sequences were compared with those in the Gold database using UCHIME (v4.2.40) for detection. The representative sequence from each OTU cluster was subsequently obtained. These sequences were taxonomically classified using the Ribosomal Database Project (RDP) Classifier (v.2.2) with a minimum confidence threshold of 0.8, and we utilized the Greengenes Database (v201305) as the reference training database. The USEARCH_global algorithm was used to compare all of the tags back to the OTUs to obtain the OTU abundance statistics for each sample. For within-sample comparisons, alpha diversity was assessed using the species richness indices (ACE and Chao1) and species diversity indices (Shannon and Simpson indices), which were calculated at the OTU level using MOTHUR software v1.35.1 (http://www.mothur.org/wiki/Download_mothur) and QIIME (v1.8.0). Analysis of similarities (ANOSIM) was performed to determine whether there was a significant difference between prefiltered and unfiltered communities by using two different beta diversity metrics: unweighted UniFrac distance and weighted UniFrac distance. Linear discriminant analysis effect size (LEfSe) was used to identify differences in microbial taxa across the three groups. Functional predictions using PICRUSt2 were performed based on a database of gene families and reference genomes, such as the Kyoto Encyclopedia of Genes and Genomes (KEGG) Orthology database. 23 The predicted pathways provided valuable insights into the functional profiles of the microbial communities. Genera showing significant differences were selected for further correlation analysis to explore potential associations between gut microbiota, brain network measures, language assessments, and NEI network-related factors.
Behavioral assessments
Prior to each MRI scan, the severity of language impairment in the patients with PSA was evaluated using the Chinese version of the WAB. The WAB consists of linguistic subtests, including spontaneous speech, auditory comprehension, repetition, and naming, as well as nonlinguistic subtests that evaluate reading, writing, praxis, and construction. In this study, only the linguistic subtests were analyzed. Additionally, the Aphasia Quotient (AQ) score, which provides an indication of the overall severity and type of aphasia, was calculated.
MRI data acquisition and preprocessing
In this study, MRI data were exclusively collected from patients with PSA and HCs. The MRI examinations were performed with a 3.0 Tesla scanner (Siemens AG, Erlangen, Germany) at Dongzhimen Hospital affiliated to Beijing University of Chinese Medicine. Sagittal structure images were acquired using a magnetization-prepared rapid gradient-echo three-dimensional (3D) T1-weighted sequence with the following parameters: repetition time (TR)/echo time (TE) = 1900 ms/2.13 ms, flip angle: 9°, inversion time = 1100 ms, voxel size: 1.0 × 1.0 × 1.0, resolution: 256 × 256, and slice thickness: 1 mm with no interslice gap. Rs-fMRI was performed using an echo-planar imaging sequence with the following parameters: TR/TE: 2000/30 ms, flip angle: 90°, field of view: 225 × 225 mm2, bandwidth = 2520, resolution: 64 × 64, slice thickness = 3.5 mm with a 0.7 mm interslice gap, 31 axial slices, and voxel size: 3.5 × 3.5 × 3.5. 24
The graph theoretical network analysis (GRETNA) toolbox (https://www.nitrc.org/projects/gretna) 25 was applied to preprocess the rs-fMRI data. The preprocessing steps included volume removal, slice timing correction, realignment, spatial normalization, and temporal filtering. Initially, the first 10 volumes from each participant were discarded to minimize the influence of magnetic disequilibrium on data quality. The remaining volumes underwent slice timing correction based on the middle slice, followed by realignment for head motion correction. Data were included only if the head motion was less than 3 mm or 3°. Subsequently, the functional images were spatially normalized to the Montreal Neurological Institute space using Advanced Normalization Tools (ANTS v1.9; https://www.nitrc.org/projects/ants) and resampled to a resolution of 3 mm × 3 mm × 3 mm. The signals were then bandpass filtered (0.01–0.08 Hz) to limit the effects of low-frequency drift and high-frequency physiological noise, with linear trends also being removed. Finally, confounding variables (including six head motion parameters, average global signals, white matter signals, and cerebrospinal fluid signals) were regressed to minimize their impact on the analysis.
Brain network construction
The topological properties of the brain networks were analyzed using the GRETNA toolbox. The Human Brainnetome Atlas was used to divide the brain into 48 regions, which were further subdivided into 246 subregions, with each subregion representing a network node. The Pearson correlation coefficients were calculated to define the edges of the brain network. A symmetric functional connectivity matrix (246 × 246) was constructed for subsequent network analysis. Using a weighted matrix, functional brain networks were constructed over the entire range of costs (0.05–0.17) at intervals of 0.01. Graph measures were calculated across this threshold range using functions implemented in the GRETNA software package.
The global network metrics analyzed in this study included clustering coefficient (Cp), Gamma, characteristic path length (Lp), Lambda, small-worldness (Sigma), global efficiency (Eg), and local efficiency (Eloc). Nodal metrics included degree centrality (DC), nodal clustering coefficient (NCC), nodal efficiency (NE), nodal local efficiency (NLE), and nodal shortest path (NLP). Cp describes the level of closeness to form a completely connected subgraph. 26 Gamma is the normalized Cp, based on random network values. Lp represents the mean shortest path length over all possible pairs of nodes, serving as a measure to quantify the functional integration level. 27 Lambda is the normalized Lp by random networks. Eg is the average efficiency of information transfer across the entire network, emphasizing parallel information transmission more directly than the shortest path length. 28 Eloc, closely related to the clustering coefficient, measures the efficiency of information flow within a node’s immediate neighborhood, reflecting local resilience or fault tolerance. 28 Small-world networks are characterized by a high clustering coefficient and a short characteristic path length relative to random networks, resulting in a small-worldness index greater than one. At the nodal level, NE is defined as the average inverse shortest path length between a specific node and all other nodes in the network. 27 NCC quantifies the likelihood of interconnection among a node’s neighbors, indicating the local cohesiveness within the network. DC, also known as degree, is the most straightforward measure of centrality. 29 NLE evaluates the efficiency of information transfer within a node’s local subnetwork. 30 NLP reflects the ability to efficiently transmit information within a network, helping to conserve system resources and reduce energy consumption. 31 To evaluate the global and nodal topological characteristics of the brain networks, the area under the curve (AUC) was computed for each parameter. In the group analysis, AUC values of global and nodal network properties are used to compare the differences in network connectivity between PSA patients and HCs.
Statistical analysis
Statistical analysis of the demographic and clinical data was performed via SPSS software (IBM SPSS Inc., Chicago, IL, USA) and GraphPad Prism 9.3.0. Categorical variables are presented as absolute frequencies and percentages, with comparisons made using Pearson’s Chi-squared test. The normality of the continuous variables are assessed via Q-Q plots and the Shapiro-Wilk test. Normally distributed continuous data are presented as the means ± standard deviations, and comparisons were conducted using either two-sample t tests or Bonferroni’s post-hoc test after analysis of variance (ANOVA). For nonnormally distributed variables, the Mann-Whitney nonparametric or Kruskal-Wallis tests were employed, and expressed as median with the interquartile range (IQR: P25, P75).
Between-group differences in network measures were determined via independent two-sample t tests in the GRETNA toolbox, with age, sex, and educational level scores included as covariates to control for their potential effects. Furthermore, correlation analyses were conducted between pairs of factors showing significant group differences, including the gut microbiota, inflammatory cytokines, brain-gut peptides, brain network measures, and language assessment scores. Pearson correlation was applied for normally distributed data, whereas Spearman correlation was utilized for nonnormally distributed data. A correlation network was constructed by using factors that exhibited significant correlations across all of the subjects. Statistical significance was defined as a p value of <0.05.
Results
Demographic and clinical information of the participants
There were no significant intergroup differences in terms of age, sex, or education level (all p > 0.05). Table 1 shows the demographic information and language assessment scores of the three groups. The lesion overlap map for all of the patients with PSA is presented in Figure 2.
Demographics, language assessments between PSA, non-PSA, and HC.

AQ, Aphasia Quotient; HC, healthy control; NA, not available; PSA, poststroke aphasia; WAB, Western Aphasia Battery.

Lesion overlap map for the 15 patients with PSA in MNI space. Color bar represents the number of patients with overlapping lesions at certain locations. The numbers refer to the MNI coordinate space in the z plane.
Analysis of the gut microbiota in patients and HCs
The differences in the α diversity of the gut microbiota between the PSA group and HC group were significant (p < 0.05) according to the Chao1 indices (Figure 3(b)), and the Shannon and Simpson indices were lower in the non-PSA patients than in the HCs (p < 0.05) (Figure 3(c) and (d)). However, no significant difference in the α diversity of the gut microbiota was found between patients with PSA and those without PSA (p > 0.05).

Analysis of the gut microbiota in patients with PSA. (a) Boxplots for comparison of the ACE index. (b) Boxplots for comparison of the Chao1 index. (c) Boxplots for comparison of the Shannon index. (d) Boxplots for comparison of the Simpson index. (e) ANOSIM based on the beta diversities between the samples (weighted UniFrac distances) revealed different distributions among the groups. (f) ANOSIM based on the beta diversities between the samples (unweighted UniFrac distances) revealed different distributions among the groups. (g) Histogram of the community composition of the gut microbiota at the phylum level. The abscissa represents the group, and the ordinate represents the relative abundance. (h) Histogram of the community composition of the gut microbiota at the genus level. The abscissa represents the group, and the ordinate represents the relative abundance. (i) Cladogram depicting the phylogenetic distribution of microbial lineages associated with the three groups. Each small circle at a different classification level represents a classification at that level, and the diameter of the small circle is proportional to the relative abundance. (j) Histogram of LDA scores to identify differentially abundant bacteria across the three groups (LDA score ⩾4.0). (k) Histogram of LDA scores to identify differentially enriched KEGG pathways across the three groups (LDA score ⩾2.0).
ANOSIM based on weighted UniFrac distances (R = 0.144, p = 0.004) and unweighted UniFrac distances (R = 0.117, p = 0.011) was performed to determine the β diversity (between-sample diversity) of the gut microbiota, which was significantly different among the groups (Figure 3(e) and (f)). Analysis at the phylum level revealed that Bacteroidetes were more abundant in PSA patient samples (11.25%) than in non-PSA samples (3.53%) and 8.14% in HCs. Actinobacteria were present at lower levels in the PSA group (24.02%) than in the non-PSA group (47.95%) (Figure 3(g)). A histogram of the community composition at the genus level revealed that g_Bifidobacterium was relatively highly abundant in the non-PSA group (38.46%), whereas the abundance of g_Blautia was relatively high in the PSA group (11.44%) (Figure 3(h)).
To identify potential bacterial taxon biomarkers for distinguishing each group, linear discriminant analysis (LDA) effect size (LEfSe) was used to identify species with abundances that significantly differed between the groups. The gut microbiome of the PSA group was mainly enriched in p_Bacteroidetes, c_Bacteroidia, o_Bacteroidales, and g_Bacteroides, whereas that of the non-PSA group was mainly enriched in p_Actinobacteria, c_Actinobacteria, and o_Bifidobacteriales (Figure 3(i) and (j)). One KEGG pathway related to butanoate metabolism was significantly enriched in the PSA patient group (Figure 3(k)).
Analysis of neuroendocrine-immune network in patients and HCs
In terms of indicators of the neuroendocrine-immune network, PSA patients exhibited higher IL-1, IL-6, TNF-α, CCK, and NPY levels than non-PSA patients and HCs (Figure 4(a),(b),(c),(f) and (h)), whereas the levels of 5-HT and ghrelin exhibited lower magnitude than the other two groups (all p < 0.01) (Figure 4 (e) and (g)).
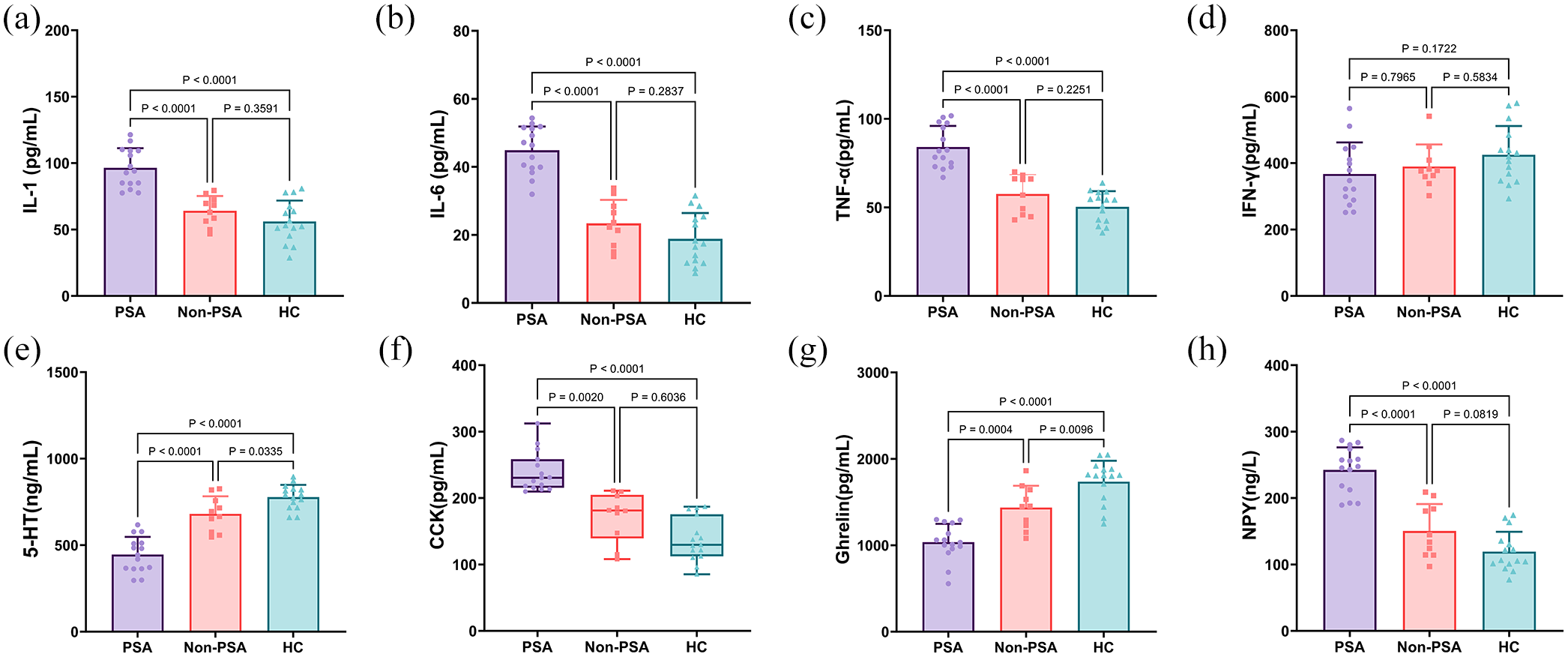
Related indexes of the neuroendocrine-immune network between PSA, non-PSA, and HC. (a) Comparison of IL-1 expression in different groups. (b) Comparison of IL-6 expression in different groups. (c) Comparison of TNF-α expression in different groups. (d) Comparison of IFN-γ expression in different groups. (e) Comparison of 5-HT expression in different groups. (f) Comparison of CCK expression in different groups. (g) Comparison of Ghrelin expression in different groups. (h) Comparison of NPY expression in different groups.
Differences between PSA patients and HCs in global network measures of the whole brain
Over the entire range of thresholds (from 0.05 to 0.17), both PSA patients and HCs exhibited small-world properties (gamma value obviously larger than 1 and lambda value close to 1) of the whole brain. However, there were no significant intergroup differences in the global network measures. The detailed global network measures are listed in Table 2.
Differences in global network metrics of the whole brain between PSA patients and HCs.

aCp, the area under curve of clustering coefficient; aEg, the area under curve of global efficiency; aEloc, the area under curve of local efficiency; aGamma, the area under curve of Gamma; aLambda, the area under curve of Lambda; aLp, the area under curve of characteristic path length; aSigma, the area under curve of Sigma; HC, healthy control; PSA, poststroke aphasia.
Differences in nodal network measures of the whole brain
To investigate the alterations in nodal strength in PSA patients, we analyzed several nodal characteristics. Compared with HCs, PSA patients exhibited decreased DC, NCC, NE, and NLE in the left middle frontal gyrus (MFG), postcentral gyrus (PoG), inferior parietal lobule (IPL), and inferior temporal gyrus (ITG) (all p < 0.001, false discovery rated-corrected). Moreover, PSA patients also exhibited increased nodal properties in the right insula (INS), amygdala (Amyg), hippocampus (Hipp), and ITG as well as in the left parahippocampal gyrus (PhG), precentral gyrus (PrG) and PoG (all p < 0.001, false discovery rated-corrected) (Figure 5).
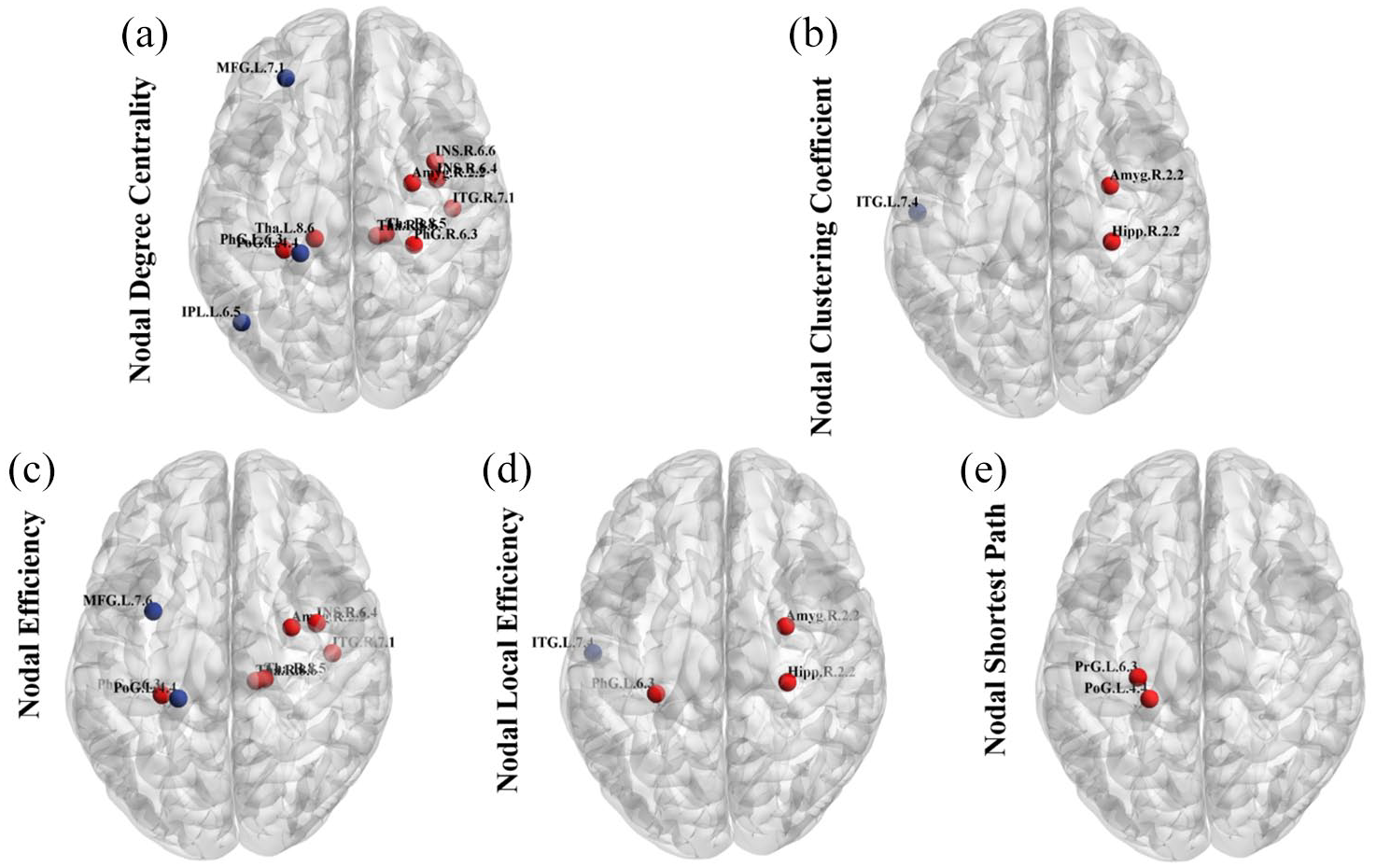
Differences in nodal network measures of the whole brain. The figures respectively display the comparisons of DC (a), NCC (b), NE (c), NLE (d) and NLP (e) of the networks across two groups of individuals. Nodes with distinct differences are exhibited in figures, and marked with their brain region abbreviations. The blue nodes indicate decreased nodal properties, while red nodes indicate increased nodal properties.
Correlation networks of the gut microbiota, neuroendocrine-immune network, brain network measures and language assessments
To explore the relationships between disease characteristics and the MGBA in greater detail, a correlation network of the gut microbiota, the neuroendocrine-immune network, brain network measures, and language assessments was constructed with the factors that exhibited significant correlations. Correlations were considered to be significant when the R value was >0.6 and the p value was <0.05 (Figure 6). The constructed network revealed intricate interconnections among these factors, thus highlighting the inherent associations within the complex dynamics of brain-gut interactions.
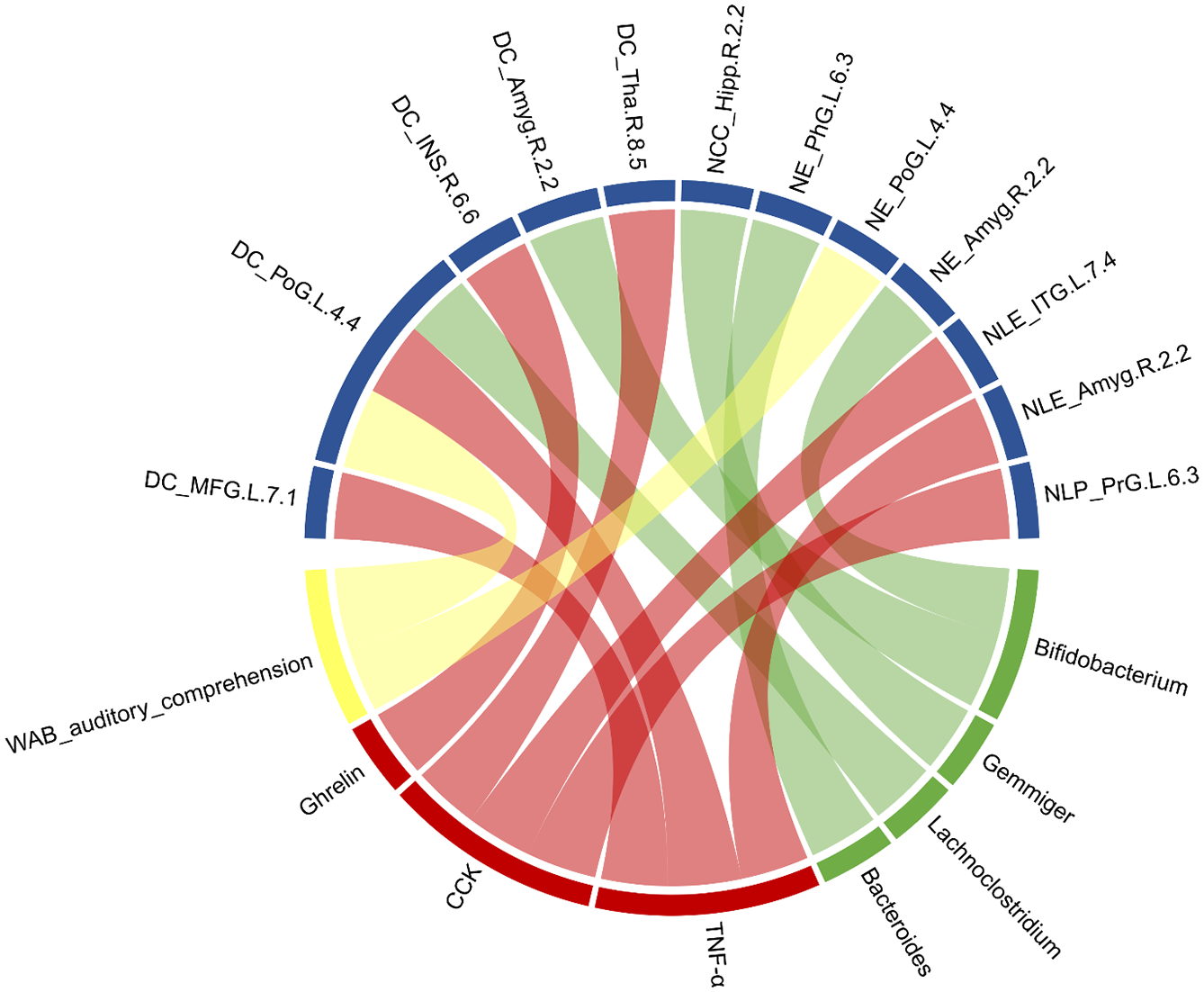
Correlation networks of gut microbiota, neuroendocrine-immune network, brain network measures, and language assessments. Green block represents gut microbiota; yellow block represents subtests of language assessment; red block represents neuroendocrine-immune network-related factors; blue block represents brain network parameters.
Discussion
Acute ischemic stroke induces dysbiosis of the microbiome, and these resulting changes in the gut microbiota correspondingly affect neuroinflammatory processes and influence stroke outcomes. As one of the common presentations of stroke, aphasia has been found to increase the length of hospital stay, incidence of inpatient complications, overall neurological disability, and mortality of patients, as well as alter the discharge disposition of patients. 32 Therefore, the accurate knowledge of the factors and mechanisms associated with aphasia is essential for providing optimal care for patients with PSA.
This study tested the hypothesis that there was an imbalance in the gut microbiota in patients with PSA, accompanied by NEI network disorders and abnormal changes in the brain network. To the best of our knowledge, this is the first study to examine the associations between functional MRI data, language function, gut microbiota, and the NEI network. These findings may expand on the potential mechanisms underlying the gut-brain interaction in PSA and provide novel insights into new therapeutic targets for PSA via the use of probiotics.
In the network analysis at the global level, although there were no significant intergroup differences in the global topological properties of the whole brain network between the two groups, our results revealed that both PSA patients and controls exhibited small-world properties in the whole brain network. Small-world organization, which reflects an optimal balance of integration and segregation, may fundamentally enhance the brain’s dynamic stability and information processing abilities. 33 Notably, compared with HCs, PSA patients exhibited an increasing trend in Gamma, Lambda, and Sigma values. The increase in the Gamma value in PSA patients was considered to indicate altered short-distance functional connections between adjacent regions. The lower Lambda value may be associated with higher efficiency among nearby brain regions. Furthermore, the increase in the Sigma value may suggest that patients with aphasia had greater local specialization and global integration. A previous study revealed that patients with PSA with courses of disease ranging from 1 to 24 months had decreased Sigma and Gamma values, 34 which indicated that the capacity for information exchange and regional information processing had been disrupted. In contrast to previous studies, in this study, the patients with PSA were in the subacute phase after stroke (within 14–30 days poststroke). During the early poststroke stage, spontaneous remodeling of neural networks is accompanied by functional recovery. 35 The upregulation of the entire language network has been regarded as being an early mechanism, that begins within hours to days after a stroke. Saur et al. reported a model with three phases of language recovery, which suggested that the early overactivation of the language network is necessary to compensate for stroke. 36 This observation is generally consistent with the result of our present study. Compared with HCs, patients with aphasia exhibited decreased node attributes in the MFG, PoG, IPL, and ITG of the left hemisphere and increased node attributes in the INS, Amyg, Hipp, and ITG of the right hemisphere. The neural mechanisms underlying language recovery following left hemisphere stroke remain poorly understood. Among the handful of acute-to-chronic longitudinal PSA studies that have been performed, several have associated better language recovery with increased activation in regions of both hemispheres. The engagement of preserved left hemisphere language areas and the recruitment of nearby perilesional tissues are thought to support aphasia recovery after stroke. Patients with smaller left hemisphere lesions and milder aphasia may be able to recruit residual left hemisphere language-capable areas 37 and thereby exhibit little right hemisphere activity. In contrast, individuals with larger left hemisphere lesions typically experience more severe aphasia due to extensive damage to the native left hemisphere language network; thus, they primarily depend on compensatory mechanisms in the right hemisphere. 38 Notably, the involvement of the right hemisphere in recovery appears to be dynamic and evolves over time, which often peaks in the early stages of recovery and gradually shifts back to the left hemisphere as the recovery process progresses. 39 This temporal pattern highlights the adaptive nature of neural mechanisms in poststroke language recovery.
Intestinal barrier dysfunction, microbial translocation, and local inflammation play crucial roles in the initiation and progression of stroke. Dysbiosis of the gut microbiota is a major contributing factor to intestinal damage. Interestingly, the dysbiosis of the intestinal microenvironment in the PSA group was characterized by increased relative abundances of Bifidobacterium, Blautia, and Bacteroides, as well as significantly lower abundances of Faecalibacterium and Eubacterium, compared to the healthy control group. Opportunistic pathogens such as Blautia, Prevotella were more abundant, whereas probiotic microbes such as Bifidobacterium were less abundant, in PSA patients than in non-PSA patients. Similar trends were also demonstrated in the LEfSe results. Many clinical and preclinical studies have established that alterations in Bacteroides lead to a wide range of neuropsychiatric disorders, such as attention-deficit/hyperactivity disorder, 40 multiple sclerosis, 41 and Alzheimer’s disease. 42 Zhang et al. 43 found that Bacteroides may play a critical role in axon development and regeneration by influencing glycerolipid and glycerophospholipid metabolism. Bifidobacterium is considered a type of beneficial bacteria in the human intestine, and it is known to metabolize lactose. A previous study confirmed that Bifidobacterium treatment successfully improved the long-term rehabilitation of mice with cerebral ischemia. 44 In terms of the metabolic pathways, butanoate metabolism was significantly enriched in the PSA patient group. Butyric acid is a short-chain fatty acid, that plays an important role in maintaining the integrity of the intestinal barrier.45,46 Butyric acid is also the major end product of lactate and acetate fermentation, and this fermentation process converts virtually all of the lactate and acetate to butyrate. 47 Our analysis revealed that Faecalibacterium and Eubacterium were relatively more abundant in HCs than in the other two groups, whereas the abundance of Bifidobacterium was relatively greater in the PSA and non-PSA group than in the HCs. Faecalibacterium and Eubacterium are known as butyrate-producing bacteria,48,49 and a decrease in these bacteria may lead to a reduction in butyrate content, as well as cause further damage to the intestinal barrier and lead to the production of proinflammatory cytokines. We hypothesize that patients with PSA experience a compensatory increase in the abundance of lactate-producing bacteria, which potentially occurs as a response to the depletion of butyrate-producing bacteria in the gut. This shift may indicate an adaptive mechanism in which a reduction in butyrate levels triggers the proliferation of lactate producers to increase lactate production, thereby promoting butyrate synthesis and restoring balance. Further correlation analysis revealed a positive association between the abundance of Bifidobacterium and both the DC and NE of Amyg.R. The amygdala, which is a critical region of the limbic system, has been extensively investigated for its role in emotion regulation and behavioral modulation. Preliminary data have suggested that the composition of the gut microbiota is associated with altered functional connectivity between the amygdala and anterior insula in young children, which is also predictive of cognitive performance at 2 years of age. 50
Brain-gut interactions reflect the functional regulation of the NEI network system based on the brain-gut axis at different levels. Inflammation is among the main pathological characteristics of stroke. In this study, compared with HCs and non-PSA patients, PSA patients exhibited significant increases in the levels of serum proinflammatory cytokines (including IL-1, IL-6, and TNF-α) in the peripheral blood, and these changes in cytokines were correlated with language function. These findings suggest that the inflammatory response could be one of the factors leading to the decline in language function. Brain-gut peptides are small-molecule polypeptide substances that are secreted by endocrine cells of the brain-gut axis and are observed at a twofold higher amount in the gastrointestinal tract and nervous system. Our results revealed that PSA patients exhibited higher CCK and NPY levels than HCs and non-PSA patients, whereas the levels of 5-HT and ghrelin were lower than those in the other groups. CCK is predominantly produced and secreted by intestinal I cells. 51 Elevated cerebral CCK levels and abnormal CCK secretion following acute stroke stimulate the enteric neurons to increase gastrointestinal CCK secretion, which affects the brain or activates the central nerves via vagal afferent fibers to induce neurological deficits. 52 5-HT is a growth factor, a paracrine factor, and an enteric neurotransmitter, 53 that is mainly observed in gut enterochromaffin cells. The modulating of 5-HT has been reported to alter motor cortex excitability and promote motor recovery in stroke patients. 54 Zhou et al. reported that ghrelin can function as a neuroprotective agent and that the peripheral administration of ghrelin can protect neurons from ischemic injury caused by middle cerebral artery occlusion, which may constitute a novel therapeutic strategy for the treatment of ischemic stroke. 55 More significantly, compared with non-PSA patients, patients with PSA had elevated levels of inflammatory cytokines, thus indicating that PSA has a more extensive inflammatory response. Moreover, the levels of brain-gut peptides in PSA patients also exhibited significantly different from those in non-PSA, suggesting that abnormal secretion of brain-gut peptides may related to impaired language function after stroke. Further correlation analysis revealed that the indicators that are involved in the NEI network were correlated with the node attributes in brain regions and the language function subitems.
The recovery of neurological deficits after ischemic stroke (particularly in the subacute phase) primarily relies on neuroplasticity, which is represented by the ability of the brain to reorganize and form new neural connections. During this critical period, stroke recovery is most effective, with neuroplasticity playing a key role in restoring function. The gut-brain axis, which represents a research area of growing interest in biomedicine and neuroscience, reveals that the gut microbiota can significantly influence brain function and behavior. Changes in the composition and diversity of gut microbes have been linked to stroke development, its sequelae, and recovery outcomes. In addition to the NEI network that was involved in this study, the gut microbiota can also promote stroke recovery by influencing neuroplasticity via the regulation of metabolites, neurotransmitter balance, and neurotrophic factor production. Language is a unique human function; although the role of the gut microbiota in overall stroke recovery has gained increased attention, studies that have specifically examined its impact on aphasia are still limited. This study suggests that gut microbiota dysbiosis in PSA patients is associated with disorder in the NEI network and abnormal alterations in brain functional networks, potentially affecting the recovery mechanisms of language function. By modulating inflammation, brain-gut peptides, and gut microbiota may help in restoring the connectivity and function of brain areas that are responsible for language. These findings may provide a valuable reference for the further exploration of the potential micromechanisms underlying language function recovery after stroke.
Although we have provided the first evidence of changes in the brain-gut axis in PSA patients, there are several limitations that should be mentioned. First, this study has a cross-sectional design. Longitudinal studies will be needed to track dynamic changes in the MGBA during aphasia recovery after stroke, providing deeper insights into how these interactions evolve over time and their potential impact on language recovery mechanisms. The second limitation is that disorders of the gut microbiota can regulate the host pathophysiological state by affecting multiple pathways, including the NEI network and metabolism. Although this study revealed that a decrease in butyric acid-producing bacteria and a compensatory increase in lactic acid-producing bacteria may be the underlying mechanism of PSA in the early stage, these metabolites were not detected; thus, no definite conclusions can be determined. Metabolomics can be introduced in future studies to provide more direct evidence for the downstream pathways of the gut microbiota. Third, this was a single-country study, limited to the Chinese population. Therefore, external validation in other countries is warranted. Fourth, as an exploratory study with a small sample size, this study can only preliminarily analyze the relationship between indicators but cannot determine whether there is a causal relationship in the correlation, which still needs further verification.
Conclusion
Language function is complex, and there is still a lack of animal models regarding this phenomenon, which limits the investigation of the pathogenesis and development mechanism of PSA. The application of fMRI and 16s rDNA provides technical support for understanding the mechanism of brain-gut interactions. We found that patients with PSA in the early stage had gut microbiota disorders, aggravated inflammatory responses, abnormal secretion of brain-gut peptides, and activation of the homologous language-related regions in the right hemisphere. Thus, we speculate that the incidence of stroke rapidly causes the disruption of the gut microbial community and that the gut microbiota may be involved in regulating the NEI network, which correspondingly induces alterations in functional brain networks. In summary, the current findings may provide important context to the growing literature on the effects of the gut microbiome on human behaviors, as well as potentially leading to the gut microbiota being utilized as a promising target for treating PSA.
Supplemental Material
sj-docx-1-tan-10.1177_17562864251319870 – Supplemental material for Crosstalk between the gut microbiota and brain network topology in poststroke aphasia patients: perspectives from neuroimaging findings
Supplemental material, sj-docx-1-tan-10.1177_17562864251319870 for Crosstalk between the gut microbiota and brain network topology in poststroke aphasia patients: perspectives from neuroimaging findings by Yun Cao, Jiaqin Huang, Danli Zhang, Jianguang Ji, Xiaojing Lei, Zhongjian Tan and Jingling Chang in Therapeutic Advances in Neurological Disorders
Footnotes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
